hfd
Evgenii O. Tretiakov
2022-06-24
Last updated: 2022-11-04
Checks: 6 1
Knit directory: 1_heteroAstrocytes/
This reproducible R Markdown analysis was created with workflowr (version 1.7.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of
the R Markdown file created these results, you’ll want to first commit
it to the Git repo. If you’re still working on the analysis, you can
ignore this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20220624) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 9cf756d. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: analysis/figure/
Ignored: data/deng2019_arc_chow.h5Seurat
Ignored: data/deng2019_arc_chow.h5ad
Ignored: data/deng2019_arc_chow_astrocytes.h5Seurat
Ignored: data/deng2019_arc_chow_astrocytes.h5ad
Ignored: data/deng2019_arc_chow_astrocytes_fin.h5Seurat
Ignored: data/deng2019_arc_chow_astrocytes_fin.h5ad
Ignored: data/deng2019_arc_chow_clusters.h5Seurat
Ignored: data/deng2019_arc_chow_clusters.h5ad
Ignored: data/deng2019_arc_hfd.h5Seurat
Ignored: data/deng2019_arc_hfd.h5ad
Ignored: data/deng2019_arc_hfd_astrocytes.h5Seurat
Ignored: data/deng2019_arc_hfd_astrocytes.h5ad
Ignored: data/deng2019_arc_hfd_astrocytes_fin.h5Seurat
Ignored: data/deng2019_arc_hfd_astrocytes_fin.h5ad
Ignored: data/deng2019_arc_hfd_clusters.h5Seurat
Ignored: data/deng2019_arc_hfd_clusters.h5ad
Ignored: data/deng2019_arc_refi_astrocytes.h5Seurat
Ignored: data/deng2019_arc_refi_astrocytes.h5ad
Ignored: output/figures/
Unstaged changes:
Modified: analysis/hfd.Rmd
Modified: output/tables/arc_ast.csv
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (analysis/hfd.Rmd) and HTML
(docs/hfd.html) files. If you’ve configured a remote Git
repository (see ?wflow_git_remote), click on the hyperlinks
in the table below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 9cf756d | Evgenii O. Tretiakov | 2022-11-03 | v2022-06-26 + small fixes |
Introduction
Here we use dataset of adult mice hypothalami Arcuate nuclei single nucleus RNA-seq from paper @dengSingleNucleusRNASequencing2020 We’re going to use both parts of data: first, we will use only astrocytes from control mice for data integration against whole hypothalamus matrix; in second, part we’re going to add another variable - normal chow vs high fat diet.
First we need to load packages that we need for processing, setup environment and load data.
Load datasets
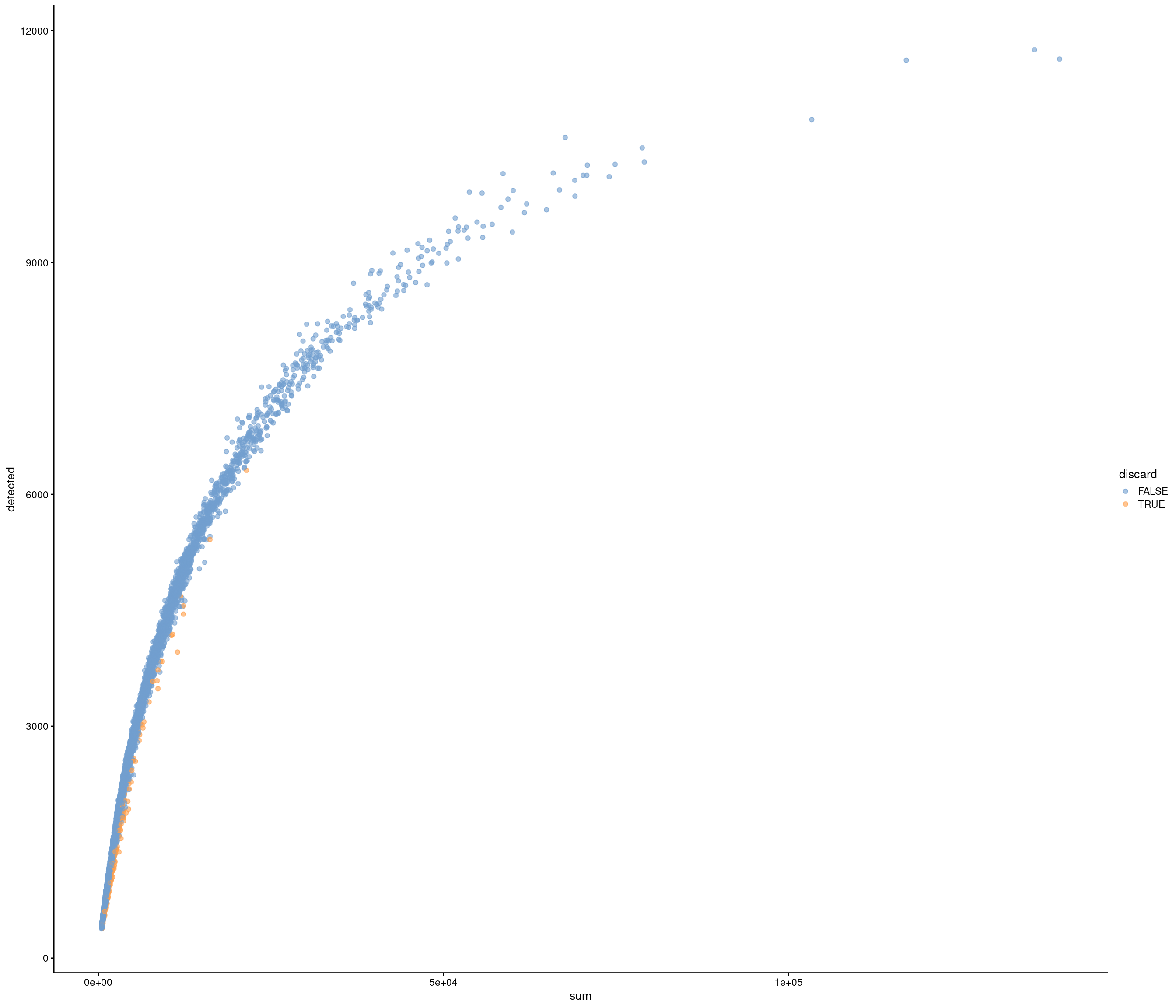
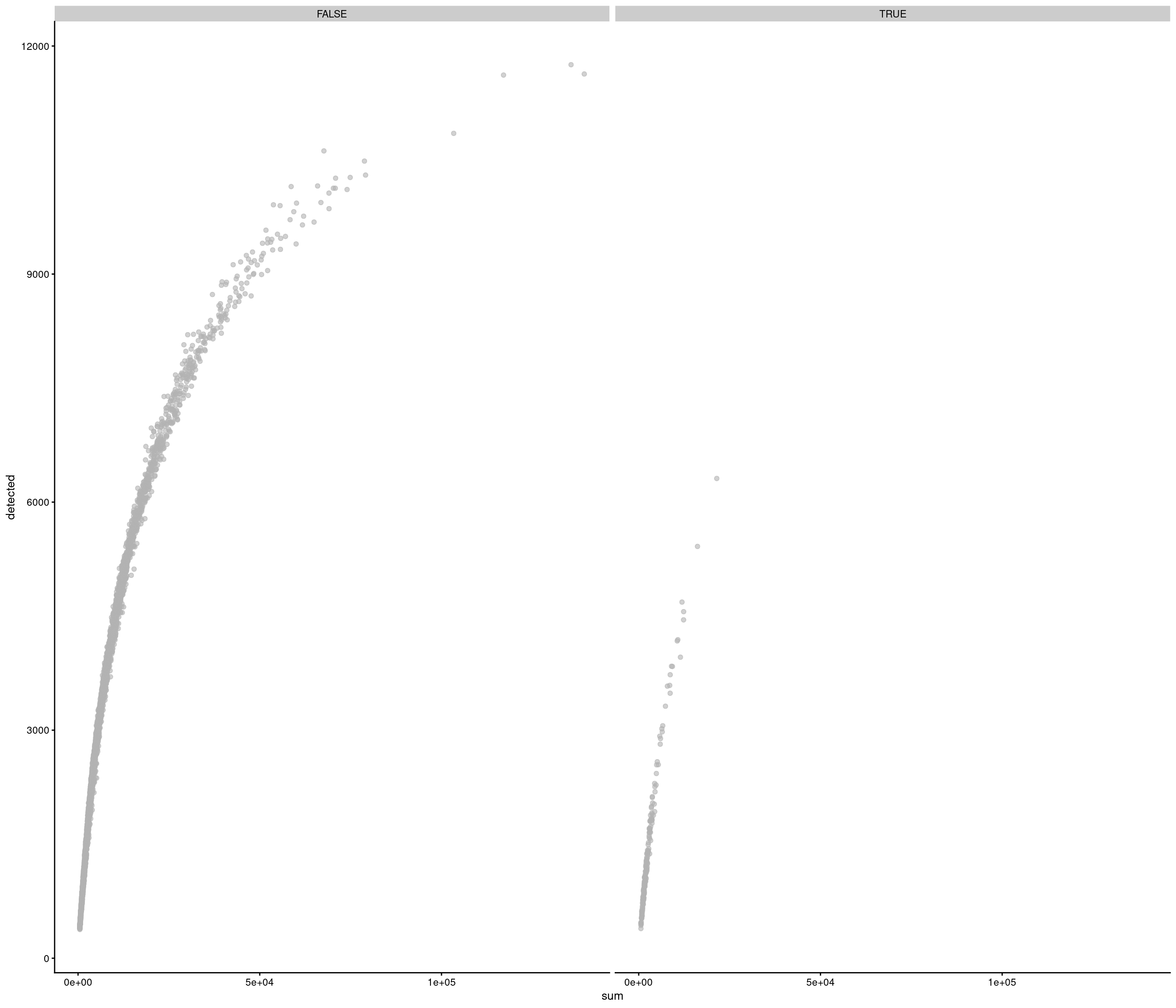
Add QC metrics and filter
Chow-diet samples
1. 536-1
$mito
[1] "mt-Nd1" "mt-Nd2" "mt-Co1" "mt-Co2" "mt-Atp8" "mt-Atp6" "mt-Co3"
[8] "mt-Nd3" "mt-Nd4l" "mt-Nd4" "mt-Nd5" "mt-Nd6" "mt-Cytb"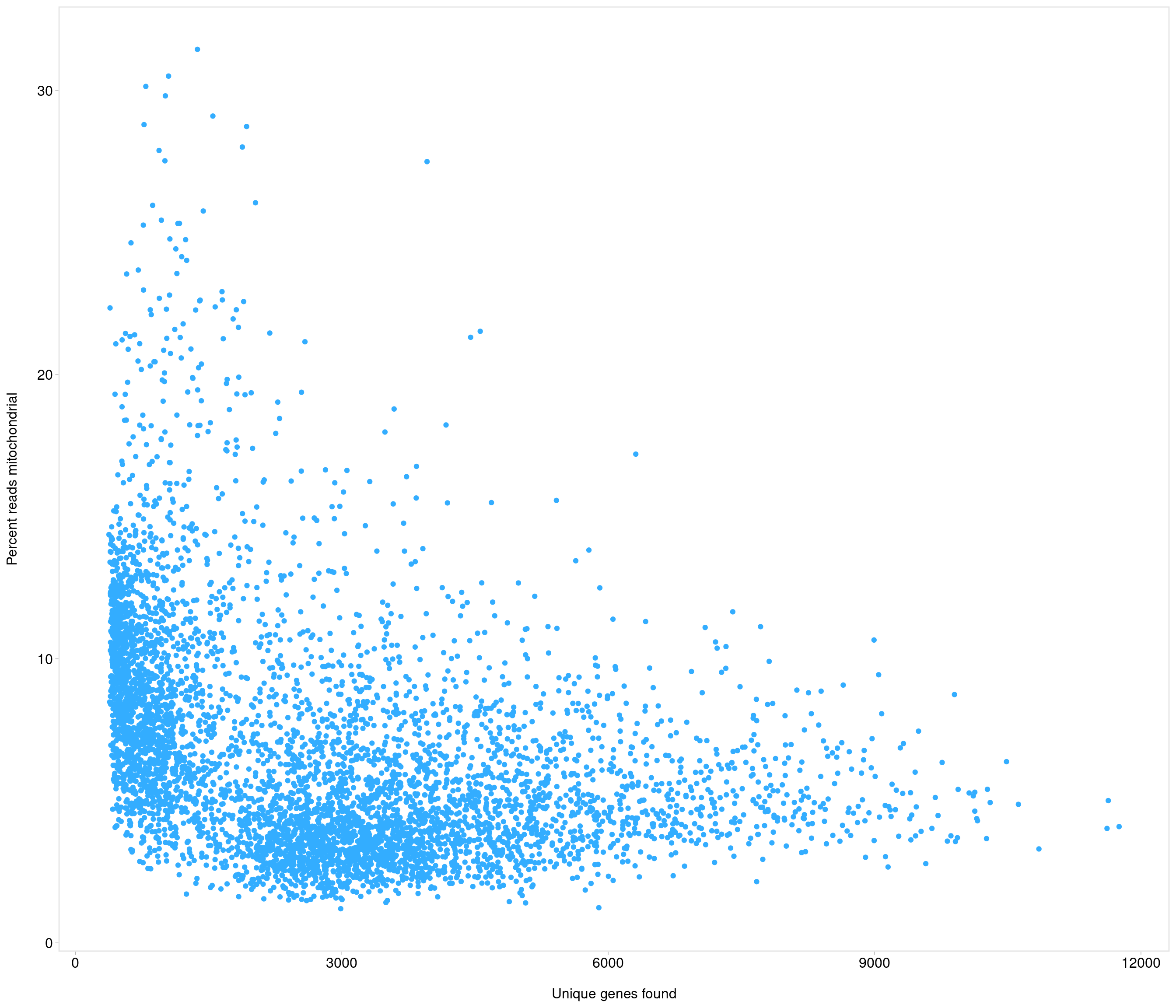
DataFrame with 6 rows and 19 columns
orig.ident nCount_RNA nFeature_RNA age
<factor> <numeric> <integer> <character>
AAACCCACAAGTCATC-1 536-1_chow-diet 5887 2916 adult
AAACCCACATACTTTC-1 536-1_chow-diet 1636 1102 adult
AAACCCAGTAAGAACT-1 536-1_chow-diet 1095 790 adult
AAACCCAGTGGTAATA-1 536-1_chow-diet 1737 1233 adult
AAACCCATCCGATGTA-1 536-1_chow-diet 4808 2598 adult
AAACCCATCGTCAAAC-1 536-1_chow-diet 7046 3498 adult
sex study_id tech hfd
<character> <character> <character> <logical>
AAACCCACAAGTCATC-1 male deng_2020 10xv3 FALSE
AAACCCACATACTTTC-1 male deng_2020 10xv3 FALSE
AAACCCAGTAAGAACT-1 male deng_2020 10xv3 FALSE
AAACCCAGTGGTAATA-1 male deng_2020 10xv3 FALSE
AAACCCATCCGATGTA-1 male deng_2020 10xv3 FALSE
AAACCCATCGTCAAAC-1 male deng_2020 10xv3 FALSE
ident sum detected subsets_mito_sum
<factor> <numeric> <integer> <numeric>
AAACCCACAAGTCATC-1 536-1_chow-diet 5887 2916 879
AAACCCACATACTTTC-1 536-1_chow-diet 1636 1102 128
AAACCCAGTAAGAACT-1 536-1_chow-diet 1095 790 123
AAACCCAGTGGTAATA-1 536-1_chow-diet 1737 1233 108
AAACCCATCCGATGTA-1 536-1_chow-diet 4808 2598 496
AAACCCATCGTCAAAC-1 536-1_chow-diet 7046 3498 298
subsets_mito_detected subsets_mito_percent total
<integer> <numeric> <numeric>
AAACCCACAAGTCATC-1 13 14.93120 5887
AAACCCACATACTTTC-1 12 7.82396 1636
AAACCCAGTAAGAACT-1 11 11.23288 1095
AAACCCAGTGGTAATA-1 11 6.21762 1737
AAACCCATCCGATGTA-1 11 10.31614 4808
AAACCCATCGTCAAAC-1 12 4.22935 7046
low_lib_size low_n_features high_subsets_mito_percent
<outlier.filter> <outlier.filter> <outlier.filter>
AAACCCACAAGTCATC-1 FALSE FALSE FALSE
AAACCCACATACTTTC-1 FALSE FALSE FALSE
AAACCCAGTAAGAACT-1 FALSE FALSE FALSE
AAACCCAGTGGTAATA-1 FALSE FALSE FALSE
AAACCCATCCGATGTA-1 FALSE FALSE FALSE
AAACCCATCGTCAAAC-1 FALSE FALSE FALSE
discard
<logical>
AAACCCACAAGTCATC-1 FALSE
AAACCCACATACTTTC-1 FALSE
AAACCCAGTAAGAACT-1 FALSE
AAACCCAGTGGTAATA-1 FALSE
AAACCCATCCGATGTA-1 FALSE
AAACCCATCGTCAAAC-1 FALSE| Name | sce$sum |
| Number of rows | 5436 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 6986.62 | 9170.88 | 500 | 1481 | 4455 | 8354.25 | 139282 | ▇▁▁▁▁ |
| Name | sce$detected |
