Chimp Total vs Nuclear
Briana Mittleman
10/7/2019
Last updated: 2019-10-07
Checks: 7 0
Knit directory: Comparative_APA/analysis/
This reproducible R Markdown analysis was created with workflowr (version 1.4.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20190902) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: code/chimp_log/
Ignored: code/human_log/
Ignored: data/metadata_HCpanel.txt.sb-f4823d1e-qihGek/
Untracked files:
Untracked: ._.DS_Store
Untracked: Chimp/
Untracked: Human/
Untracked: code/._Config_chimp.yaml
Untracked: code/._Config_human.yaml
Untracked: code/._LiftOrthoPAS2chimp.sh
Untracked: code/._Snakefile
Untracked: code/._SnakefilePAS
Untracked: code/._SnakefilePASfilt
Untracked: code/._bed215upbed.py
Untracked: code/._bed2SAF_gen.py
Untracked: code/._buildStarIndex.sh
Untracked: code/._cleanbed2saf.py
Untracked: code/._cluster.json
Untracked: code/._extraSnakefiltpas
Untracked: code/._filterPASforMP.py
Untracked: code/._filterPostLift.py
Untracked: code/._fixUTRexonanno.py
Untracked: code/._formathg38Anno.py
Untracked: code/._formatpantro6Anno.py
Untracked: code/._intersectLiftedPAS.sh
Untracked: code/._liftPAS19to38.sh
Untracked: code/._makeSamplyGroupsHuman_TvN.py
Untracked: code/._maphg19.sh
Untracked: code/._maphg19_subjunc.sh
Untracked: code/._overlapapaQTLPAS.sh
Untracked: code/._prepareCleanLiftedFC_5perc4LC.py
Untracked: code/._preparePAS4lift.py
Untracked: code/._primaryLift.sh
Untracked: code/._recLiftchim2human.sh
Untracked: code/._revLiftPAShg38to19.sh
Untracked: code/._reverseLift.sh
Untracked: code/._runChimpDiffIso.sh
Untracked: code/._runHumanDiffIso.sh
Untracked: code/._snakemake.batch
Untracked: code/._snakemakePAS.batch
Untracked: code/._snakemakePASchimp.batch
Untracked: code/._snakemakePAShuman.batch
Untracked: code/._snakemake_chimp.batch
Untracked: code/._snakemake_human.batch
Untracked: code/._snakemakefiltPAS.batch
Untracked: code/._snakemakefiltPAS_chimp
Untracked: code/._snakemakefiltPAS_chimp.sh
Untracked: code/._snakemakefiltPAS_human.sh
Untracked: code/._submit-snakemake-chimp.sh
Untracked: code/._submit-snakemake-human.sh
Untracked: code/._submit-snakemakePAS-chimp.sh
Untracked: code/._submit-snakemakePAS-human.sh
Untracked: code/._submit-snakemakefiltPAS-chimp.sh
Untracked: code/._submit-snakemakefiltPAS-human.sh
Untracked: code/.snakemake/
Untracked: code/Config_chimp.yaml
Untracked: code/Config_human.yaml
Untracked: code/LiftOrthoPAS2chimp.sh
Untracked: code/LiftorthoPAS.err
Untracked: code/LiftorthoPASt.out
Untracked: code/Log.out
Untracked: code/Rev_liftoverPAShg19to38.err
Untracked: code/Rev_liftoverPAShg19to38.out
Untracked: code/SAF215upbed_gen.py
Untracked: code/Snakefile
Untracked: code/SnakefilePAS
Untracked: code/SnakefilePASfilt
Untracked: code/Upstream10Bases_general.py
Untracked: code/apaQTLsnake.err
Untracked: code/apaQTLsnake.out
Untracked: code/apaQTLsnakePAS.err
Untracked: code/apaQTLsnakePAS.out
Untracked: code/apaQTLsnakePAShuman.err
Untracked: code/bed215upbed.py
Untracked: code/bed2SAF_gen.py
Untracked: code/bed2saf.py
