Summary DE genes
almut
14 November 2019
Last updated: 2019-11-18
Checks: 7 0
Knit directory: transcriptome_cll/
This reproducible R Markdown analysis was created with workflowr (version 1.4.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20190511) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Untracked files:
Untracked: data/2018-03-05_IGHV.RData
Untracked: data/patmeta_170324.RData
Untracked: output/IGHV1_69.svg
Untracked: output/cluster1000exprgenes.pdf
Untracked: output/cluster500exprgenes.pdf
Untracked: output/desRes_15112019.RData
Untracked: output/figures/hist_mutations.svg
Untracked: output/figures/overview_mutations.pdf
Untracked: output/figures/r_objects/
Untracked: output/figures/sum_diffGenes_0.05_2.pdf
Untracked: output/figures/sum_diffGenes_noTsig.pdf
Untracked: output/figures/sum_diffGenes_noTsig_IGHVTri12.pdf
Unstaged changes:
Modified: analysis/trisomy12.Rmd
Modified: output/diff_genes/ATM_diffGenes.csv
Modified: output/diff_genes/BRAF_diffGenes.csv
Modified: output/diff_genes/MED12_diffGenes.csv
Modified: output/diff_genes/NOTCH1_diffGenes.csv
Modified: output/diff_genes/SF3B1_diffGenes.csv
Modified: output/diff_genes/TP53_diffGenes.csv
Modified: output/diff_genes/del11q22.3_diffGenes.csv
Modified: output/diff_genes/del13q14_diffGenes.csv
Modified: output/diff_genes/del17p13_diffGenes.csv
Modified: output/diff_genes/del8p12_diffGenes.csv
Modified: output/diff_genes/gain8q24_diffGenes.csv
Modified: output/diff_genes/trisomy12_diffGenes.csv
Modified: output/figures/pca_Meth_top150.svg
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the R Markdown and HTML files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view them.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| html | e20c885 | aluetge | 2019-11-14 | Build site. |
| Rmd | 7a81840 | aluetge | 2019-11-14 | wflow_publish(“analysis/summary_de_genes.Rmd”) |
| html | 794365a | aluetge | 2019-11-14 | Build site. |
| Rmd | e8e7a32 | aluetge | 2019-11-14 | wflow_publish(“analysis/summary_de_genes.Rmd”) |
| html | f9c1040 | aluetge | 2019-11-14 | Build site. |
| Rmd | 225f170 | aluetge | 2019-11-14 | wflow_publish(“analysis/summary_de_genes.Rmd”) |
Summary DE genes by variants
Aim: Generate a summary plot giving an overview on the number of differntially expressed genes / number upreagulated / number downregulated genes and chromosomal distribution for CNVs.
libraries
suppressPackageStartupMessages({
library(tidyverse)
library(ggplot2)
library(here)
library(ggpubr)
library(ggsci)
library(RColorBrewer)
library(circlize)
library(reshape2)
library(ggsci)
library(DESeq2)
})data
data_dir <- here("data")
output_dir <- here("output")
figure_dir <- here("output/figures")
#dds data set. gene expression data + patmetadata
load(paste0(data_dir, "/ddsrnaCLL_150218.RData"))
variants <- c( "trisomy12", "del13q14", "del8p12", "gain8q24", "del11q22.3", "del17p13", "BRAF", "NOTCH1", "SF3B1","TP53", "ATM", "MED12","IGHV")Summarize de genes in a table
#Function to summarize de genes
variant_stats <- function(variant, chr){
diff_genes <-read.csv(file=paste0(output_dir, "/diff_genes/", variant, "_diffGenes.csv"))
sig_genes <- as.tibble(diff_genes) %>% filter(padj < 0.01)
up_genes <- sig_genes %>% filter(log2FoldChange > 0) %>% mutate(chrom = rowData(ddsCLL)$chromosome[which(rownames(ddsCLL) %in% X)])
dn_genes <- sig_genes %>% filter(log2FoldChange < 0) %>% mutate(chrom = rowData(ddsCLL)$chromosome[which(rownames(ddsCLL) %in% X)])
chr_up <- ifelse(!is.na(chr), length(which(up_genes$chrom %in% chr)), 0)
chr_dn <- ifelse(!is.na(chr), length(which(dn_genes$chrom %in% chr)), 0)
summary <- tibble(
variant = variant,
n_genes = nrow(sig_genes),
up_genes = nrow(up_genes),
dn_genes = nrow(dn_genes),
nr_up = chr_up,
nr_dn = chr_dn,
nr_chrom = chr_dn + chr_up,
nr_other = n_genes - chr_dn - chr_up
)
}
# Variant chromosomal location
chromo <- c( "12", "13", "8", "8", "11", "17", NA, NA, NA, NA, NA, NA, NA)
input <- cbind(variants, chromo)
summary <- mapply(variant_stats, variants, chromo)Warning: `as.tibble()` is deprecated, use `as_tibble()` (but mind the new semantics).