| Number of rows | 5436 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 2855.79 | 2043.79 | 378 | 1015.5 | 2617 | 3956 | 11754 | ▇▆▂▁▁ |
| Name | sce$subsets_mito_percent |
| Number of rows | 5436 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 6.68 | 3.91 | 1.2 | 3.84 | 5.64 | 8.53 | 31.45 | ▇▃▁▁▁ |
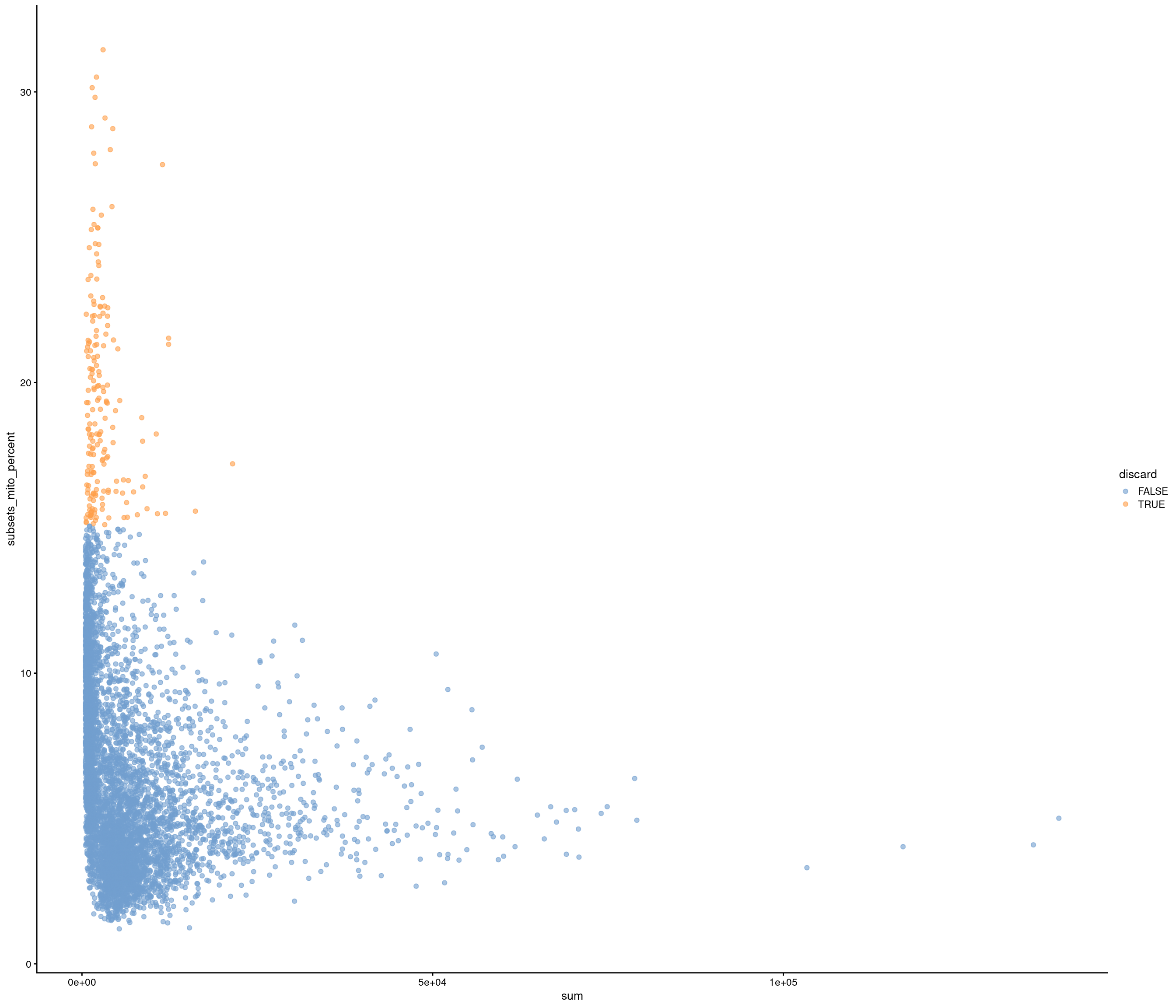
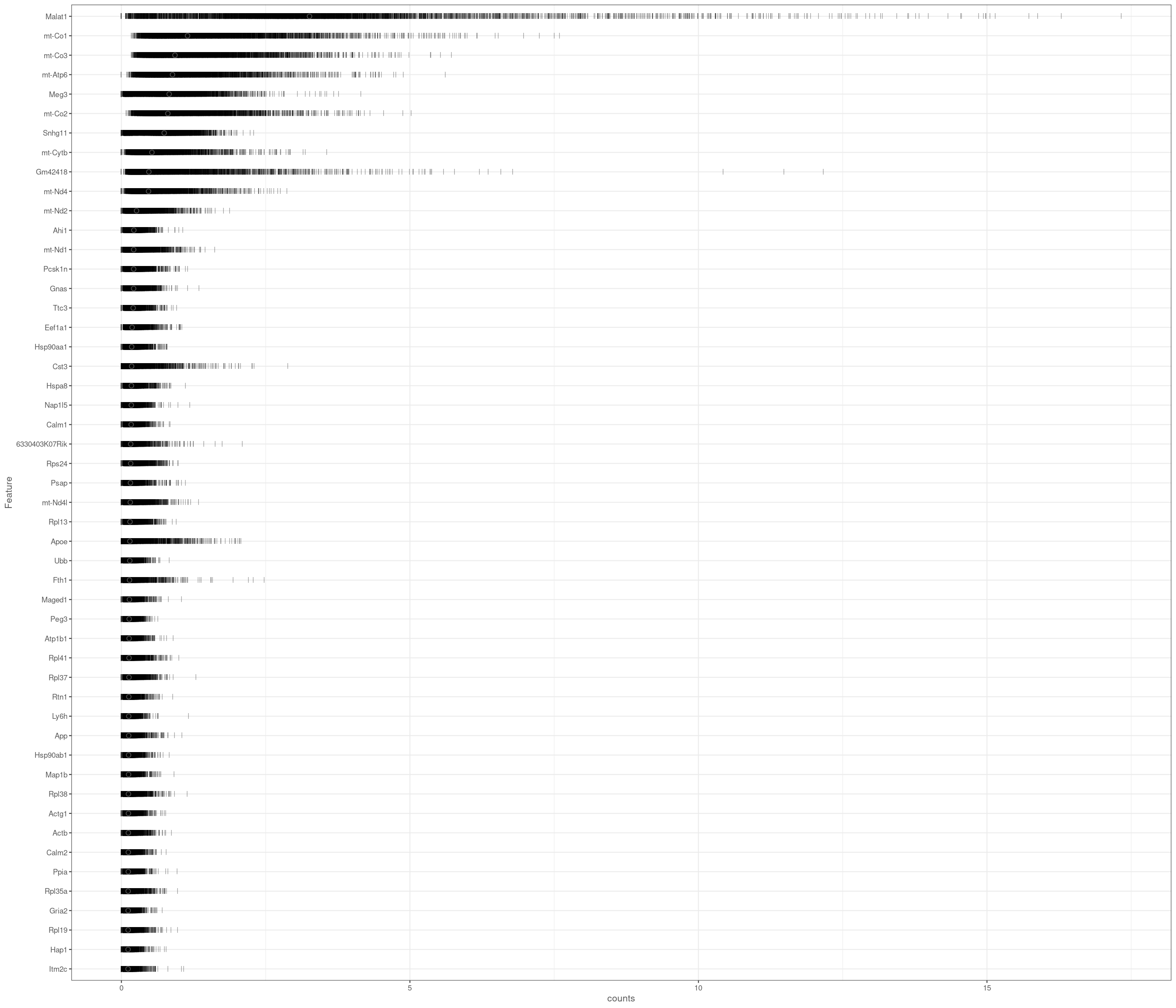
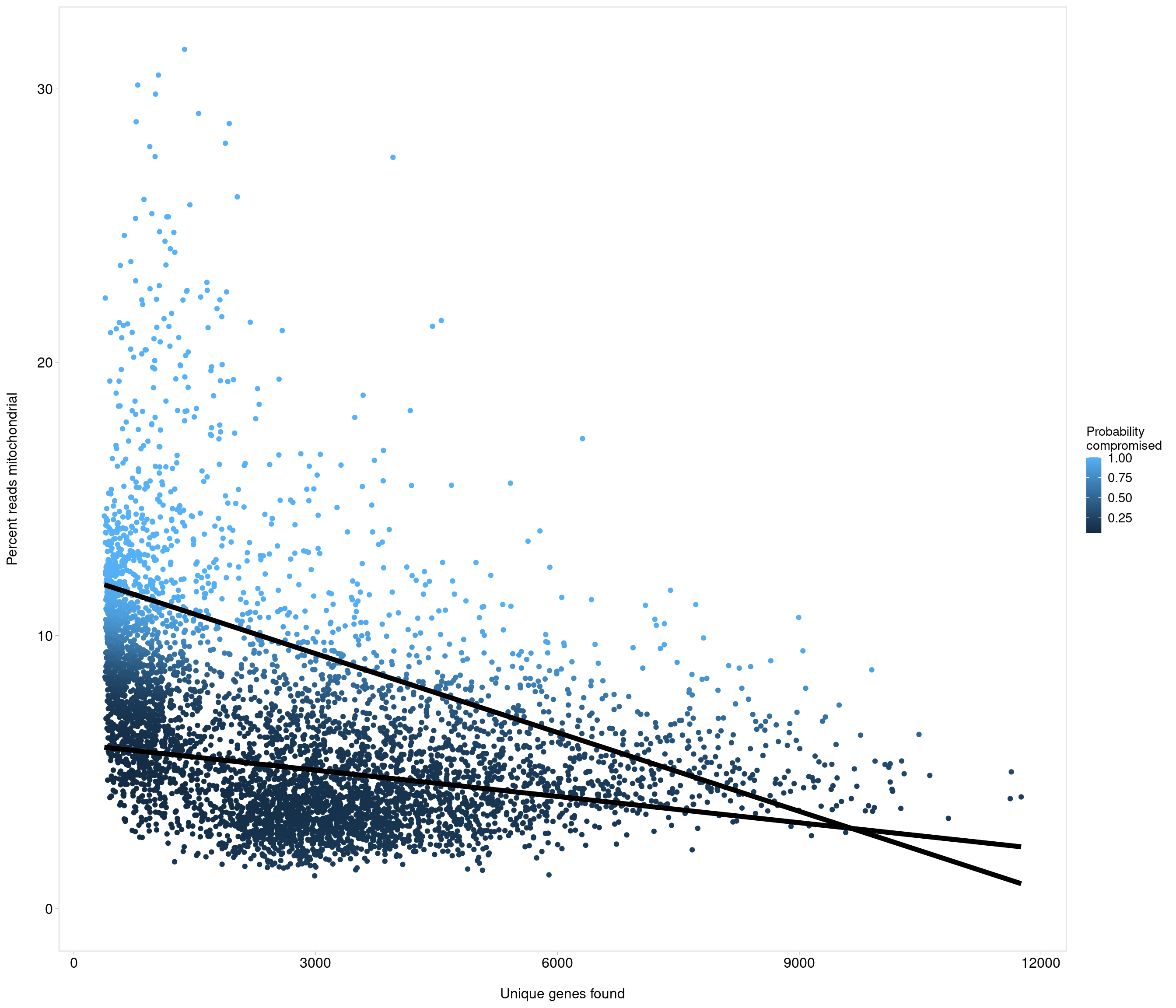
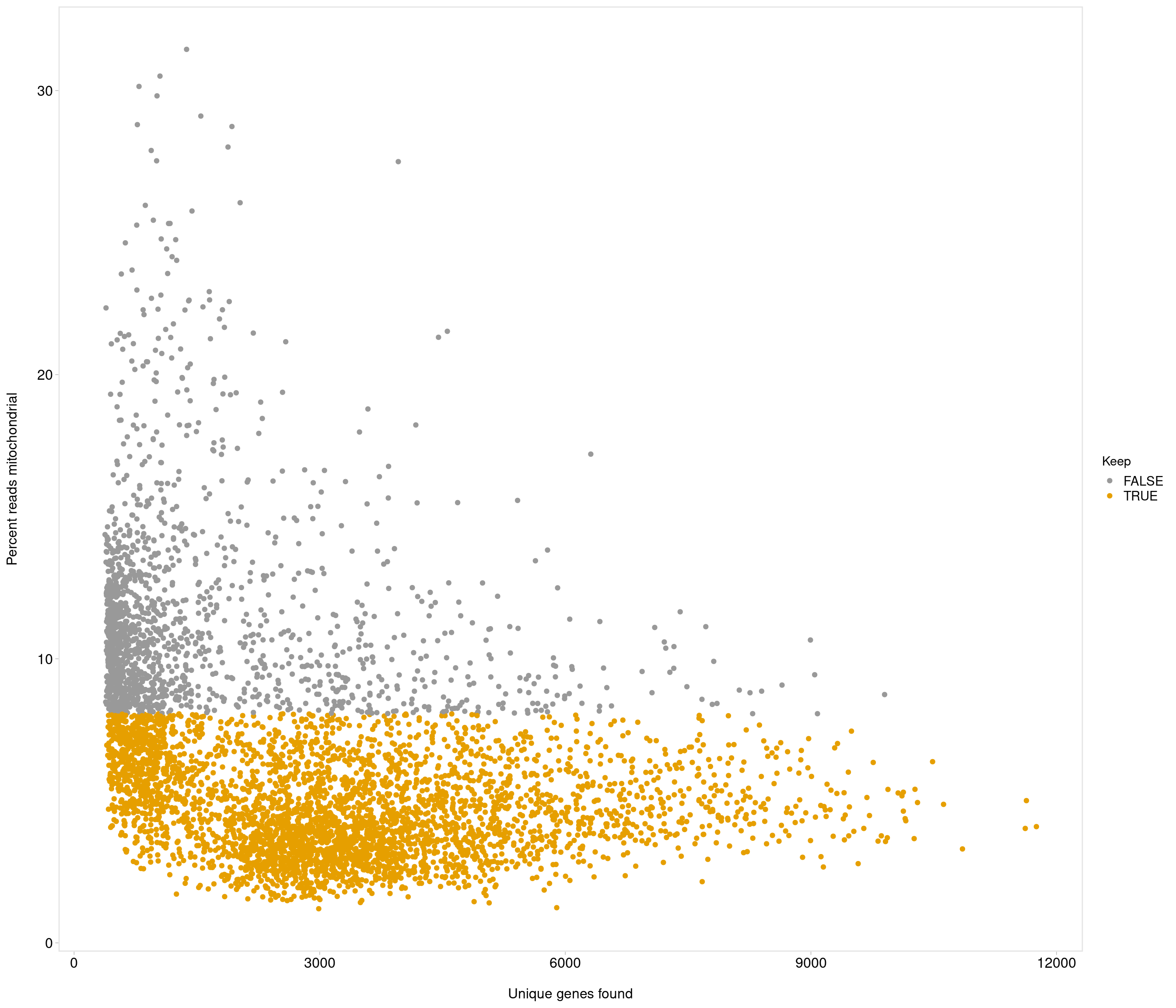
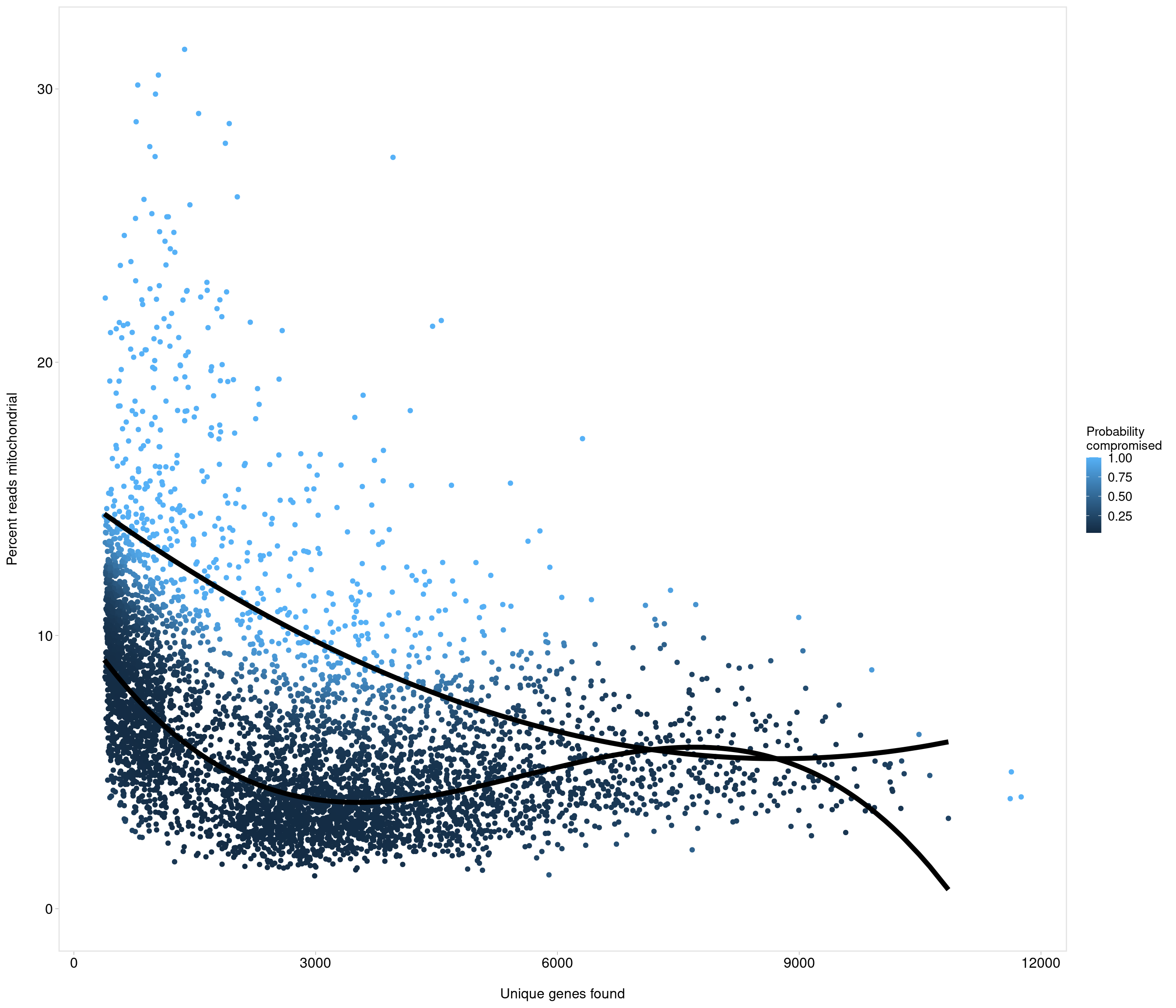
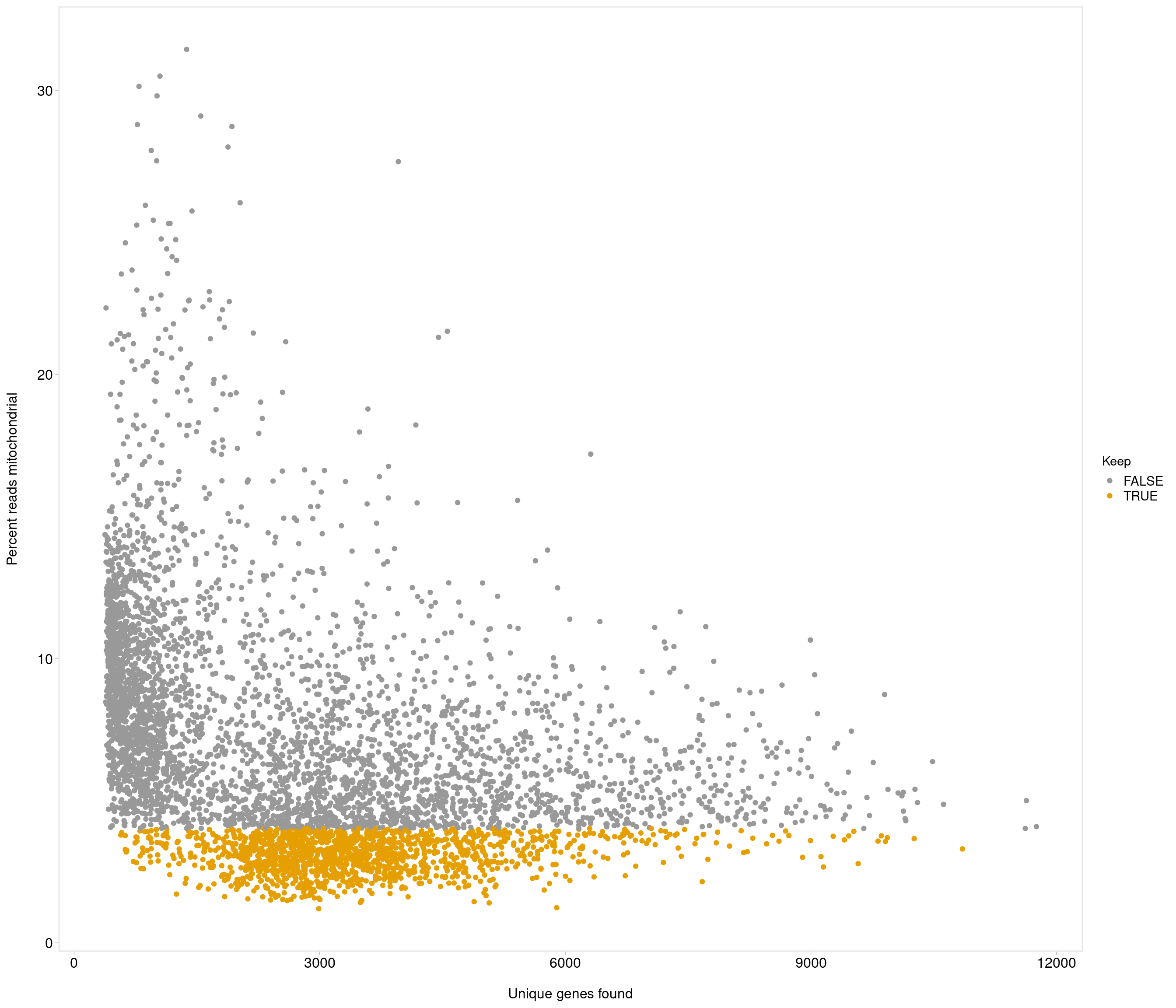
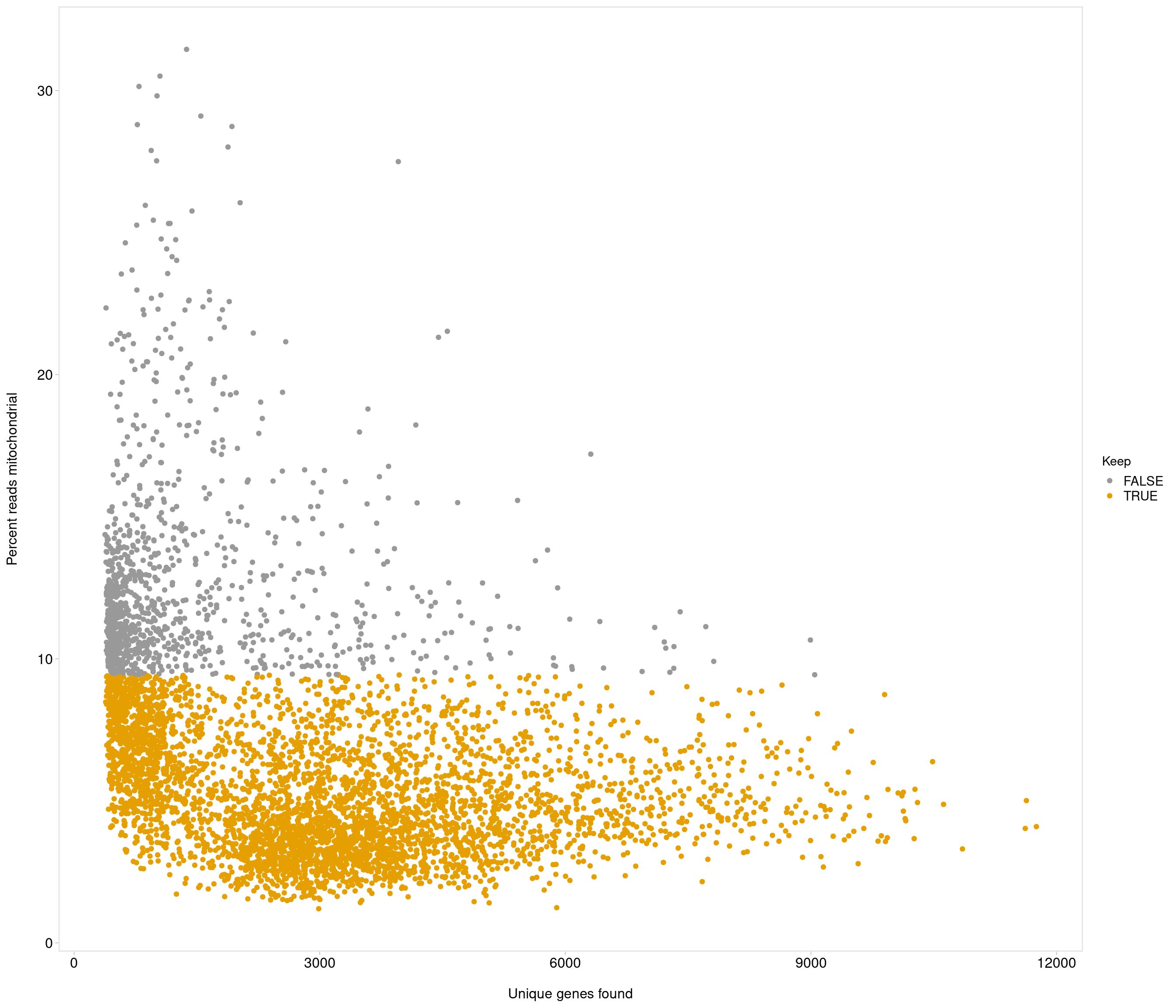
Removing 1053 out of 5436 cells. used (Mb) gc trigger (Mb) max used (Mb)
Ncells 9220665 492.5 13872300 740.9 13872300 740.9
Vcells 178820996 1364.3 298101996 2274.4 248350937 1894.82. 536-3
$mito
[1] "mt-Nd1" "mt-Nd2" "mt-Co1" "mt-Co2" "mt-Atp8" "mt-Atp6" "mt-Co3"
[8] "mt-Nd3" "mt-Nd4l" "mt-Nd4" "mt-Nd5" "mt-Nd6" "mt-Cytb"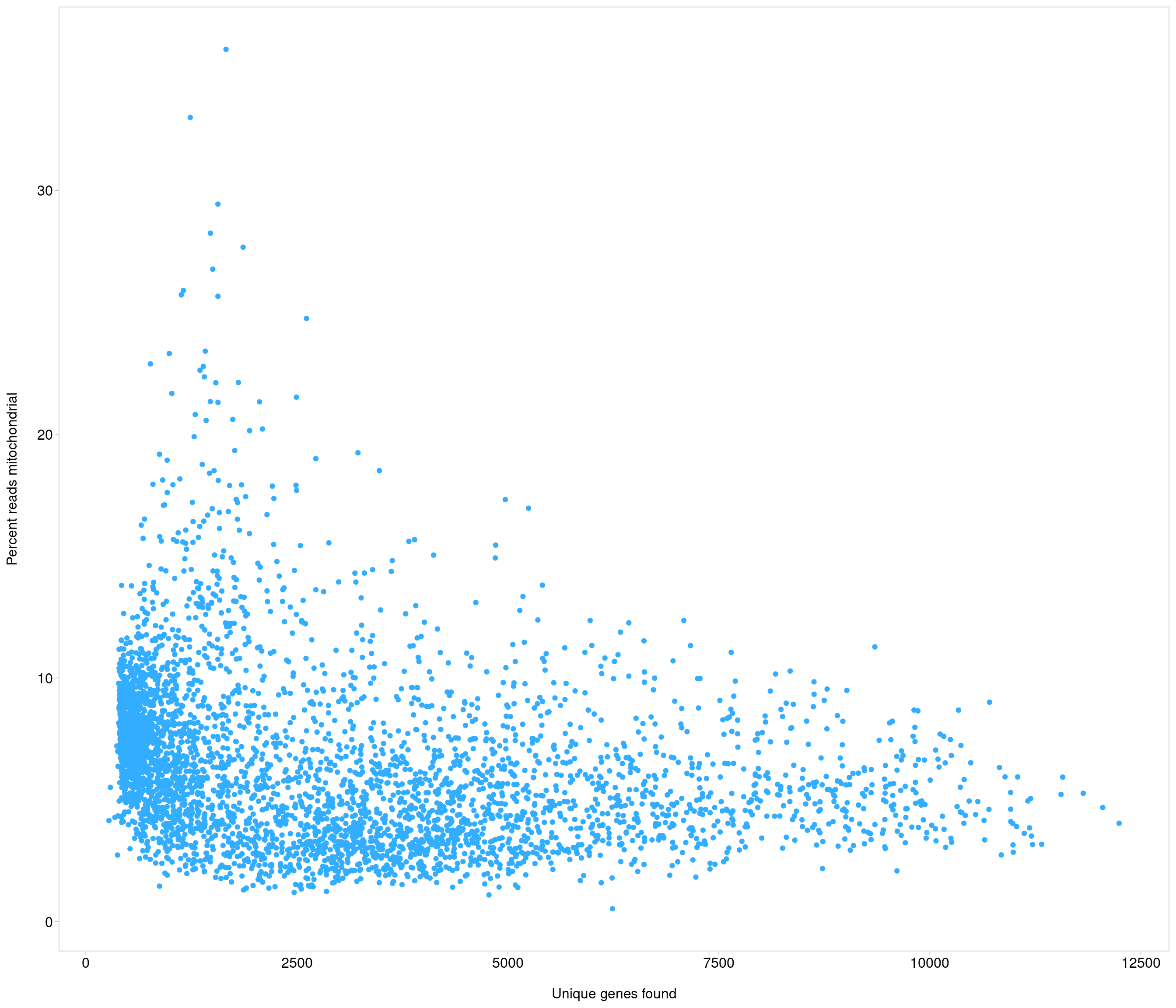
DataFrame with 6 rows and 19 columns
orig.ident nCount_RNA nFeature_RNA age
<factor> <numeric> <integer> <character>
AAACCCAAGACAACAT-1 536-3_chow-diet 13504 5296 adult
AAACCCACATATAGCC-1 536-3_chow-diet 10733 4354 adult
AAACCCATCACTGGTA-1 536-3_chow-diet 1248 907 adult
AAACCCATCTAATTCC-1 536-3_chow-diet 1996 1139 adult
AAACGAAAGTTTCGAC-1 536-3_chow-diet 2378 1497 adult
AAACGAACATTCGATG-1 536-3_chow-diet 2332 1603 adult
sex study_id tech hfd
<character> <character> <character> <logical>
AAACCCAAGACAACAT-1 male deng_2020 10xv3 FALSE
AAACCCACATATAGCC-1 male deng_2020 10xv3 FALSE
AAACCCATCACTGGTA-1 male deng_2020 10xv3 FALSE
AAACCCATCTAATTCC-1 male deng_2020 10xv3 FALSE
AAACGAAAGTTTCGAC-1 male deng_2020 10xv3 FALSE
AAACGAACATTCGATG-1 male deng_2020 10xv3 FALSE
ident sum detected subsets_mito_sum
<factor> <numeric> <integer> <numeric>
AAACCCAAGACAACAT-1 536-3_chow-diet 13504 5296 602
AAACCCACATATAGCC-1 536-3_chow-diet 10733 4354 903
AAACCCATCACTGGTA-1 536-3_chow-diet 1248 907 108
AAACCCATCTAATTCC-1 536-3_chow-diet 1996 1139 82
AAACGAAAGTTTCGAC-1 536-3_chow-diet 2378 1497 269
AAACGAACATTCGATG-1 536-3_chow-diet 2332 1603 158
subsets_mito_detected subsets_mito_percent total
<integer> <numeric> <numeric>
AAACCCAAGACAACAT-1 12 4.45794 13504
AAACCCACATATAGCC-1 13 8.41330 10733
AAACCCATCACTGGTA-1 11 8.65385 1248
AAACCCATCTAATTCC-1 11 4.10822 1996
AAACGAAAGTTTCGAC-1 12 11.31203 2378
AAACGAACATTCGATG-1 10 6.77530 2332
low_lib_size low_n_features high_subsets_mito_percent
<outlier.filter> <outlier.filter> <outlier.filter>
AAACCCAAGACAACAT-1 FALSE FALSE FALSE
AAACCCACATATAGCC-1 FALSE FALSE FALSE
AAACCCATCACTGGTA-1 FALSE FALSE FALSE
AAACCCATCTAATTCC-1 FALSE FALSE FALSE
AAACGAAAGTTTCGAC-1 FALSE FALSE FALSE
AAACGAACATTCGATG-1 FALSE FALSE FALSE
discard
<logical>
AAACCCAAGACAACAT-1 FALSE
AAACCCACATATAGCC-1 FALSE
AAACCCATCACTGGTA-1 FALSE
AAACCCATCTAATTCC-1 FALSE
AAACGAAAGTTTCGAC-1 FALSE
AAACGAACATTCGATG-1 FALSE| Name | sce$sum |
| Number of rows | 4083 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 9370.84 | 15380.62 | 500 | 1273.5 | 3964 | 10163 | 216216 | ▇▁▁▁▁ |
| Name | sce$detected |
| Number of rows | 4083 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 3007.93 | 2536.25 | 274 | 888.5 | 2227 | 4403 | 12242 | ▇▃▂▁▁ |
| Name | sce$subsets_mito_percent |
| Number of rows | 4083 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 6.4 | 3.39 | 0.53 | 3.96 | 5.8 | 8.06 | 35.79 | ▇▃▁▁▁ |
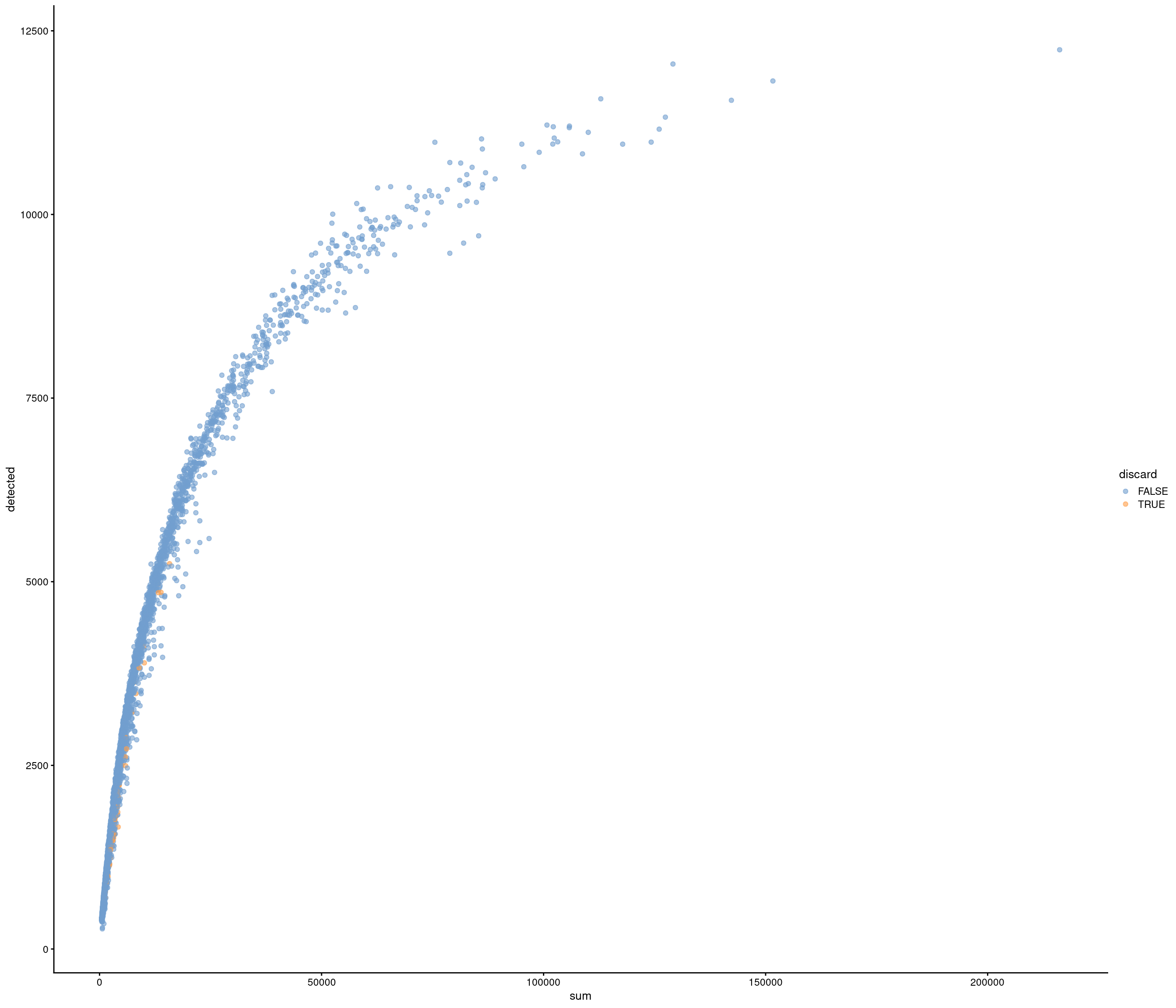
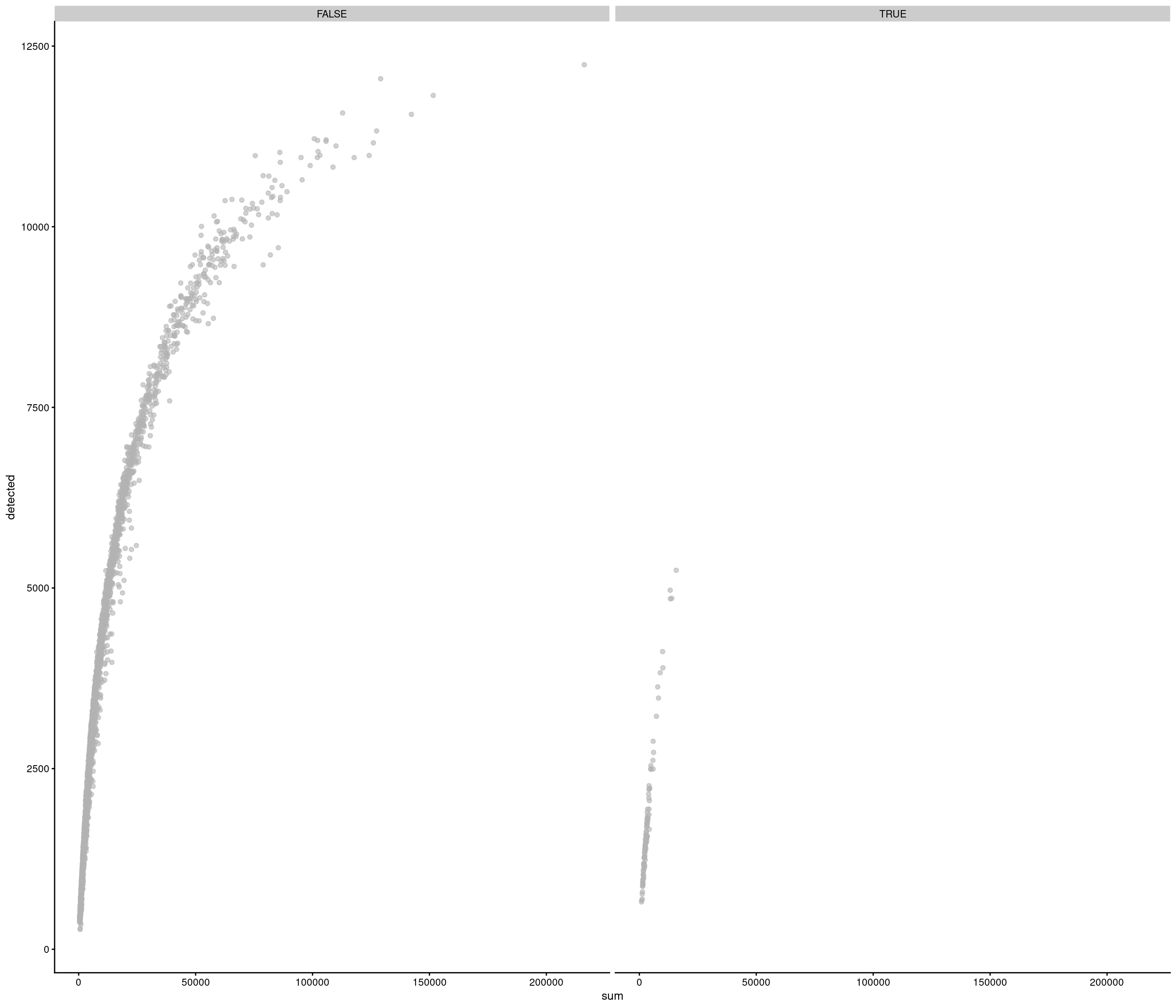
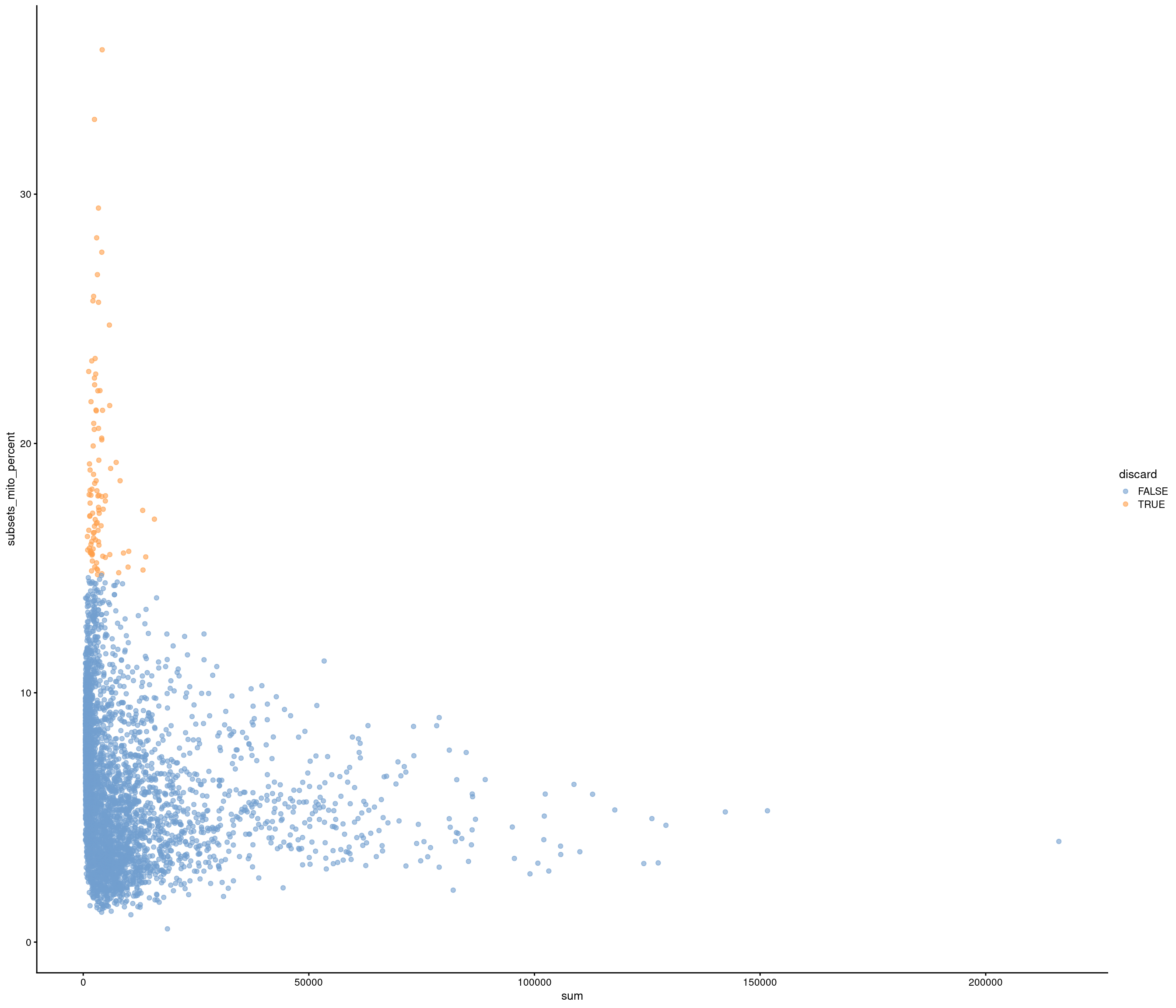
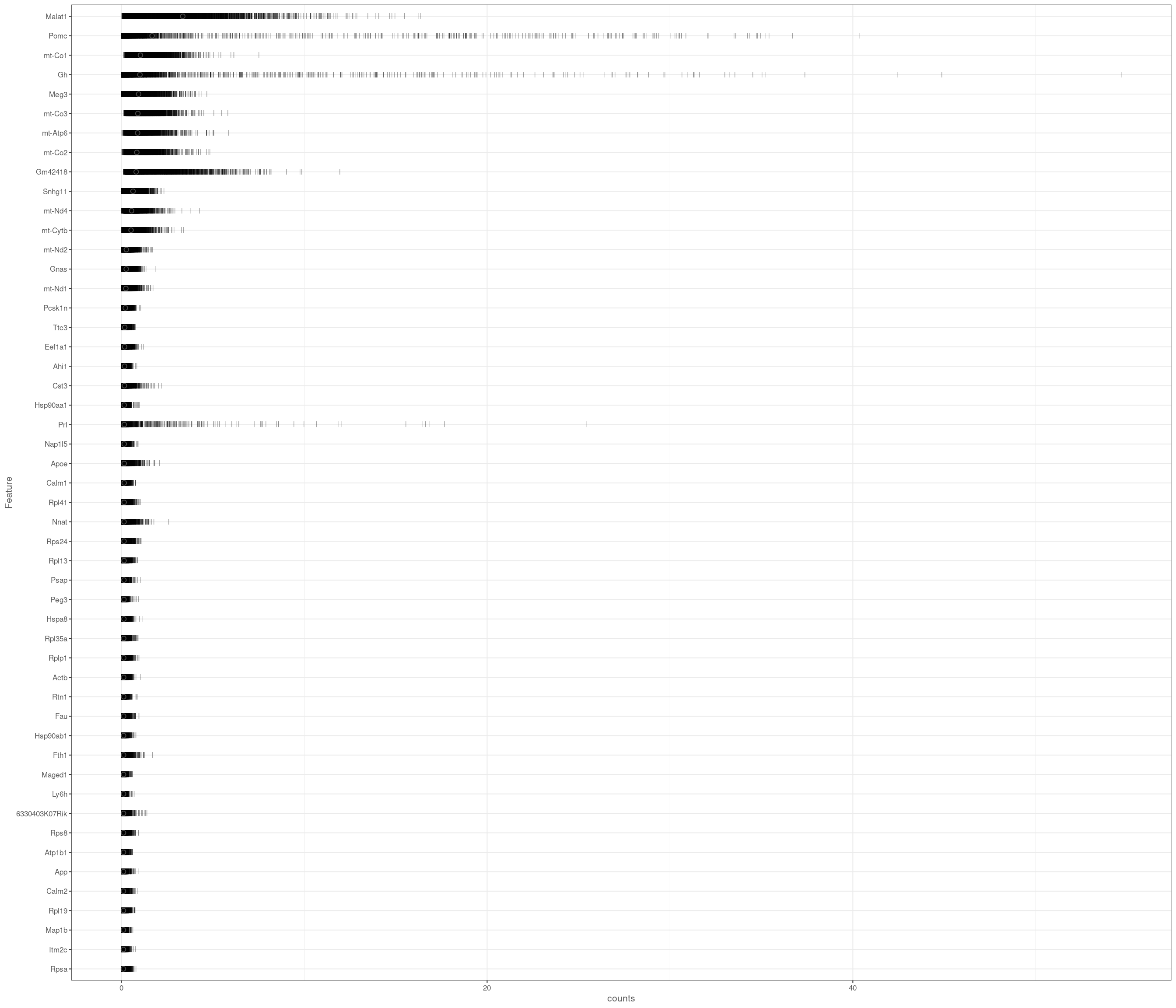
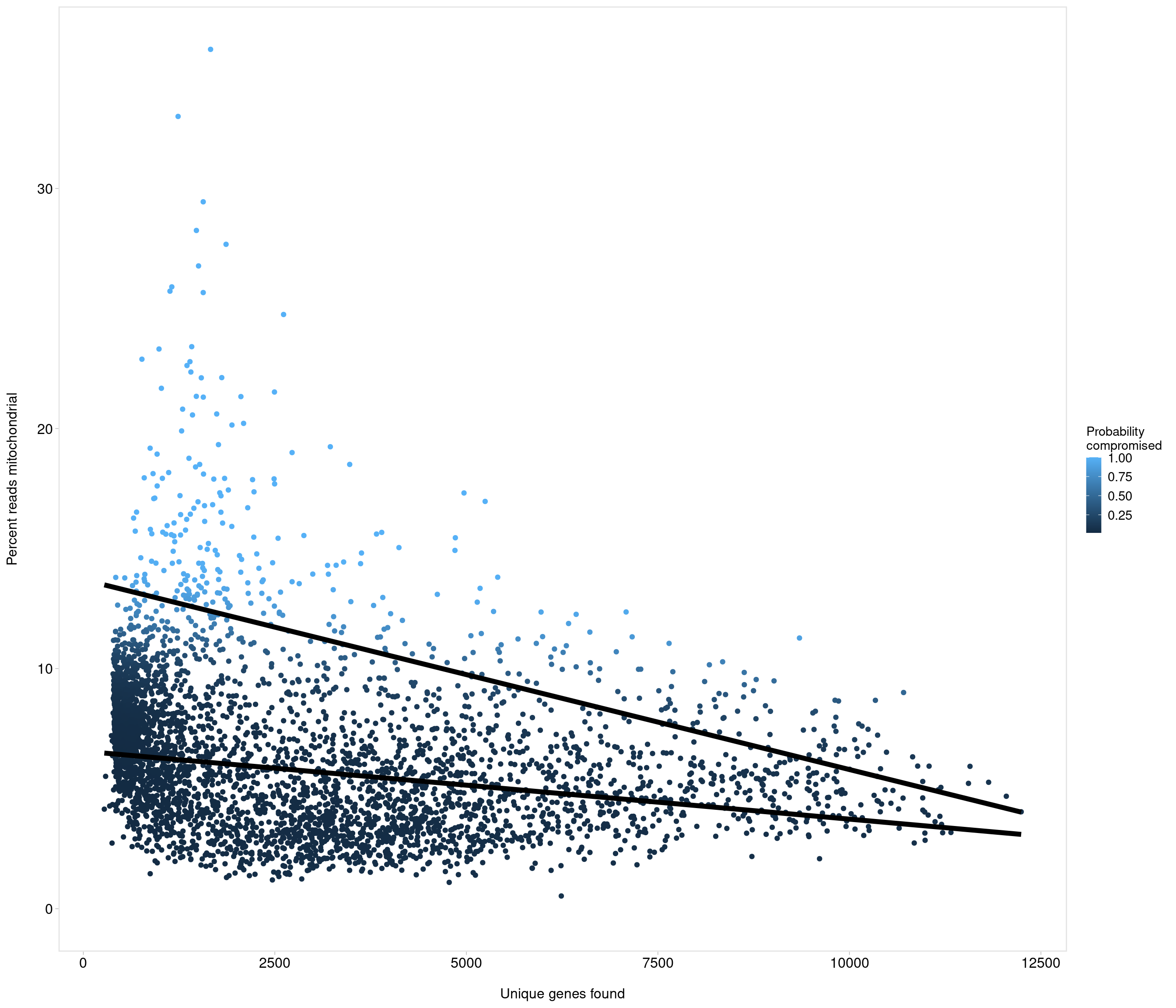
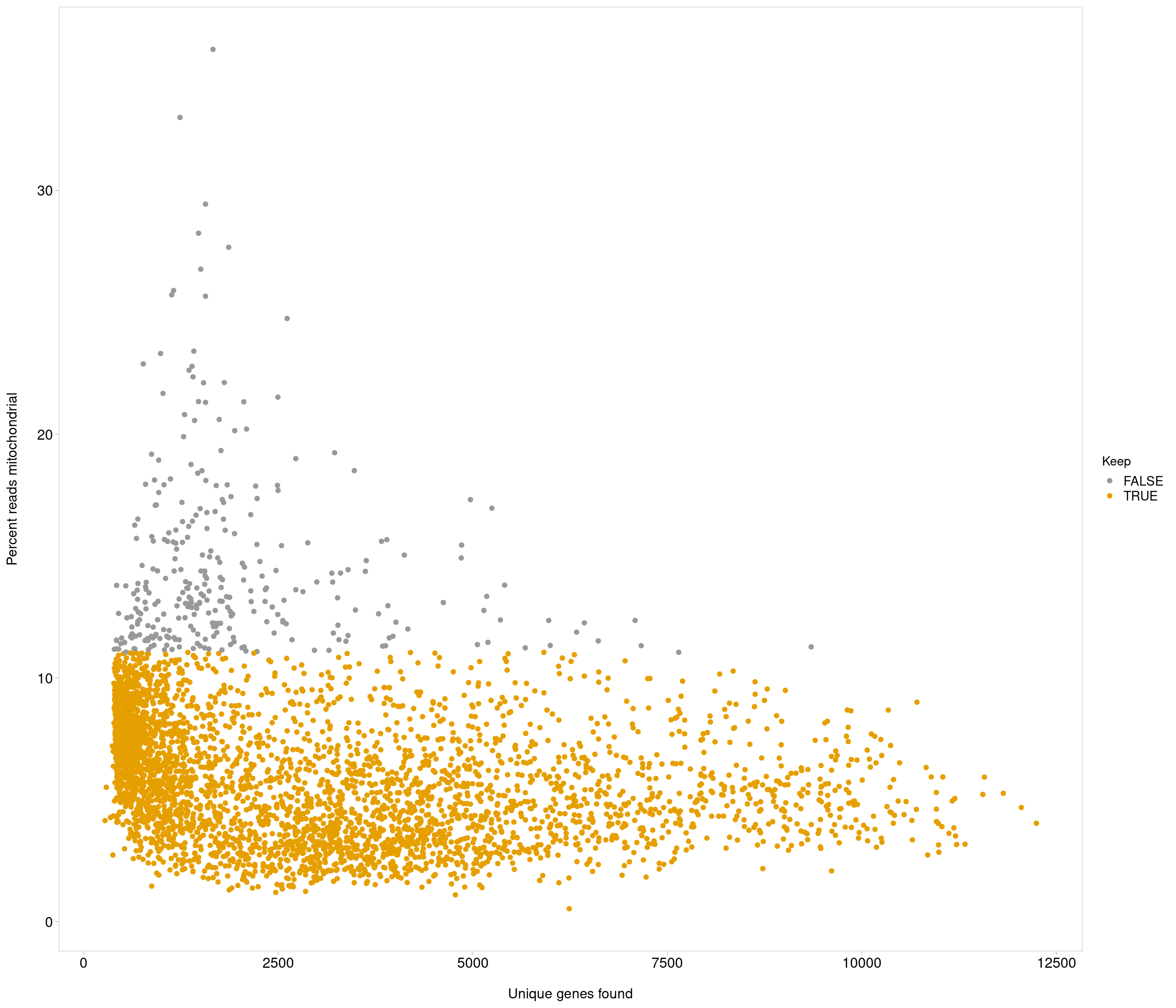
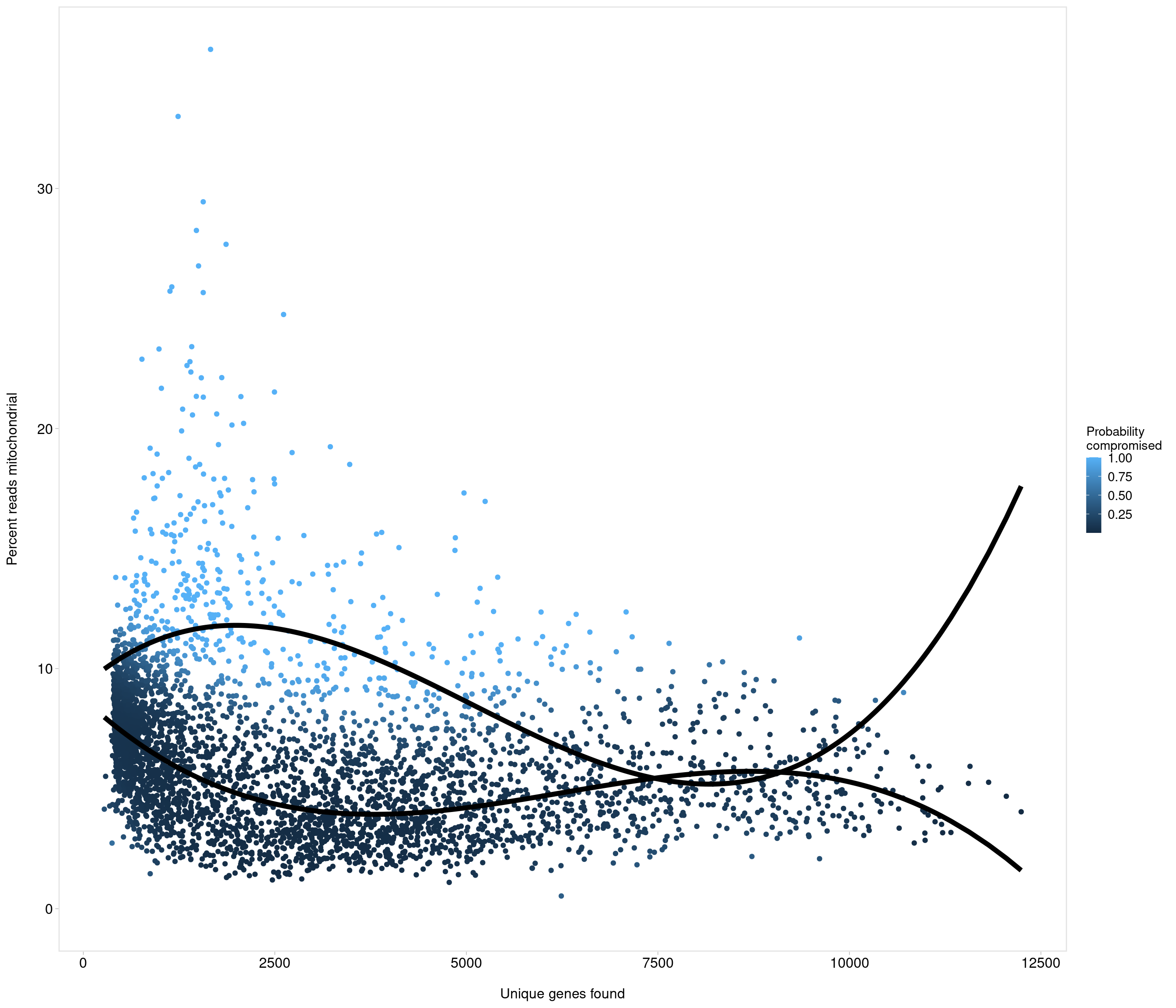
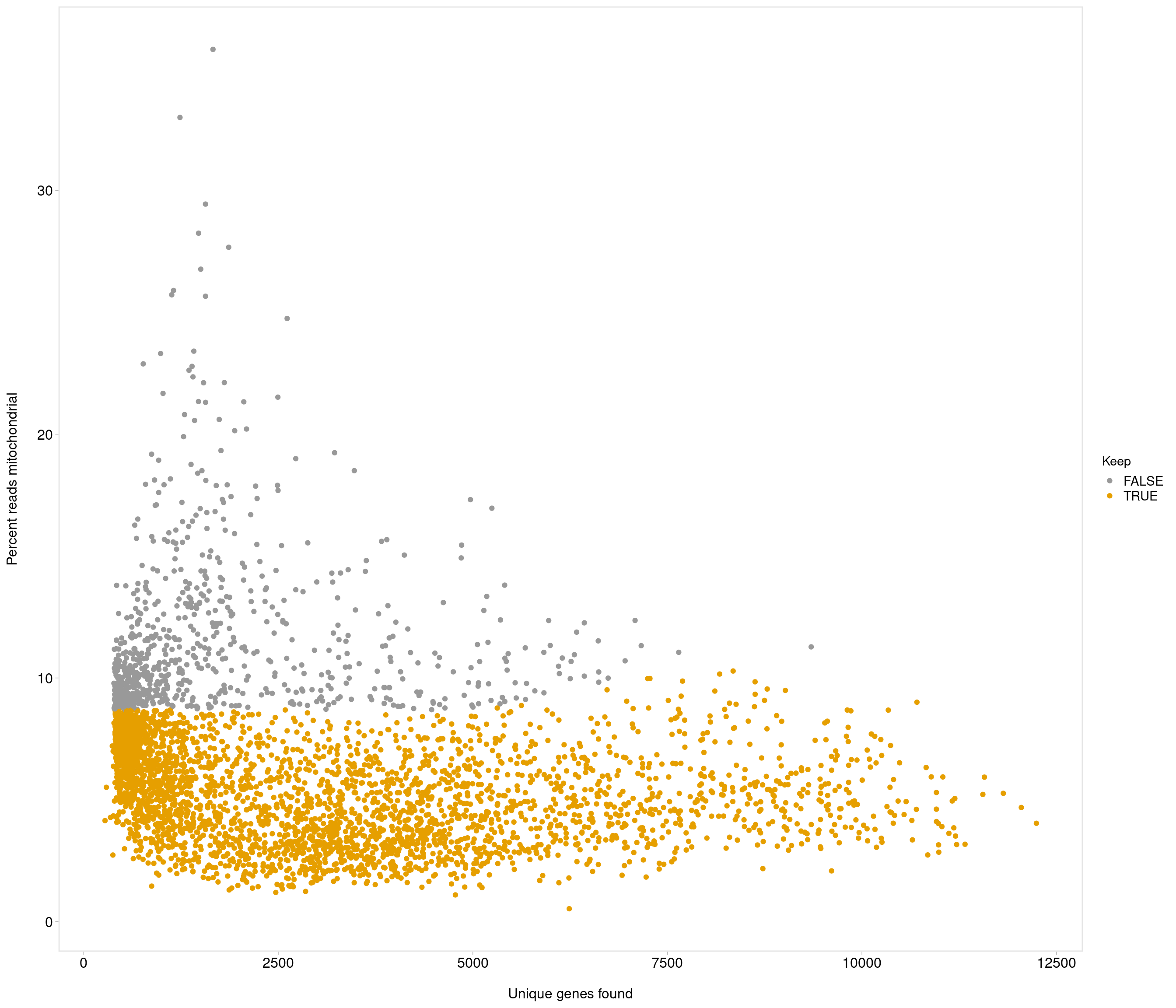
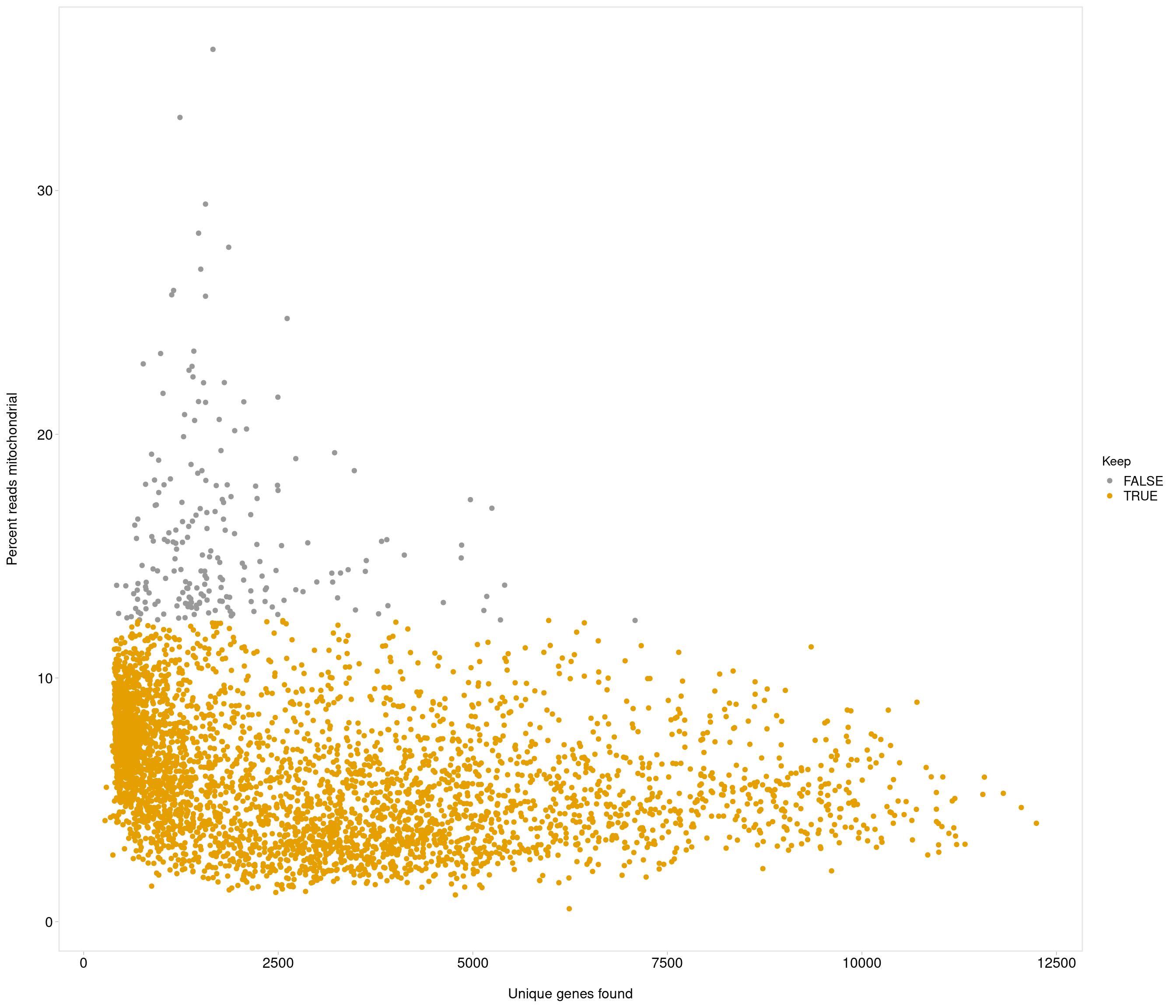
Removing 205 out of 4083 cells. used (Mb) gc trigger (Mb) max used (Mb)
Ncells 9234509 493.2 13872300 740.9 13872300 740.9
Vcells 191279147 1459.4 298101996 2274.4 273421233 2086.13. 536-5
$mito
[1] "mt-Nd1" "mt-Nd2" "mt-Co1" "mt-Co2" "mt-Atp8" "mt-Atp6" "mt-Co3"
[8] "mt-Nd3" "mt-Nd4l" "mt-Nd4" "mt-Nd5" "mt-Nd6" "mt-Cytb"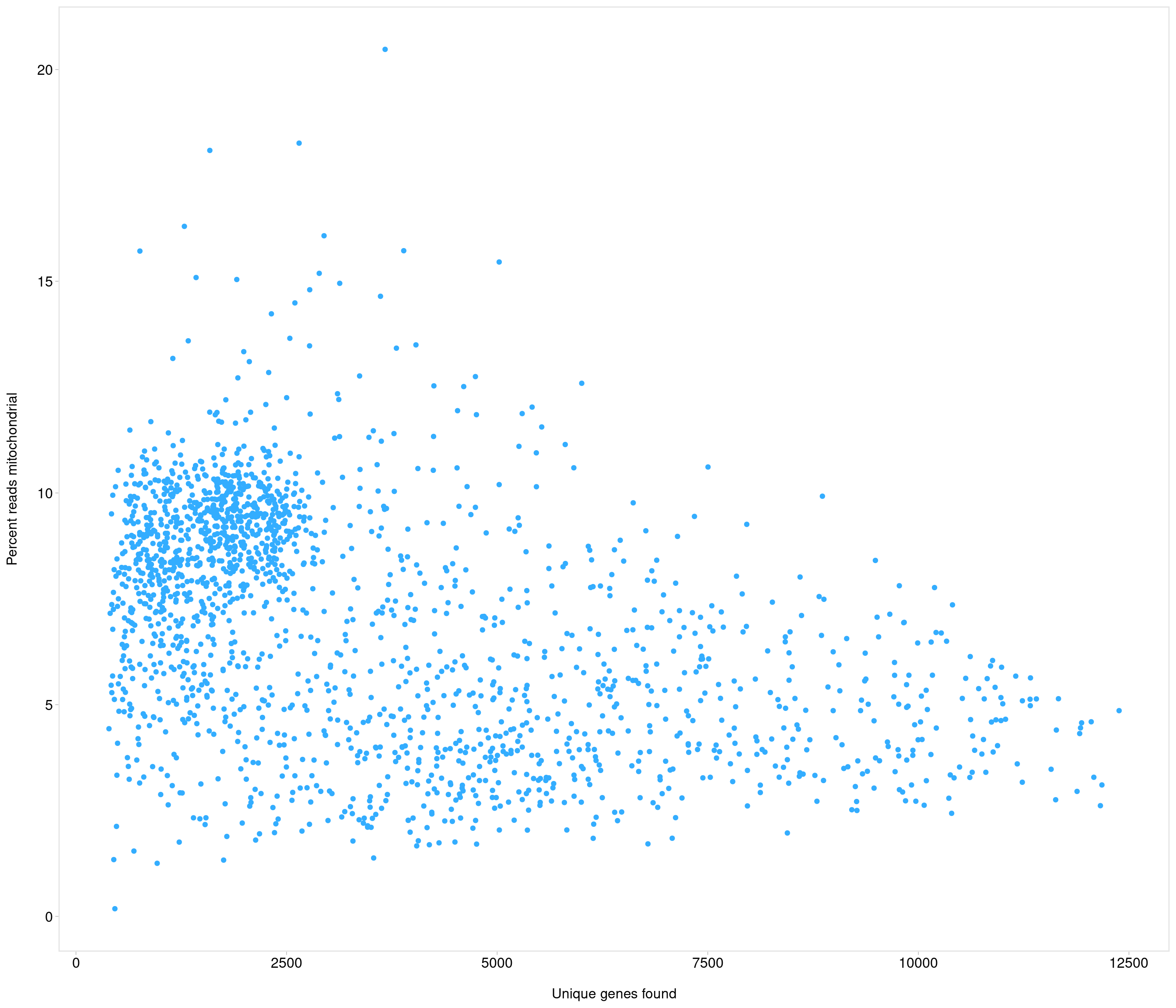
DataFrame with 6 rows and 19 columns
orig.ident nCount_RNA nFeature_RNA age
<factor> <numeric> <integer> <character>
AAACCCAAGCTACTAC-1 536-5_chow-diet 6654 3164 adult
AAACCCACACATTCTT-1 536-5_chow-diet 11880 4671 adult
AAACCCAGTGAATAAC-1 536-5_chow-diet 4700 2485 adult
AAACCCATCGCCAATA-1 536-5_chow-diet 5104 2590 adult
AAACGAAGTTCCTAGA-1 536-5_chow-diet 1807 1268 adult
AAACGCTGTTCTCAGA-1 536-5_chow-diet 49217 8927 adult
sex study_id tech hfd
<character> <character> <character> <logical>
AAACCCAAGCTACTAC-1 male deng_2020 10xv3 FALSE
AAACCCACACATTCTT-1 male deng_2020 10xv3 FALSE
AAACCCAGTGAATAAC-1 male deng_2020 10xv3 FALSE
AAACCCATCGCCAATA-1 male deng_2020 10xv3 FALSE
AAACGAAGTTCCTAGA-1 male deng_2020 10xv3 FALSE
AAACGCTGTTCTCAGA-1 male deng_2020 10xv3 FALSE
ident sum detected subsets_mito_sum