Untracked: code/bg_to_cov.py
Untracked: code/buildStarIndex.sh
Untracked: code/callPeaksYL.py
Untracked: code/chooseAnno2Bed.py
Untracked: code/chooseAnno2SAF.py
Untracked: code/cleanbed2saf.py
Untracked: code/cluster.json
Untracked: code/clusterPAS.json
Untracked: code/clusterfiltPAS.json
Untracked: code/convertNumeric.py
Untracked: code/extraSnakefiltpas
Untracked: code/filter5perc.R
Untracked: code/filter5percPheno.py
Untracked: code/filterBamforMP.pysam2_gen.py
Untracked: code/filterMissprimingInNuc10_gen.py
Untracked: code/filterPASforMP.py
Untracked: code/filterPostLift.py
Untracked: code/filterSAFforMP_gen.py
Untracked: code/filterSortBedbyCleanedBed_gen.R
Untracked: code/filterpeaks.py
Untracked: code/fixFChead.py
Untracked: code/fixFChead_bothfrac.py
Untracked: code/fixUTRexonanno.py
Untracked: code/formathg38Anno.py
Untracked: code/generateStarIndex.err
Untracked: code/generateStarIndex.out
Untracked: code/intersectAnno.err
Untracked: code/intersectAnno.out
Untracked: code/intersectLiftedPAS.sh
Untracked: code/liftPAS19to38.sh
Untracked: code/liftoverPAShg19to38.err
Untracked: code/liftoverPAShg19to38.out
Untracked: code/log/
Untracked: code/make5percPeakbed.py
Untracked: code/makeFileID.py
Untracked: code/makePheno.py
Untracked: code/makeSamplyGroupsChimp_TvN.py
Untracked: code/makeSamplyGroupsHuman_TvN.py
Untracked: code/maphg19.err
Untracked: code/maphg19.out
Untracked: code/maphg19.sh
Untracked: code/maphg19_sub.err
Untracked: code/maphg19_sub.out
Untracked: code/maphg19_subjunc.sh
Untracked: code/namePeaks.py
Untracked: code/overlapPAS.err
Untracked: code/overlapPAS.out
Untracked: code/overlapapaQTLPAS.sh
Untracked: code/peak2PAS.py
Untracked: code/pheno2countonly.R
Untracked: code/prepareCleanLiftedFC_5perc4LC.py
Untracked: code/preparePAS4lift.py
Untracked: code/primaryLift.err
Untracked: code/primaryLift.out
Untracked: code/primaryLift.sh
Untracked: code/quantLiftedPAS.err
Untracked: code/quantLiftedPAS.out
Untracked: code/quantLiftedPAS.sh
Untracked: code/recChimpback2Human.err
Untracked: code/recChimpback2Human.out
Untracked: code/recLiftchim2human.sh
Untracked: code/revLift.err
Untracked: code/revLift.out
Untracked: code/revLiftPAShg38to19.sh
Untracked: code/reverseLift.sh
Untracked: code/runChimpDiffIso.sh
Untracked: code/runHumanDiffIso.sh
Untracked: code/run_Chimpleafcutter_ds.err
Untracked: code/run_Chimpleafcutter_ds.out
Untracked: code/run_Humanleafcutter_ds.err
Untracked: code/run_Humanleafcutter_ds.out
Untracked: code/slurm-62824013.out
Untracked: code/slurm-62825841.out
Untracked: code/slurm-62826116.out
Untracked: code/snakePASChimp.err
Untracked: code/snakePASChimp.out
Untracked: code/snakePAShuman.out
Untracked: code/snakemake.batch
Untracked: code/snakemakePAS.batch
Untracked: code/snakemakePASFiltChimp.err
Untracked: code/snakemakePASFiltChimp.out
Untracked: code/snakemakePASFiltHuman.err
Untracked: code/snakemakePASFiltHuman.out
Untracked: code/snakemakePASchimp.batch
Untracked: code/snakemakePAShuman.batch
Untracked: code/snakemake_chimp.batch
