Investigate Rank and Inverse-Wishart fit of N3finemapping data
Last updated: 2025-12-02
Checks: 7 0
Knit directory: Improved_LD_SuSiE/
This reproducible R Markdown analysis was created with workflowr (version 1.7.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20250821) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 153ece7. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .DS_Store
Unstaged changes:
Modified: code_push.R
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown
(analysis/genotype-matrix-rank-investigation.Rmd) and HTML
(docs/genotype-matrix-rank-investigation.html) files. If
you’ve configured a remote Git repository (see
?wflow_git_remote), click on the hyperlinks in the table
below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 153ece7 | dodat97 | 2025-12-02 | wflow_publish(c("analysis/nu-matrix-F-distribution.Rmd", "analysis/genotype-matrix-rank-investigation.Rmd")) |
| html | fca02ee | dodat97 | 2025-12-01 | Build site. |
| html | 691542f | dodat97 | 2025-12-01 | Build site. |
| Rmd | 317d89b | dodat97 | 2025-12-01 | wflow_publish(c("analysis/genotype-matrix-rank-investigation.Rmd")) |
library(susieR)
library(Matrix)
data(N3finemapping)
attach(N3finemapping)
X0 = N3finemapping$X
## getting covariance matrix from the whole sample
## and examine the eigendecomposition to estimate numerical rank
R = cov(X0)
eig <- eigen(R)
plot(eig$values,
main = "Eigenvalues of covariance matrix calculated using all samples",
ylab = "Value",
xlab = "Eigenvalue index")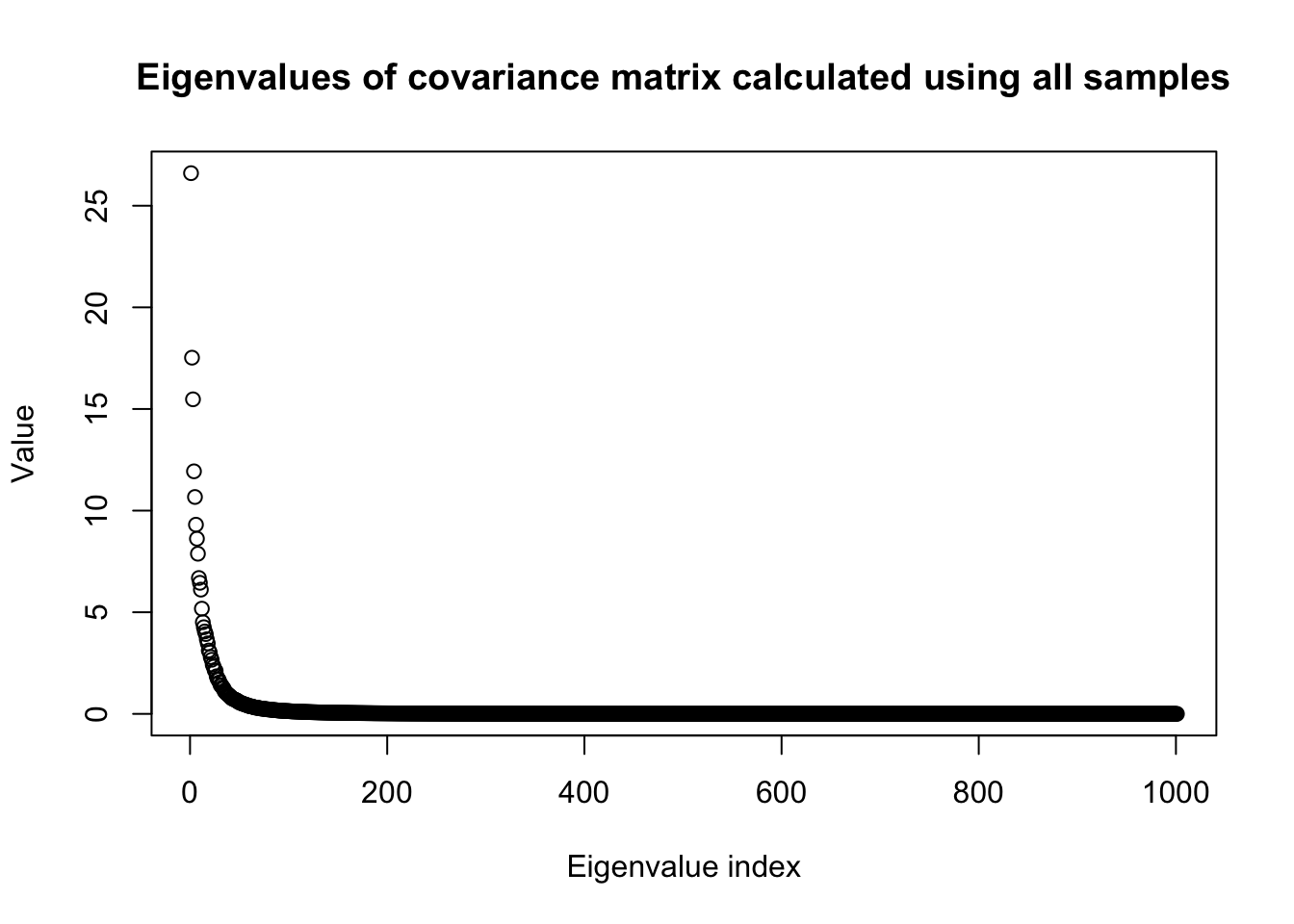
| Version | Author | Date |
|---|---|---|
| 691542f | dodat97 | 2025-12-01 |
n0 = dim(X0)[1]
p0 = dim(X0)[2]
percent_explained = .95
eig_cumsum = cumsum(eig$values)
r_p = sum(eig_cumsum < percent_explained * eig_cumsum[p0]) ## percentage variance explained
sprintf("%d first principle components explain %.1f percent of variance", r_p, percent_explained*100)[1] "82 first principle components explain 95.0 percent of variance"snp_total = p0
sprintf("Total number of SNPs is %d", p0)[1] "Total number of SNPs is 1001"sprintf("Sample size %d", n0)[1] "Sample size 574"Now we proceed to split the data into half and look at the heatmap of the covariance matrices of two sub-samples.
Later we want to examine the Inverse-Wishart likelihood so I also sub-sample the SNPs here to make sure that the number of SNPs is less than the sample size.
#### randomly split the data into half
#### randomly select p consecutive SNPs where p < n so IW is proper
seed = 10
p = 50