<factor> <numeric> <integer> <numeric>
AAACCCAAGCTACTAC-1 536-5_chow-diet 6654 3164 690
AAACCCACACATTCTT-1 536-5_chow-diet 11880 4671 621
AAACCCAGTGAATAAC-1 536-5_chow-diet 4700 2485 363
AAACCCATCGCCAATA-1 536-5_chow-diet 5104 2590 494
AAACGAAGTTCCTAGA-1 536-5_chow-diet 1807 1268 84
AAACGCTGTTCTCAGA-1 536-5_chow-diet 49217 8927 2666
subsets_mito_detected subsets_mito_percent total
<integer> <numeric> <numeric>
AAACCCAAGCTACTAC-1 13 10.36970 6654
AAACCCACACATTCTT-1 13 5.22727 11880
AAACCCAGTGAATAAC-1 12 7.72340 4700
AAACCCATCGCCAATA-1 12 9.67868 5104
AAACGAAGTTCCTAGA-1 12 4.64859 1807
AAACGCTGTTCTCAGA-1 13 5.41683 49217
low_lib_size low_n_features high_subsets_mito_percent
<outlier.filter> <outlier.filter> <outlier.filter>
AAACCCAAGCTACTAC-1 FALSE FALSE FALSE
AAACCCACACATTCTT-1 FALSE FALSE FALSE
AAACCCAGTGAATAAC-1 FALSE FALSE FALSE
AAACCCATCGCCAATA-1 FALSE FALSE FALSE
AAACGAAGTTCCTAGA-1 FALSE FALSE FALSE
AAACGCTGTTCTCAGA-1 FALSE FALSE FALSE
discard
<logical>
AAACCCAAGCTACTAC-1 FALSE
AAACCCACACATTCTT-1 FALSE
AAACCCAGTGAATAAC-1 FALSE
AAACCCATCGCCAATA-1 FALSE
AAACGAAGTTCCTAGA-1 FALSE
AAACGCTGTTCTCAGA-1 FALSE| Name | sce$sum |
| Number of rows | 1871 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 12991.02 | 22423.98 | 511 | 2335 | 4332 | 12648.5 | 209773 | ▇▁▁▁▁ |
| Name | sce$detected |
| Number of rows | 1871 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 3503.85 | 2741.39 | 392 | 1491 | 2357 | 4985.5 | 12383 | ▇▃▂▁▁ |
| Name | sce$subsets_mito_percent |
| Number of rows | 1871 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 6.99 | 2.76 | 0.18 | 4.66 | 7.24 | 9.19 | 20.48 | ▅▇▇▁▁ |
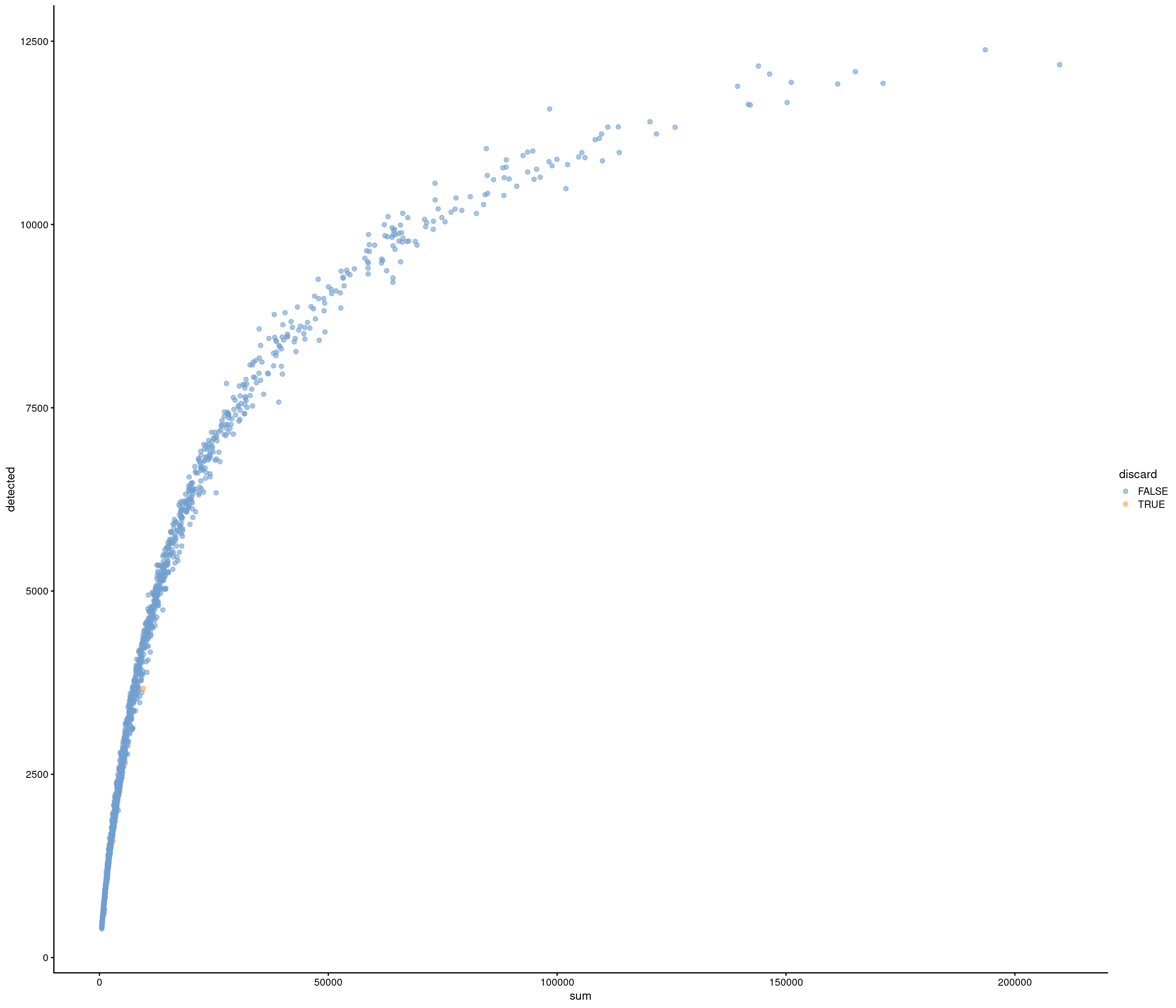
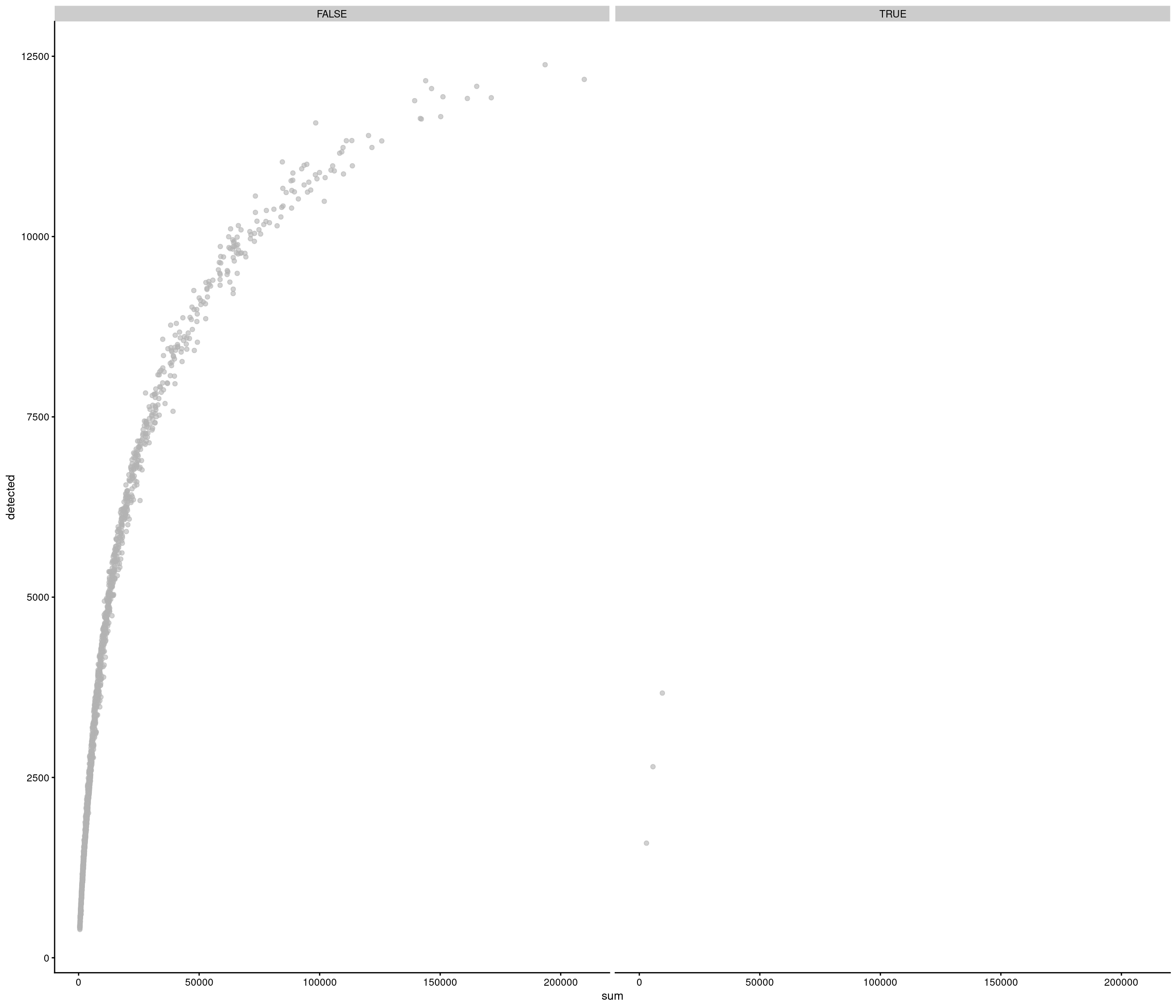
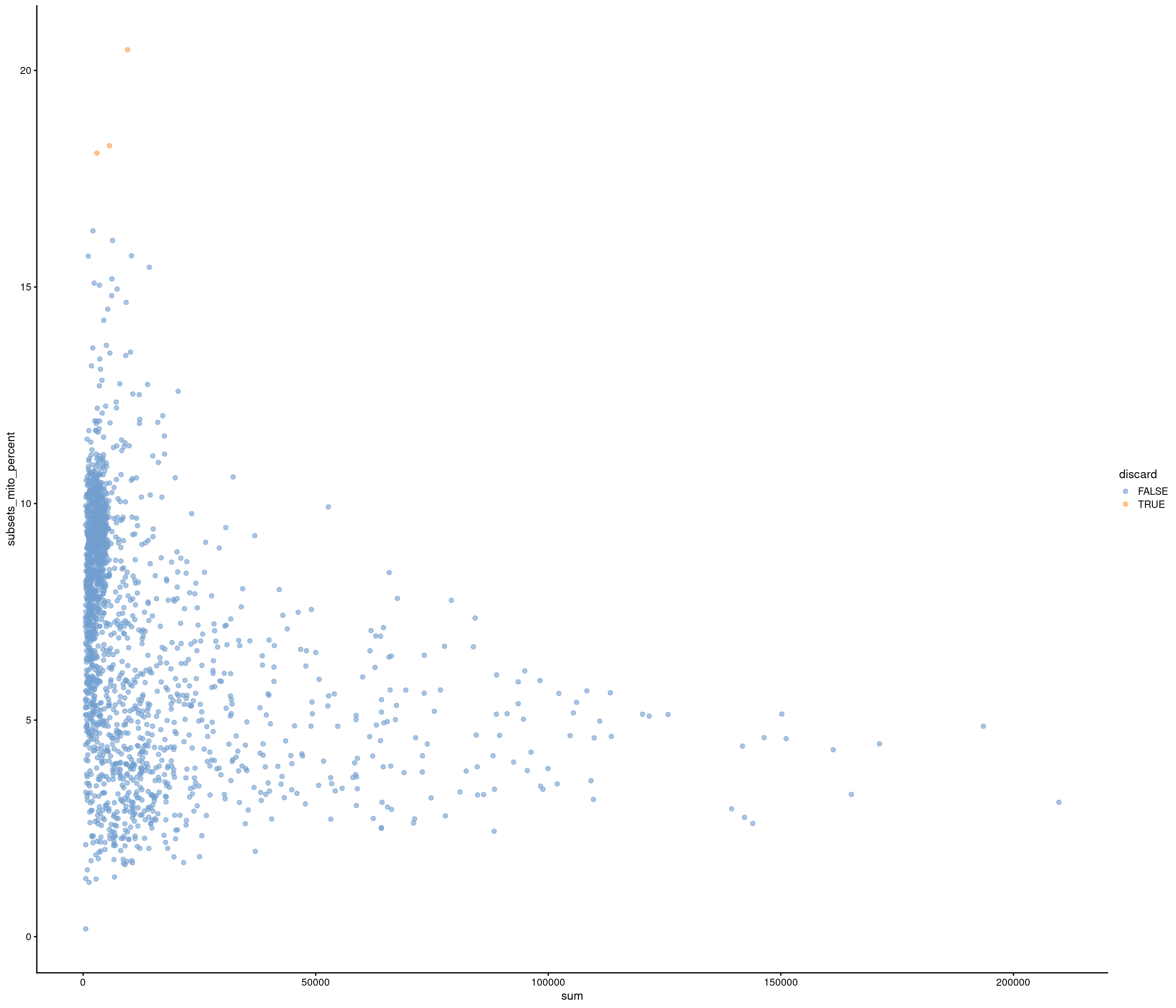
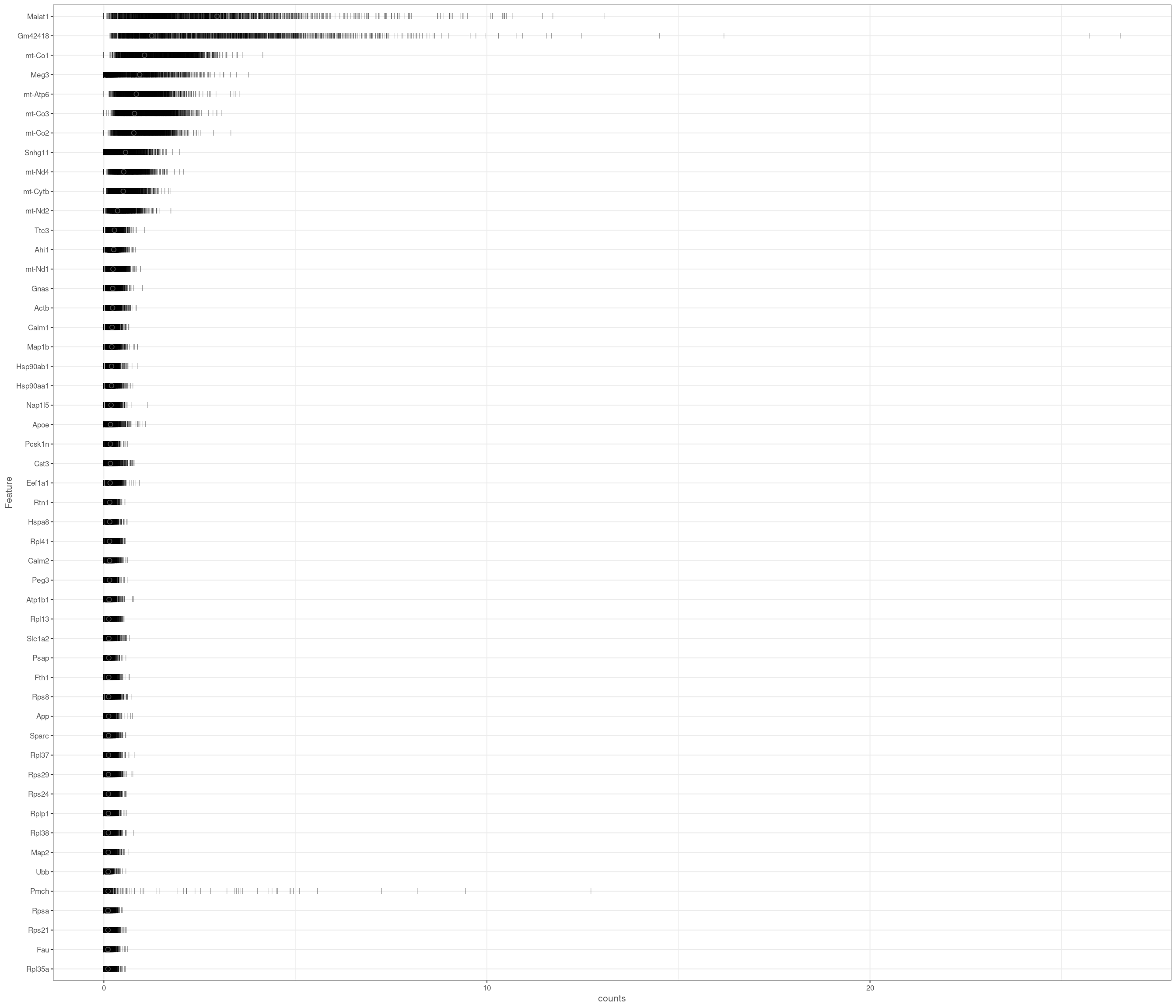
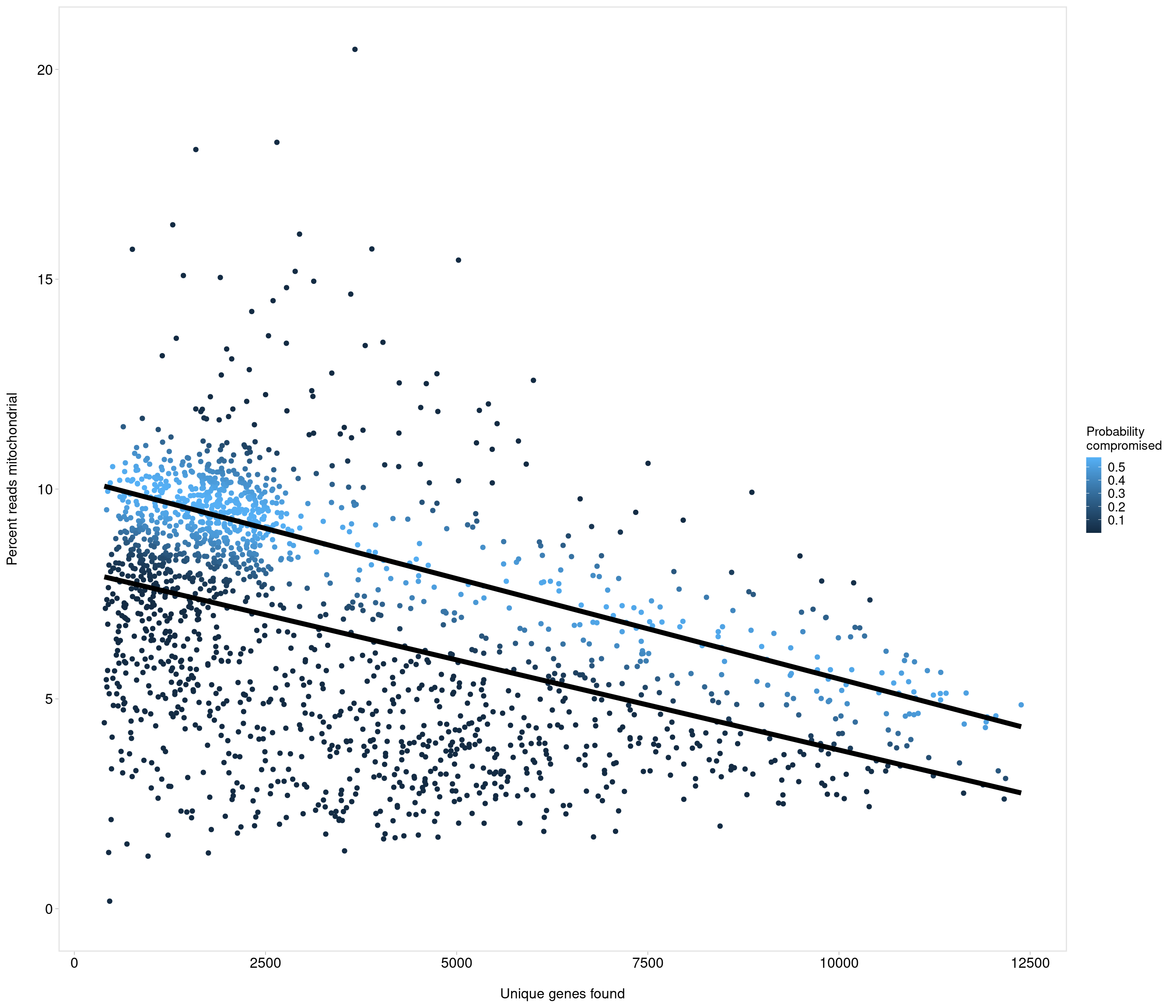
the model above gives error
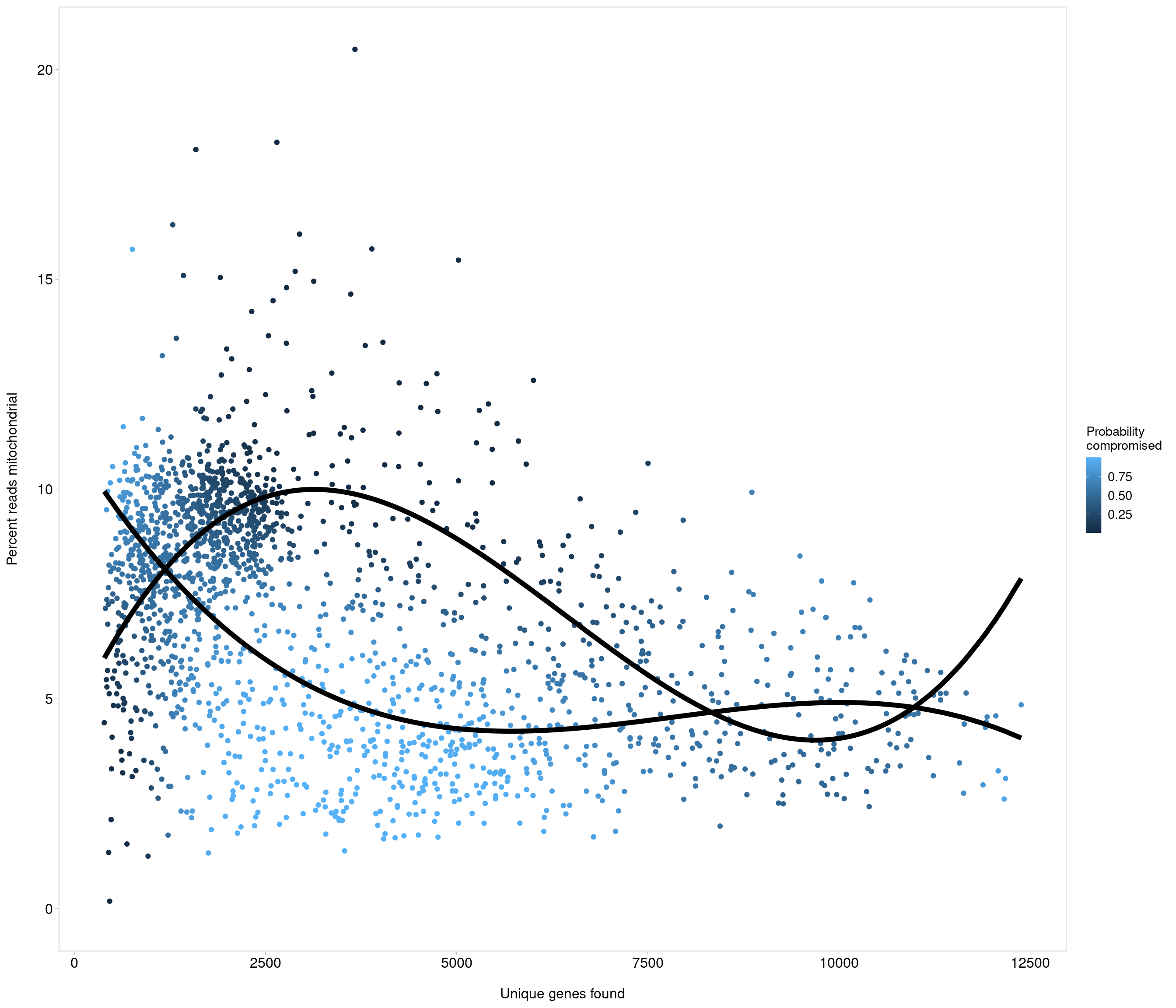
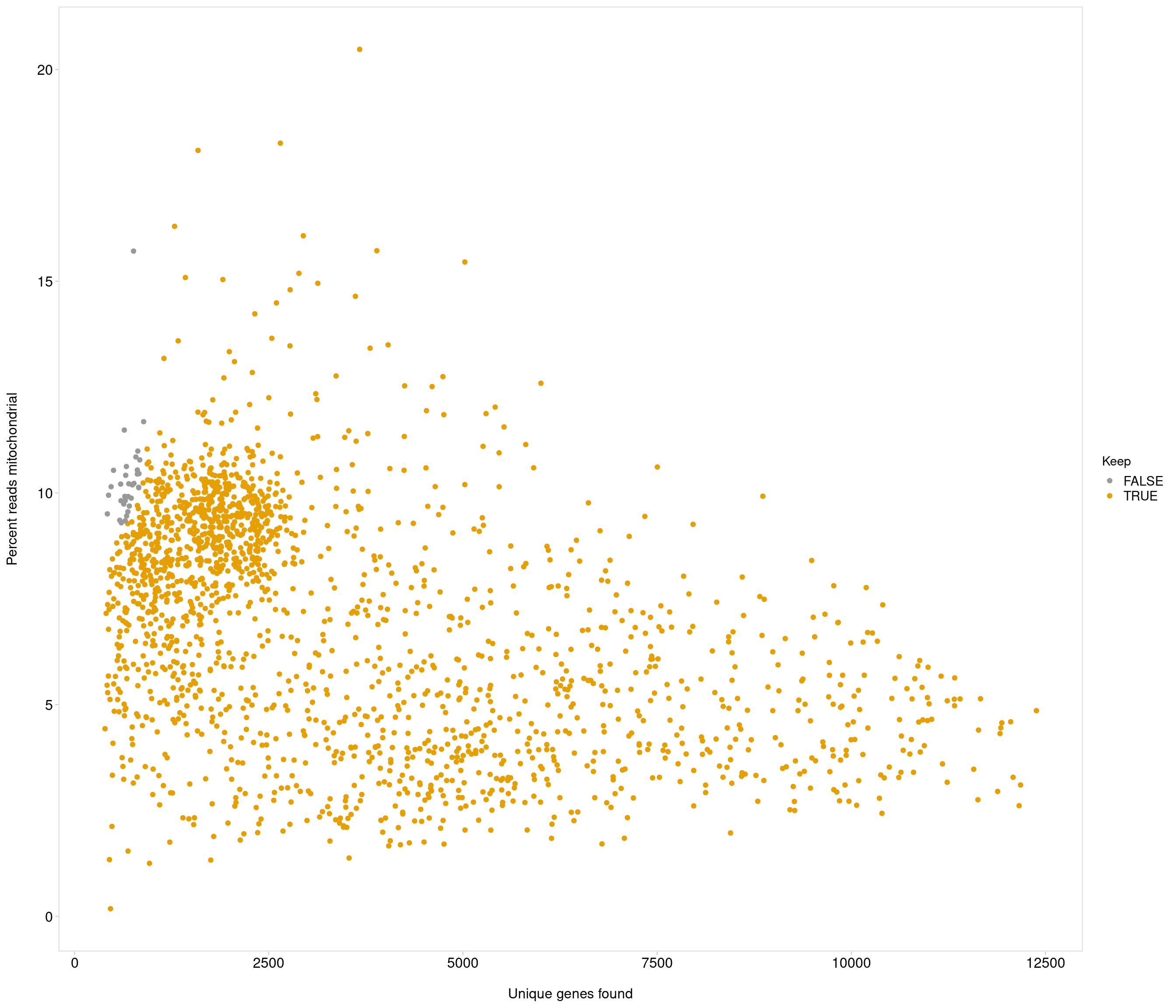
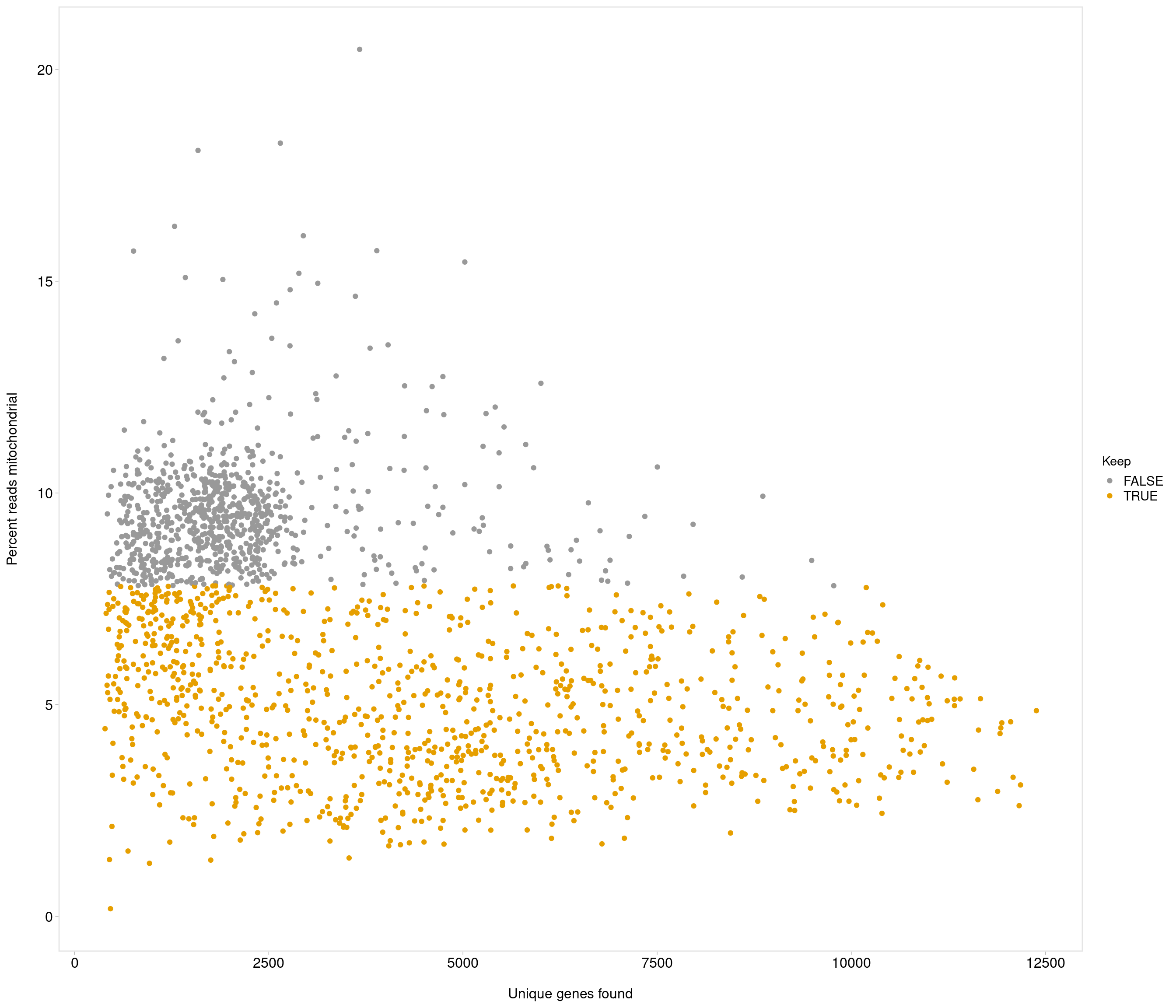
use strong criteria due to strength data
Removing 833 out of 1871 cells. used (Mb) gc trigger (Mb) max used (Mb)
Ncells 9235040 493.3 13872300 740.9 13872300 740.9
Vcells 187236121 1428.5 298101996 2274.4 273421233 2086.1HFD samples
1. 536-2_537-4
$mito
[1] "mt-Nd1" "mt-Nd2" "mt-Co1" "mt-Co2" "mt-Atp8" "mt-Atp6" "mt-Co3"
[8] "mt-Nd3" "mt-Nd4l" "mt-Nd4" "mt-Nd5" "mt-Nd6" "mt-Cytb"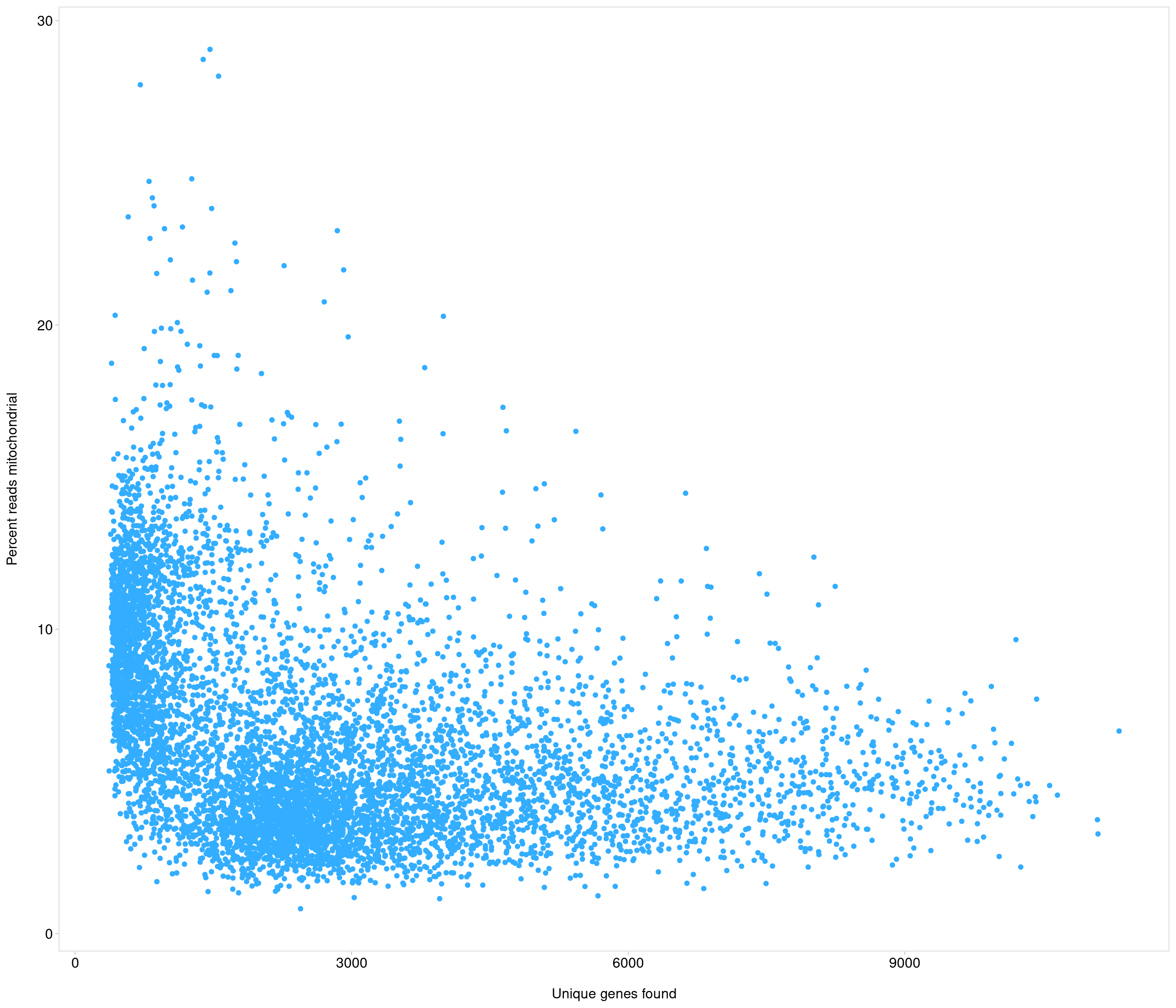
DataFrame with 6 rows and 19 columns
orig.ident nCount_RNA nFeature_RNA
<factor> <numeric> <integer>
AAACCCAAGAGTGACC-1 536-2_537-4_high-fat-diet 2405 1565
AAACCCAAGTCACTGT-1 536-2_537-4_high-fat-diet 3172 2100
AAACCCACAACGGGTA-1 536-2_537-4_high-fat-diet 21420 6324
AAACCCAGTCGTACTA-1 536-2_537-4_high-fat-diet 4907 2767
AAACCCAGTTCTGACA-1 536-2_537-4_high-fat-diet 5381 3058
AAACCCATCACCTACC-1 536-2_537-4_high-fat-diet 4366 2644
age sex study_id tech hfd
<character> <character> <character> <character> <logical>
AAACCCAAGAGTGACC-1 adult male deng_2020 10xv3 TRUE
AAACCCAAGTCACTGT-1 adult male deng_2020 10xv3 TRUE
AAACCCACAACGGGTA-1 adult male deng_2020 10xv3 TRUE
AAACCCAGTCGTACTA-1 adult male deng_2020 10xv3 TRUE
AAACCCAGTTCTGACA-1 adult male deng_2020 10xv3 TRUE
AAACCCATCACCTACC-1 adult male deng_2020 10xv3 TRUE
ident sum detected
<factor> <numeric> <integer>
AAACCCAAGAGTGACC-1 536-2_537-4_high-fat-diet 2405 1565
AAACCCAAGTCACTGT-1 536-2_537-4_high-fat-diet 3172 2100
AAACCCACAACGGGTA-1 536-2_537-4_high-fat-diet 21420 6324
AAACCCAGTCGTACTA-1 536-2_537-4_high-fat-diet 4907 2767
AAACCCAGTTCTGACA-1 536-2_537-4_high-fat-diet 5381 3058
AAACCCATCACCTACC-1 536-2_537-4_high-fat-diet 4366 2644
subsets_mito_sum subsets_mito_detected subsets_mito_percent
<numeric> <integer> <numeric>
AAACCCAAGAGTGACC-1 137 11 5.69647
AAACCCAAGTCACTGT-1 140 12 4.41362
AAACCCACAACGGGTA-1 1352 13 6.31186
AAACCCAGTCGTACTA-1 300 12 6.11372
AAACCCAGTTCTGACA-1 213 12 3.95837
AAACCCATCACCTACC-1 259 13 5.93220
total low_lib_size low_n_features
<numeric> <outlier.filter> <outlier.filter>
AAACCCAAGAGTGACC-1 2405 FALSE FALSE
AAACCCAAGTCACTGT-1 3172 FALSE FALSE
AAACCCACAACGGGTA-1 21420 FALSE FALSE
AAACCCAGTCGTACTA-1 4907 FALSE FALSE
AAACCCAGTTCTGACA-1 5381 FALSE FALSE
AAACCCATCACCTACC-1 4366 FALSE FALSE
high_subsets_mito_percent discard
<outlier.filter> <logical>
AAACCCAAGAGTGACC-1 FALSE FALSE
AAACCCAAGTCACTGT-1 FALSE FALSE
AAACCCACAACGGGTA-1 FALSE FALSE
AAACCCAGTCGTACTA-1 FALSE FALSE
AAACCCAGTTCTGACA-1 FALSE FALSE
AAACCCATCACCTACC-1 FALSE FALSE| Name | sce$sum |
| Number of rows | 7087 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 6958.85 | 9730.65 | 500 | 1554 | 3756 | 7573 | 96629 | ▇▁▁▁▁ |
| Name | sce$detected |
| Number of rows | 7087 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 2795.17 | 2122.27 | 368 | 1076 | 2309 | 3737 | 11326 | ▇▃▂▁▁ |
| Name | sce$subsets_mito_percent |