Untracked: code/snakemake_human.batch
Untracked: code/snakemakefiltPAS.batch
Untracked: code/snakemakefiltPAS_chimp.sh
Untracked: code/snakemakefiltPAS_human.sh
Untracked: code/submit-snakemake-chimp.sh
Untracked: code/submit-snakemake-human.sh
Untracked: code/submit-snakemakePAS-chimp.sh
Untracked: code/submit-snakemakePAS-human.sh
Untracked: code/submit-snakemakefiltPAS-chimp.sh
Untracked: code/submit-snakemakefiltPAS-human.sh
Untracked: code/subset_diffisopheno.py
Untracked: code/subset_diffisopheno_Chimp_tvN.py
Untracked: code/subset_diffisopheno_Huma_tvN.py
Untracked: data/._metadata_HCpanel.txt
Untracked: data/._metadata_HCpanel.txt.sb-f4823d1e-qihGek
Untracked: data/._metadata_HCpanel.xlsx
Untracked: data/CompapaQTLpas/
Untracked: data/Peaks_5perc/
Untracked: data/Pheno_5perc/
Untracked: data/chainFiles/
Untracked: data/cleanPeaks_anno/
Untracked: data/cleanPeaks_byspecies/
Untracked: data/cleanPeaks_lifted/
Untracked: data/liftover_files/
Untracked: data/metadata_HCpanel.txt
Untracked: data/metadata_HCpanel.xlsx
Untracked: data/primaryLift/
Untracked: data/reverseLift/
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the R Markdown and HTML files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view them.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 5759e7e | brimittleman | 2019-10-07 | fix name |
| html | bf8c7fc | brimittleman | 2019-10-07 | Build site. |
| Rmd | 4263687 | brimittleman | 2019-10-07 | fix graph label |
| html | 62b5285 | brimittleman | 2019-10-07 | Build site. |
| Rmd | 8466418 | brimittleman | 2019-10-07 | add human and chimp tvN res |
| html | c3aa529 | brimittleman | 2019-10-07 | Build site. |
| Rmd | da85ad8 | brimittleman | 2019-10-07 | add code to prepare human and chimp TvN |
library(tidyverse)── Attaching packages ───────────────────────────────────────────────────────────── tidyverse 1.2.1 ──✔ ggplot2 3.1.1 ✔ purrr 0.3.2
✔ tibble 2.1.1 ✔ dplyr 0.8.0.1
✔ tidyr 0.8.3 ✔ stringr 1.3.1
✔ readr 1.3.1 ✔ forcats 0.3.0 ── Conflicts ──────────────────────────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()library(reshape2)
Attaching package: 'reshape2'The following object is masked from 'package:tidyr':
smithsIn this anaylsis I will complete the Chimp total vs nuclear analysis. This analysis is similar to the analysis in the apaQTL project
I need the human 5% both fraction feature counts.
Thee feature counts are in ../Chimp/data/CleanLiftedPeaks_FC/ALLPAS_postLift_LocParsed_Chimp_fixed.fc. I need to subset these for those in the annotations. Keep the PAS in this file: ../data/Peaks_5perc/Peaks_5perc_either_HumanCoordHummanUsage.bed. I will do this with a python script.
mkdir ../Chimp/data/CleanLiftedPeaks4LC/
python prepareCleanLiftedFC_5perc4LC.py ../Chimp/data/CleanLiftedPeaks_FC/ALLPAS_postLift_LocParsed_Chimp_fixed.fc ../data/Peaks_5perc/Peaks_5perc_either_HumanCoordHummanUsage.bed ../Chimp/data/CleanLiftedPeaks4LC/ALLPAS_postLift_LocParsed_Chimp_fixed4LC.fcThis will only look at PAS on chromosomes 1-22 no extra haplotpyes.