# Start from a random point on the genome
indx_start = sample(1: (snp_total - p), 1)
X = X0[, indx_start:(indx_start + p -1)]
# View(cor(X)[1:10, 1:10])
## sub-sample into two
out_sample_size = n0 / 2
out_sample = sample(1:n0, out_sample_size)
X_out = X[out_sample, ]
X_in = X[setdiff(1:n0, out_sample), ]
rm_p = c(which(diag(cov(X_in))==0), which(diag(cov(X_out))==0))
indx_p = setdiff(1:p, rm_p)
X_in = X_in[, indx_p]
X_out = X_out[, indx_p]
## out-sample LD matrix
p = length(indx_p)
Rp = cov(X_out)
R0 = cov(X_in)
library(ggplot2)
library(reshape2)
df1 <- melt(R0)
df2 <- melt(Rp)
N_in = nrow(X_in)
N_out = nrow(X_out)
p1 <- ggplot(df1, aes(Var1, Var2, fill = value)) +
geom_tile() +
scale_fill_gradient2(low="blue", mid="white", high="red") +
coord_fixed() +
ggtitle(sprintf("In-sample Cov, %d samples", nrow(X_in)))
p2 <- ggplot(df2, aes(Var1, Var2, fill = value)) +
geom_tile() +
scale_fill_gradient2(low="blue", mid="white", high="red") +
coord_fixed() +
ggtitle(sprintf("Out-of-sample Cov, %d samples", nrow(X_out)))
library(gridExtra)
grid.arrange(p1, p2, ncol = 2)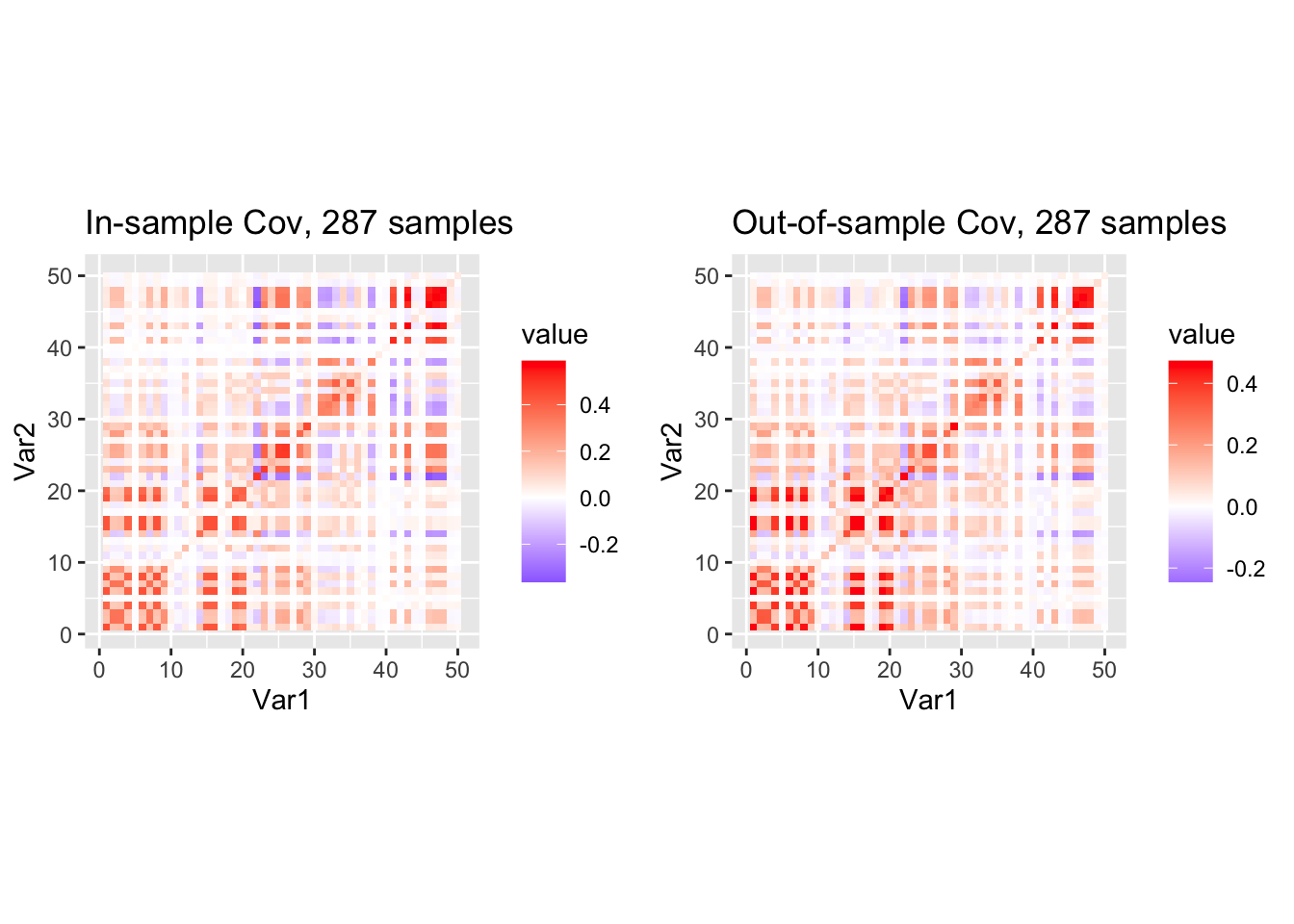
| Version | Author | Date |
|---|---|---|
| 691542f | dodat97 | 2025-12-01 |
They look pretty similar, let us see what is the MLE of \(\nu_0\) in the likelihood \(IW(R_0 | mean = R', df=\nu_0 + p + 1)\).