| Number of rows | 7087 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 6.42 | 3.43 | 0.82 | 3.82 | 5.47 | 8.3 | 29.05 | ▇▅▁▁▁ |
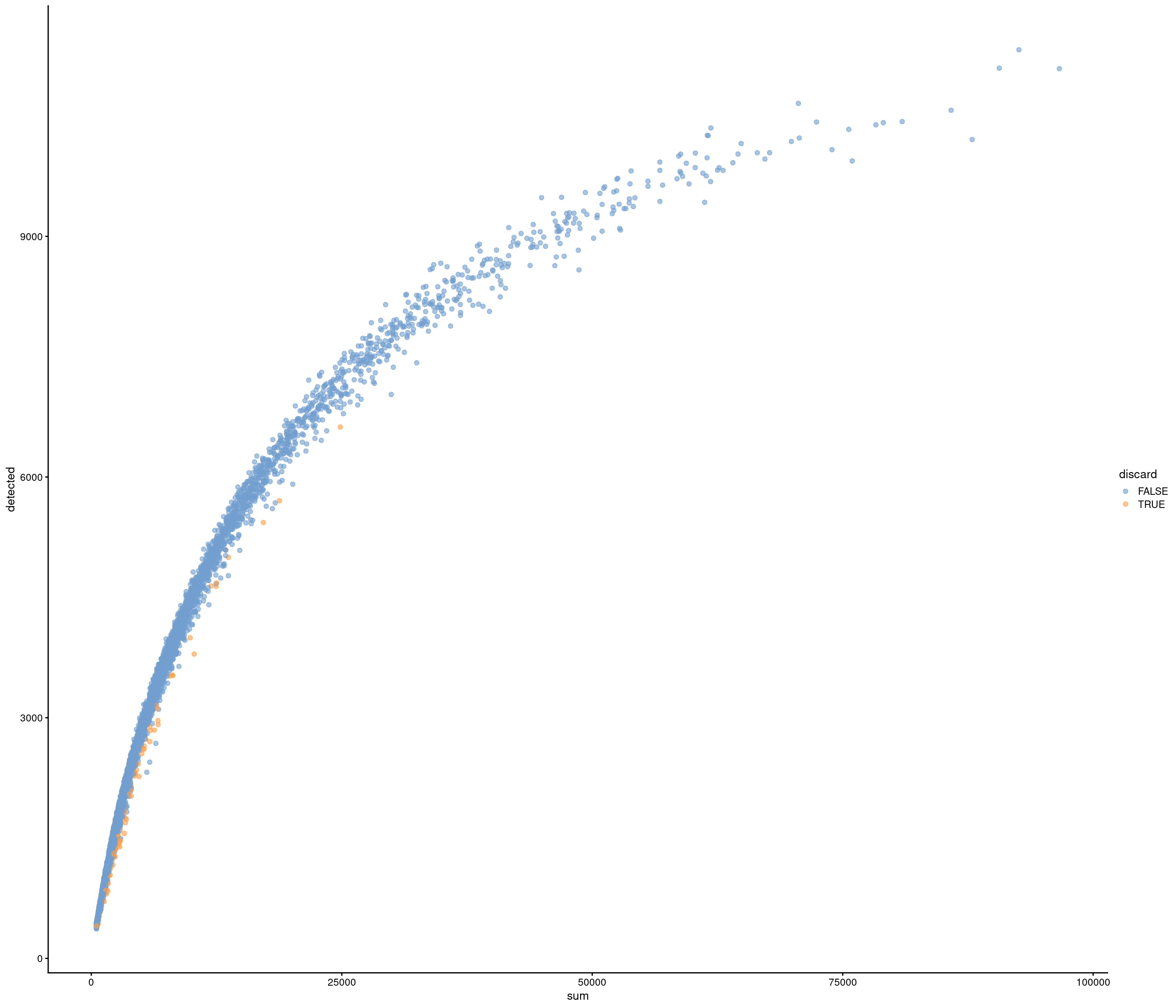
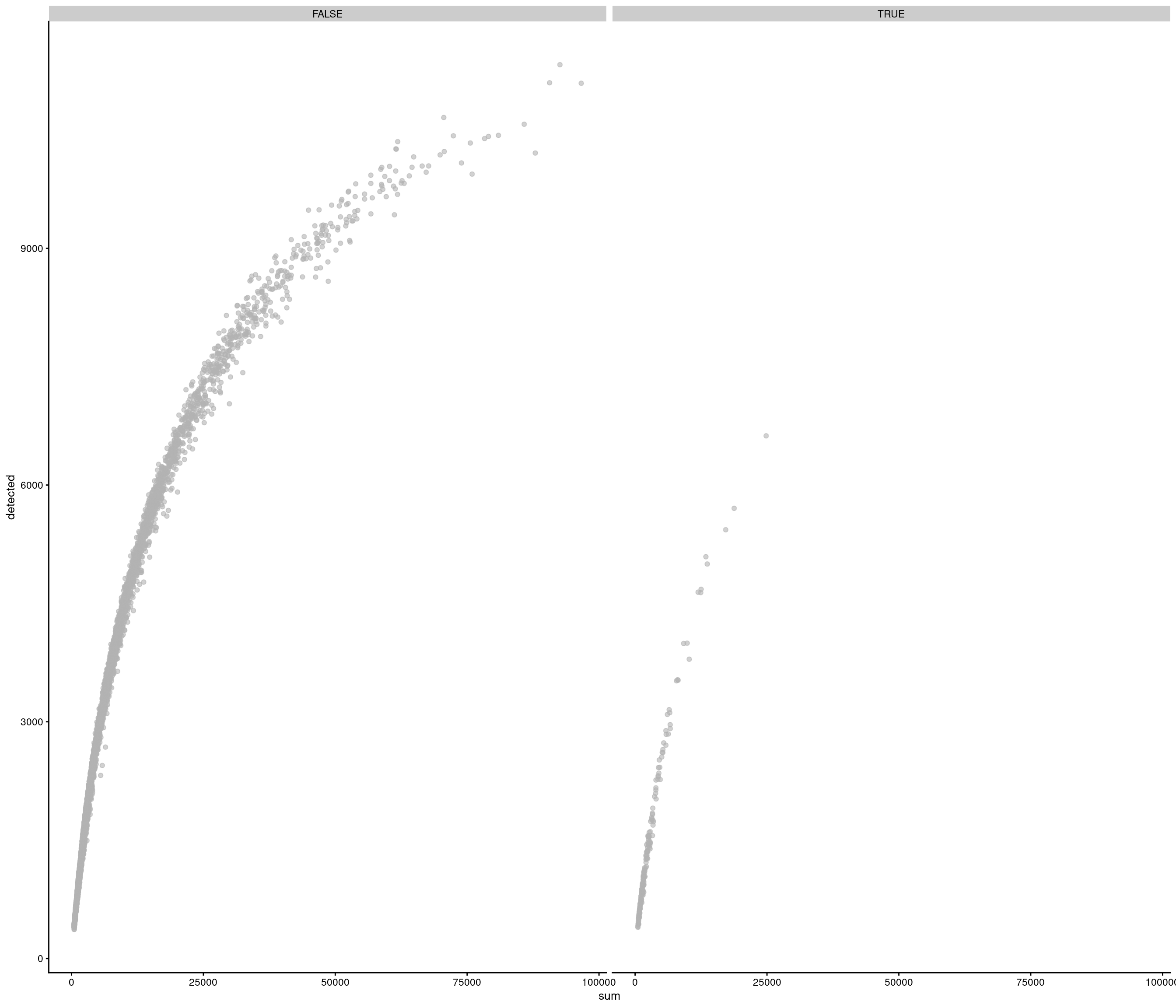
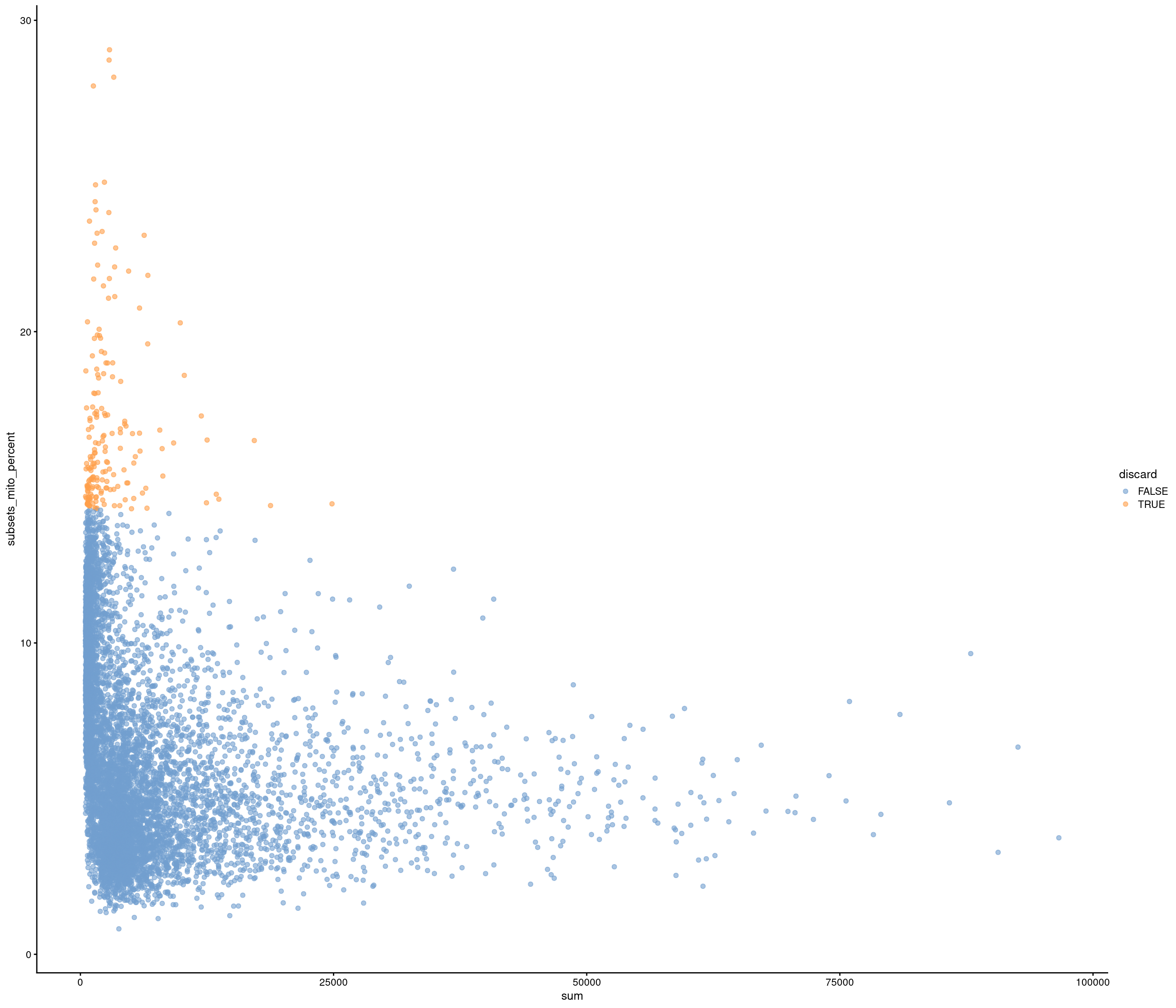
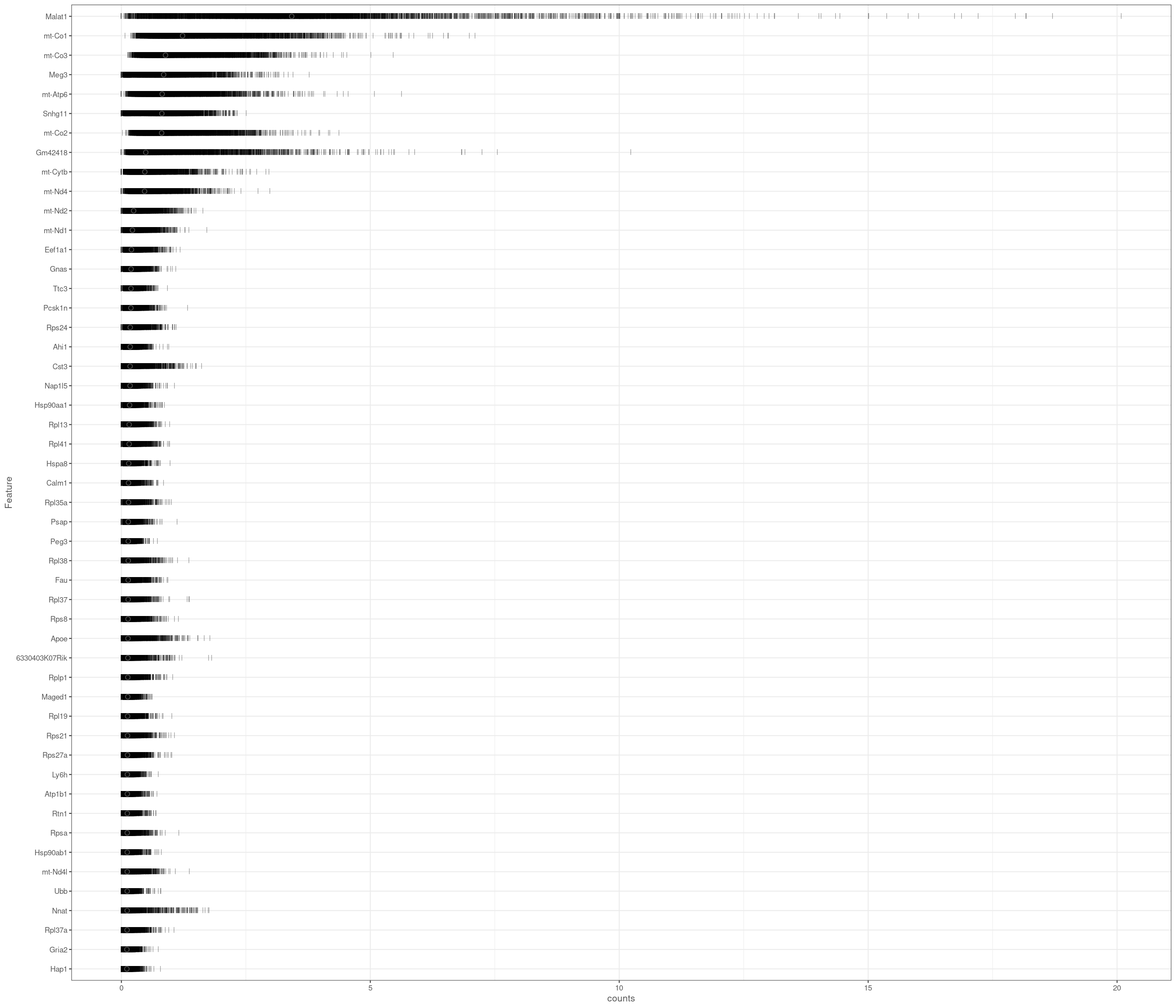
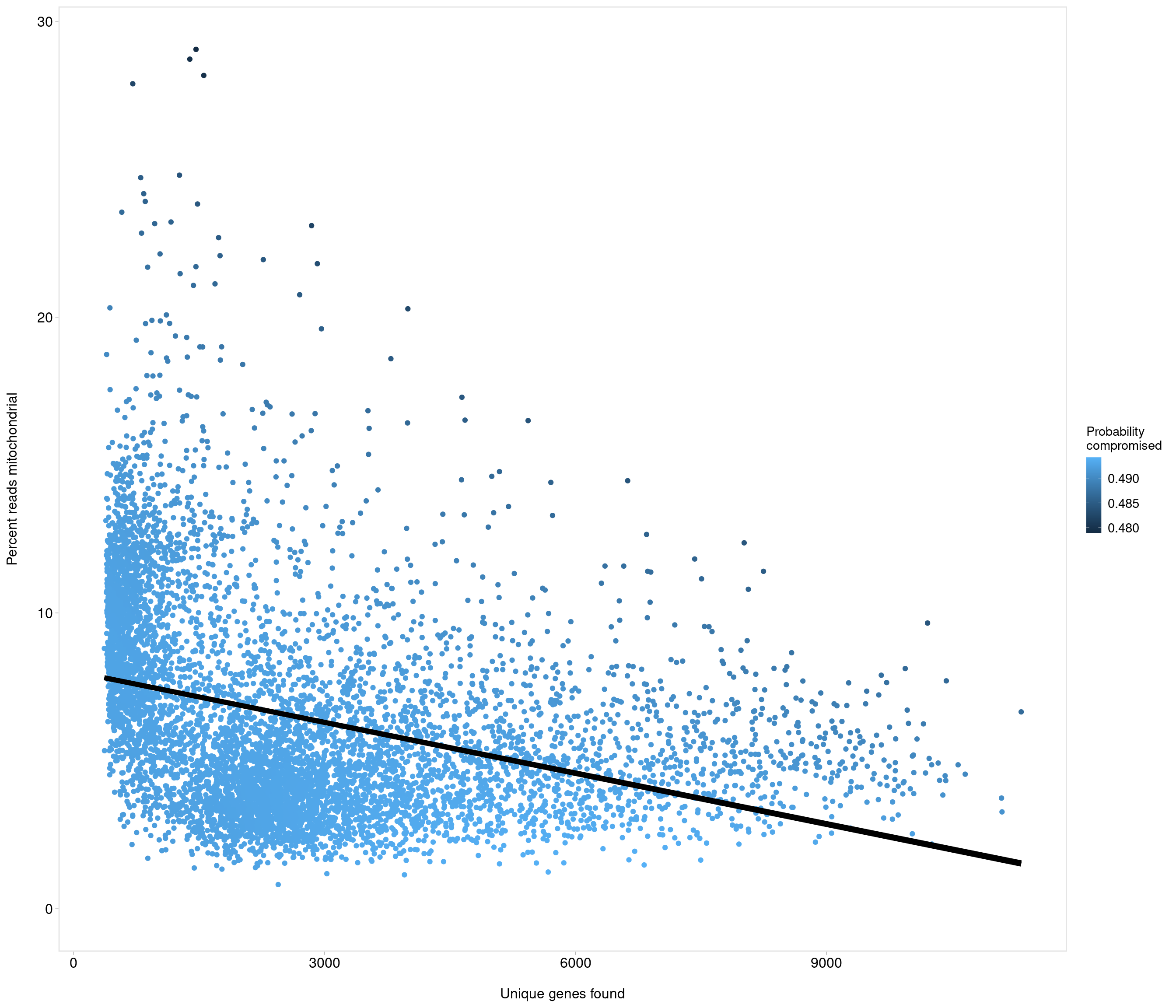
accept all?
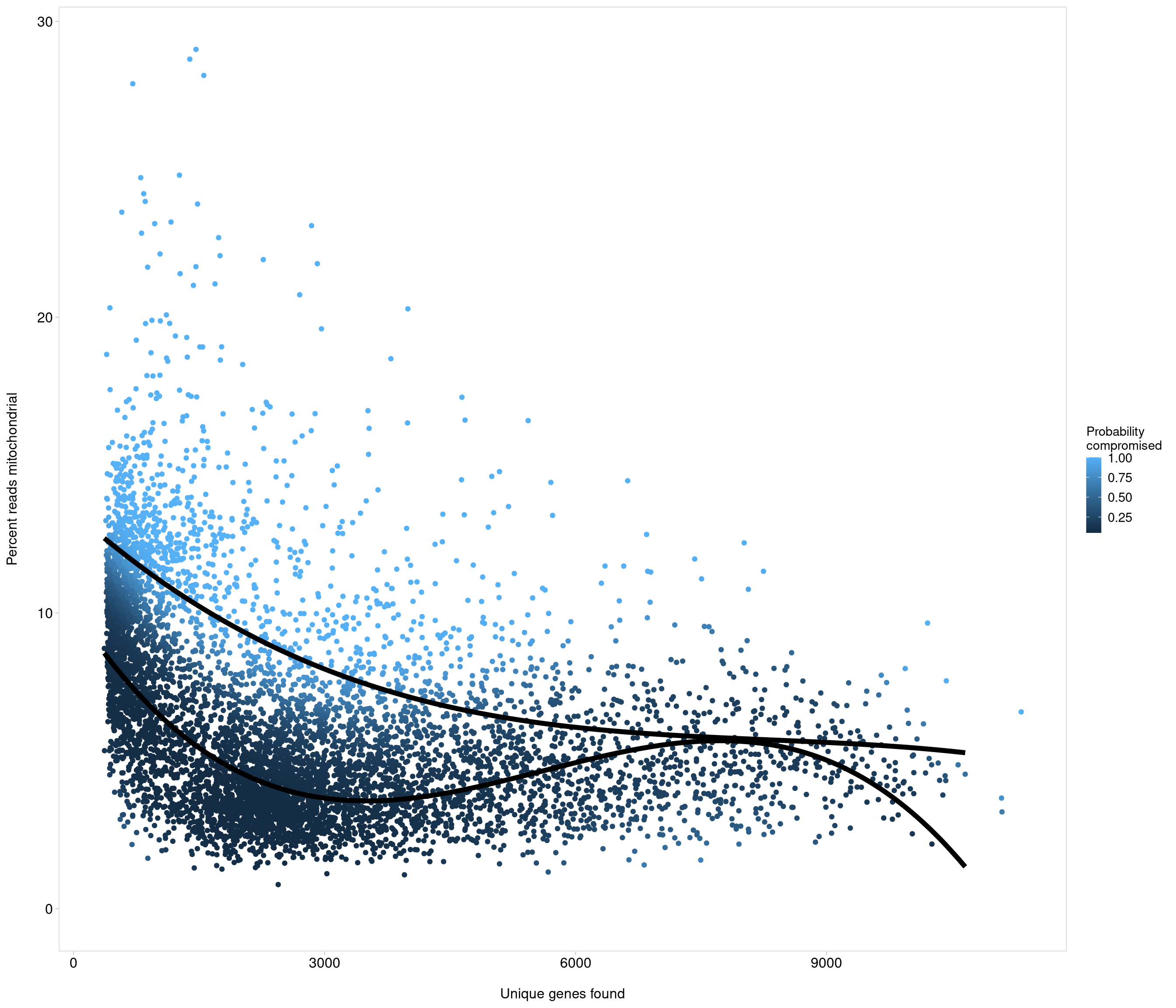
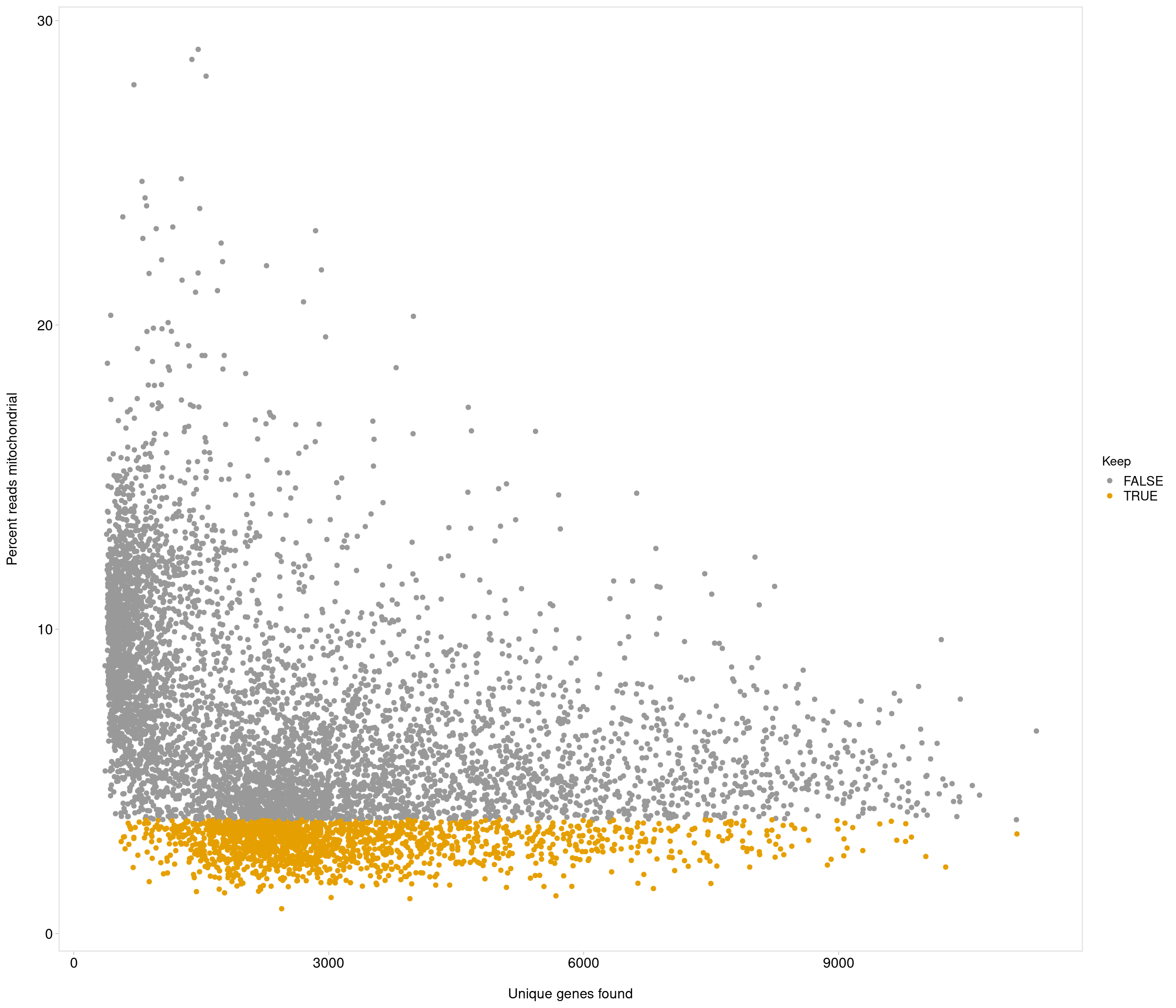
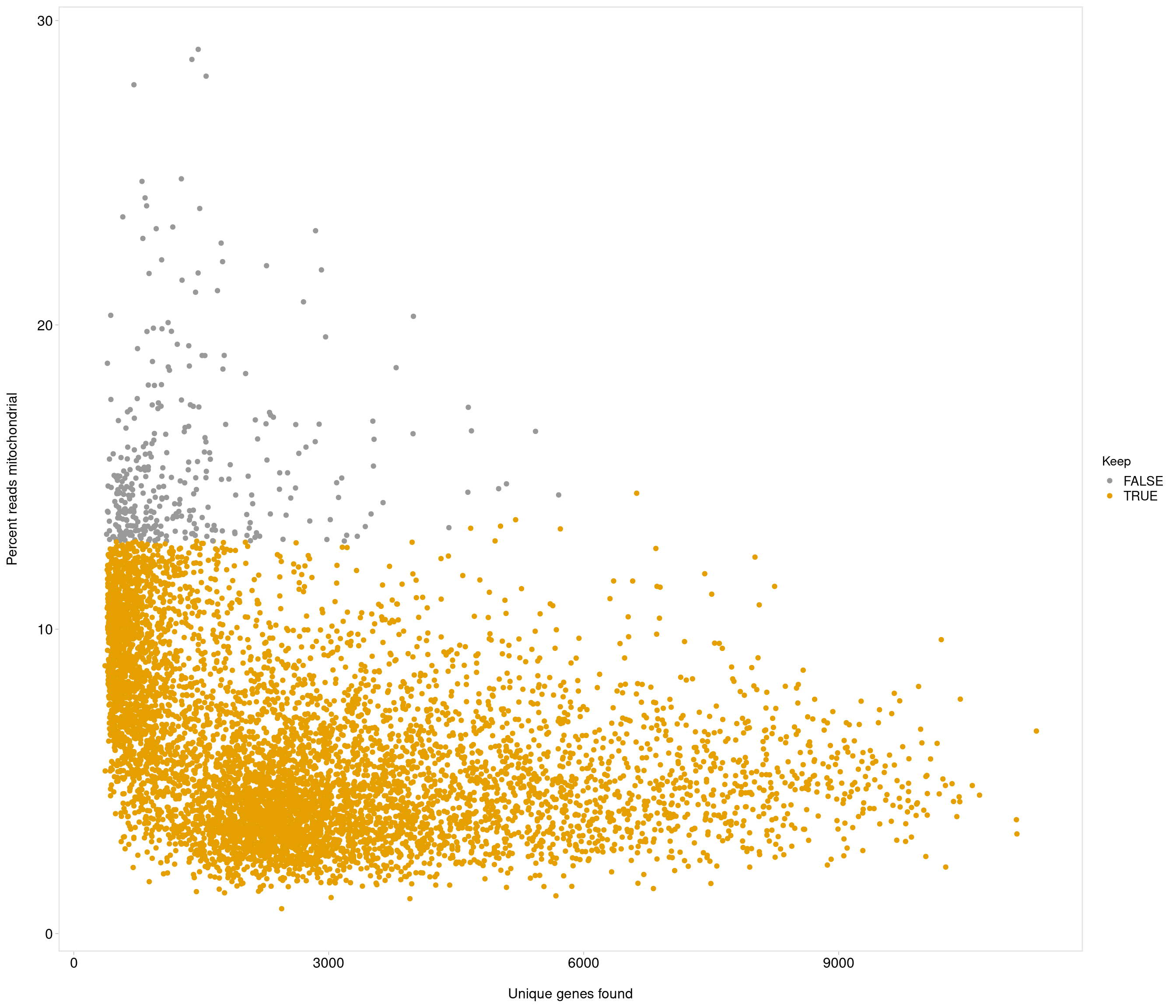
Removing 336 out of 7087 cells. used (Mb) gc trigger (Mb) max used (Mb)
Ncells 9234750 493.2 13872300 740.9 13872300 740.9
Vcells 235846119 1799.4 357802395 2729.9 323639064 2469.22. 537-1_537-3
$mito
[1] "mt-Nd1" "mt-Nd2" "mt-Co1" "mt-Co2" "mt-Atp8" "mt-Atp6" "mt-Co3"
[8] "mt-Nd3" "mt-Nd4l" "mt-Nd4" "mt-Nd5" "mt-Nd6" "mt-Cytb"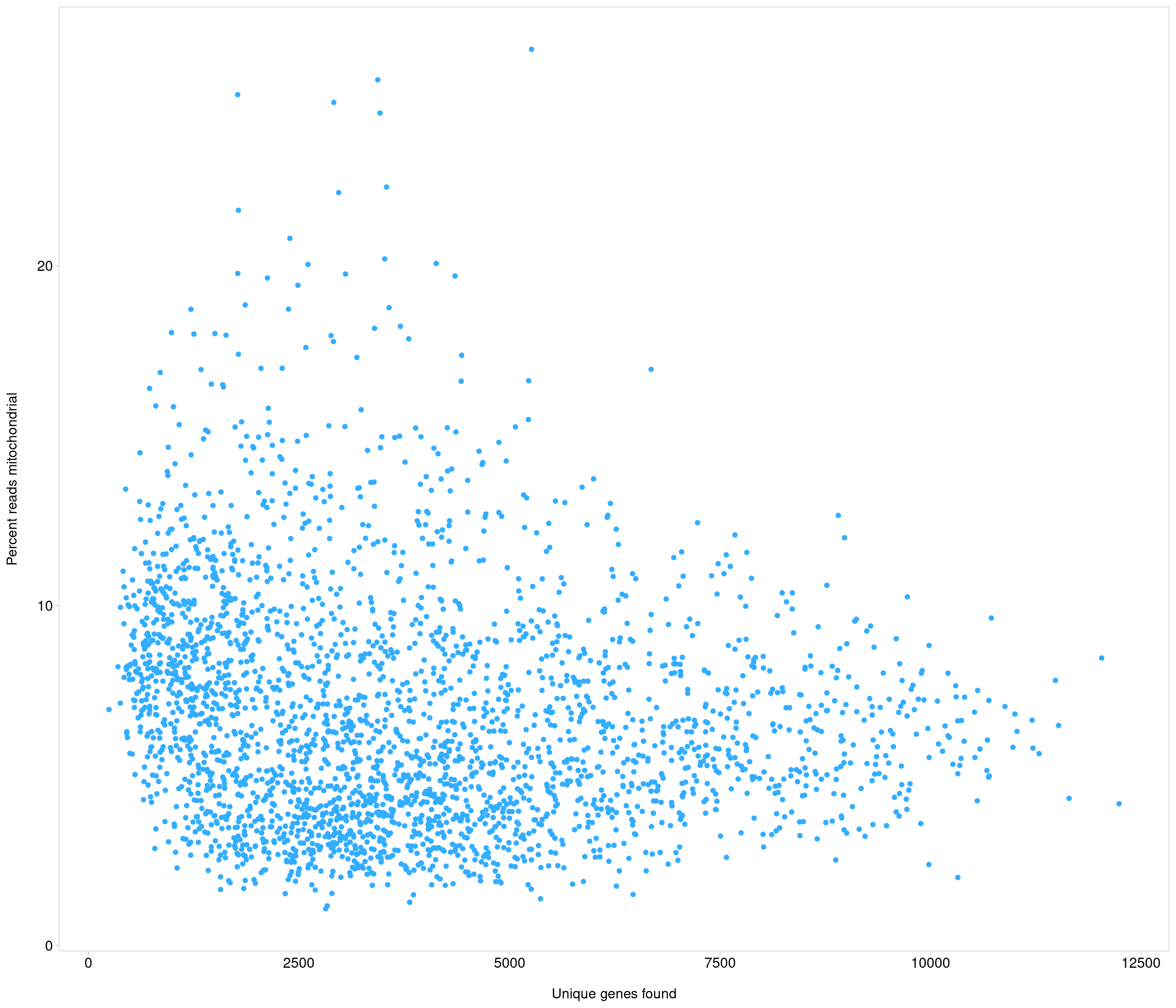
DataFrame with 6 rows and 19 columns
orig.ident nCount_RNA nFeature_RNA
<factor> <numeric> <integer>
AAACCCAAGGGAGTTC-1 537-1_537-3_high-fat-diet 16200 5741
AAACGAAAGCACTCAT-1 537-1_537-3_high-fat-diet 1009 749
AAACGAACATCCCGTT-1 537-1_537-3_high-fat-diet 2222 1595
AAACGAATCAATCTCT-1 537-1_537-3_high-fat-diet 15833 5450
AAACGAATCACGAGGA-1 537-1_537-3_high-fat-diet 2396 1556
AAACGAATCTTGATTC-1 537-1_537-3_high-fat-diet 30315 7033
age sex study_id tech hfd
<character> <character> <character> <character> <logical>
AAACCCAAGGGAGTTC-1 adult male deng_2020 10xv3 TRUE
AAACGAAAGCACTCAT-1 adult male deng_2020 10xv3 TRUE
AAACGAACATCCCGTT-1 adult male deng_2020 10xv3 TRUE
AAACGAATCAATCTCT-1 adult male deng_2020 10xv3 TRUE
AAACGAATCACGAGGA-1 adult male deng_2020 10xv3 TRUE
AAACGAATCTTGATTC-1 adult male deng_2020 10xv3 TRUE
ident sum detected
<factor> <numeric> <integer>
AAACCCAAGGGAGTTC-1 537-1_537-3_high-fat-diet 16200 5741
AAACGAAAGCACTCAT-1 537-1_537-3_high-fat-diet 1009 749
AAACGAACATCCCGTT-1 537-1_537-3_high-fat-diet 2222 1595
AAACGAATCAATCTCT-1 537-1_537-3_high-fat-diet 15833 5450
AAACGAATCACGAGGA-1 537-1_537-3_high-fat-diet 2396 1556
AAACGAATCTTGATTC-1 537-1_537-3_high-fat-diet 30315 7033
subsets_mito_sum subsets_mito_detected subsets_mito_percent
<numeric> <integer> <numeric>
AAACCCAAGGGAGTTC-1 887 12 5.47531
AAACGAAAGCACTCAT-1 80 12 7.92864
AAACGAACATCCCGTT-1 110 12 4.95050
AAACGAATCAATCTCT-1 739 12 4.66747
AAACGAATCACGAGGA-1 112 12 4.67446
AAACGAATCTTGATTC-1 2406 12 7.93667
total low_lib_size low_n_features
<numeric> <outlier.filter> <outlier.filter>
AAACCCAAGGGAGTTC-1 16200 FALSE FALSE
AAACGAAAGCACTCAT-1 1009 FALSE FALSE
AAACGAACATCCCGTT-1 2222 FALSE FALSE
AAACGAATCAATCTCT-1 15833 FALSE FALSE
AAACGAATCACGAGGA-1 2396 FALSE FALSE
AAACGAATCTTGATTC-1 30315 FALSE FALSE
high_subsets_mito_percent discard
<outlier.filter> <logical>
AAACCCAAGGGAGTTC-1 FALSE FALSE
AAACGAAAGCACTCAT-1 FALSE FALSE
AAACGAACATCCCGTT-1 FALSE FALSE
AAACGAATCAATCTCT-1 FALSE FALSE
AAACGAATCACGAGGA-1 FALSE FALSE
AAACGAATCTTGATTC-1 FALSE FALSE| Name | sce$sum |
| Number of rows | 2842 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 12191.45 | 15744.52 | 512 | 3088 | 6722 | 14008.75 | 180047 | ▇▁▁▁▁ |
| Name | sce$detected |
| Number of rows | 2842 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 3856.69 | 2427.68 | 243 | 1862.25 | 3391 | 5264.25 | 12240 | ▇▇▃▂▁ |
| Name | sce$subsets_mito_percent |
| Number of rows | 2842 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 6.78 | 3.38 | 1.08 | 4.22 | 6.2 | 8.63 | 26.36 | ▇▆▁▁▁ |
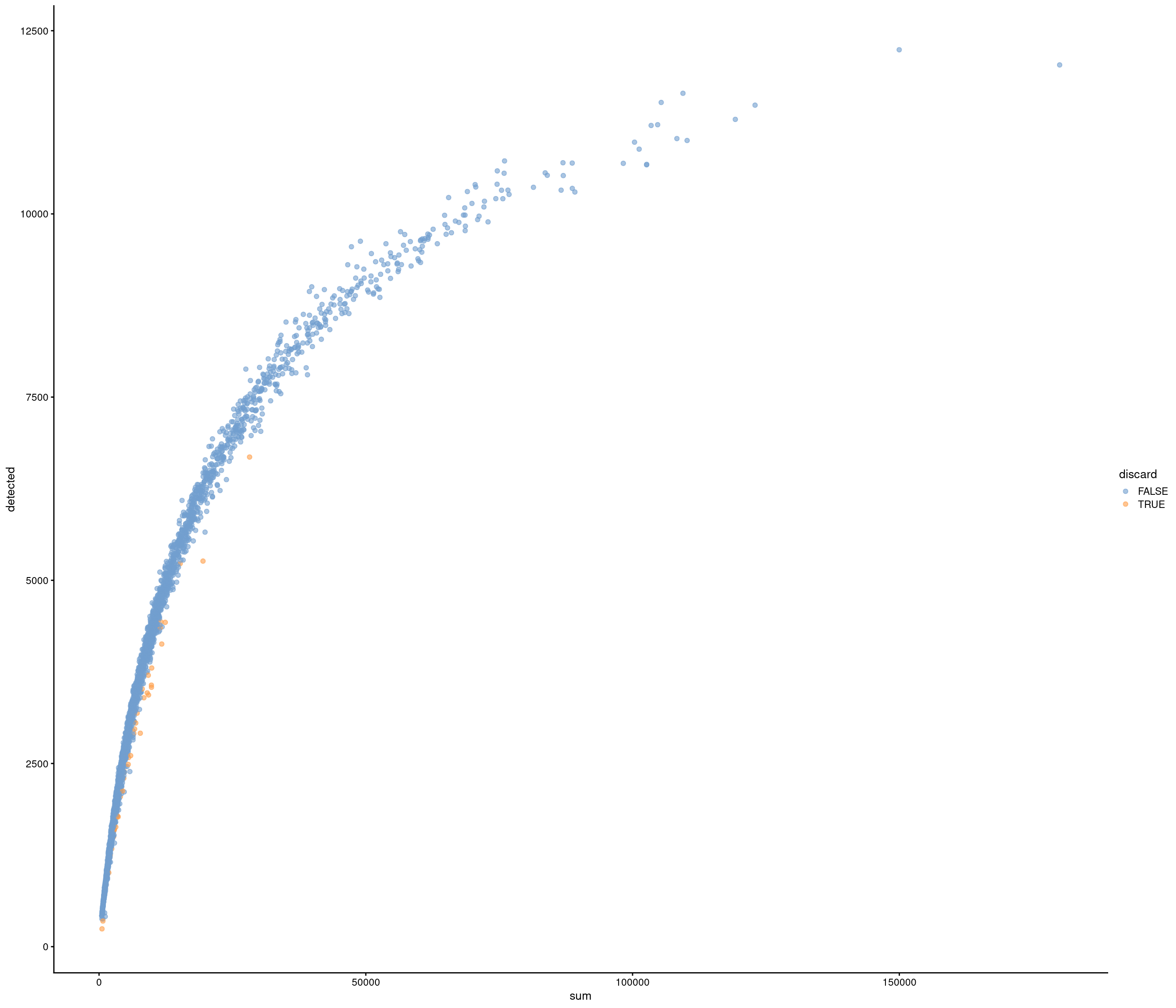
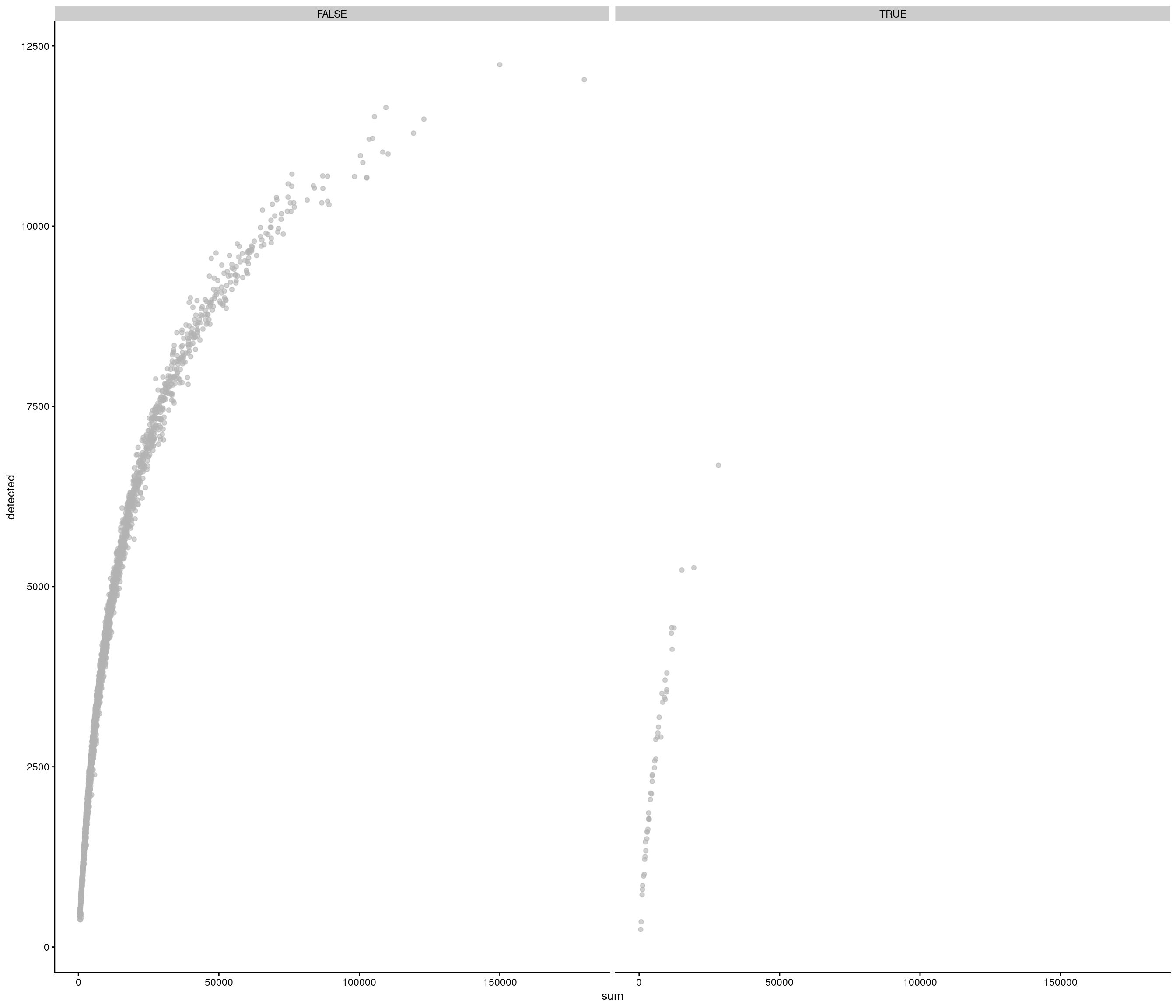
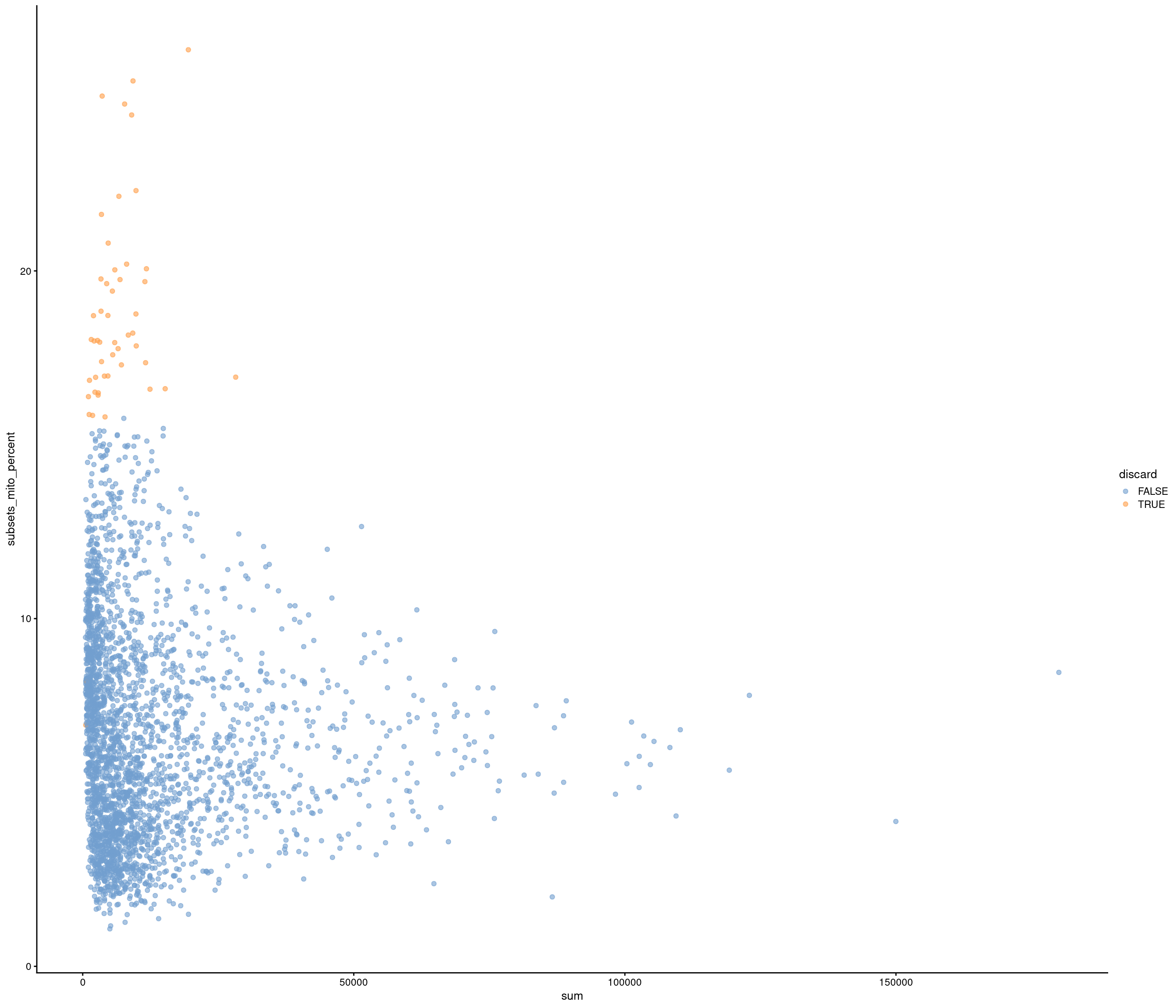
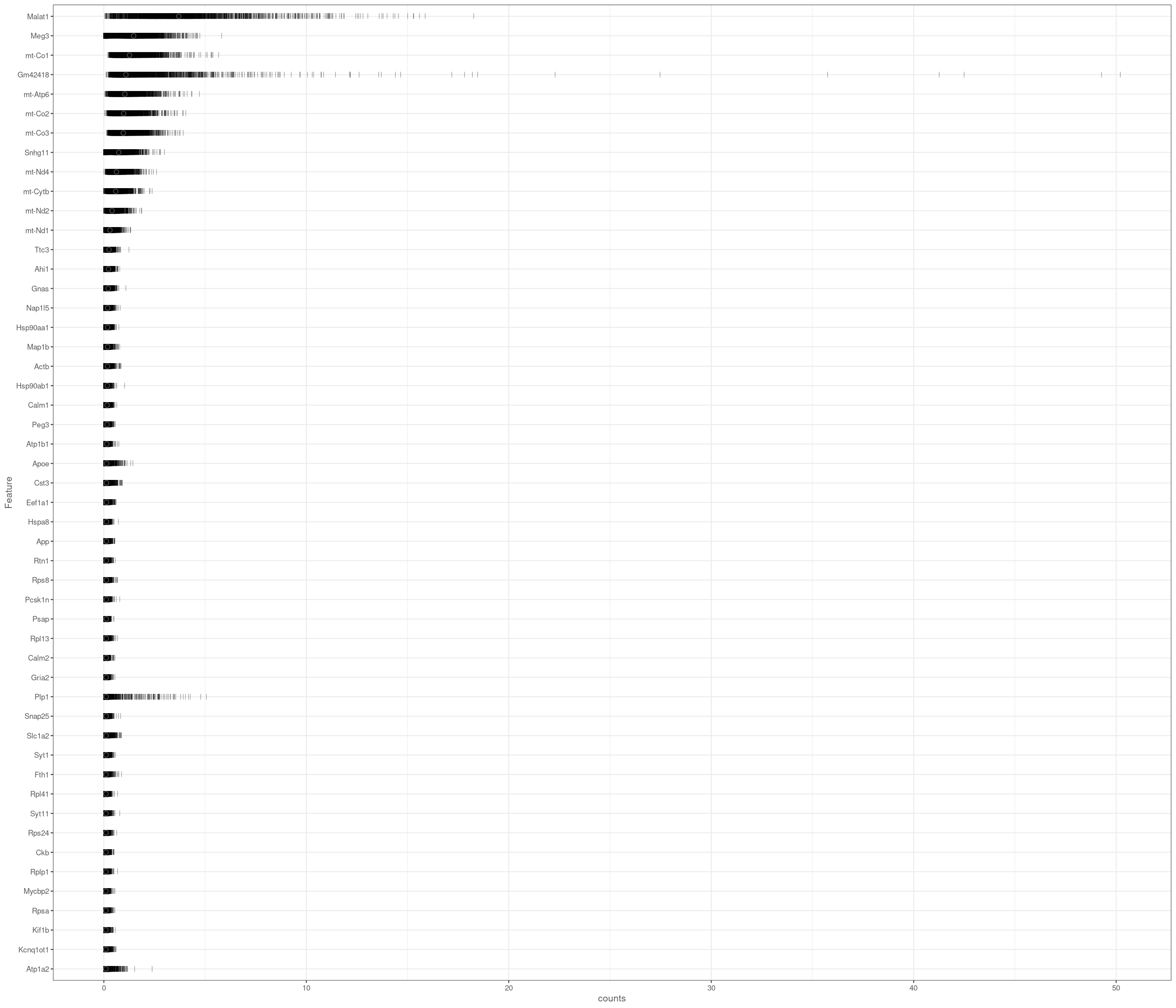
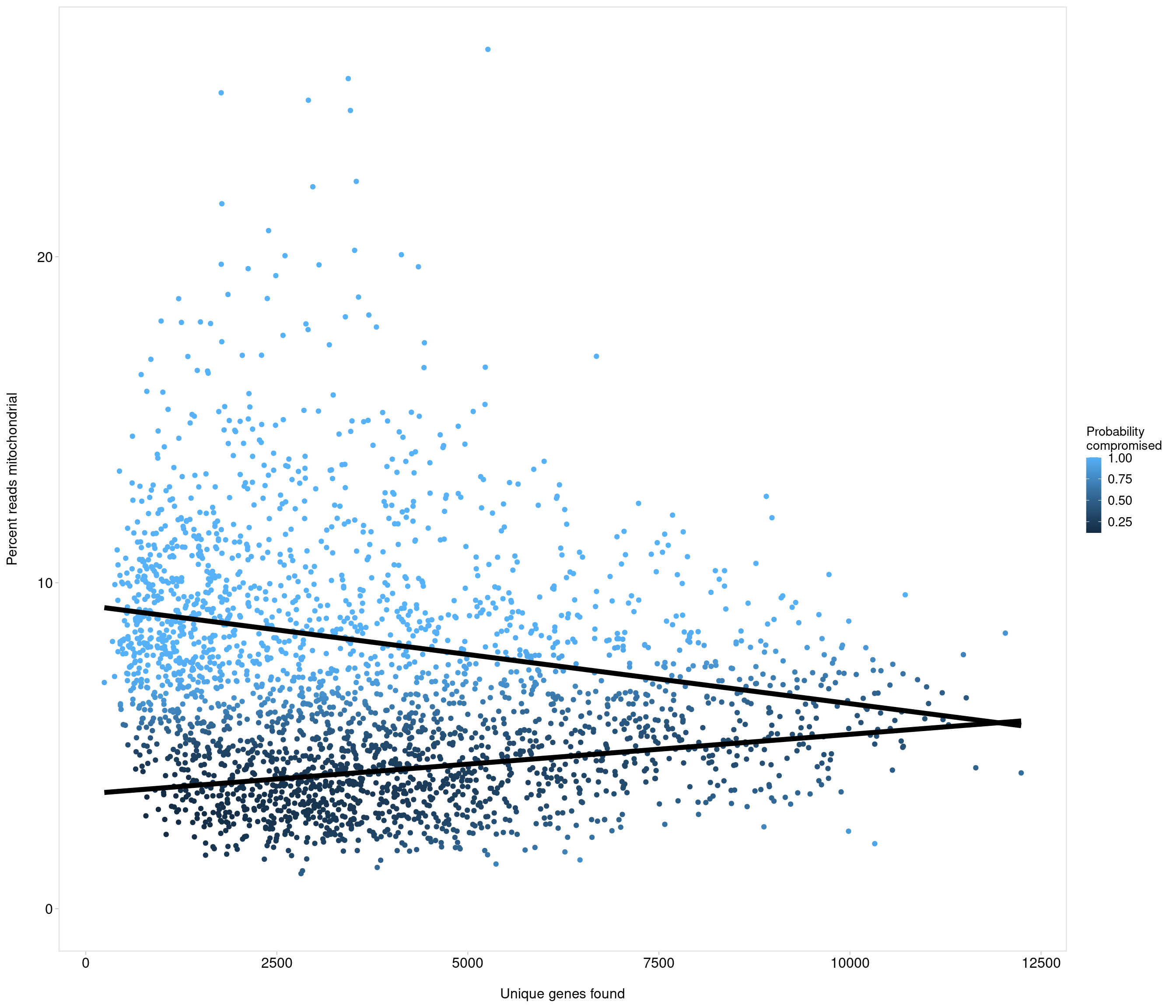
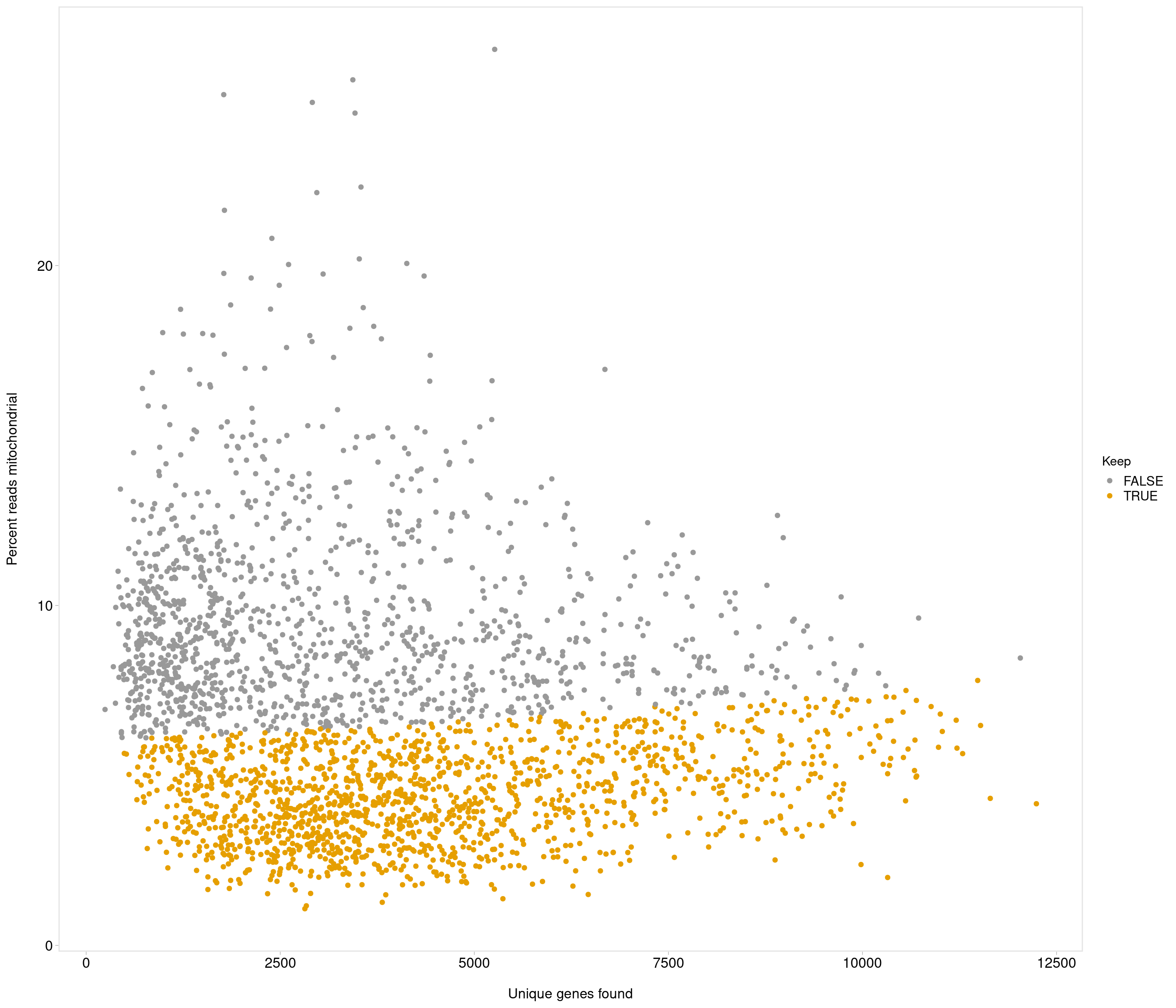
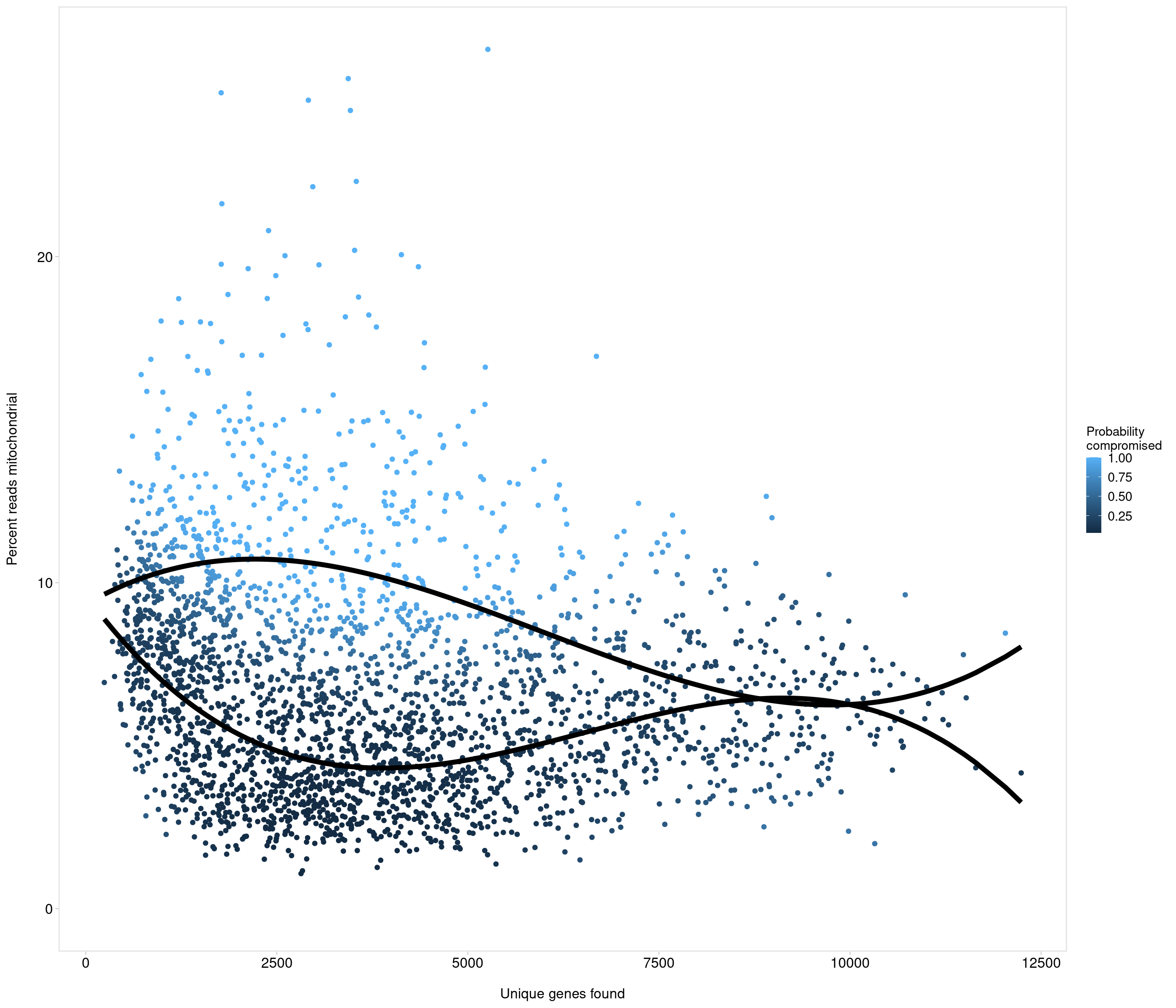
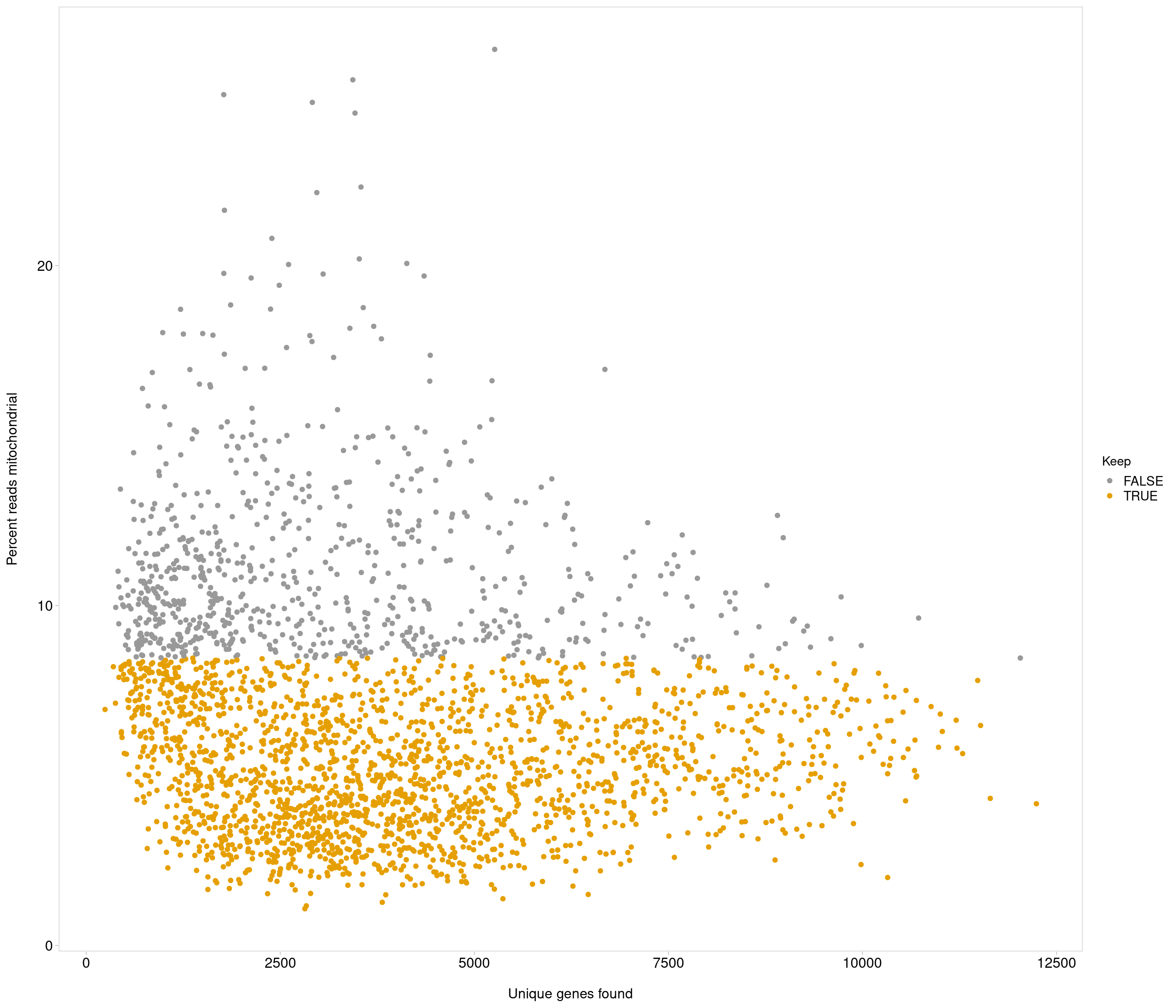
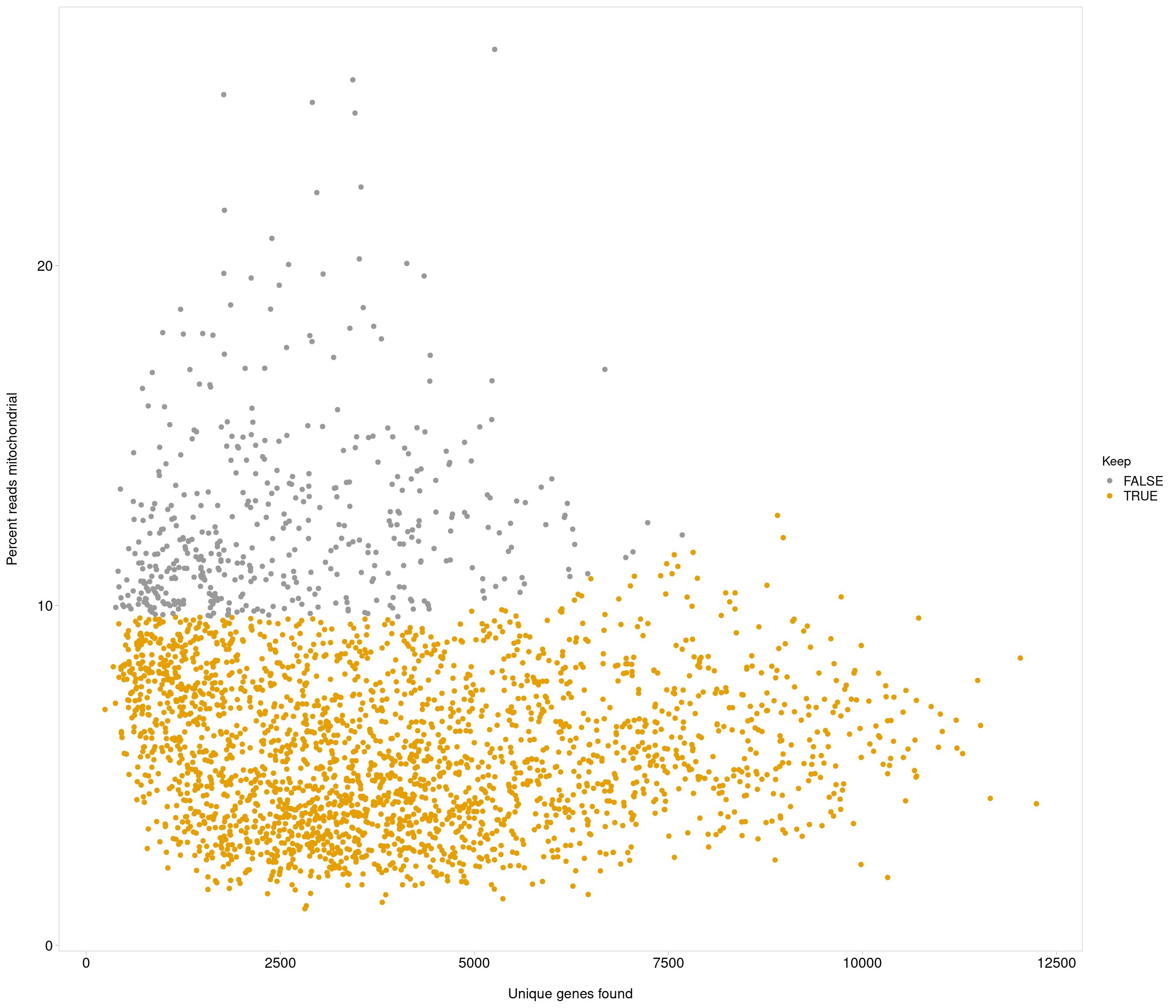
Removing 453 out of 2842 cells. used (Mb) gc trigger (Mb) max used (Mb)
Ncells 9235080 493.3 13872300 740.9 13872300 740.9
Vcells 235395270 1796.0 357802395 2729.9 323639064 2469.23. 537-5_538-2
$mito
[1] "mt-Nd1" "mt-Nd2" "mt-Co1" "mt-Co2" "mt-Atp8" "mt-Atp6" "mt-Co3"
[8] "mt-Nd3" "mt-Nd4l" "mt-Nd4" "mt-Nd5" "mt-Nd6" "mt-Cytb"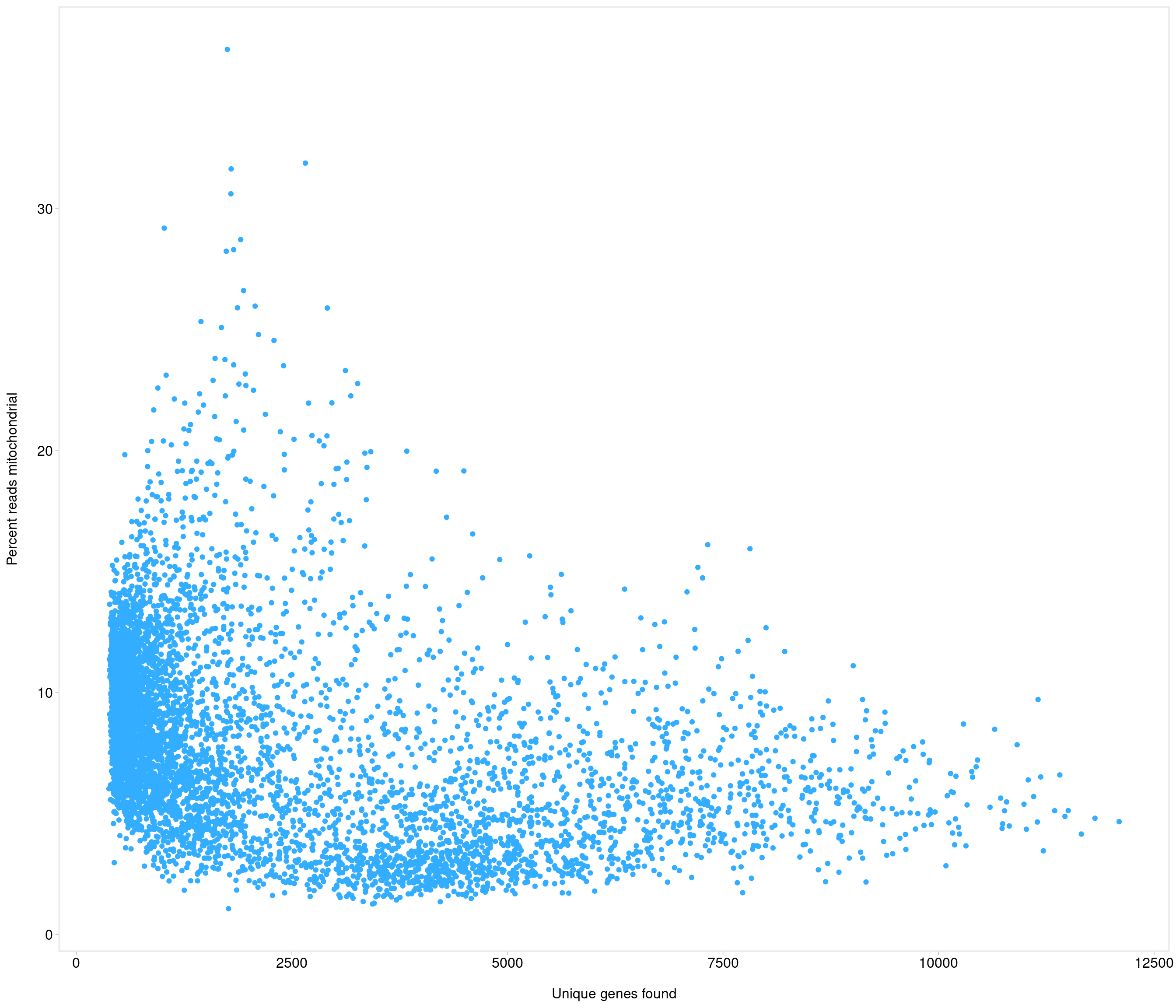
DataFrame with 6 rows and 19 columns
orig.ident nCount_RNA nFeature_RNA
<factor> <numeric> <integer>
AAACCCACACACGTGC-1 537-5_538-2_high-fat-diet 6371 3407
AAACGAAAGCGTGCCT-1 537-5_538-2_high-fat-diet 4798 2587
AAACGCTAGACCATAA-1 537-5_538-2_high-fat-diet 750 588
AAACGCTAGCAGGCAT-1 537-5_538-2_high-fat-diet 553 426
AAACGCTAGCCTTGAT-1 537-5_538-2_high-fat-diet 1052 739
AAACGCTAGGTATCTC-1 537-5_538-2_high-fat-diet 709 561
age sex study_id tech hfd
<character> <character> <character> <character> <logical>
AAACCCACACACGTGC-1 adult male deng_2020 10xv3 TRUE
AAACGAAAGCGTGCCT-1 adult male deng_2020 10xv3 TRUE
AAACGCTAGACCATAA-1 adult male deng_2020 10xv3 TRUE
AAACGCTAGCAGGCAT-1 adult male deng_2020 10xv3 TRUE
AAACGCTAGCCTTGAT-1 adult male deng_2020 10xv3 TRUE