This warning is displayed once per session.sum_dat <- data.frame(matrix(unlist(summary), nrow=13, byrow=T),stringsAsFactors=FALSE)
colnames(sum_dat) <- c("variant", "n_genes", "up_genes", "dn_genes", "nr_up", "nr_dn", "nr_chrom", "nr_other")
sum_order <- as.tibble(sum_dat) %>% arrange(desc(-1*as.numeric(n_genes))) %>% dplyr::select(variant)
#sum_dat_long <- melt(sum_dat[,c("variant","up_genes", "dn_genes")], id.vars = c("variant")) #, variable_name = c("up_genes", "dn_genes"))
sum_dat_long <- melt(sum_dat[,c("variant","nr_other", "nr_chrom")], id.vars = c("variant")) #, variable_name = c("up_genes", "dn_genes"))
colnames(sum_dat_long) <- c("variant", "location", "genes")
sum_dat_long$dir_color <-paste0(sum_dat_long$variant, "_", sum_dat_long$location)
sum_dat_long$genes <- as.numeric(as.character(sum_dat_long$genes))
sum_dat_long <- as.tibble(sum_dat_long) %>% arrange(desc(genes)) %>% mutate(group=ifelse(variant %in% c("trisomy12", "IGHV"), 1, 2))Plots
Summary all variants
col_pal1 <- pal_igv(palette = c("default"), alpha = 1)(13)
col_pal2 <- pal_igv(palette = c("default"), alpha = 0.4)(13)
color <- c(col_pal1, col_pal2)
color <- sort(color)
#pdf(file=paste0(figure_dir, "/sum_diffGenes_0.05_2.pdf"), width=8, height=5)
ggbarplot(sum_dat_long[order(match(sum_dat_long$variant, sum_order$variant)),], "variant", "genes",
fill = "dir_color", palette = color,
label = FALSE, font.x =23 ,font.y = 23, font.tickslab = 18, x.text.angle = 90, legend = "none") 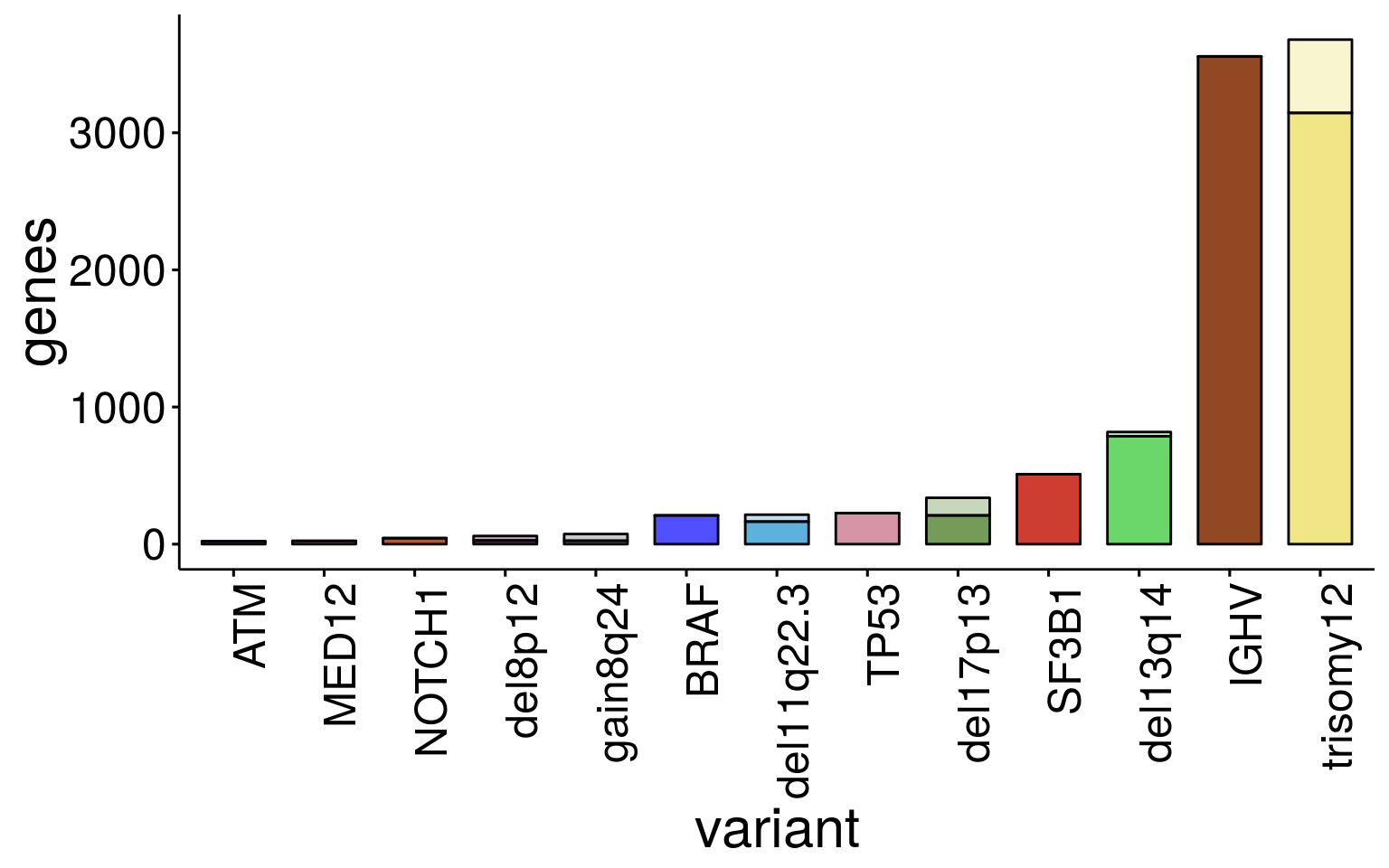