## IW and examine IW likelihood
#### log IW(R0 | nu0 * Rp, nu0 + J + 1)
log_multigamma_vec <- function(a, p) {
# vectorized multivariate gamma
j <- 1:p
# sum over j, but broadcasting a over j
(p*(p-1)/4)*log(pi) +
rowSums(matrix(lgamma(a), nrow=length(a), ncol=p, byrow=FALSE) +
matrix((1 - j)/2, nrow=length(a), ncol=p, byrow=TRUE))
}
log_iw <- function(R0, Rp, nu_vec) {
p <- nrow(R0)
jitter = 1e-8
R0 = R0 + jitter * diag(rep(1, p))
Rp = Rp + jitter * diag(rep(1, p))
# Precompute expensive shared quantities
logdet_nu_Rp <- determinant(Rp, logarithm = TRUE)$modulus + p * log(nu_vec)
logdetR0 <- determinant(R0, logarithm = TRUE)$modulus
tr_term <- nu_vec * sum(t(Rp) * solve(R0))
llhs = (.5 * (nu_vec + p + 1) * logdet_nu_Rp
- .5 * (nu_vec + p + 1) * p * log(2)
- log_multigamma_vec((nu_vec + p + 1) / 2, p)
- .5 * (nu_vec + 2 * (p + 1)) * logdetR0
- .5 * tr_term)
as.numeric(llhs)
}
nu_vec = c(1:100)
llhs = log_iw(R0, Rp, nu_vec)
plot(nu_vec, llhs, xlab = "nu value", ylab = "log-likelihood")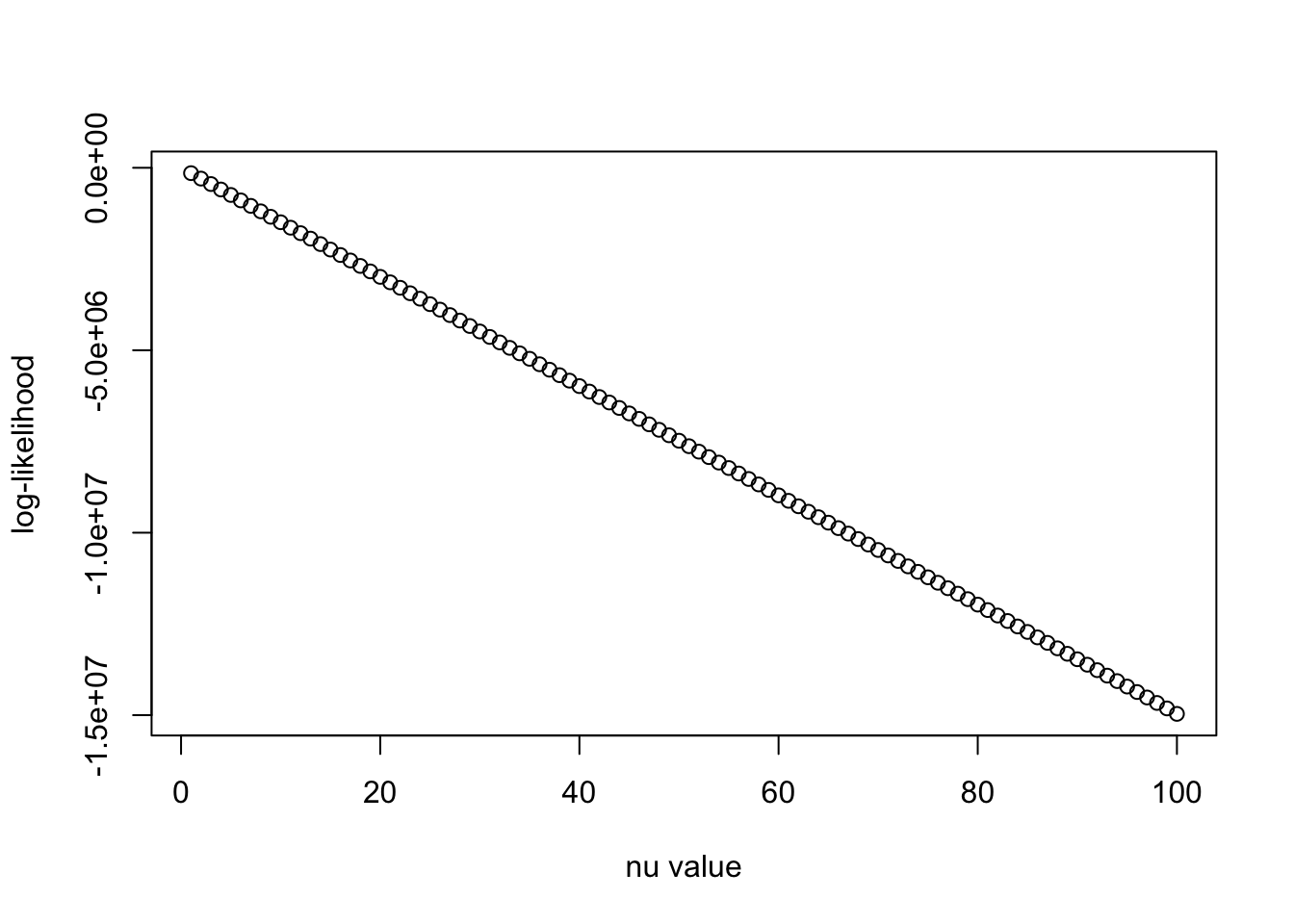