AAACGCTAGGTATCTC-1 adult male deng_2020 10xv3 TRUE
ident sum detected
<factor> <numeric> <integer>
AAACCCACACACGTGC-1 537-5_538-2_high-fat-diet 6371 3407
AAACGAAAGCGTGCCT-1 537-5_538-2_high-fat-diet 4798 2587
AAACGCTAGACCATAA-1 537-5_538-2_high-fat-diet 750 588
AAACGCTAGCAGGCAT-1 537-5_538-2_high-fat-diet 553 426
AAACGCTAGCCTTGAT-1 537-5_538-2_high-fat-diet 1052 739
AAACGCTAGGTATCTC-1 537-5_538-2_high-fat-diet 709 561
subsets_mito_sum subsets_mito_detected subsets_mito_percent
<numeric> <integer> <numeric>
AAACCCACACACGTGC-1 149 12 2.33872
AAACGAAAGCGTGCCT-1 397 12 8.27428
AAACGCTAGACCATAA-1 52 10 6.93333
AAACGCTAGCAGGCAT-1 73 12 13.20072
AAACGCTAGCCTTGAT-1 126 10 11.97719
AAACGCTAGGTATCTC-1 44 10 6.20592
total low_lib_size low_n_features
<numeric> <outlier.filter> <outlier.filter>
AAACCCACACACGTGC-1 6371 FALSE FALSE
AAACGAAAGCGTGCCT-1 4798 FALSE FALSE
AAACGCTAGACCATAA-1 750 FALSE FALSE
AAACGCTAGCAGGCAT-1 553 FALSE FALSE
AAACGCTAGCCTTGAT-1 1052 FALSE FALSE
AAACGCTAGGTATCTC-1 709 FALSE FALSE
high_subsets_mito_percent discard
<outlier.filter> <logical>
AAACCCACACACGTGC-1 FALSE FALSE
AAACGAAAGCGTGCCT-1 FALSE FALSE
AAACGCTAGACCATAA-1 FALSE FALSE
AAACGCTAGCAGGCAT-1 FALSE FALSE
AAACGCTAGCCTTGAT-1 FALSE FALSE
AAACGCTAGGTATCTC-1 FALSE FALSE| Name | sce$sum |
| Number of rows | 6238 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 6846.18 | 12830.54 | 500 | 836 | 1838 | 7668.75 | 175743 | ▇▁▁▁▁ |
| Name | sce$detected |
| Number of rows | 6238 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 2385.75 | 2348.67 | 379 | 636 | 1225.5 | 3736 | 12094 | ▇▂▁▁▁ |
| Name | sce$subsets_mito_percent |
| Number of rows | 6238 |
| Number of columns | 1 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| data | 0 | 1 | 7.91 | 3.85 | 1.07 | 5.04 | 7.62 | 10.18 | 36.59 | ▇▆▁▁▁ |
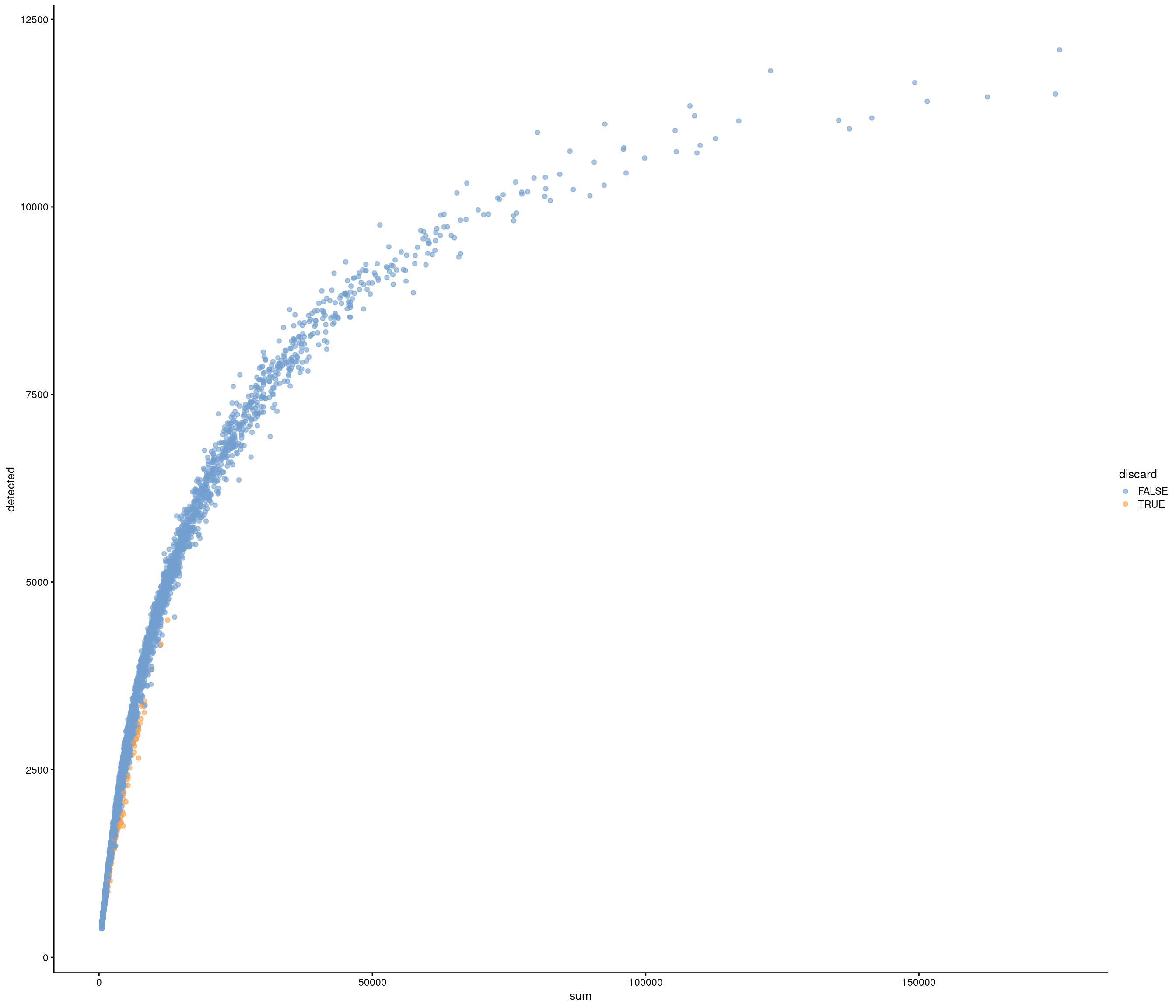
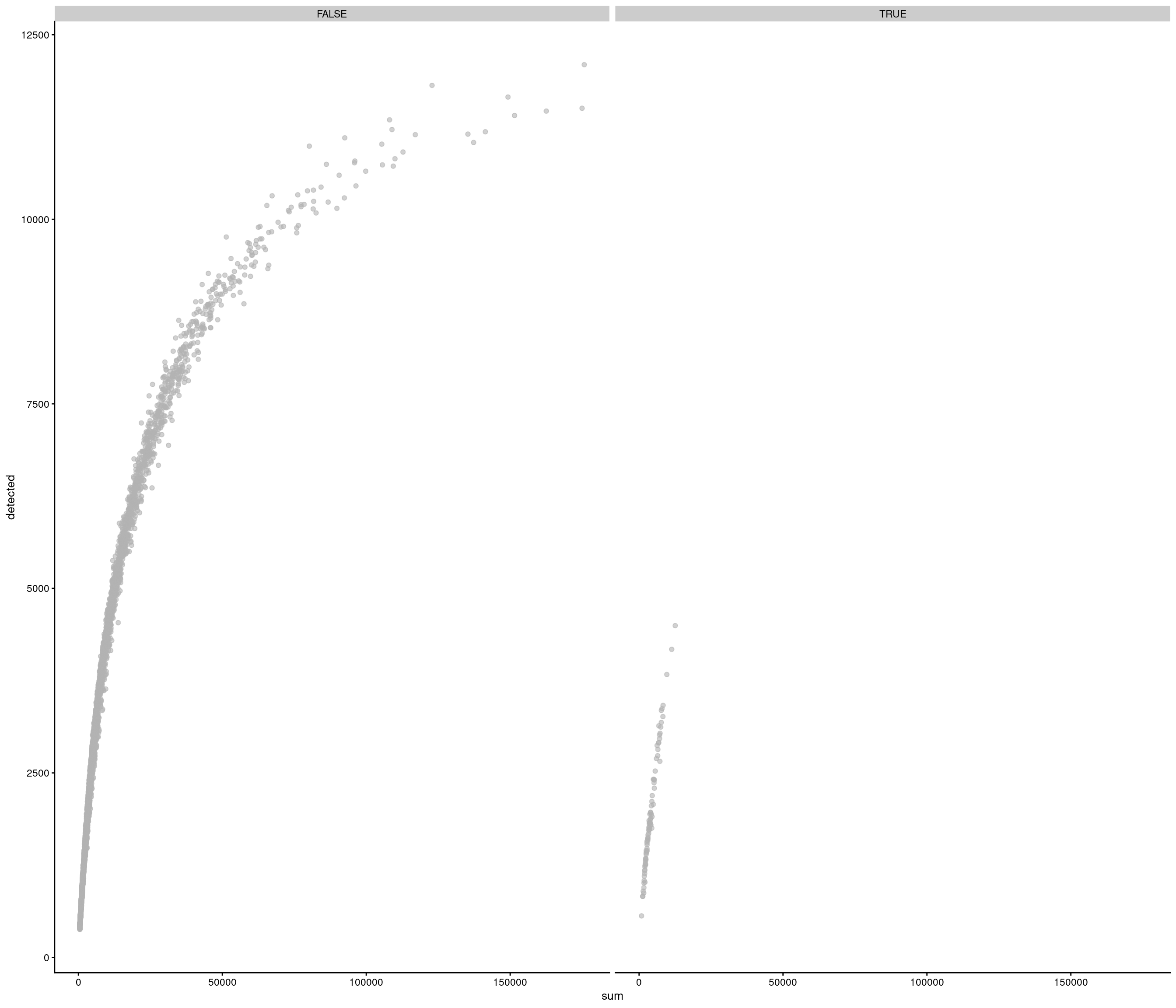
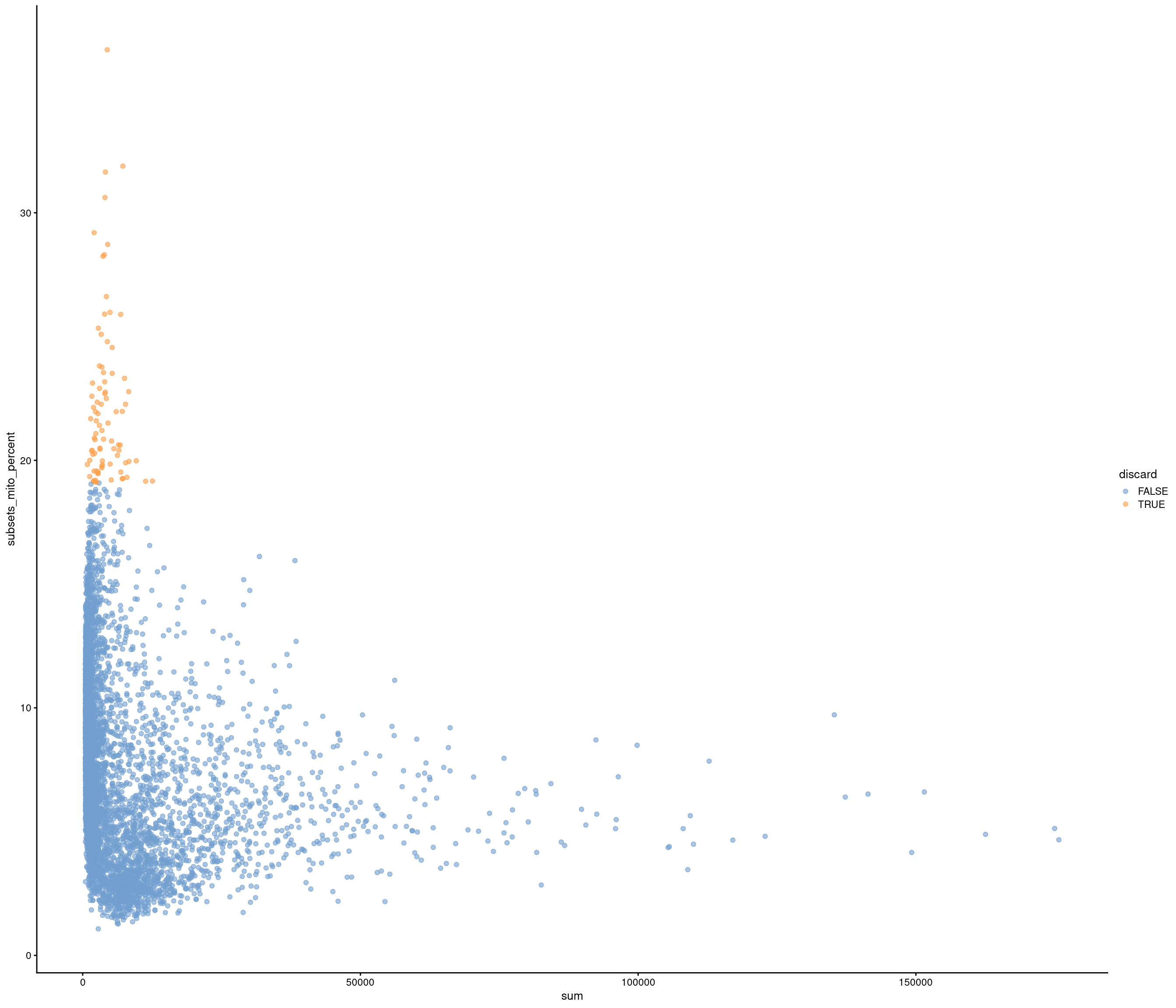
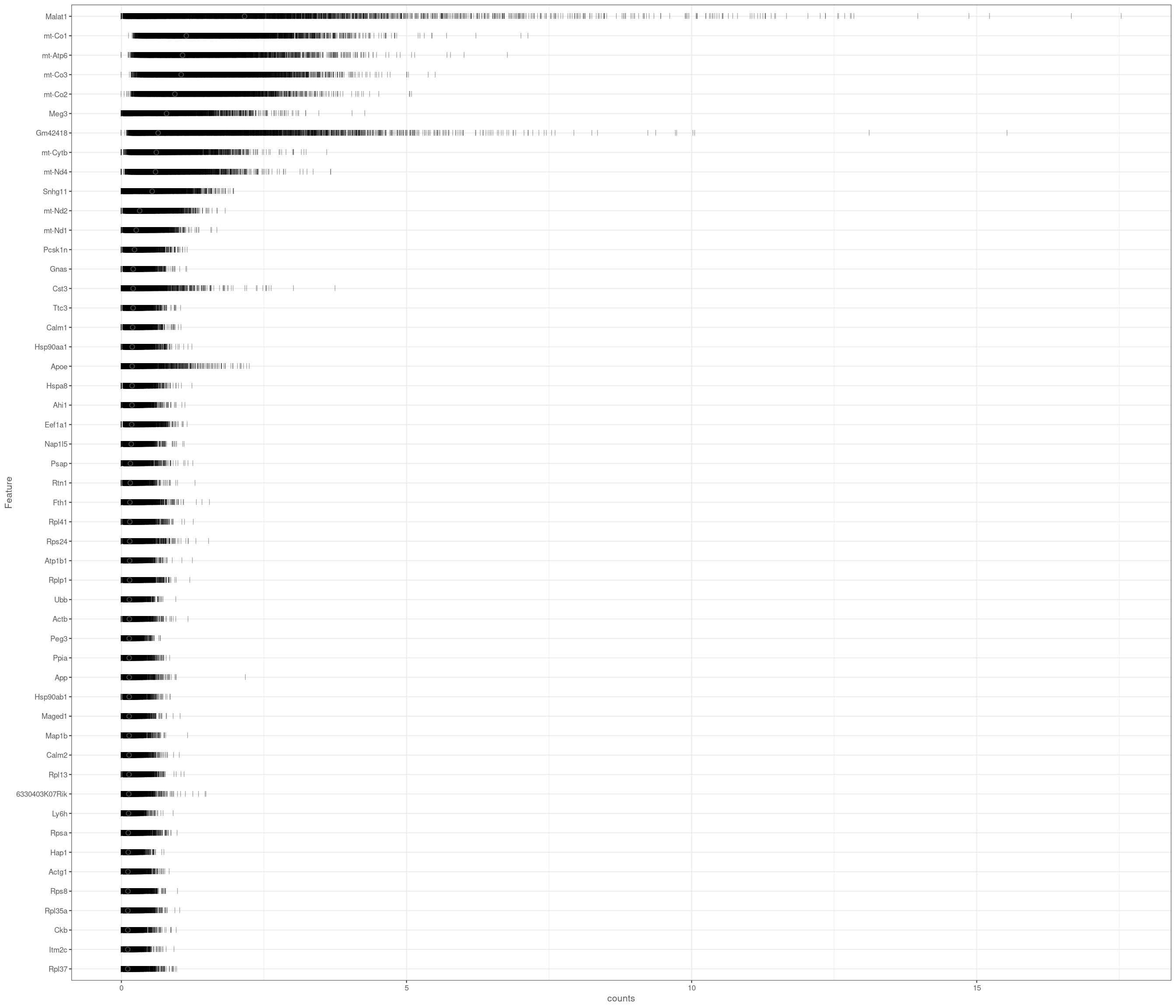
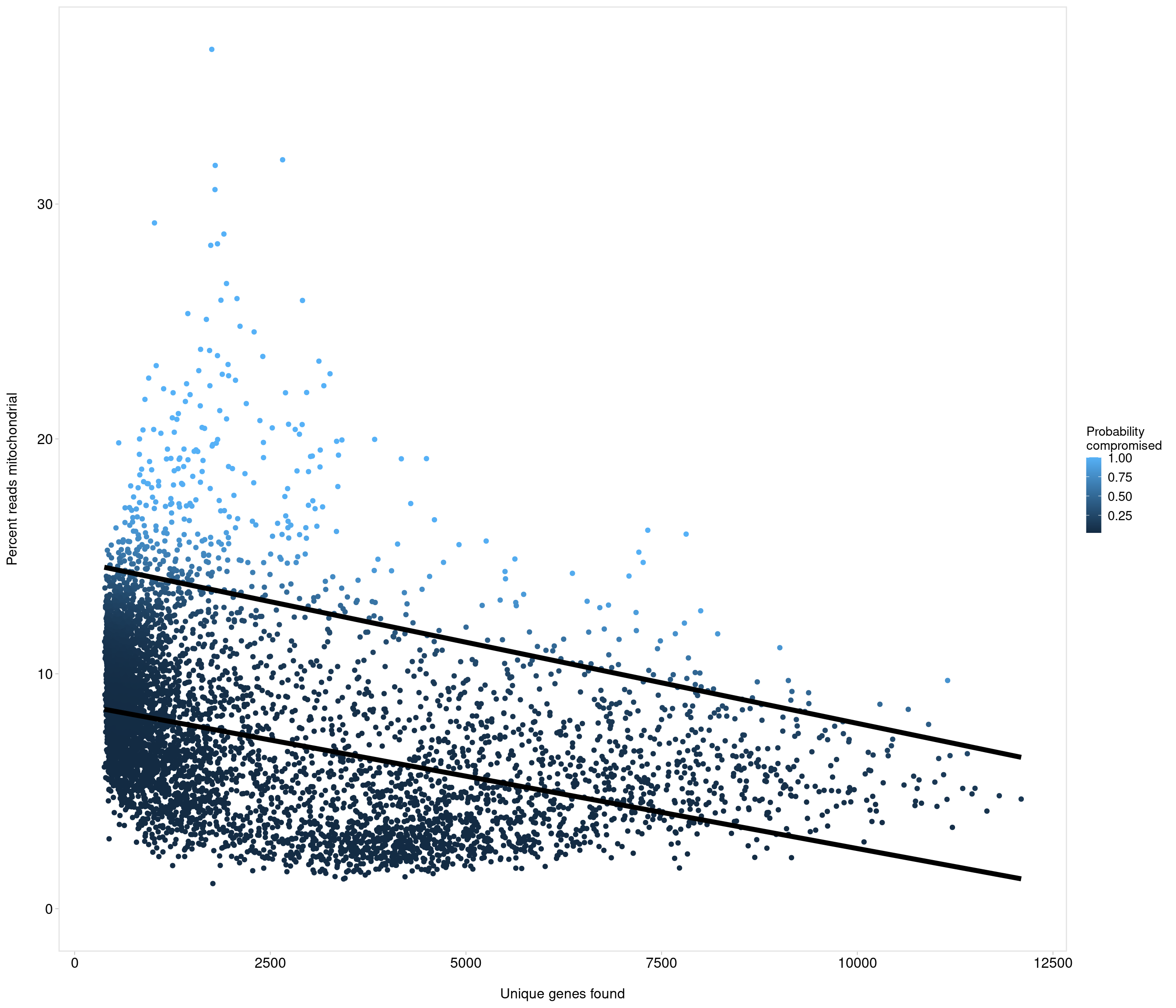
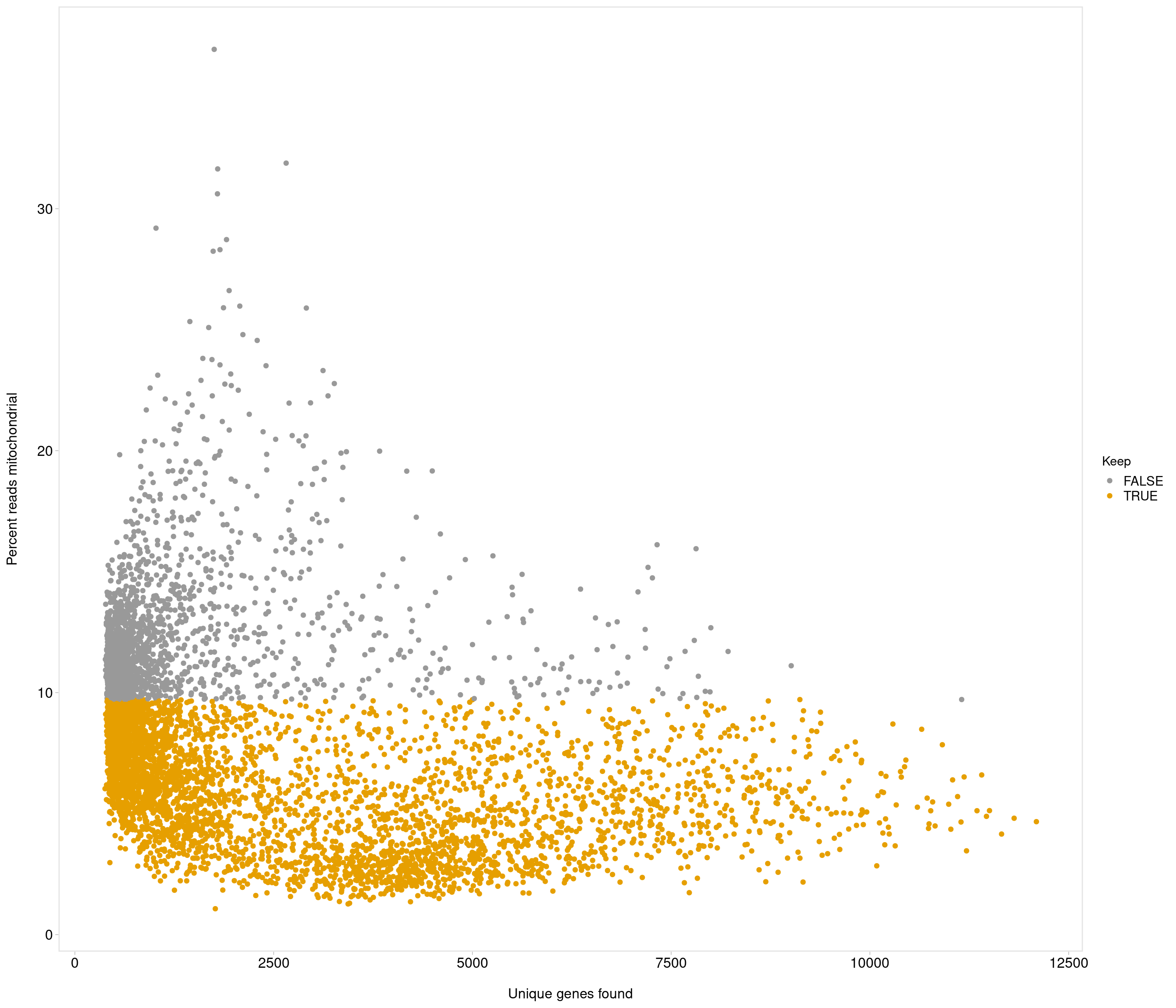
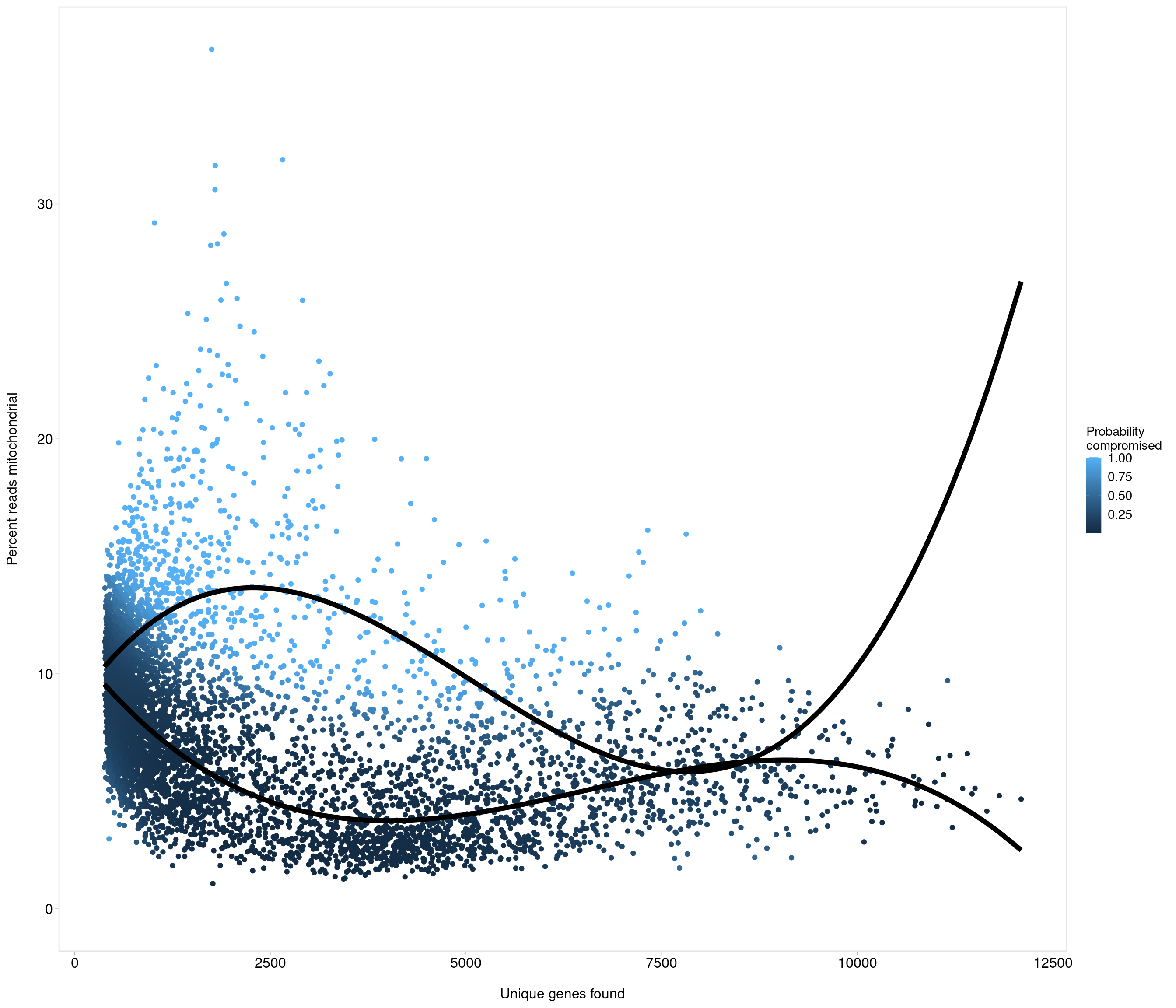
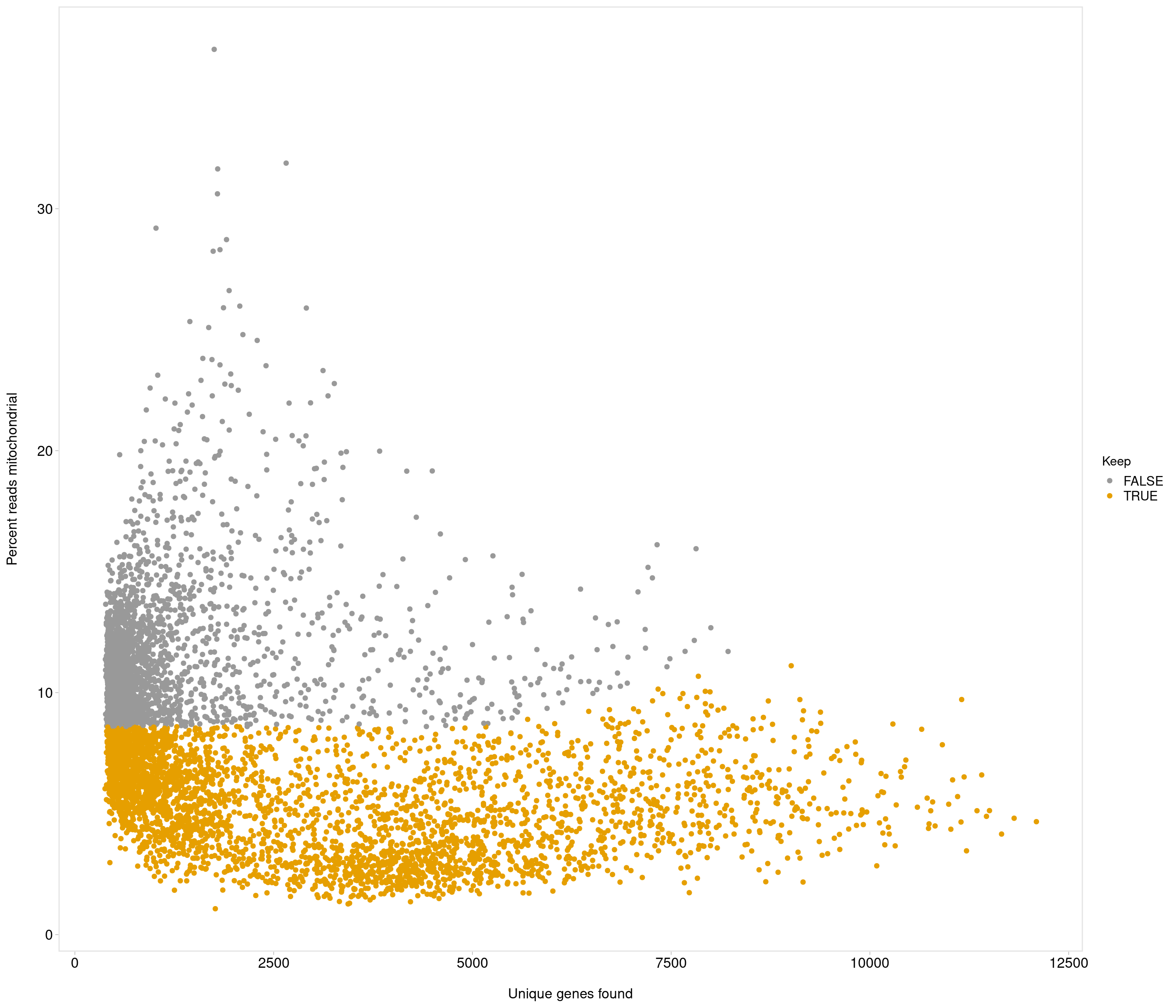
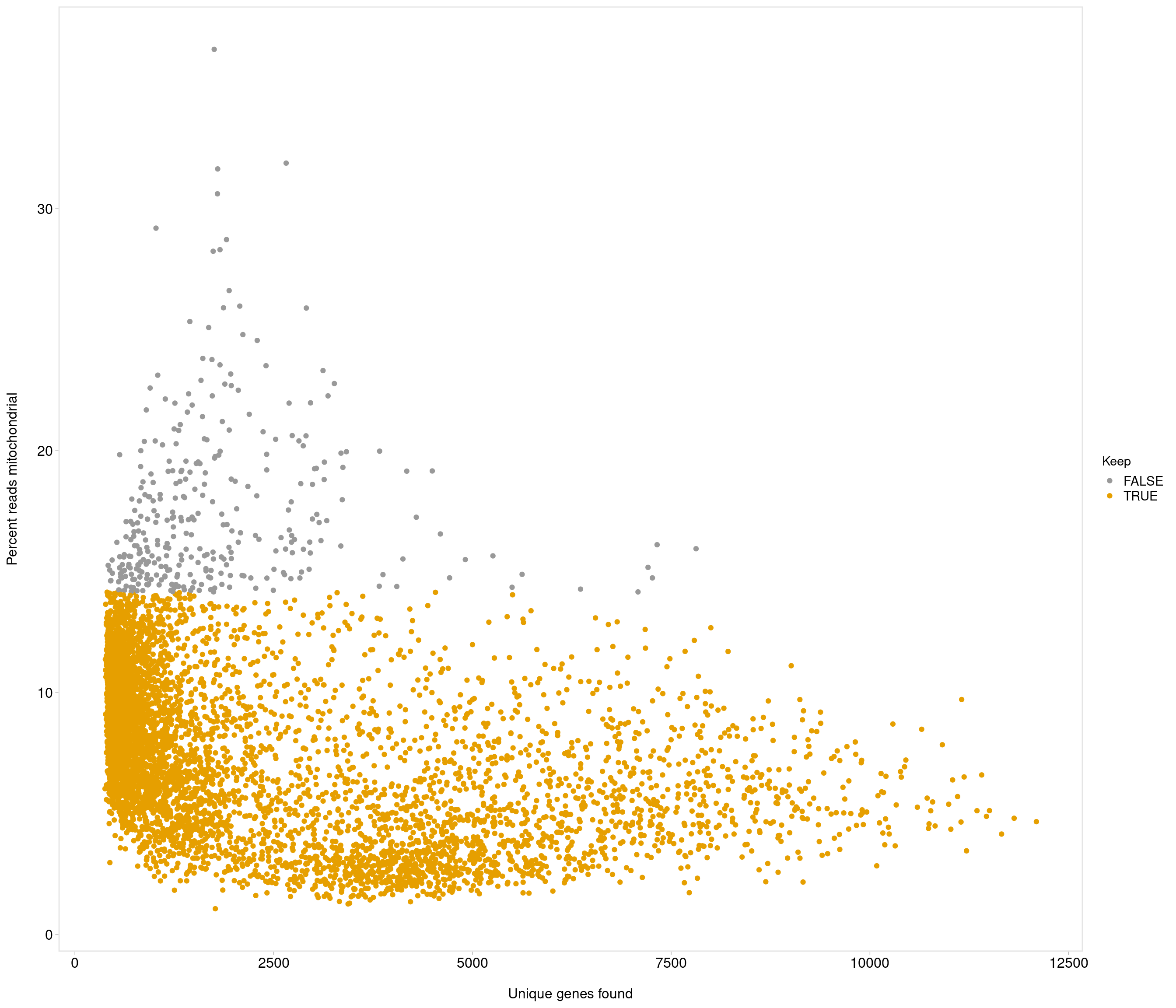
Removing 337 out of 6238 cells. used (Mb) gc trigger (Mb) max used (Mb)
Ncells 9235275 493.3 13872300 740.9 13872300 740.9
Vcells 262073048 1999.5 429442874 3276.4 353580218 2697.7Save initial data
Rows: 9,299
Columns: 20
$ orig.ident <chr> "536-1_chow-diet", "536-1_chow-diet", "536-1…
$ nCount_RNA <dbl> 1636, 1737, 7046, 70254, 24516, 5240, 17145,…
$ nFeature_RNA <int> 1102, 1233, 3498, 10129, 7050, 2832, 6144, 7…
$ age <chr> "adult", "adult", "adult", "adult", "adult",…
$ sex <chr> "male", "male", "male", "male", "male", "mal…
$ study_id <chr> "deng_2020", "deng_2020", "deng_2020", "deng…
$ tech <chr> "10xv3", "10xv3", "10xv3", "10xv3", "10xv3",…
$ hfd <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ ident <chr> "536-1_chow-diet", "536-1_chow-diet", "536-1…
$ sum <dbl> 1636, 1737, 7046, 70254, 24516, 5240, 17145,…
$ detected <int> 1102, 1233, 3498, 10129, 7050, 2832, 6144, 7…
$ subsets_mito_sum <dbl> 128, 108, 298, 3726, 1254, 169, 806, 998, 25…
$ subsets_mito_detected <int> 12, 11, 12, 13, 13, 12, 12, 12, 13, 12, 9, 1…
$ subsets_mito_percent <dbl> 7.823961, 6.217617, 4.229350, 5.303613, 5.11…
$ total <dbl> 1636, 1737, 7046, 70254, 24516, 5240, 17145,…
$ low_lib_size <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ low_n_features <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ high_subsets_mito_percent <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ discard <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ prob_compromised <dbl> 0.24285756, 0.11109518, 0.11795886, 0.309056…
536-1_chow-diet 536-3_chow-diet 536-5_chow-diet
4383 3878 1038 Rows: 15,041
Columns: 20
$ orig.ident <chr> "536-2_537-4_high-fat-diet", "536-2_537-4_hi…
$ nCount_RNA <dbl> 2405, 3172, 21420, 4907, 5381, 4366, 4434, 6…
$ nFeature_RNA <int> 1565, 2100, 6324, 2767, 3058, 2644, 2560, 34…
$ age <chr> "adult", "adult", "adult", "adult", "adult",…
$ sex <chr> "male", "male", "male", "male", "male", "mal…
$ study_id <chr> "deng_2020", "deng_2020", "deng_2020", "deng…
$ tech <chr> "10xv3", "10xv3", "10xv3", "10xv3", "10xv3",…
$ hfd <lgl> TRUE, TRUE, TRUE, TRUE, TRUE, TRUE, TRUE, TR…
$ ident <chr> "536-2_537-4_high-fat-diet", "536-2_537-4_hi…
$ sum <dbl> 2405, 3172, 21420, 4907, 5381, 4366, 4434, 6…
$ detected <int> 1565, 2100, 6324, 2767, 3058, 2644, 2560, 34…
$ subsets_mito_sum <dbl> 137, 140, 1352, 300, 213, 259, 279, 195, 634…
$ subsets_mito_detected <int> 11, 12, 13, 12, 12, 13, 11, 11, 12, 8, 11, 1…
$ subsets_mito_percent <dbl> 5.696466, 4.413619, 6.311858, 6.113715, 3.95…
$ total <dbl> 2405, 3172, 21420, 4907, 5381, 4366, 4434, 6…