mkdir ../Chimp/data/DiffIso_Chimp/
python subset_diffisopheno_Chimp_tvN.py 1
python subset_diffisopheno_Chimp_tvN.py 2
python subset_diffisopheno_Chimp_tvN.py 3
python subset_diffisopheno_Chimp_tvN.py 4
python subset_diffisopheno_Chimp_tvN.py 5
python subset_diffisopheno_Chimp_tvN.py 6
python subset_diffisopheno_Chimp_tvN.py 7
python subset_diffisopheno_Chimp_tvN.py 8
python subset_diffisopheno_Chimp_tvN.py 9
python subset_diffisopheno_Chimp_tvN.py 10
python subset_diffisopheno_Chimp_tvN.py 11
python subset_diffisopheno_Chimp_tvN.py 12
python subset_diffisopheno_Chimp_tvN.py 13
python subset_diffisopheno_Chimp_tvN.py 14
python subset_diffisopheno_Chimp_tvN.py 15
python subset_diffisopheno_Chimp_tvN.py 16
python subset_diffisopheno_Chimp_tvN.py 18
python subset_diffisopheno_Chimp_tvN.py 19
python subset_diffisopheno_Chimp_tvN.py 20
python subset_diffisopheno_Chimp_tvN.py 21
python subset_diffisopheno_Chimp_tvN.py 22Make sample groups:
python makeSamplyGroupsChimp_TvN.pyRun Leafcutter:
sbatch runChimpDiffIso.shConcatinate results:
awk '{if(NR>1)print}' ../Chimp/data/DiffIso_Chimp/TN_diff_isoform_chr*.txt_effect_sizes.txt > ../Chimp/data/DiffIso_Chimp/TN_diff_isoform_allChrom.txt_effect_sizes.txt
awk '{if(NR>1)print}' ../Chimp/data/DiffIso_Chimp/TN_diff_isoform_chr*.txt_cluster_significance.txt > ../Chimp/data/DiffIso_Chimp/TN_diff_isoform_allChrom.txt_significance.txt
Significant clusters:
sig=read.table("../Chimp/data/DiffIso_Chimp/TN_diff_isoform_allChrom.txt_significance.txt",sep="\t" ,col.names = c('status','loglr','df','p','cluster','p.adjust'),stringsAsFactors = F) %>% filter(status=="Success")
sig$p.adjust=as.numeric(as.character(sig$p.adjust))qqplot(-log10(runif(nrow(sig))), -log10(sig$p.adjust),ylab="-log10 Total Adjusted Leafcutter pvalue", xlab="-log 10 Uniform expectation", main="Chimp: Leafcutter differencial isoform analysis between fractions")