| Version | Author | Date |
|---|---|---|
| 691542f | dodat97 | 2025-12-01 |
The likelihood just decreases linearly as \(\nu_0\) increases. It is because of the term \(\mathrm{trace}((R_0)^{-1} R')\) that dominates the log det term, and \(\nu_0\) is multiplied into this trace term in the log-likelihood. This numerical behavior is similar to when calculating \(v^\top R^{-1} v\) in the presentation in the group meeting we saw earlier (curse of dimensionality).
jitter = 1e-8
R0_ = R0 + jitter * diag(rep(1, p))
Rp_ = Rp + jitter * diag(rep(1, p))
trace_term = sum(t(Rp_) * solve(R0_))
logdet_nu_Rp <- determinant(Rp_, logarithm = TRUE)$modulus
logdetR0 <- determinant(R0_, logarithm = TRUE)$modulus
print(trace_term)[1] 299475.8print(logdetR0)[1] -227.7098
attr(,"logarithm")
[1] TRUEprint(logdet_nu_Rp)[1] -217.3115
attr(,"logarithm")
[1] TRUE## now let's try to turn it into the PCA-like form and learn nu again
eig <- eigen(Rp)
eig_cumsum = cumsum(eig$values)
percent_explained = .99
eig_cumsum = cumsum(eig$values)
q = sum(eig_cumsum < percent_explained * eig_cumsum[p])
sprintf("%d first principle components explain %.1f percent of variance", q, percent_explained*100)[1] "26 first principle components explain 99.0 percent of variance"lambda <- eig$values
U <- eig$vectors
sigma2_est <- mean(lambda[(q+1):p])
L_diag <- sqrt(lambda[1:q] - sigma2_est)
# L_diag <- sqrt(lambda[1:q])
W_ppca <- U[,1:q] %*% diag(L_diag)
Rp_PCA <- W_ppca %*% t(W_ppca) + sigma2_est * diag(p)
eig <- eigen(R0)
lambda <- eig$values
U <- eig$vectors
sigma2_est <- mean(lambda[(q+1):p])
L_diag <- sqrt(lambda[1:q] - sigma2_est)
# L_diag <- sqrt(lambda[1:q])
W_ppca <- U[,1:q] %*% diag(L_diag)
R0_PCA <- W_ppca %*% t(W_ppca) + sigma2_est * diag(p)
df3 <- melt(R0_PCA)
df4 <- melt(Rp_PCA)
p3 <- ggplot(df3, aes(Var1, Var2, fill = value)) +
geom_tile() +
scale_fill_gradient2(low="blue", mid="white", high="red") +
coord_fixed() +
ggtitle(paste0("In-sample PCA"))
p4 <- ggplot(df4, aes(Var1, Var2, fill = value)) +