$ low_lib_size <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ low_n_features <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ high_subsets_mito_percent <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ discard <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ prob_compromised <dbl> 0.08177799, 0.06766935, 0.21001466, 0.309050…
536-2_537-4_high-fat-diet 537-1_537-3_high-fat-diet 537-5_538-2_high-fat-diet
6751 2389 5901 Perform normalization and dimensionality reduction
Chow-diet
Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 9299
Number of edges: 380090
Running Louvain algorithm...
Maximum modularity in 10 random starts: 0.8927
Number of communities: 21
Elapsed time: 0 seconds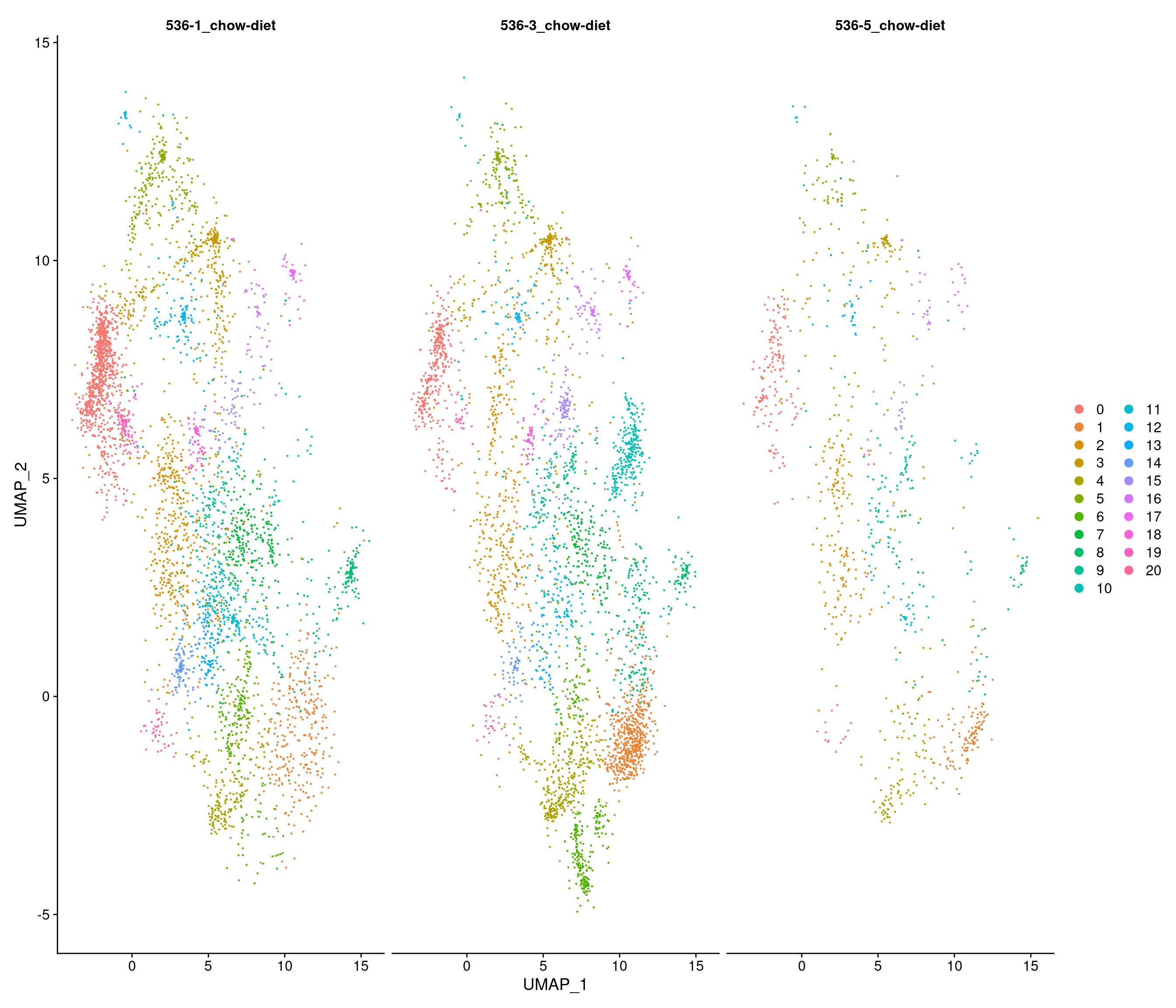
HFD
Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 15041
Number of edges: 603608
Running Louvain algorithm...
Maximum modularity in 10 random starts: 0.8914
Number of communities: 22
Elapsed time: 2 seconds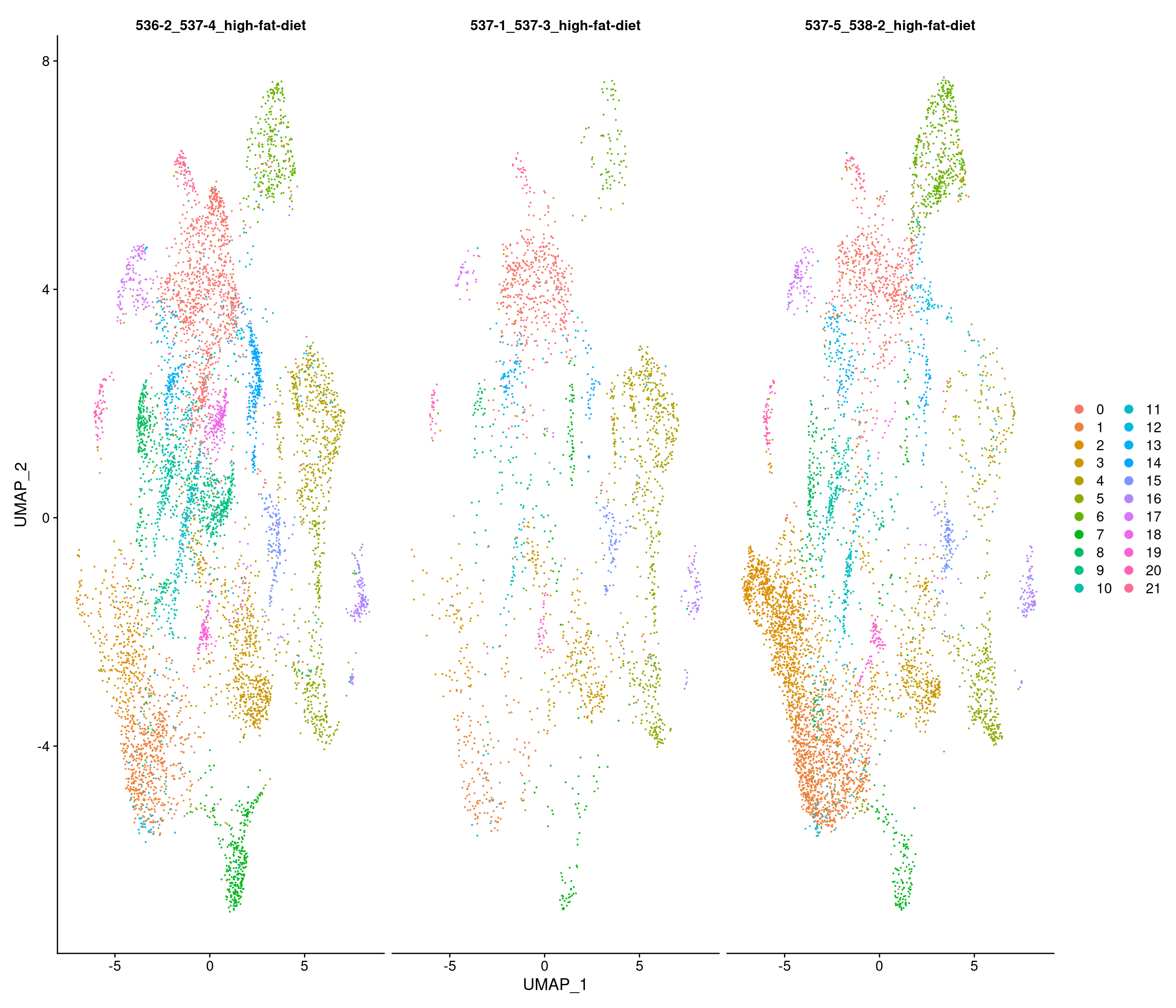
Identify Clusters
Chow-diet
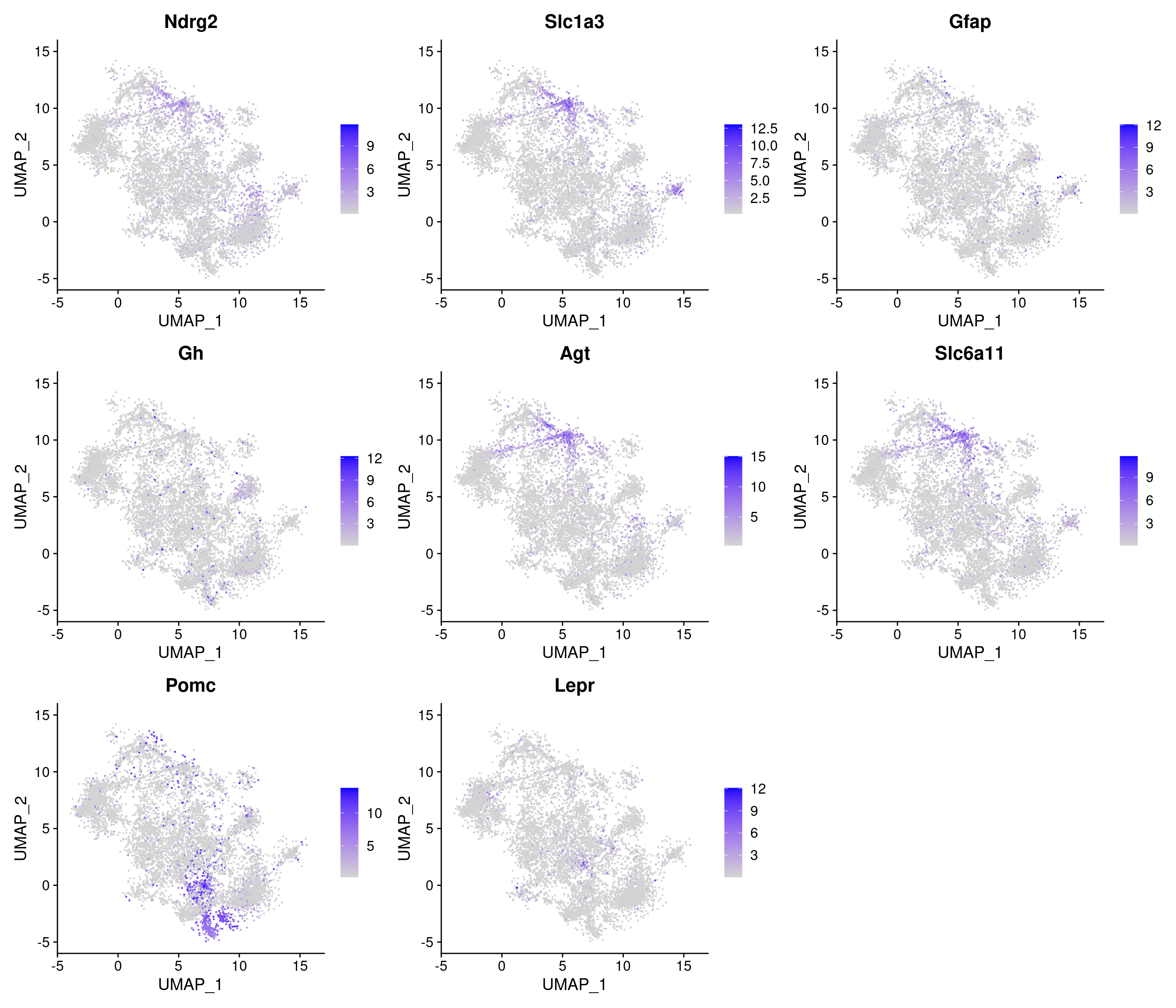
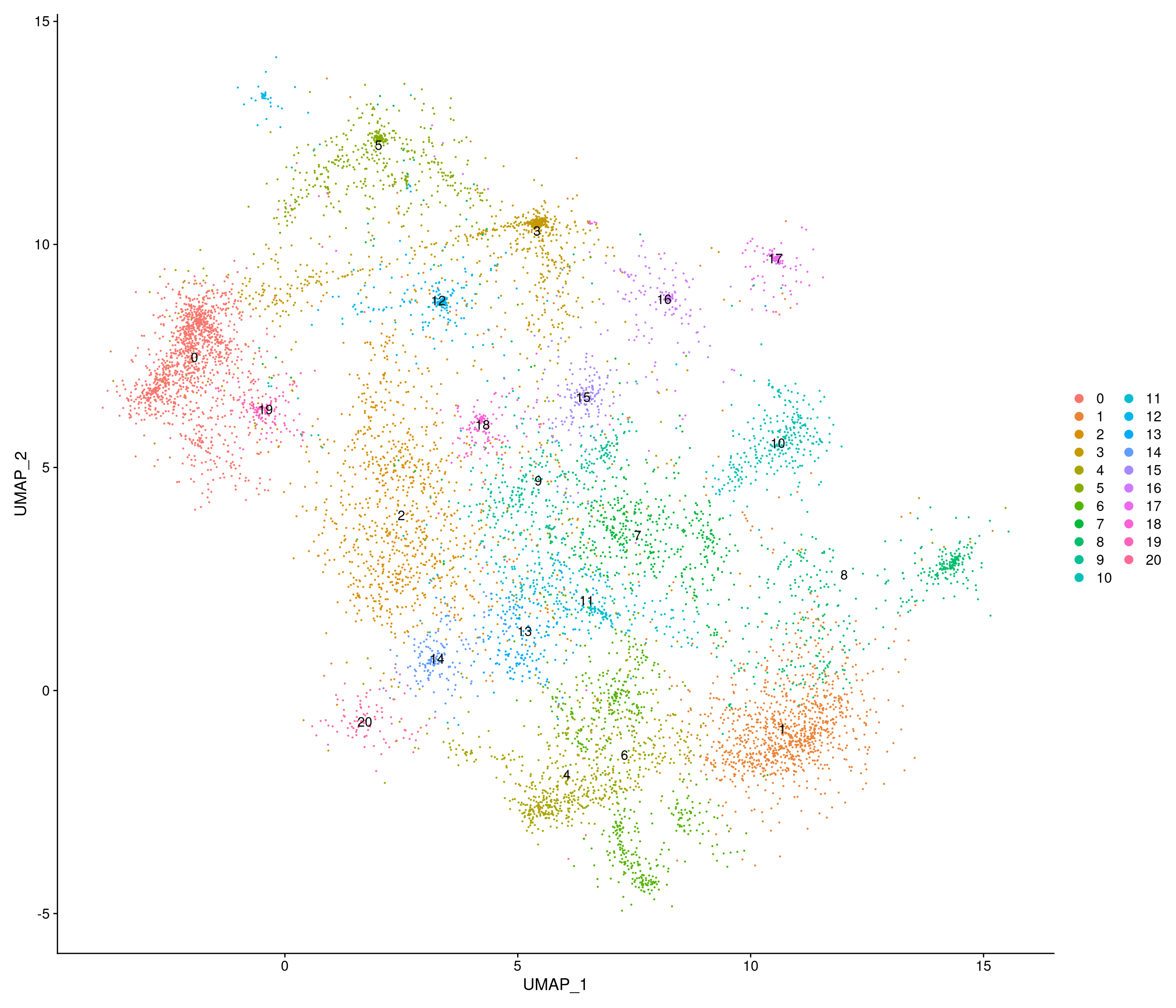
[1] "4602 variable genes to use"
Initial stress : 0.23396
stress after 10 iters: 0.09707, magic = 0.092
stress after 20 iters: 0.06484, magic = 0.500
stress after 30 iters: 0.06054, magic = 0.500
stress after 40 iters: 0.06004, magic = 0.500
stress after 50 iters: 0.05999, magic = 0.500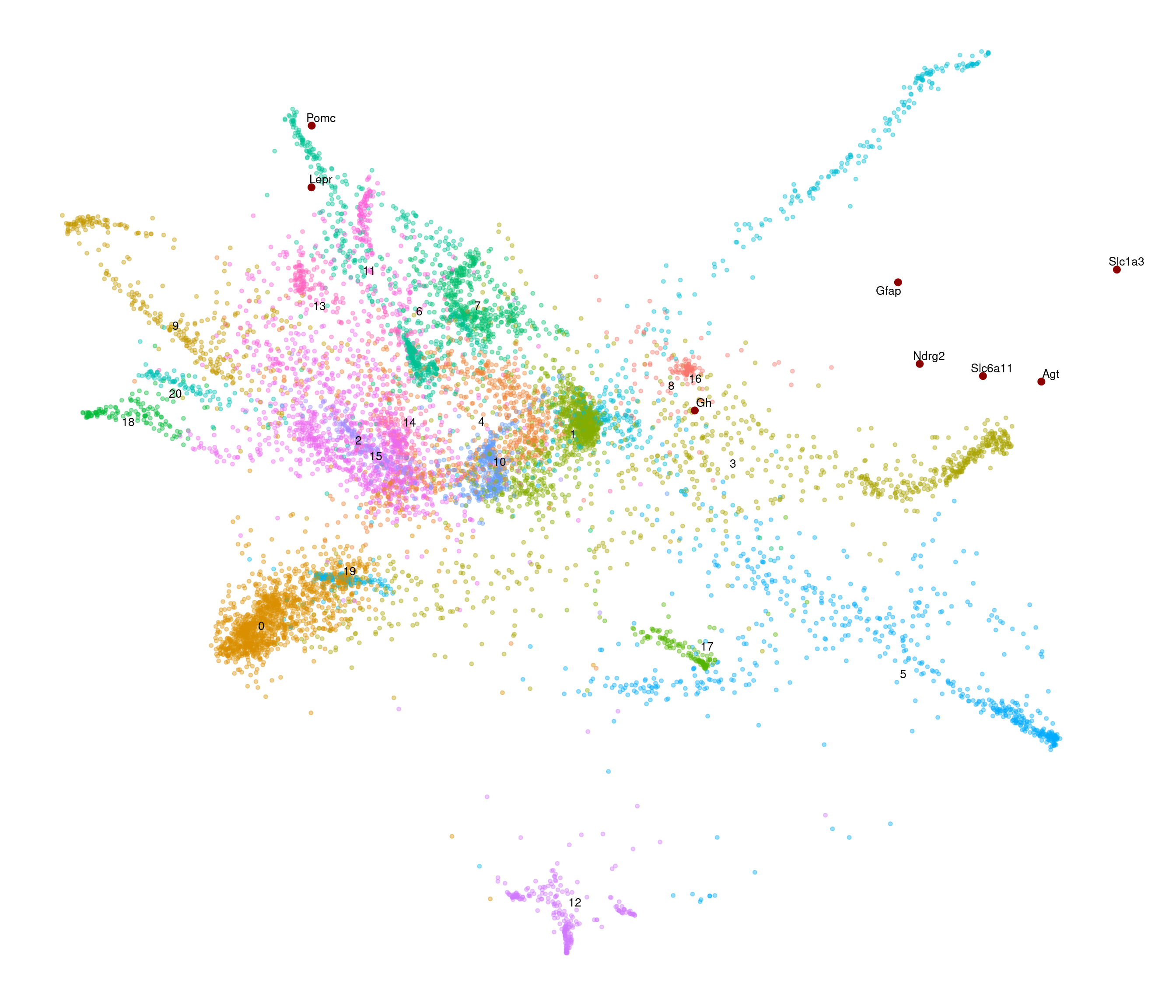
used (Mb) gc trigger (Mb) max used (Mb)
Ncells 9553530 510.3 13872301 740.9 13872301 740.9
Vcells 1757632913 13409.7 3889224212 29672.5 3880321403 29604.6 used (Mb) gc trigger (Mb) max used (Mb)
Ncells 9550562 510.1 13872301 740.9 13872301 740.9
Vcells 1787823922 13640.1 3889224212 29672.5 3880321403 29604.6HFD
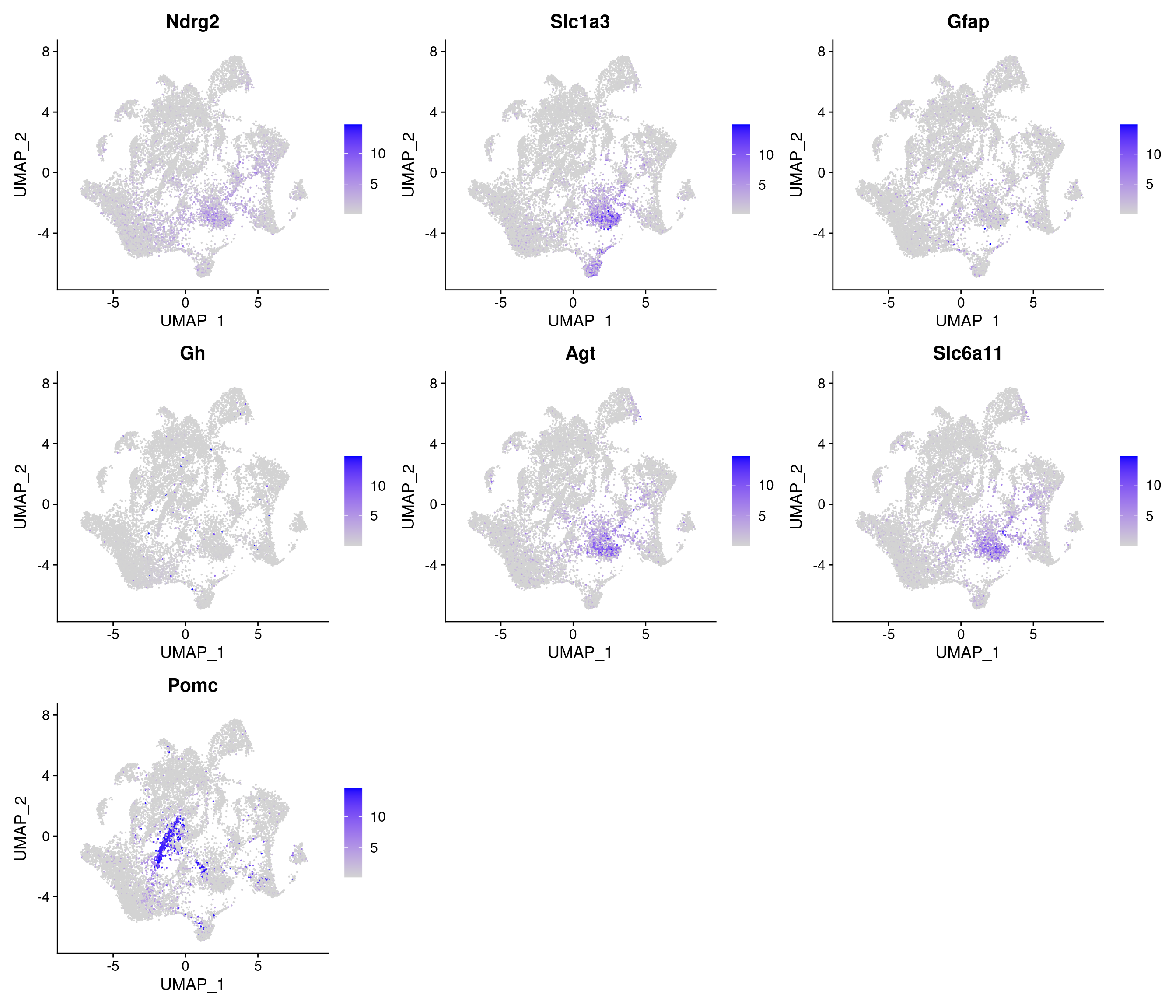
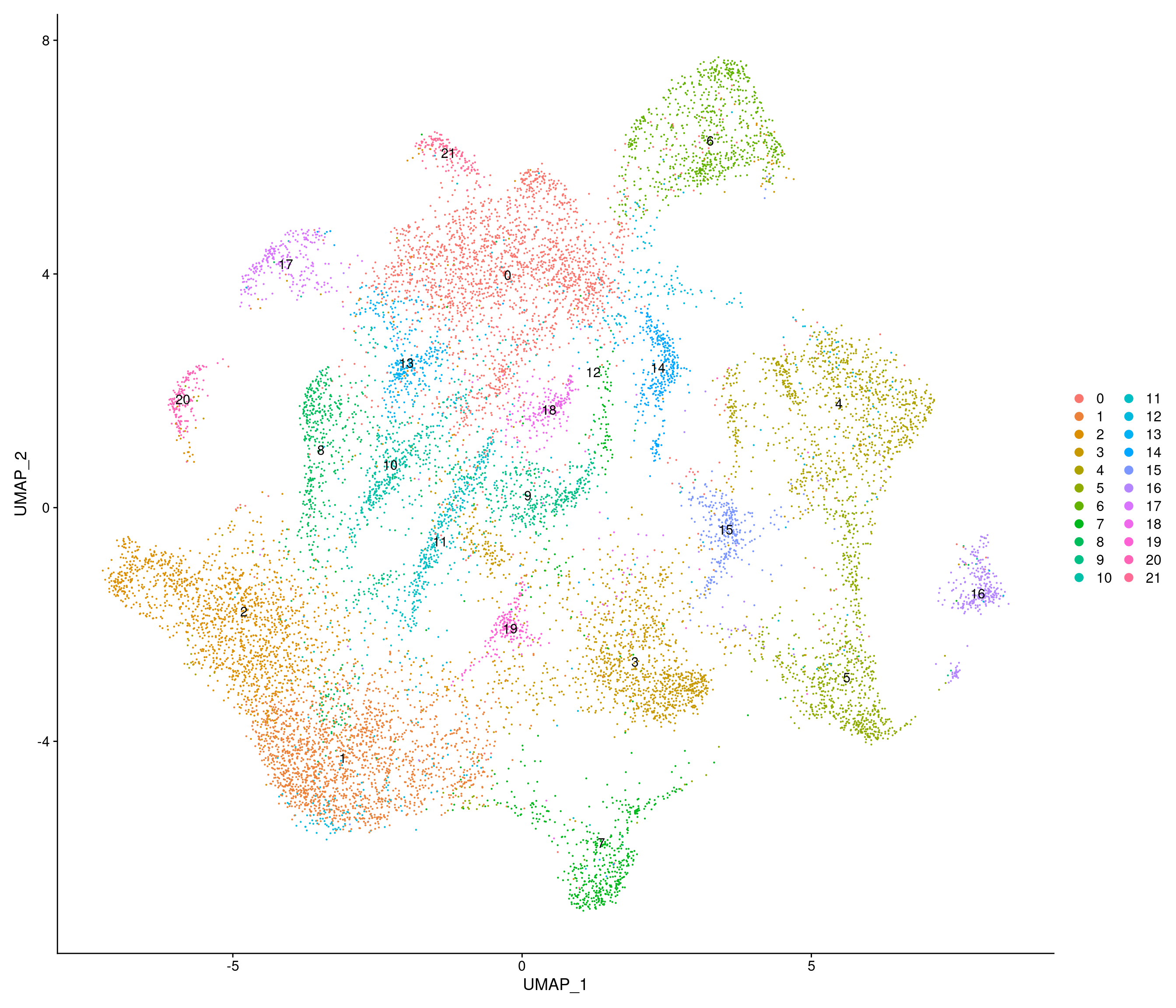
[1] "4783 variable genes to use"
Initial stress : 0.27148
stress after 10 iters: 0.07338, magic = 0.461
stress after 20 iters: 0.06400, magic = 0.500
stress after 30 iters: 0.06383, magic = 0.500
stress after 40 iters: 0.06378, magic = 0.500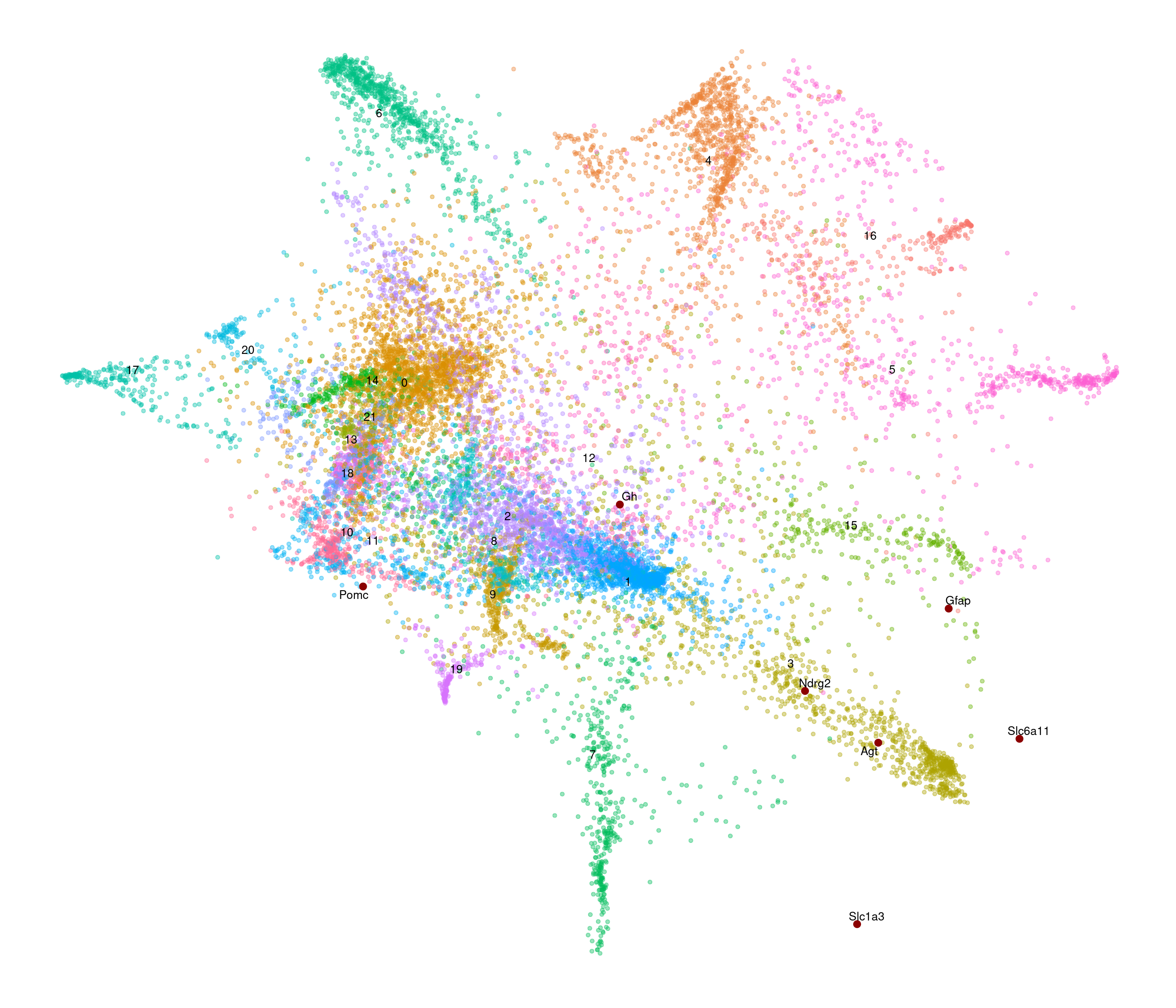
used (Mb) gc trigger (Mb) max used (Mb)
Ncells 9555390 510.4 13872301 740.9 13872301 740.9
Vcells 1788123593 13642.4 3889224212 29672.5 3880321403 29604.6 used (Mb) gc trigger (Mb) max used (Mb)
Ncells 9551548 510.2 13872301 740.9 13872301 740.9
Vcells 1834251550 13994.3 3889224212 29672.5 3880321403 29604.6Save initial clusters and subset astrocytes
Rows: 9,299
Columns: 24
$ orig.ident <chr> "536-1_chow-diet", "536-1_chow-diet", "536-1…
$ nCount_RNA <dbl> 1636, 1737, 7046, 70254, 24516, 5240, 17145,…
$ nFeature_RNA <int> 1102, 1233, 3498, 10129, 7050, 2832, 6144, 7…
$ age <chr> "adult", "adult", "adult", "adult", "adult",…
$ sex <chr> "male", "male", "male", "male", "male", "mal…
$ study_id <chr> "deng_2020", "deng_2020", "deng_2020", "deng…
$ tech <chr> "10xv3", "10xv3", "10xv3", "10xv3", "10xv3",…
$ hfd <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ ident <chr> "536-1_chow-diet", "536-1_chow-diet", "536-1…
$ sum <dbl> 1636, 1737, 7046, 70254, 24516, 5240, 17145,…
$ detected <int> 1102, 1233, 3498, 10129, 7050, 2832, 6144, 7…
$ subsets_mito_sum <dbl> 128, 108, 298, 3726, 1254, 169, 806, 998, 25…
$ subsets_mito_detected <int> 12, 11, 12, 13, 13, 12, 12, 12, 13, 12, 9, 1…
$ subsets_mito_percent <dbl> 7.823961, 6.217617, 4.229350, 5.303613, 5.11…
$ total <dbl> 1636, 1737, 7046, 70254, 24516, 5240, 17145,…
$ low_lib_size <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ low_n_features <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ high_subsets_mito_percent <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ discard <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ prob_compromised <dbl> 0.24285756, 0.11109518, 0.11795886, 0.309056…
$ nCount_SCT <dbl> 4112, 4165, 6189, 5708, 5779, 5233, 5496, 59…
$ nFeature_SCT <int> 1531, 1553, 3476, 3178, 3302, 2822, 3129, 35…
$ integrated_snn_res.0.7 <fct> 5, 4, 0, 6, 5, 11, 11, 7, 0, 3, 5, 0, 1, 9, …
$ seurat_clusters <fct> 5, 4, 0, 6, 5, 11, 11, 7, 0, 3, 5, 0, 1, 9, …
0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15
1337 1106 975 826 662 600 577 457 430 322 321 292 233 192 170 167
16 17 18 19 20
144 136 132 131 89 Rows: 15,041
Columns: 24
$ orig.ident <chr> "536-2_537-4_high-fat-diet", "536-2_537-4_hi…
$ nCount_RNA <dbl> 2405, 3172, 21420, 4907, 5381, 4366, 4434, 6…
$ nFeature_RNA <int> 1565, 2100, 6324, 2767, 3058, 2644, 2560, 34…
$ age <chr> "adult", "adult", "adult", "adult", "adult",…
$ sex <chr> "male", "male", "male", "male", "male", "mal…
$ study_id <chr> "deng_2020", "deng_2020", "deng_2020", "deng…
$ tech <chr> "10xv3", "10xv3", "10xv3", "10xv3", "10xv3",…
$ hfd <lgl> TRUE, TRUE, TRUE, TRUE, TRUE, TRUE, TRUE, TR…
$ ident <chr> "536-2_537-4_high-fat-diet", "536-2_537-4_hi…
$ sum <dbl> 2405, 3172, 21420, 4907, 5381, 4366, 4434, 6…
$ detected <int> 1565, 2100, 6324, 2767, 3058, 2644, 2560, 34…
$ subsets_mito_sum <dbl> 137, 140, 1352, 300, 213, 259, 279, 195, 634…