| Version | Author | Date |
|---|---|---|
| f9c1040 | aluetge | 2019-11-14 |
#dev.off()IGHV and trisomy12
sum_dat_long1 <- as.tibble(sum_dat_long) %>% filter(variant %in% c("trisomy12", "IGHV"))
#pdf(file=paste0(figure_dir, "/sum_diffGenes_noTsig_IGHVTri12.pdf"), width=5, height=2.3)
p1 <- ggbarplot(sum_dat_long1[order(match(sum_dat_long1$variant, sum_order$variant)),], "variant", "genes",
fill = "dir_color", palette = color[c(1:4)],
label = FALSE, font.x =23 ,font.y = 23, font.tickslab = 18, x.text.angle = 0, legend = "none", rotate = T)
#dev.off()
p1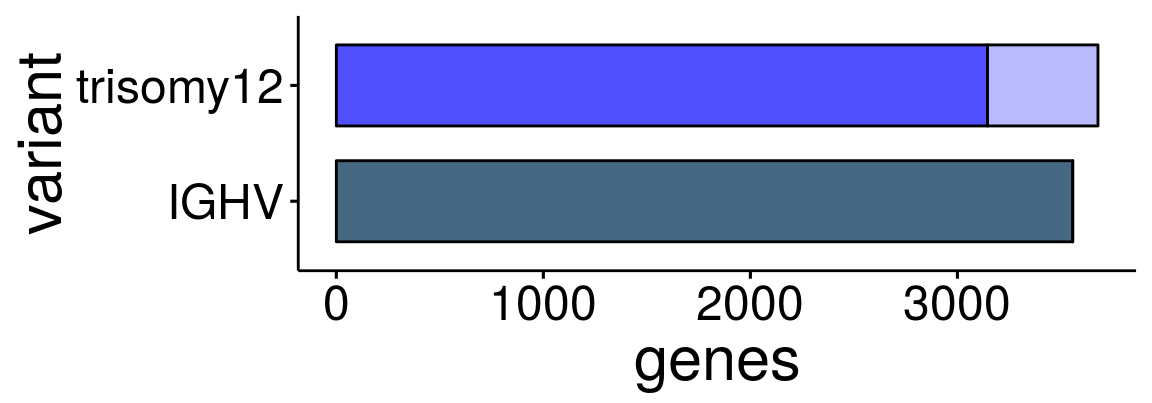
| Version | Author | Date |
|---|---|---|
| f9c1040 | aluetge | 2019-11-14 |
saveRDS(p1, file = paste0(output_dir, "/figures/r_objects/summary_de_genes_IGHV_tri12.rds"))All other
sum_dat_long2 <- as.tibble(sum_dat_long) %>% filter(!variant %in% c("trisomy12", "IGHV"))
#pdf(file=paste0(figure_dir, "/sum_diffGenes_noTsig.pdf"), width=6, height=7.5)
p2 <- ggbarplot(sum_dat_long2[order(match(sum_dat_long2$variant, sum_order$variant)),], "variant", "genes",
fill = "dir_color", palette = color[c(5:26)],
label = FALSE, font.x =23 ,font.y = 23, font.tickslab = 18, x.text.angle = 0, legend = "none", rotate = T)
#dev.off()
p2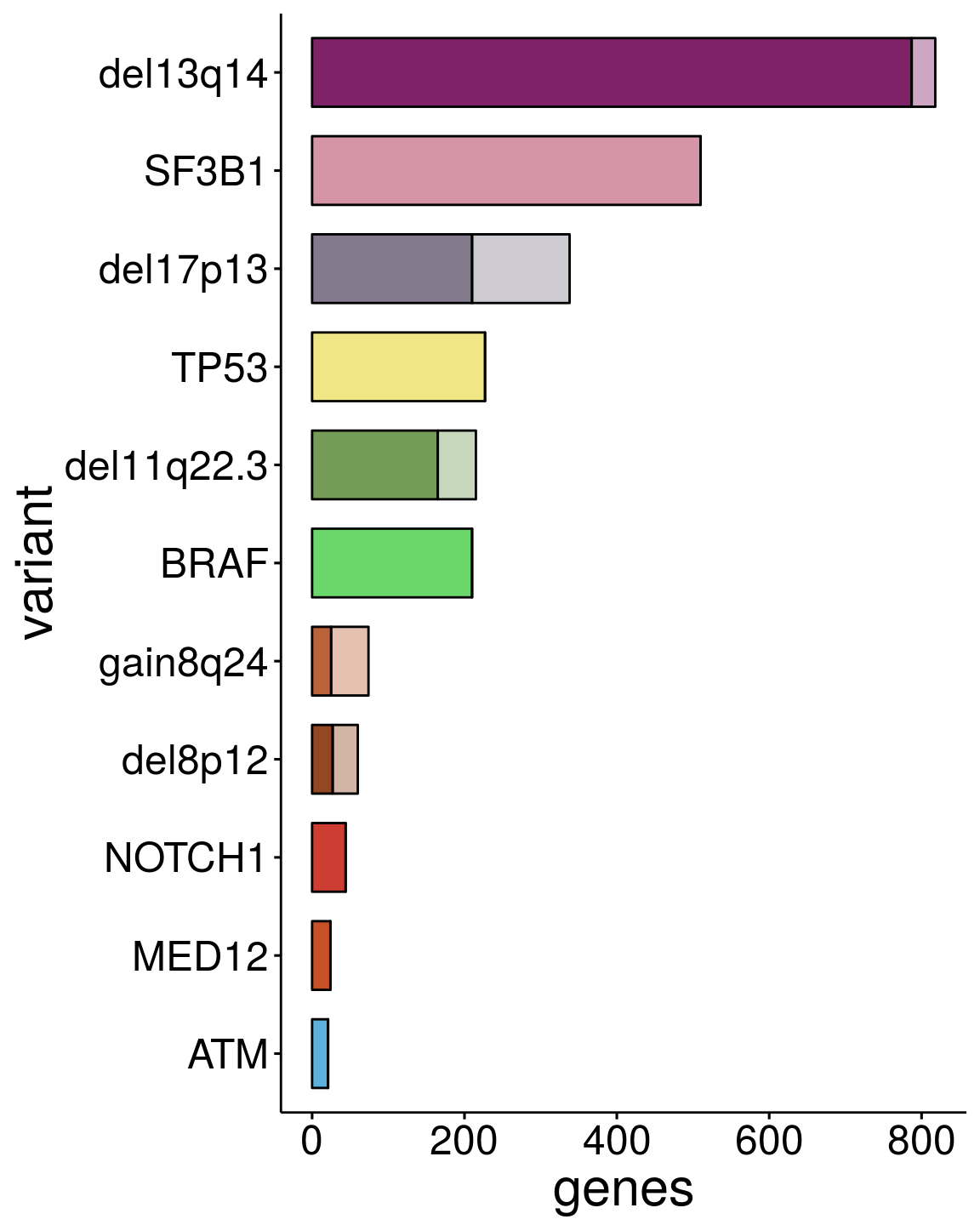
| Version | Author | Date |
|---|---|---|
| f9c1040 | aluetge | 2019-11-14 |
saveRDS(p2, file = paste0(output_dir, "/figures/r_objects/summary_de_genes_all.rds"))summary table
#pdf(file=paste0(output_dir,"/sum_diffGenes_table.pdf"), width=5, height=7.5)