geom_tile() +
scale_fill_gradient2(low="blue", mid="white", high="red") +
coord_fixed() +
ggtitle(paste0("Out-of-sample PCA"))
grid.arrange(p1, p2, p3, p4, ncol = 2, nrow = 2)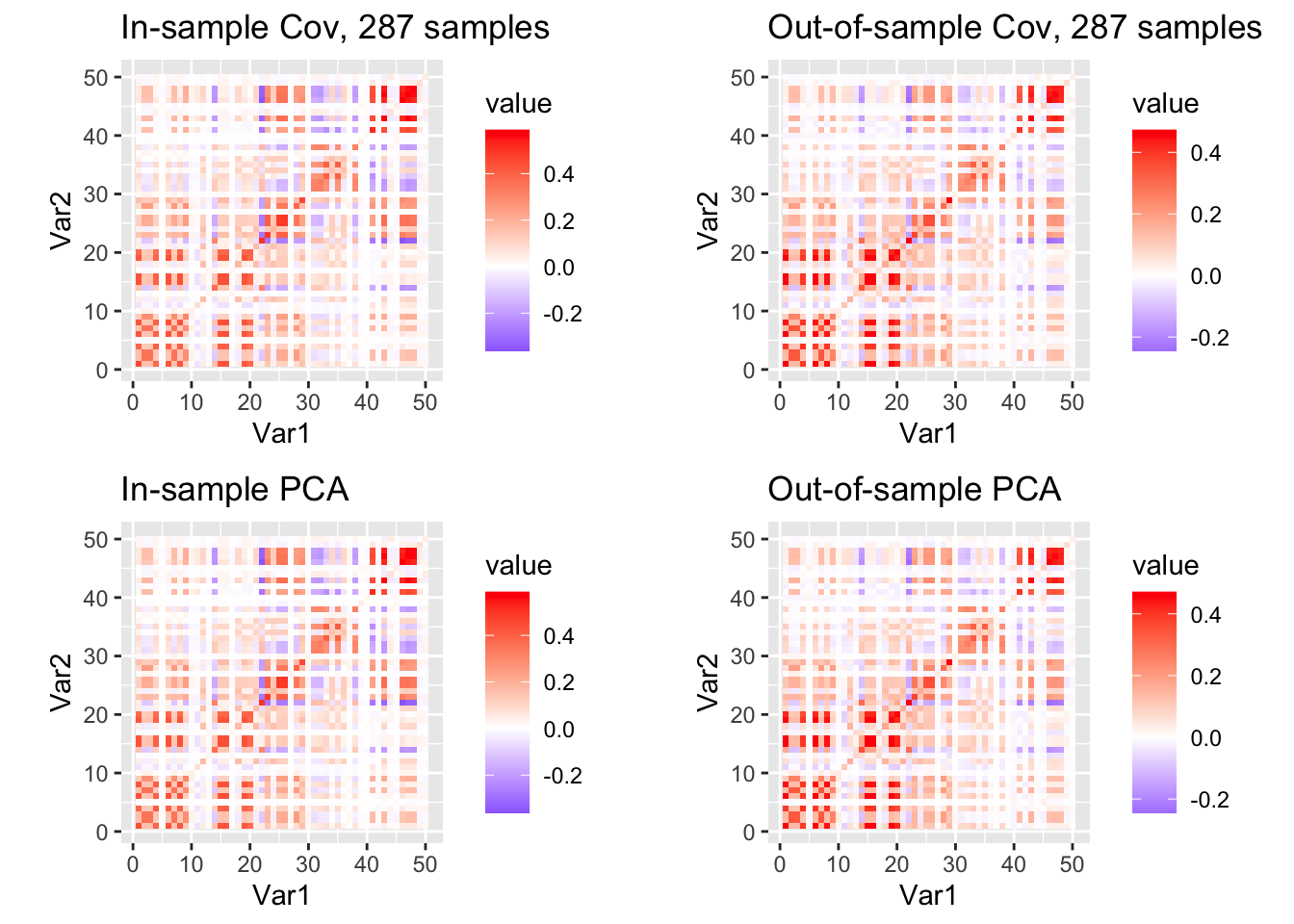
| Version | Author | Date |
|---|---|---|
| 691542f | dodat97 | 2025-12-01 |
nu_vec = c(1:200)
llhs = log_iw(R0_PCA, Rp_PCA, nu_vec)
# llhs = log_iw(R0_PCA, Rp, nu_vec)
# llhs = log_iw(R0, Rp_PCA, nu_vec)
plot(nu_vec, llhs, xlab = "nu value", ylab = "log-likelihood")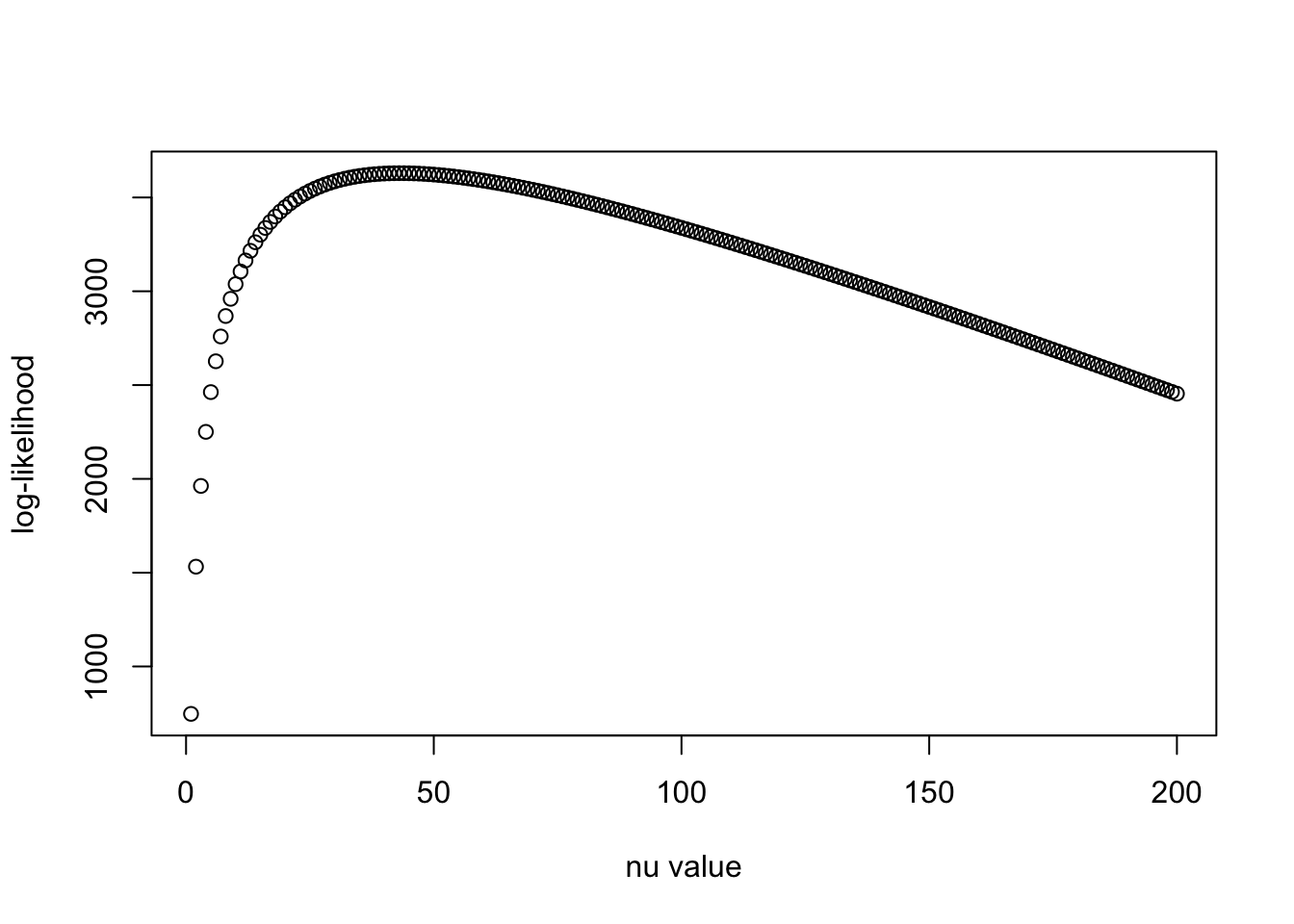
| Version | Author | Date |
|---|---|---|
| 691542f | dodat97 | 2025-12-01 |
print(nu_vec[which.max(llhs)])[1] 43Seems like we get more reasonable estimate of \(\nu_0\) with 99% variance-explained PCA version of the covariance matrix.
sessionInfo()R version 4.5.1 (2025-06-13)
Platform: aarch64-apple-darwin20
Running under: macOS Sequoia 15.0
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.1
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
time zone: America/Chicago
tzcode source: internal