$ subsets_mito_detected <int> 11, 12, 13, 12, 12, 13, 11, 11, 12, 8, 11, 1…
$ subsets_mito_percent <dbl> 5.696466, 4.413619, 6.311858, 6.113715, 3.95…
$ total <dbl> 2405, 3172, 21420, 4907, 5381, 4366, 4434, 6…
$ low_lib_size <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ low_n_features <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ high_subsets_mito_percent <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ discard <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FA…
$ prob_compromised <dbl> 0.08177799, 0.06766935, 0.21001466, 0.309050…
$ nCount_SCT <dbl> 3101, 3351, 4517, 4506, 4715, 4282, 4295, 47…
$ nFeature_SCT <int> 1566, 2094, 2695, 2744, 3035, 2635, 2542, 33…
$ integrated_snn_res.0.7 <fct> 14, 10, 4, 0, 0, 0, 9, 14, 5, 17, 10, 20, 1,…
$ seurat_clusters <fct> 14, 10, 4, 0, 0, 0, 9, 14, 5, 17, 10, 20, 1,…
0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15
2204 1843 1749 1317 1069 838 799 626 531 485 463 426 379 351 346 320
16 17 18 19 20 21
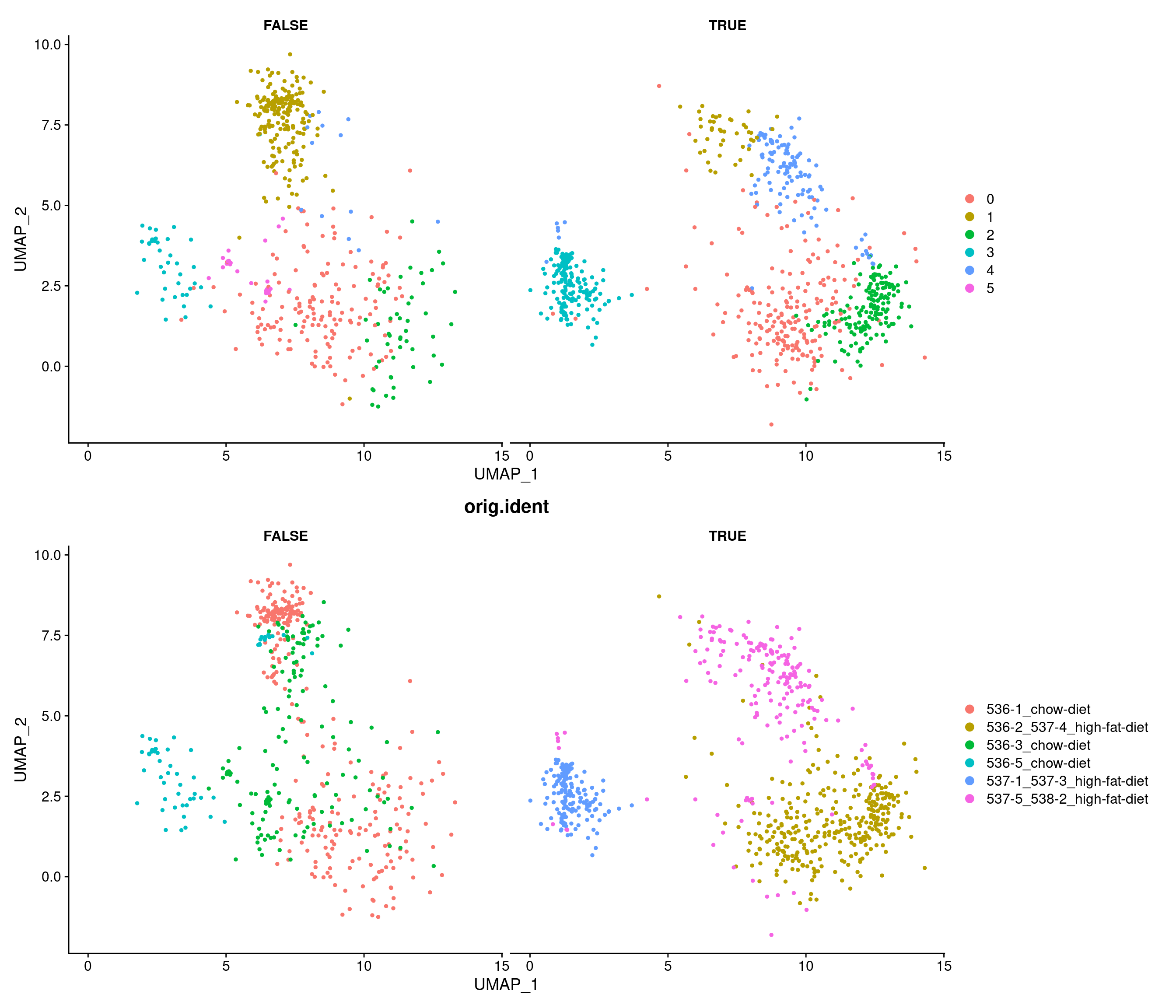
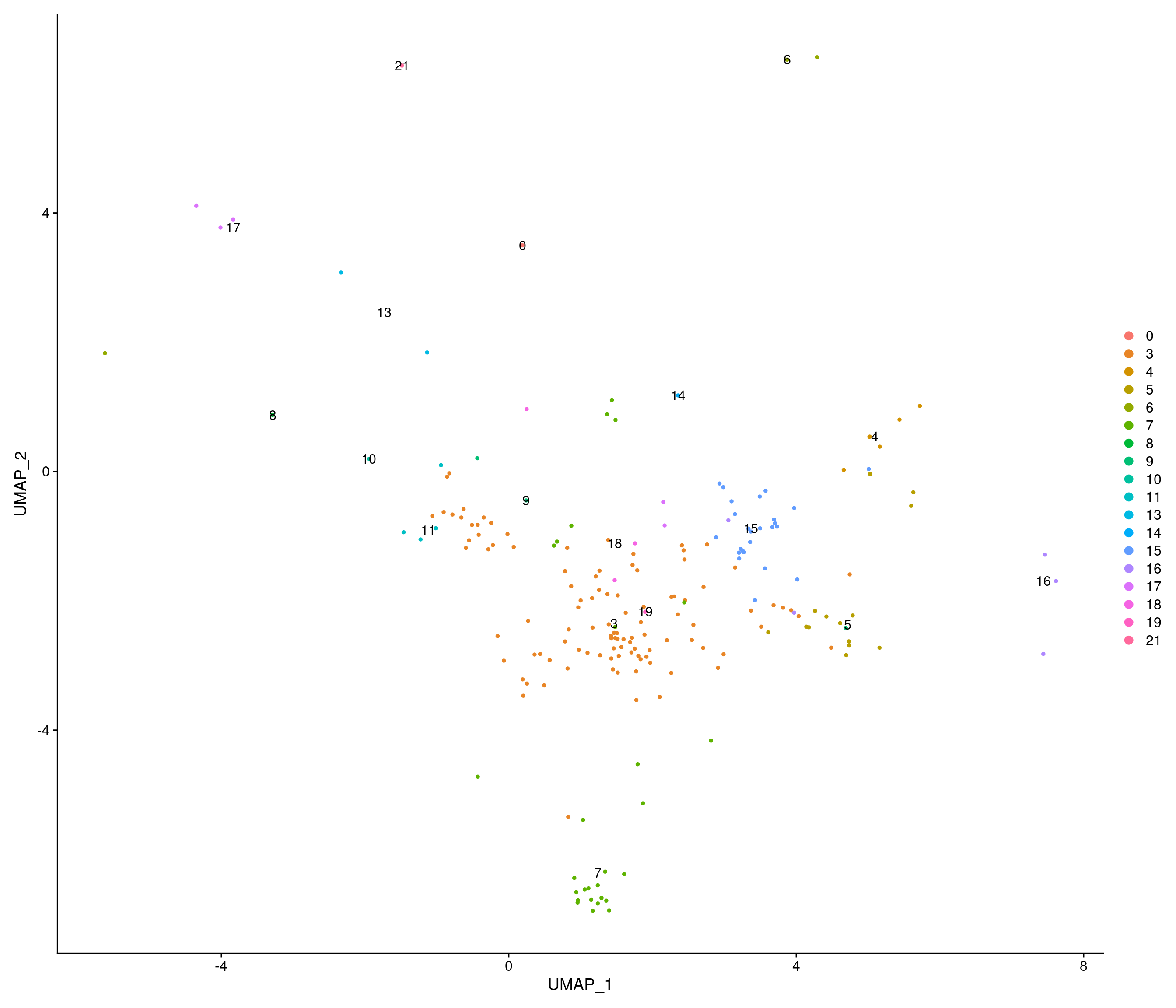
301 256 230 205 153 150 Test astrocytes difference
Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 2143
Number of edges: 134788
Running Louvain algorithm...
Maximum modularity in 10 random starts: 0.7628
Number of communities: 4
Elapsed time: 0 seconds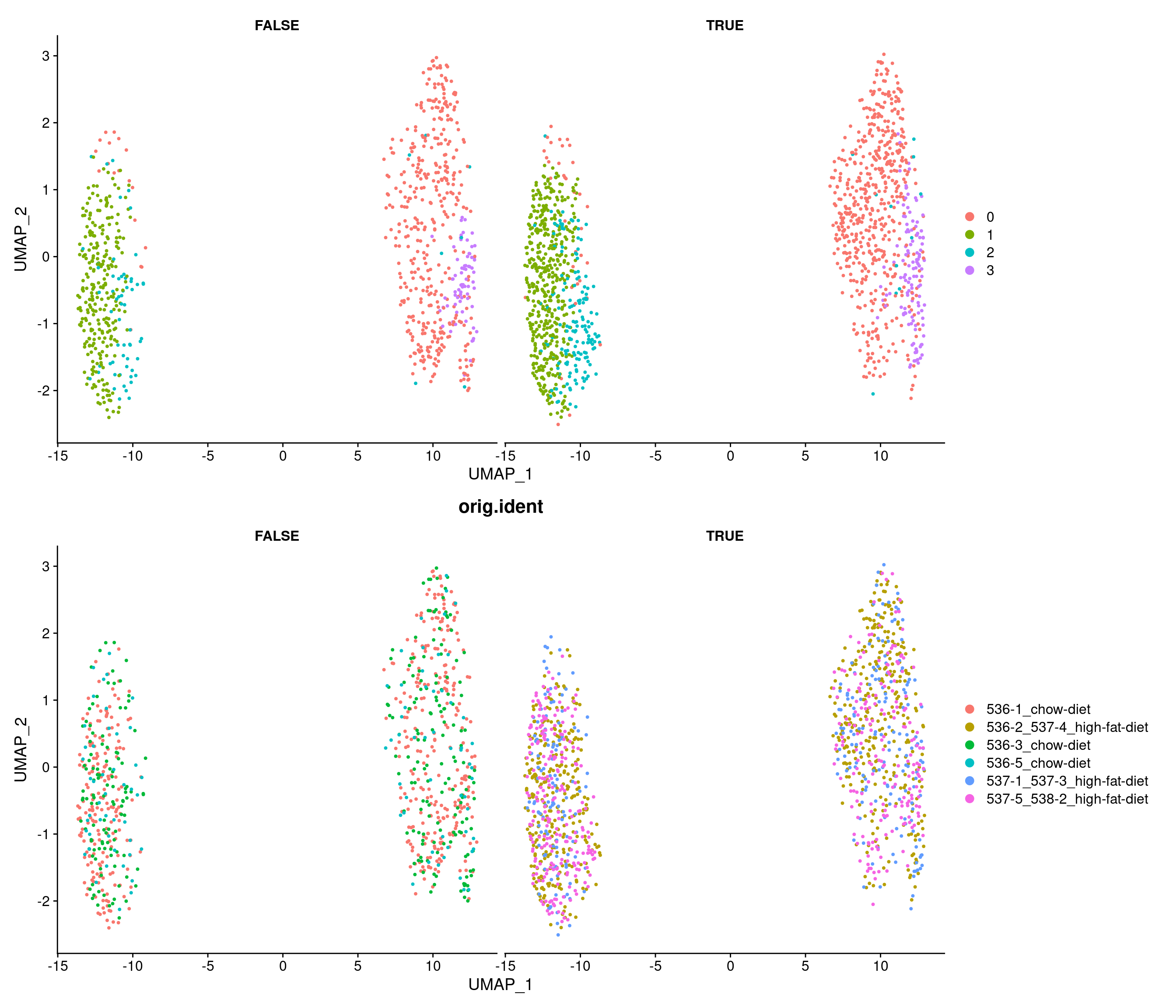
# A tibble: 7 × 7
# Groups: cluster [4]
p_val avg_log2FC pct.1 pct.2 p_val_adj cluster gene
<dbl> <dbl> <dbl> <dbl> <dbl> <fct> <chr>
1 1.62e-218 0.502 0.996 0.978 2.54e-214 0 Snhg11
2 2.42e-164 0.494 0.999 0.988 3.81e-160 0 Meg3
3 1.82e-225 0.788 1 0.634 2.86e-221 1 Gm3764
4 3.39e-193 0.700 0.994 0.618 5.33e-189 1 Ntsr2
5 5.19e- 12 0.262 0.838 0.646 8.16e- 8 2 Mbp
6 6.21e- 62 0.802 0.898 0.529 9.76e- 58 3 Plp1
7 8.83e- 91 0.791 0.801 0.196 1.39e- 86 3 Ptgds # A tibble: 16 × 7
# Groups: cluster [8]
p_val avg_log2FC pct.1 pct.2 p_val_adj cluster gene
<dbl> <dbl> <dbl> <dbl> <dbl> <fct> <chr>
1 5.99e- 31 0.467 0.294 0.082 9.42e- 27 0_0 Gh
2 9.19e- 16 0.328 0.504 0.289 1.45e- 11 0_0 Pomc
3 8.23e- 25 0.635 0.303 0.097 1.29e- 20 1_0 Gh
4 1.39e- 65 0.552 1 0.723 2.19e- 61 1_0 Gm3764
5 1.13e- 22 0.732 0.904 0.551 1.78e- 18 3_0 Plp1
6 2.24e- 24 0.700 0.877 0.406 3.52e- 20 3_0 Nrgn
7 8.02e- 12 0.636 0.388 0.114 1.26e- 7 2_0 Gh
8 7.15e- 7 0.536 0.582 0.323 1.13e- 2 2_0 Pomc
9 7.32e- 43 0.390 0.74 0.482 1.15e- 38 0_1 Gad2
10 2.15e-107 0.364 0.998 0.982 3.37e-103 0_1 Snhg11
11 4.68e-119 0.629 1 0.693 7.36e-115 1_1 Gm3764
12 6.75e-123 0.559 1 0.777 1.06e-118 1_1 Slc1a3
13 4.47e- 69 0.844 0.854 0.215 7.02e- 65 3_1 Ptgds
14 2.71e- 38 0.769 0.894 0.543 4.26e- 34 3_1 Plp1
15 2.73e- 9 0.262 0.844 0.652 4.29e- 5 2_1 Mbp
16 1.62e- 10 -0.254 0.365 0.598 2.55e- 6 2_1 Ewsr1 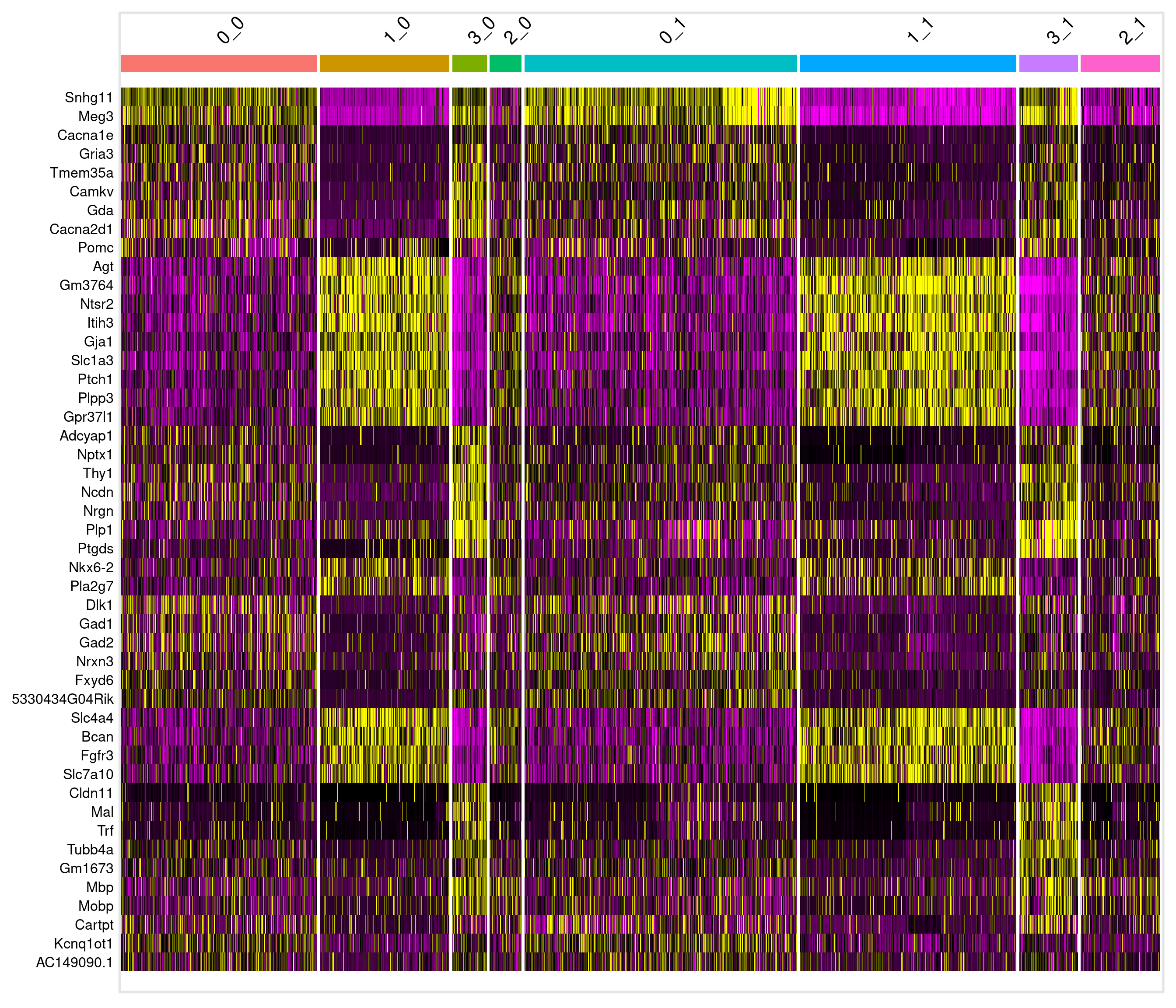
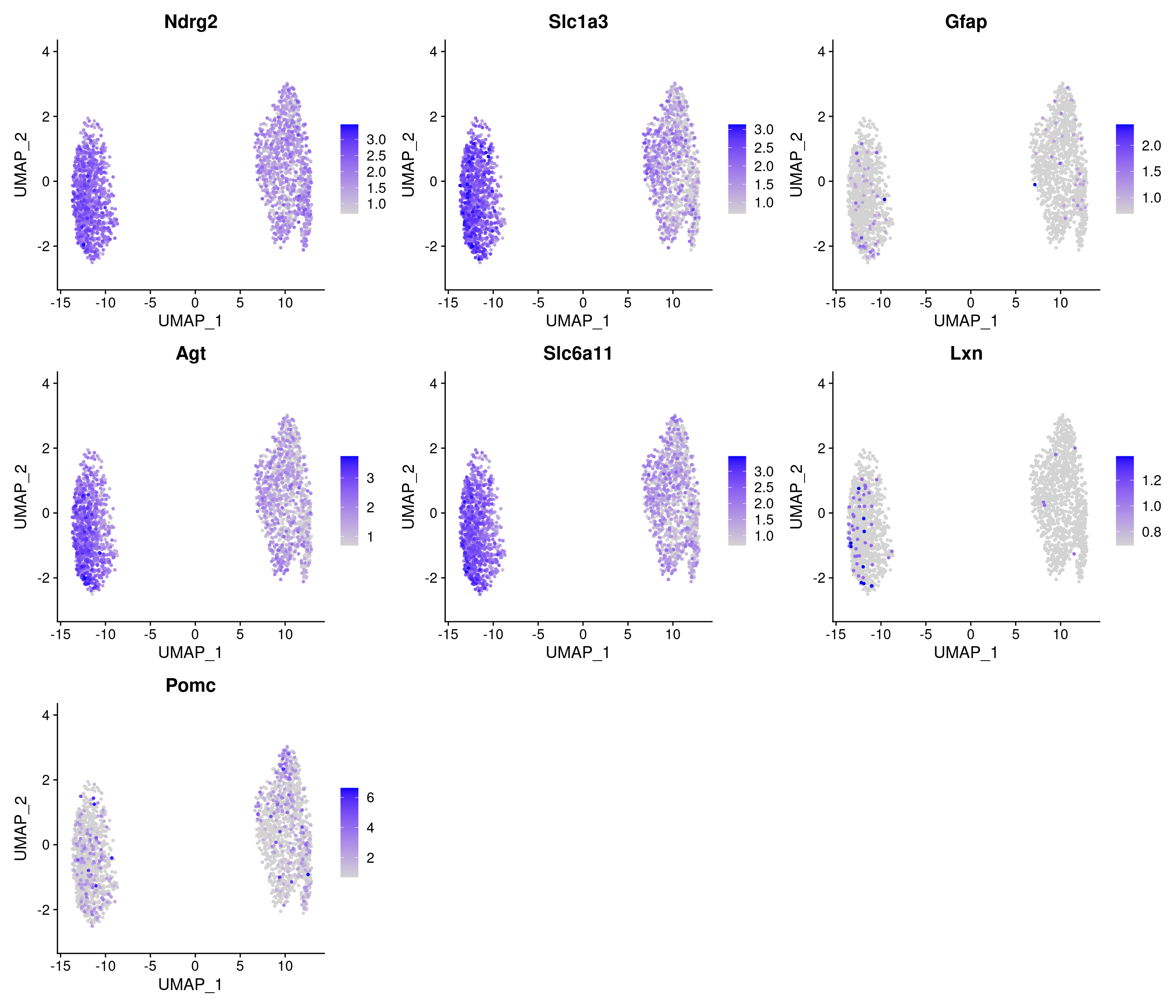
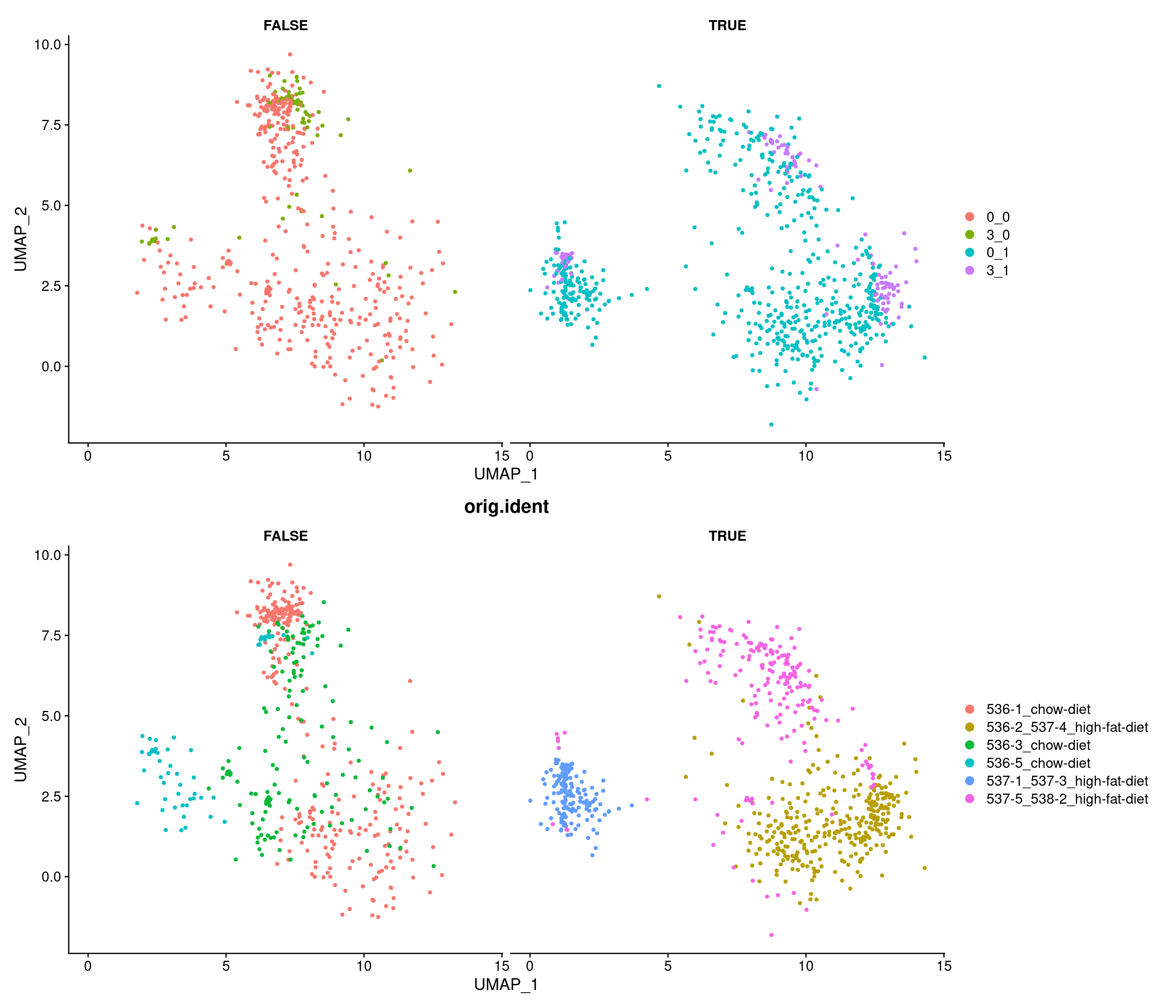
# A tibble: 8 × 7
# Groups: cluster [4]
p_val avg_log2FC pct.1 pct.2 p_val_adj cluster gene
<dbl> <dbl> <dbl> <dbl> <dbl> <fct> <chr>
1 2.64e-36 0.862 0.325 0.06 4.51e-32 0_0 Gh
2 3.40e-34 0.465 0.275 0.035 5.82e-30 0_0 Prl
3 1.45e-23 0.642 0.753 0.246 2.49e-19 3_0 Fezf1
4 1.21e-23 0.631 0.877 0.359 2.06e-19 3_0 Adcyap1
5 3.81e-12 0.285 0.243 0.1 6.52e- 8 0_1 Zic1
6 5.31e-23 0.282 0.955 0.843 9.09e-19 0_1 Gad2
7 1.20e-48 0.900 0.902 0.399 2.05e-44 3_1 Ptgds
8 5.04e-41 0.692 0.724 0.197 8.63e-37 3_1 Mal 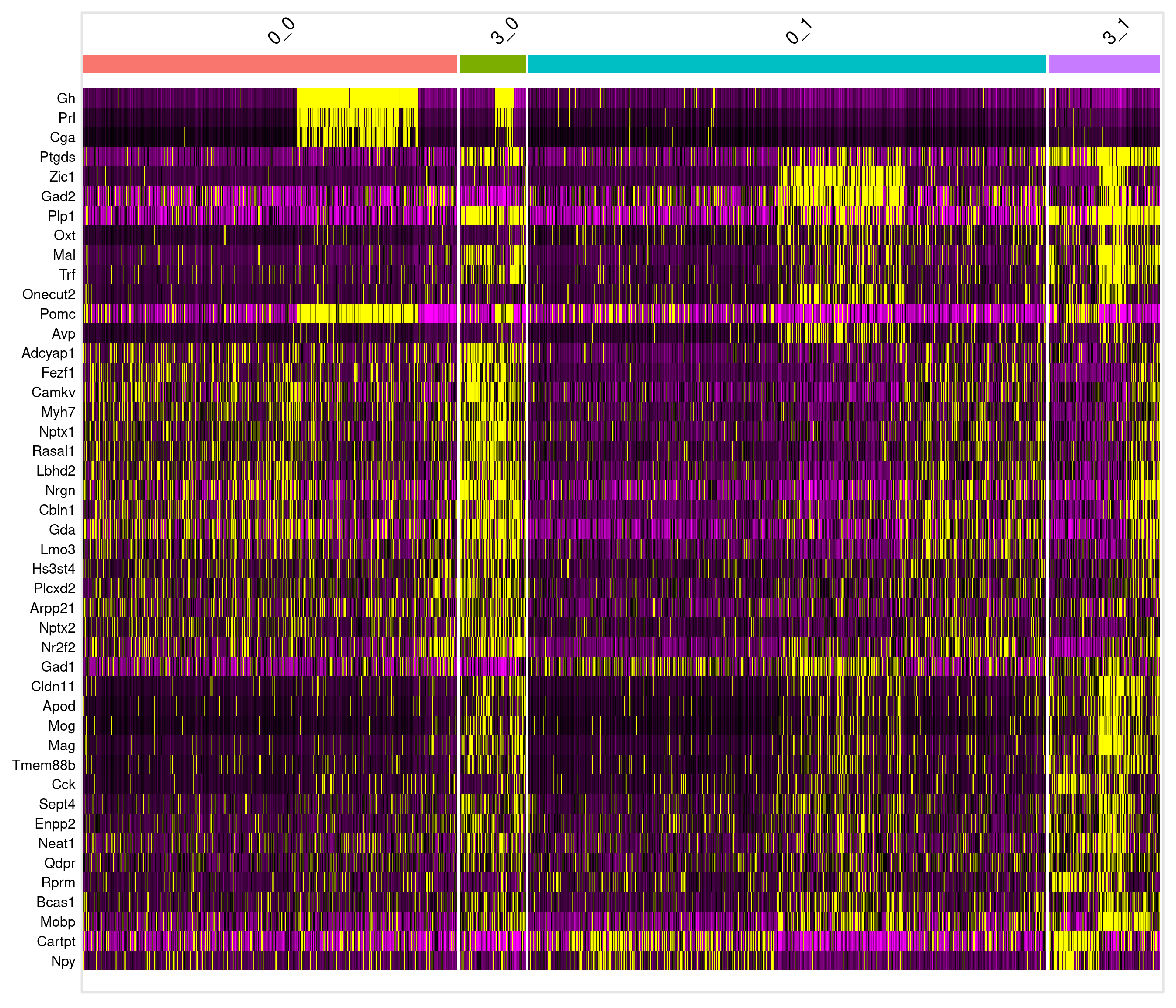
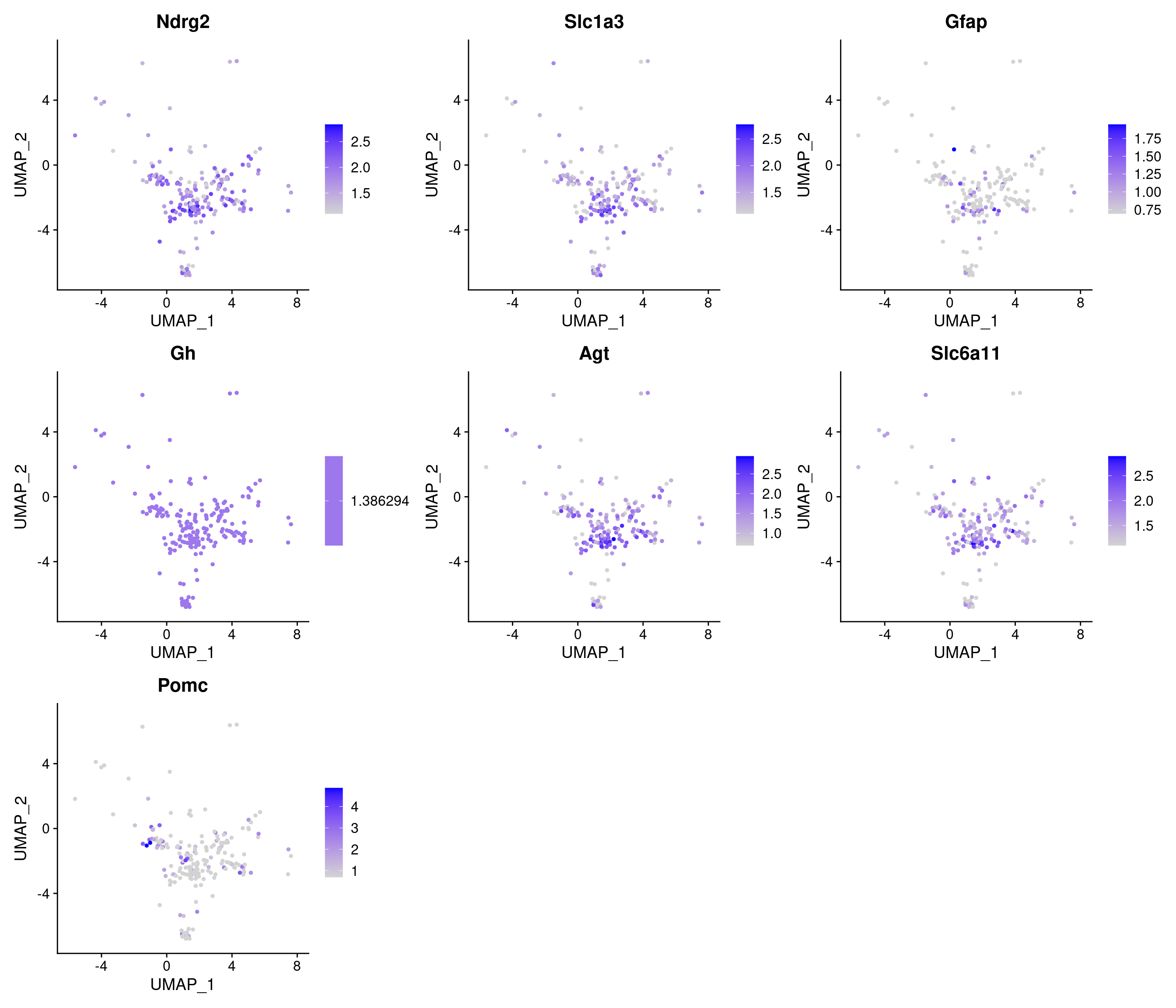
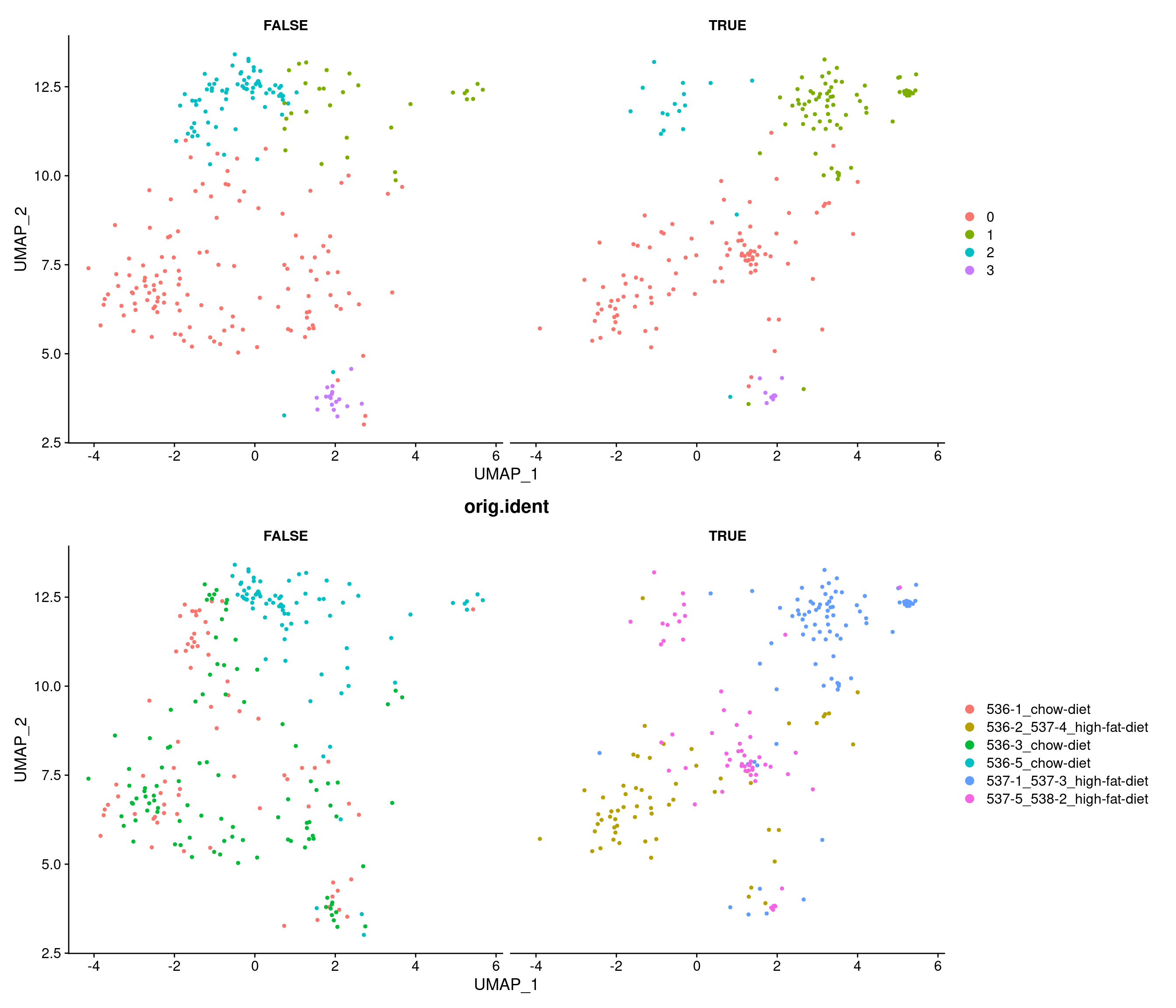
# A tibble: 8 × 7
# Groups: cluster [4]
p_val avg_log2FC pct.1 pct.2 p_val_adj cluster gene
<dbl> <dbl> <dbl> <dbl> <dbl> <fct> <chr>
1 1.67e-25 0.766 0.948 0.632 2.83e-21 0 Pomc
2 2.40e-16 0.728 0.561 0.177 4.06e-12 0 Gh
3 2.48e-19 1.22 0.616 0.246 4.20e-15 1 Avp
4 3.28e-44 1.13 0.821 0.206 5.56e-40 1 Zic1
5 4.80e-48 1.01 0.946 0.242 8.12e-44 2 Fezf1
6 3.65e-34 0.748 0.978 0.603 6.18e-30 2 Lbhd2
7 4.61e-28 1.48 1 0.209 7.79e-24 3 Spef2
8 8.96e-30 1.42 1 0.191 1.51e-25 3 Cfap100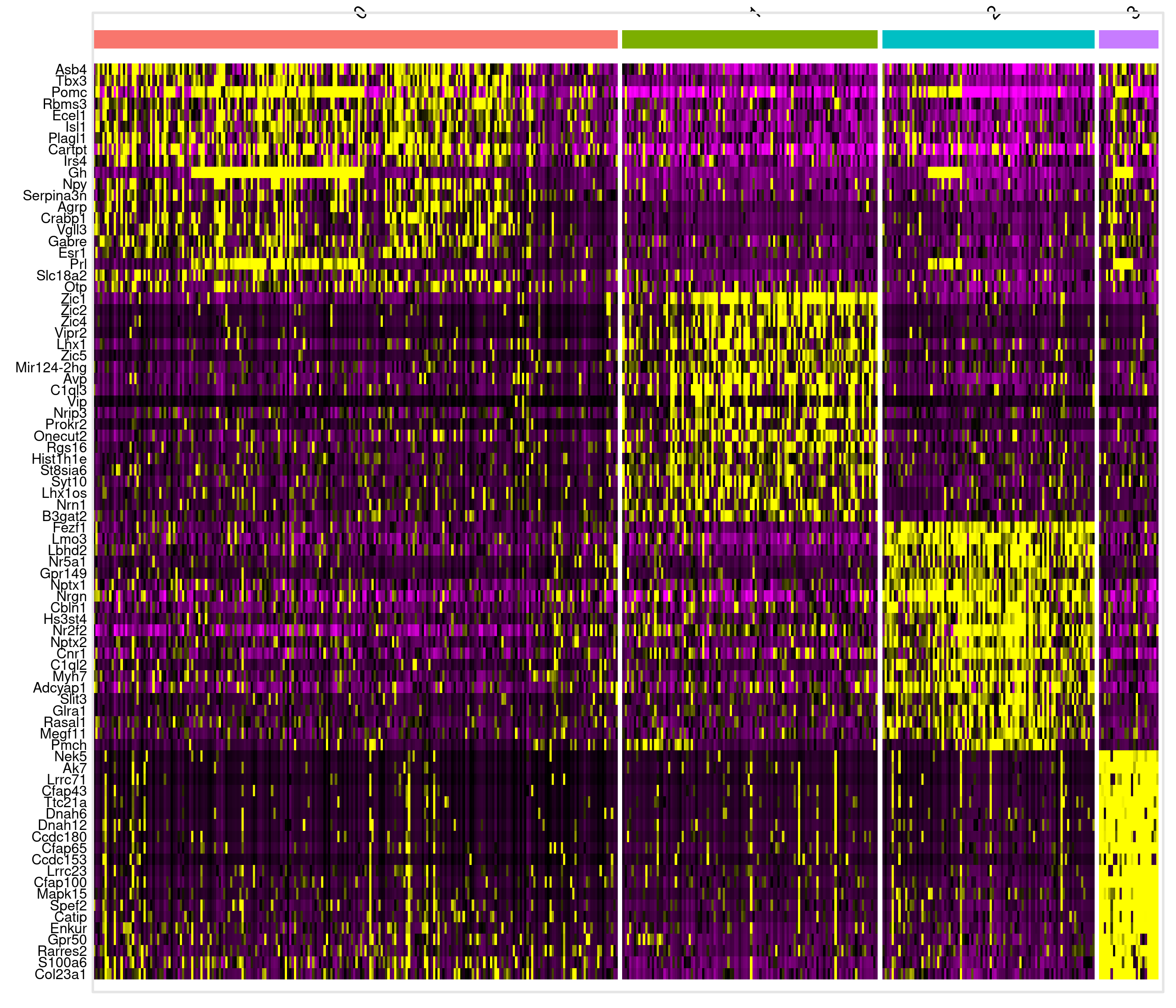
# A tibble: 17 × 8
# Groups: cluster [8]
myAUC avg_diff power avg_log2FC pct.1 pct.2 cluster gene
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <fct> <chr>
1 0.788 2.02 0.576 0.802 0.953 0.728 0_0 Pomc
2 0.775 2.68 0.55 1.24 0.701 0.243 0_0 Gh
3 0.928 0.967 0.856 0.412 1 0.99 2_0 Cacna2d1
4 0.913 0.796 0.826 0.352 1 0.995 2_0 Ncdn
5 0.98 1.96 0.96 1.36 1 0.207 3_0 Cfap100
6 0.978 1.61 0.956 1.26 1 0.095 3_0 Ak7
7 0.978 1.78 0.956 1.33 1 0.171 3_0 Cfap65
8 0.818 0.568 0.636 0.285 1 0.979 1_0 mt-Atp8
9 0.8 0.496 0.6 0.275 1 0.984 1_0 Pcsk2
10 0.759 0.617 0.518 0.369 0.961 0.872 0_1 Fgfr3
11 0.754 0.594 0.508 0.279 0.99 0.98 0_1 Ntsr2
12 0.901 0.970 0.802 0.857 1 0.36 2_1 Fezf1
13 0.889 0.624 0.778 0.293 1 0.995 2_1 Rasgrf2
14 0.988 2.21 0.976 1.54 1 0.239 3_1 Spef2