sum_dat variant n_genes up_genes dn_genes nr_up nr_dn nr_chrom nr_other
1 trisomy12 3679 1806 1873 520 14 534 3145
2 del13q14 818 253 565 6 25 31 787
3 del8p12 60 19 41 4 29 33 27
4 gain8q24 74 56 18 38 11 49 25
5 del11q22.3 215 74 141 3 47 50 165
6 del17p13 338 161 177 20 108 128 210
7 BRAF 210 170 40 0 0 0 210
8 NOTCH1 44 12 32 0 0 0 44
9 SF3B1 510 308 202 0 0 0 510
10 TP53 227 110 117 0 0 0 227
11 ATM 21 14 7 0 0 0 21
12 MED12 24 14 10 0 0 0 24
13 IGHV 3557 2108 1449 0 0 0 3557#dev.off()
sessionInfo()R version 3.6.0 (2019-04-26)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Ubuntu 16.04.6 LTS
Matrix products: default
BLAS: /usr/lib/libblas/libblas.so.3.6.0
LAPACK: /usr/lib/lapack/liblapack.so.3.6.0
locale:
[1] LC_CTYPE=de_DE.UTF-8 LC_NUMERIC=C
[3] LC_TIME=de_DE.UTF-8 LC_COLLATE=de_DE.UTF-8
[5] LC_MONETARY=de_DE.UTF-8 LC_MESSAGES=de_DE.UTF-8
[7] LC_PAPER=de_DE.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=de_DE.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] parallel stats4 stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] DESeq2_1.24.0 SummarizedExperiment_1.14.0
[3] DelayedArray_0.10.0 BiocParallel_1.18.0
[5] matrixStats_0.54.0 Biobase_2.44.0
[7] GenomicRanges_1.36.0 GenomeInfoDb_1.20.0
[9] IRanges_2.18.1 S4Vectors_0.22.0
[11] BiocGenerics_0.30.0 reshape2_1.4.3
[13] circlize_0.4.6 RColorBrewer_1.1-2
[15] ggsci_2.9 ggpubr_0.2
[17] magrittr_1.5 here_0.1
[19] forcats_0.4.0 stringr_1.4.0
[21] dplyr_0.8.1 purrr_0.3.2