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] gridExtra_2.3 reshape2_1.4.4 ggplot2_3.5.2 Matrix_1.7-3
[5] susieR_0.14.2 workflowr_1.7.2
loaded via a namespace (and not attached):
[1] sass_0.4.10 stringi_1.8.7 lattice_0.22-7 digest_0.6.37
[5] magrittr_2.0.3 evaluate_1.0.5 grid_4.5.1 RColorBrewer_1.1-3
[9] fastmap_1.2.0 rprojroot_2.1.1 plyr_1.8.9 jsonlite_2.0.0
[13] processx_3.8.6 whisker_0.4.1 reshape_0.8.10 ps_1.9.1
[17] mixsqp_0.3-54 promises_1.5.0 httr_1.4.7 scales_1.4.0
[21] jquerylib_0.1.4 cli_3.6.5 rlang_1.1.6 crayon_1.5.3
[25] withr_3.0.2 cachem_1.1.0 yaml_2.3.10 otel_0.2.0
[29] tools_4.5.1 httpuv_1.6.16 vctrs_0.6.5 R6_2.6.1
[33] matrixStats_1.5.0 lifecycle_1.0.4 git2r_0.36.2 stringr_1.5.2
[37] fs_1.6.6 irlba_2.3.5.1 pkgconfig_2.0.3 callr_3.7.6
[41] pillar_1.11.0 bslib_0.9.0 later_1.4.4 gtable_0.3.6
[45] glue_1.8.0 Rcpp_1.1.0 xfun_0.53 tibble_3.3.0
[49] rstudioapi_0.17.1 knitr_1.50 farver_2.1.2 htmltools_0.5.8.1
[53] labeling_0.4.3 rmarkdown_2.29 compiler_4.5.1 getPass_0.2-4