15 0.988 1.67 0.976 1.27 1 0.166 3_1 Crb2
16 0.936 1.71 0.872 1.23 0.938 0.232 1_1 Zic1
17 0.915 0.852 0.83 0.400 1 0.984 1_1 Zfhx3 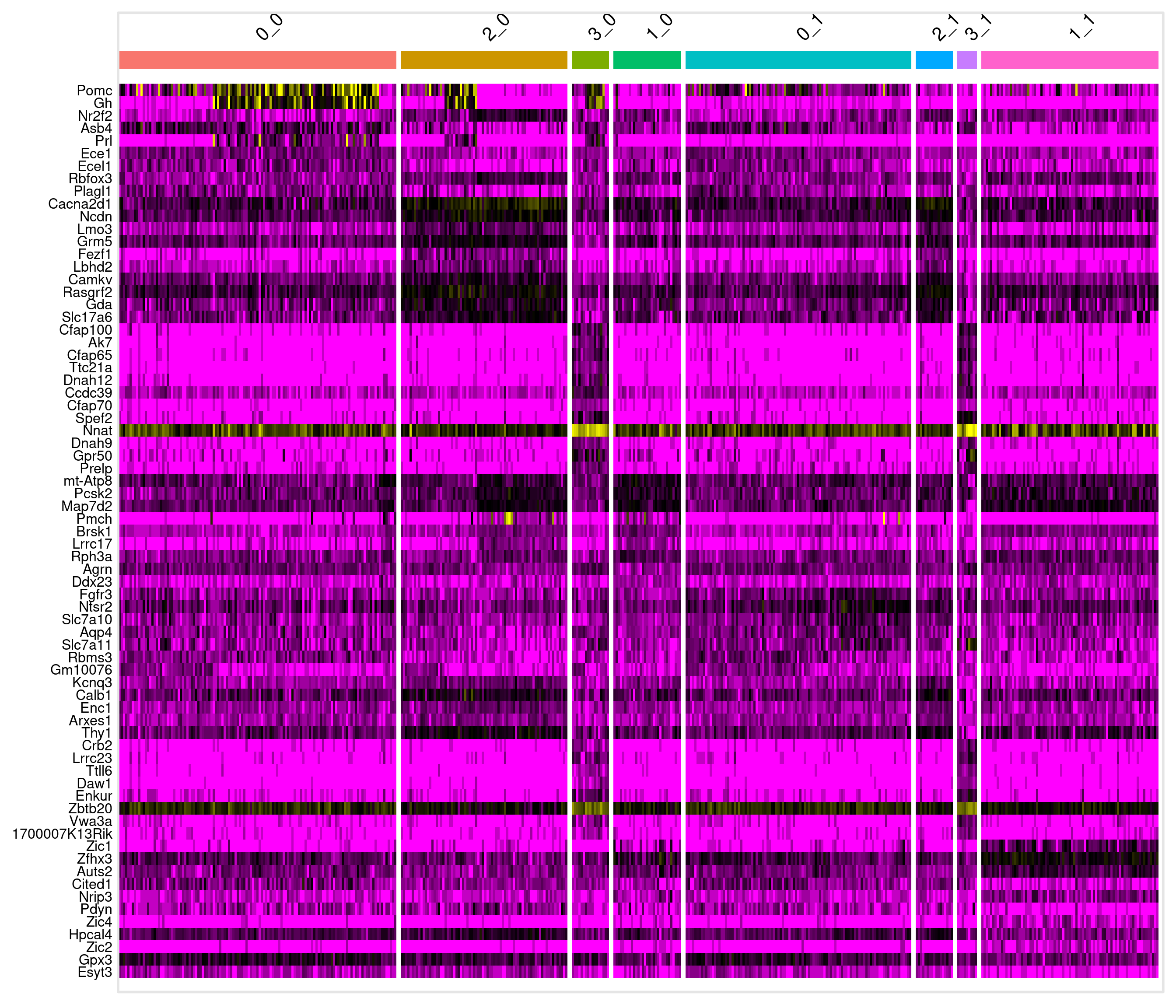
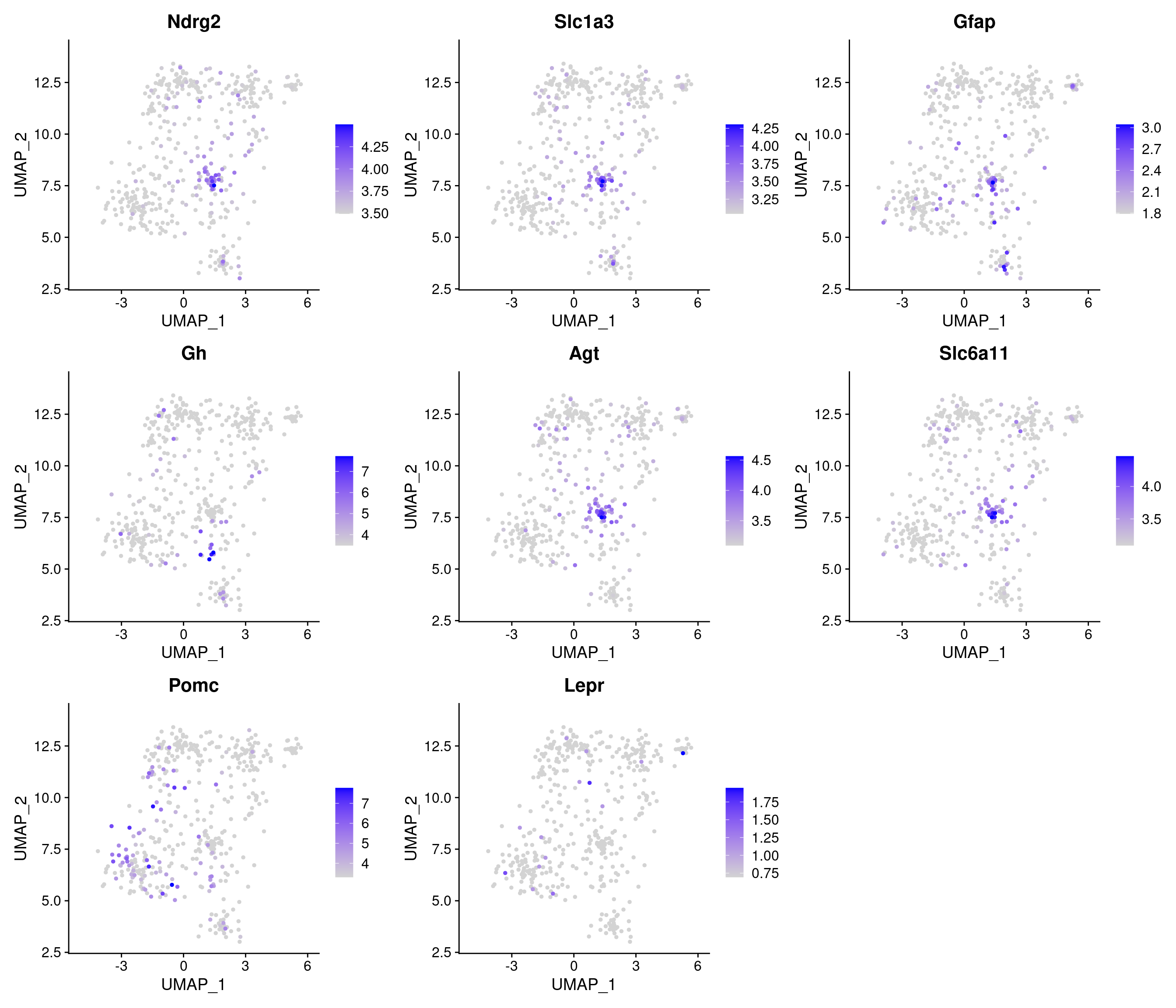
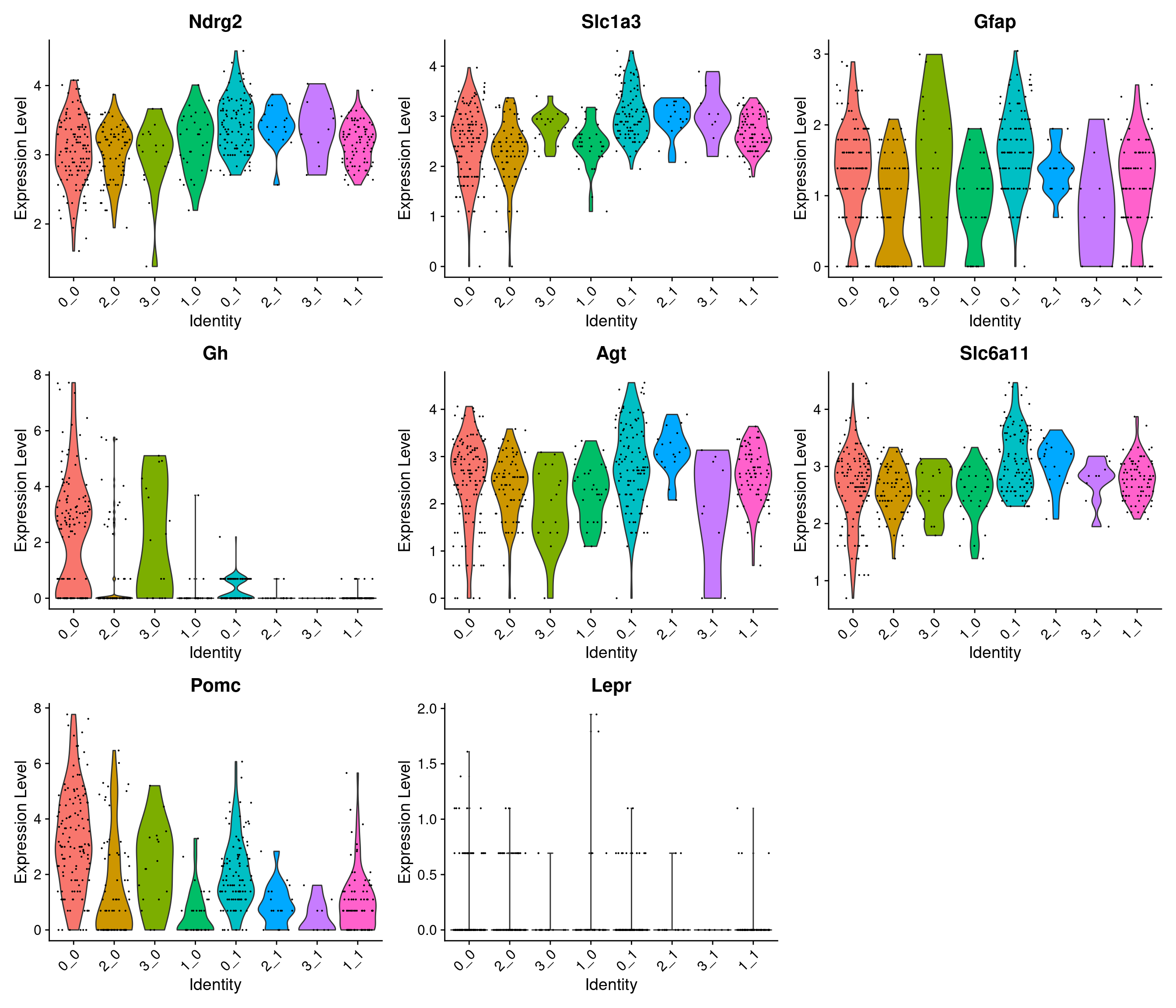
SWNE
calculating variance fit ... using gam [1] "3000 variable genes to use"
Initial stress : 0.14646
stress after 10 iters: 0.03191, magic = 0.500
stress after 20 iters: 0.02948, magic = 0.500
stress after 30 iters: 0.02881, magic = 0.500
stress after 40 iters: 0.02867, magic = 0.500
stress after 50 iters: 0.02863, magic = 0.500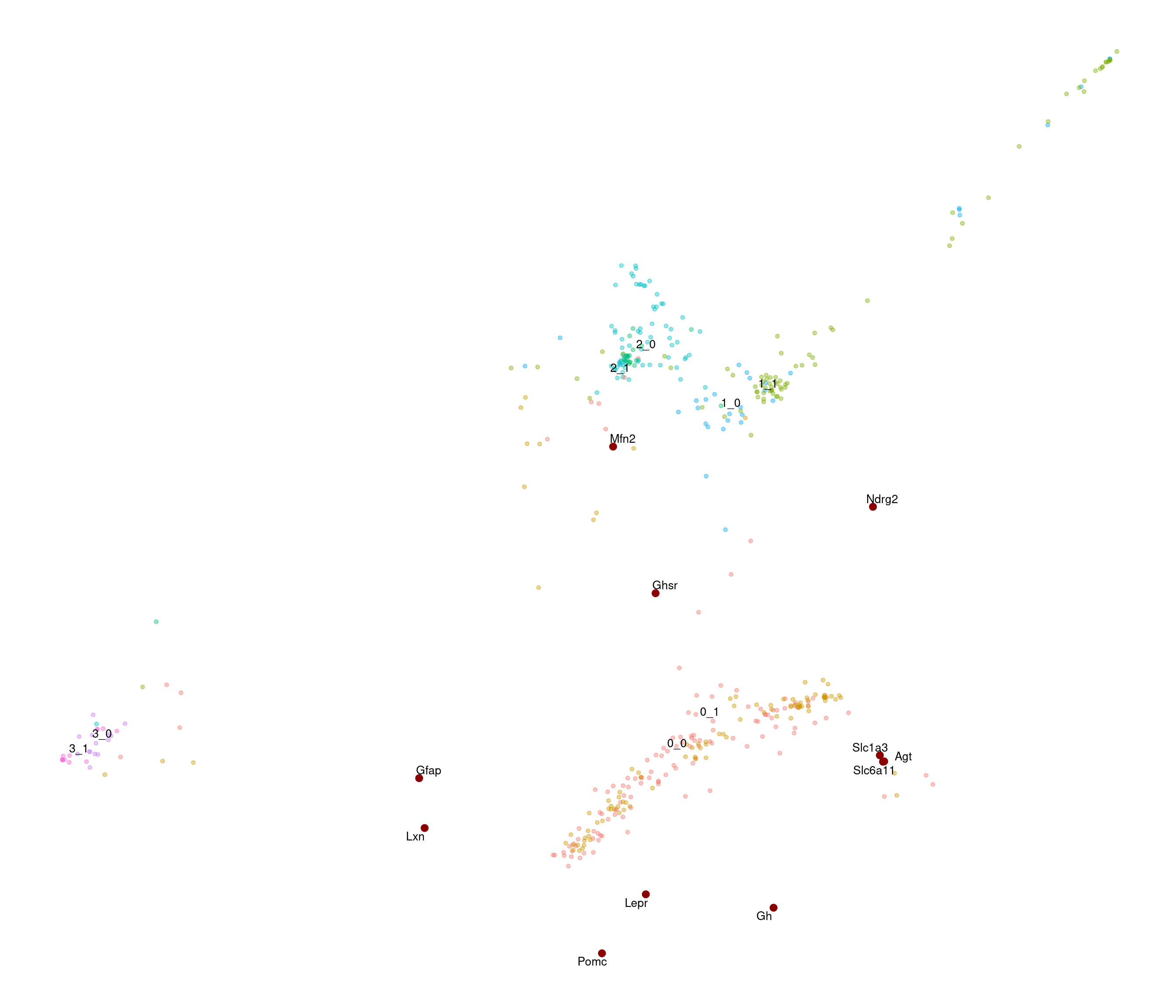
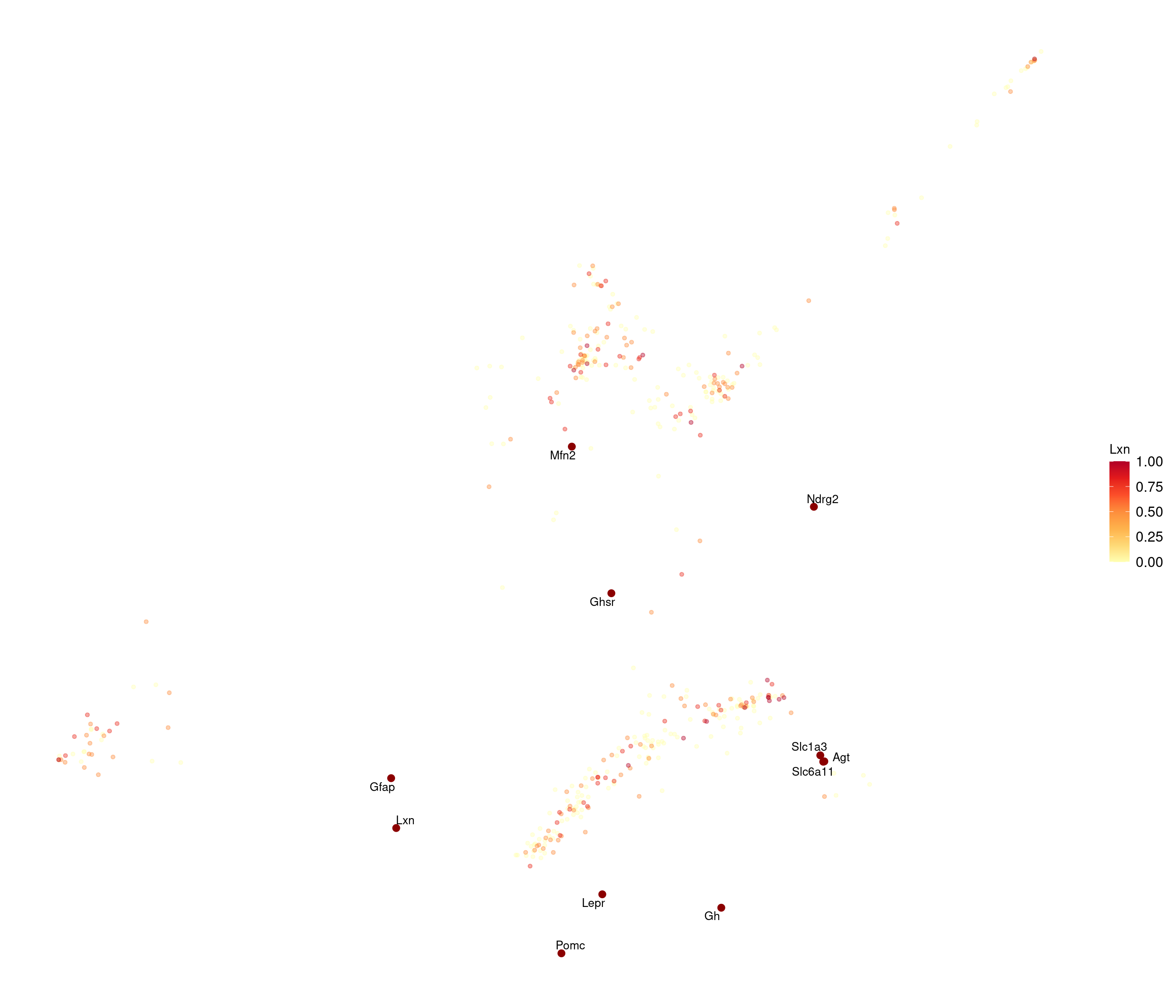
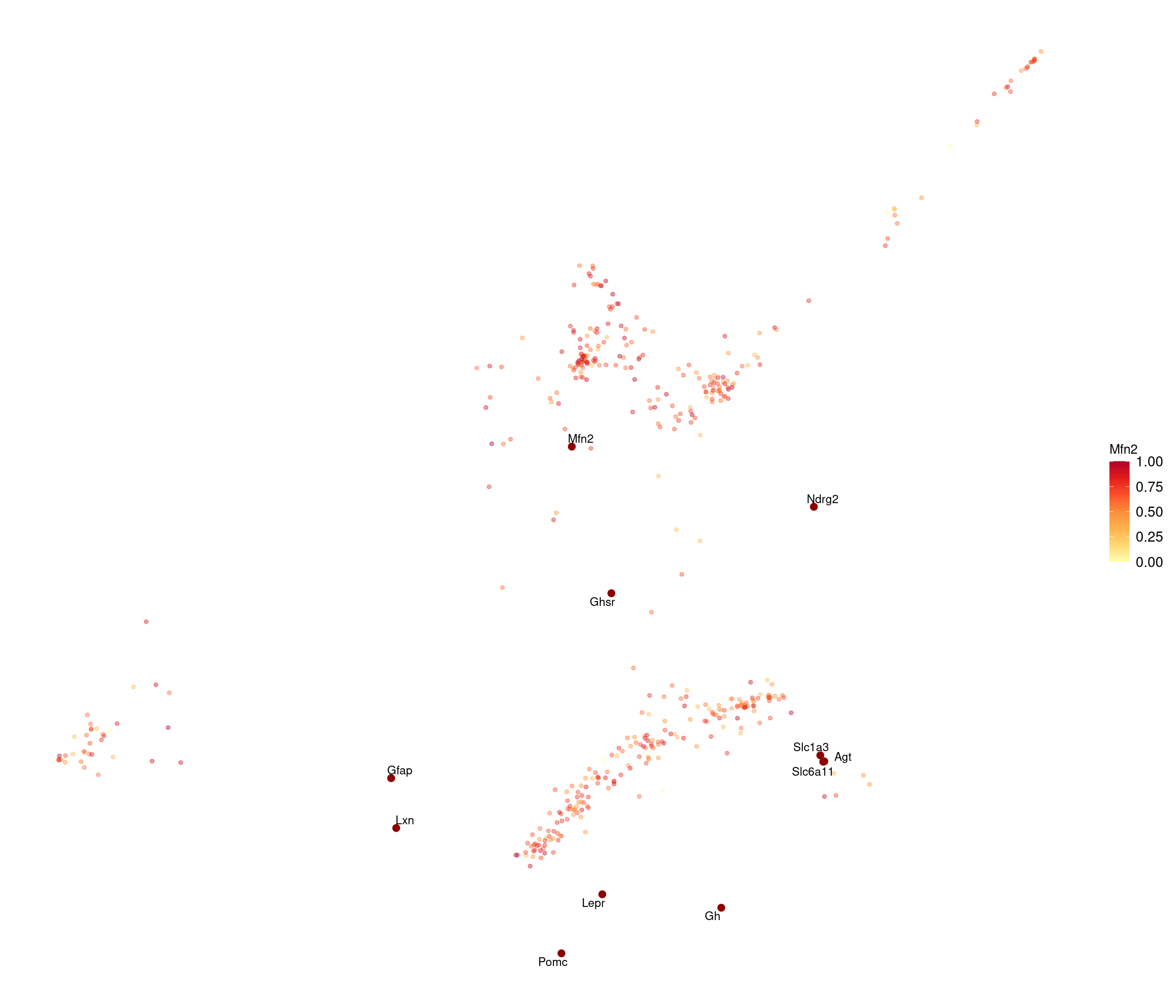


R version 4.2.2 (2022-10-31)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Ubuntu 22.04.1 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.20.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] splines stats4 stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] Nebulosa_1.6.0 swne_0.6.20
[3] patchwork_1.1.2 UpSetR_1.4.0
[5] glmGamPoi_1.8.0 sctransform_0.3.5
[7] SeuratDisk_0.0.0.9020 SeuratWrappers_0.3.0
[9] sp_1.5-0 SeuratObject_4.1.2
[11] Seurat_4.2.0 miQC_1.5.1
[13] flexmix_2.3-18 lattice_0.20-45
[15] scater_1.24.0 scuttle_1.6.3
[17] SingleCellExperiment_1.18.1 SummarizedExperiment_1.26.1
[19] Biobase_2.56.0 GenomicRanges_1.48.0
[21] GenomeInfoDb_1.32.4 IRanges_2.30.1
[23] S4Vectors_0.34.0 BiocGenerics_0.42.0
[25] MatrixGenerics_1.8.1 matrixStats_0.62.0
[27] kableExtra_1.3.4 future_1.28.0
[29] skimr_2.1.4 magrittr_2.0.3
[31] forcats_0.5.2 stringr_1.4.1
[33] dplyr_1.0.10 purrr_0.3.5
[35] readr_2.1.3 tidyr_1.2.1
[37] tibble_3.1.8 ggplot2_3.3.6
[39] tidyverse_1.3.2 here_1.0.1
[41] workflowr_1.7.0
loaded via a namespace (and not attached):
[1] scattermore_0.8 R.methodsS3_1.8.2
[3] bit64_4.0.5 knitr_1.40
[5] irlba_2.3.5.1 DelayedArray_0.22.0
[7] R.utils_2.12.0 data.table_1.14.4
[9] rpart_4.1.19 RCurl_1.98-1.9
[11] generics_0.1.3 snow_0.4-4
[13] ScaledMatrix_1.4.1 callr_3.7.2
[15] cowplot_1.1.1 RANN_2.6.1
[17] proxy_0.4-27 bit_4.0.4
[19] tzdb_0.3.0 spatstat.data_3.0-0
[21] webshot_0.5.4 xml2_1.3.3
[23] lubridate_1.8.0 httpuv_1.6.6
[25] assertthat_0.2.1 viridis_0.6.2
[27] gargle_1.2.1 xfun_0.34
[29] hms_1.1.2 jquerylib_0.1.4
[31] evaluate_0.17 promises_1.2.0.1
[33] fansi_1.0.3 dbplyr_2.2.1
[35] readxl_1.4.1 igraph_1.3.5
[37] DBI_1.1.3 htmlwidgets_1.5.4
[39] spatstat.geom_3.0-3 googledrive_2.0.0
[41] ellipsis_0.3.2 ks_1.13.5
[43] RSpectra_0.16-1 backports_1.4.1
[45] deldir_1.0-6 sparseMatrixStats_1.8.0
[47] vctrs_0.5.0 remotes_2.4.2
[49] ROCR_1.0-11 abind_1.4-5
[51] cachem_1.0.6 withr_2.5.0
[53] progressr_0.11.0 vroom_1.6.0
[55] rgdal_1.5-32 mclust_5.4.10
[57] goftest_1.2-3 svglite_2.1.0
[59] cluster_2.1.4 lazyeval_0.2.2
[61] crayon_1.5.2 NNLM_0.4.4
[63] hdf5r_1.3.7 labeling_0.4.2
[65] pkgconfig_2.0.3 nlme_3.1-160
[67] vipor_0.4.5 nnet_7.3-18
[69] rlang_1.0.6 globals_0.16.1
[71] lifecycle_1.0.3 miniUI_0.1.1.1
[73] modelr_0.1.9 rsvd_1.0.5
[75] ggrastr_1.0.1 cellranger_1.1.0
[77] rprojroot_2.0.3 polyclip_1.10-4
[79] lmtest_0.9-40 Matrix_1.5-1
[81] zoo_1.8-11 reprex_2.0.2
[83] base64enc_0.1-3 beeswarm_0.4.0
[85] whisker_0.4 ggridges_0.5.4
[87] processx_3.8.0 googlesheets4_1.0.1
[89] png_0.1-7 viridisLite_0.4.1
[91] bitops_1.0-7 getPass_0.2-2
[93] R.oo_1.25.0 KernSmooth_2.23-20
[95] DelayedMatrixStats_1.18.1 parallelly_1.32.1
[97] spatstat.random_2.2-0 beachmat_2.12.0
[99] scales_1.2.1 usedist_0.4.0
[101] plyr_1.8.7 ica_1.0-3
[103] zlibbioc_1.42.0 compiler_4.2.2
[105] RColorBrewer_1.1-3 fitdistrplus_1.1-8
[107] cli_3.4.1 XVector_0.36.0
[109] listenv_0.8.0 pbapply_1.5-0
[111] ps_1.7.1 MASS_7.3-58.1
[113] mgcv_1.8-41 tidyselect_1.2.0
[115] stringi_1.7.8 highr_0.9
[117] yaml_2.3.6 askpass_1.1
[119] BiocSingular_1.12.0 ggrepel_0.9.1.9999
[121] grid_4.2.2 sass_0.4.2
[123] tools_4.2.2 future.apply_1.9.1
[125] parallel_4.2.2 rstudioapi_0.14
[127] git2r_0.30.1 liger_2.0.1
[129] gridExtra_2.3 farver_2.1.1
[131] Rtsne_0.16 digest_0.6.30
[133] BiocManager_1.30.18 rgeos_0.5-9
[135] pracma_2.4.2 FNN_1.1.3.1
[137] shiny_1.7.3 Rcpp_1.0.9
[139] broom_1.0.1 later_1.3.0
[141] RcppAnnoy_0.0.19 httr_1.4.4
[143] colorspace_2.0-3 rvest_1.0.3
[145] fs_1.5.2 tensor_1.5
[147] reticulate_1.26 umap_0.2.9.0
[149] uwot_0.1.14 spatstat.utils_3.0-1
[151] plotly_4.10.0 systemfonts_1.0.4
[153] xtable_1.8-4 jsonlite_1.8.3
[155] modeltools_0.2-23 R6_2.5.1
[157] pillar_1.8.1 htmltools_0.5.3
[159] mime_0.12 glue_1.6.2
[161] fastmap_1.1.0 BiocParallel_1.30.3
[163] BiocNeighbors_1.14.0 ggmin_0.0.0.9000
[165] codetools_0.2-18 mvtnorm_1.1-3
[167] utf8_1.2.2 bslib_0.4.0
[169] spatstat.sparse_3.0-0 ggbeeswarm_0.6.0
[171] leiden_0.4.3 openssl_2.0.4
[173] limma_3.52.4 survival_3.4-0
[175] rmarkdown_2.17 repr_1.1.4
[177] munsell_0.5.0 GenomeInfoDbData_1.2.8
[179] haven_2.5.1 reshape2_1.4.4
[181] gtable_0.3.1 spatstat.core_2.4-4