[23] readr_1.3.1 tidyr_0.8.3
[25] tibble_2.1.3 ggplot2_3.1.1
[27] tidyverse_1.2.1
loaded via a namespace (and not attached):
[1] colorspace_1.4-1 rprojroot_1.3-2 htmlTable_1.13.1
[4] XVector_0.24.0 GlobalOptions_0.1.0 base64enc_0.1-3
[7] fs_1.3.1 rstudioapi_0.10 bit64_0.9-7
[10] AnnotationDbi_1.46.0 lubridate_1.7.4 xml2_1.2.0
[13] splines_3.6.0 geneplotter_1.62.0 knitr_1.23
[16] Formula_1.2-3 jsonlite_1.6 workflowr_1.4.0
[19] broom_0.5.2 annotate_1.62.0 cluster_2.1.0
[22] compiler_3.6.0 httr_1.4.0 backports_1.1.4
[25] assertthat_0.2.1 Matrix_1.2-17 lazyeval_0.2.2
[28] cli_1.1.0 acepack_1.4.1 htmltools_0.3.6
[31] tools_3.6.0 gtable_0.3.0 glue_1.3.1
[34] GenomeInfoDbData_1.2.1 Rcpp_1.0.1 cellranger_1.1.0
[37] nlme_3.1-140 xfun_0.7 rvest_0.3.4
[40] XML_3.98-1.20 zlibbioc_1.30.0 scales_1.0.0
[43] hms_0.4.2 yaml_2.2.0 memoise_1.1.0
[46] gridExtra_2.3 rpart_4.1-15 latticeExtra_0.6-28
[49] stringi_1.4.3 RSQLite_2.1.1 genefilter_1.66.0
[52] checkmate_1.9.3 shape_1.4.4 rlang_0.3.4
[55] pkgconfig_2.0.2 bitops_1.0-6 evaluate_0.14
[58] lattice_0.20-38 labeling_0.3 htmlwidgets_1.3
[61] bit_1.1-14 tidyselect_0.2.5 plyr_1.8.4
[64] R6_2.4.0 generics_0.0.2 Hmisc_4.2-0
[67] DBI_1.0.0 pillar_1.4.1 haven_2.1.0
[70] whisker_0.3-2 foreign_0.8-71 withr_2.1.2
[73] survival_2.44-1.1 RCurl_1.95-4.12 nnet_7.3-12
[76] modelr_0.1.4 crayon_1.3.4 rmarkdown_1.13
[79] locfit_1.5-9.1 grid_3.6.0 readxl_1.3.1
[82] data.table_1.12.2 blob_1.1.1 git2r_0.25.2
[85] digest_0.6.19 xtable_1.8-4 munsell_0.5.0