abline(0,1)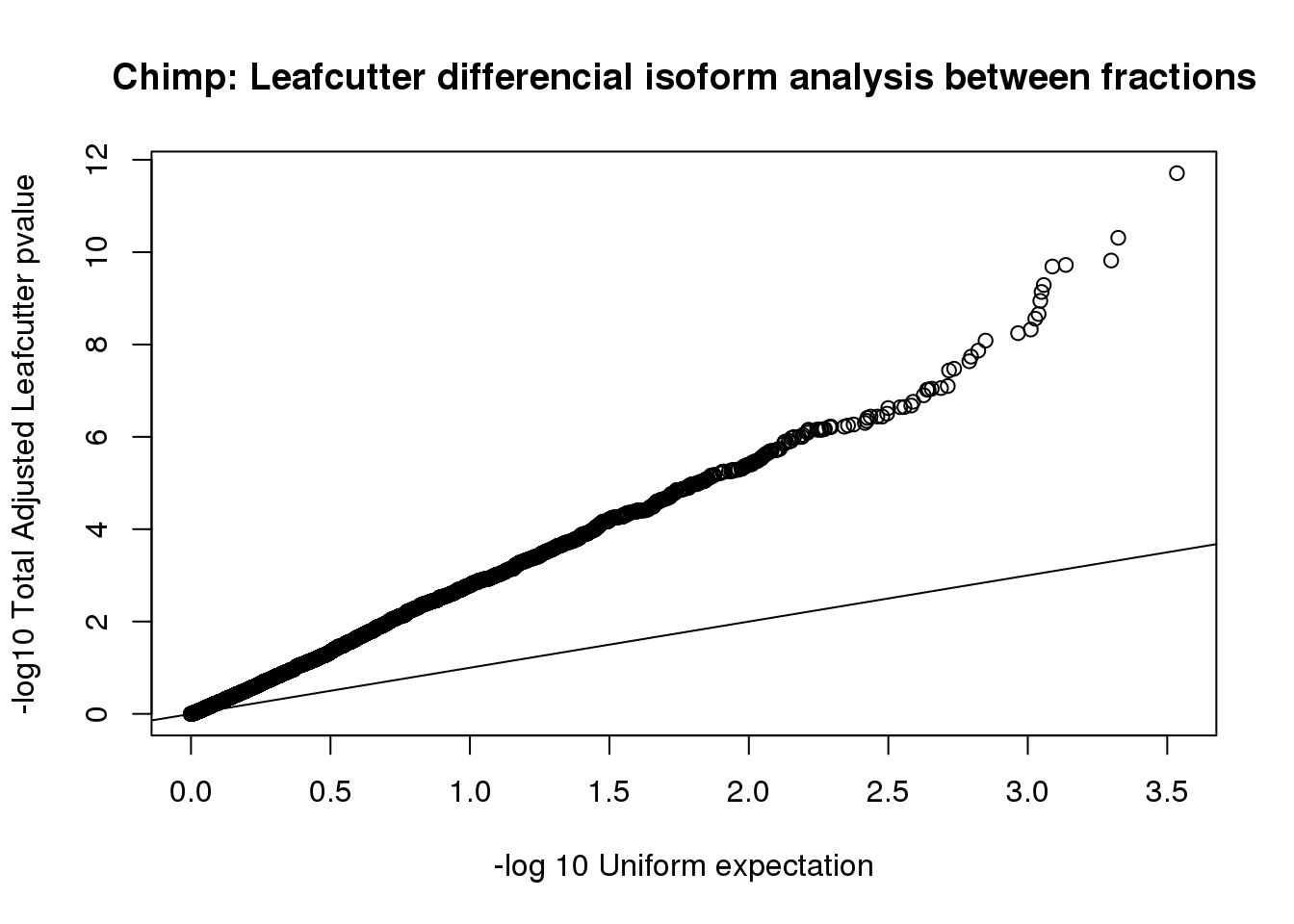
| Version | Author | Date |
|---|---|---|
| 62b5285 | brimittleman | 2019-10-07 |
tested_genes=nrow(sig)
tested_genes[1] 7571sig_genes=sig %>% filter(p.adjust<.05)
number_sig_genes=nrow(sig_genes)
number_sig_genes[1] 2528Effect Sizes
effectsize=read.table("../Chimp/data/DiffIso_Chimp/TN_diff_isoform_allChrom.txt_effect_sizes.txt", stringsAsFactors = F, col.names=c('intron', 'logef' ,'Nuclear', 'Total','deltaPAU')) %>% filter(intron != "intron")
effectsize$deltaPAU=as.numeric(as.character(effectsize$deltaPAU))
effectsize$logef=as.numeric(as.character(effectsize$logef))plot(sort(effectsize$deltaPAU),main="Chimp: Leafcutter delta PAU", ylab="Delta PAU", xlab="PAS Index")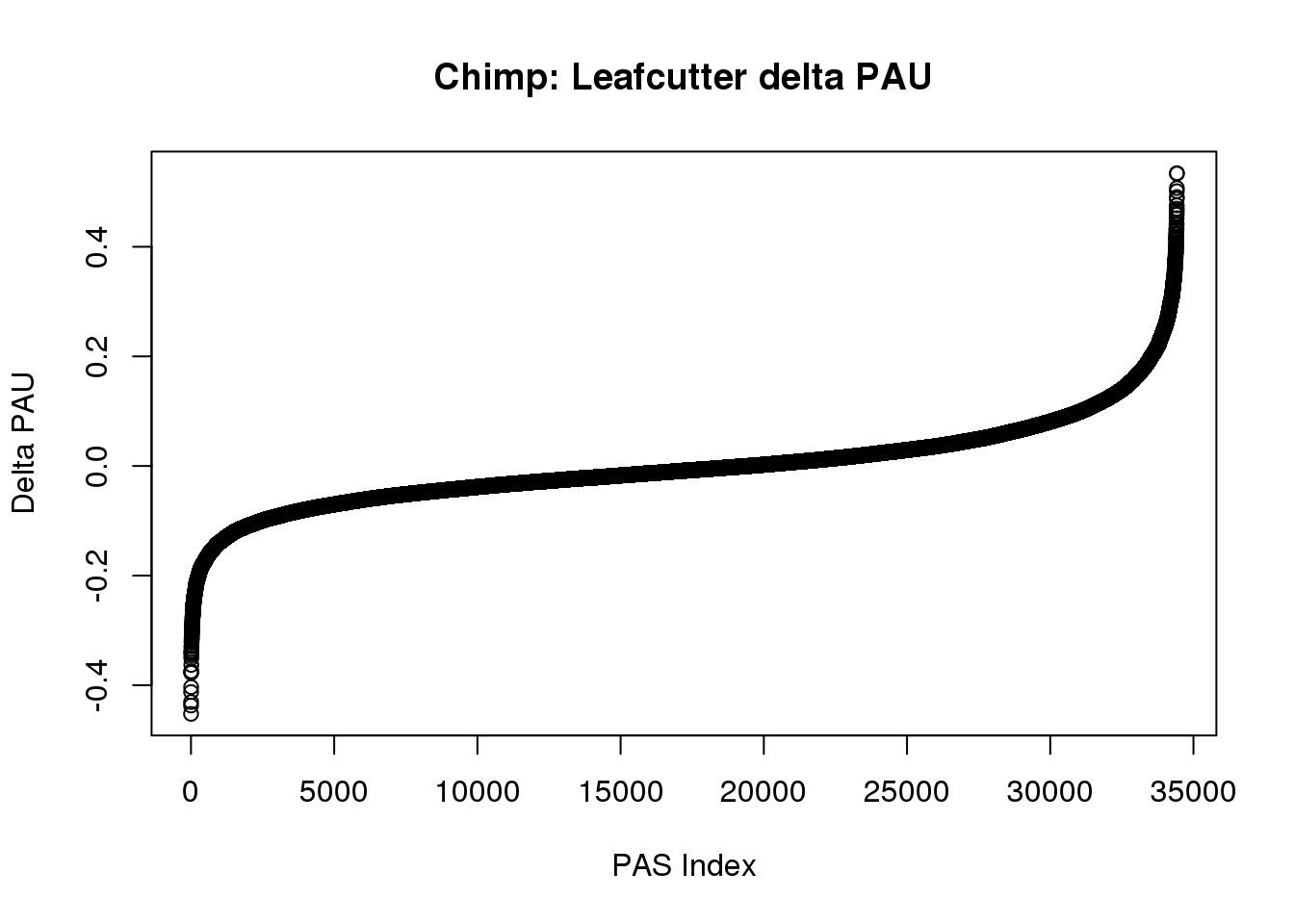
| Version | Author | Date |
|---|---|---|
| 62b5285 | brimittleman | 2019-10-07 |
effectsize_deltaPAU= effectsize %>% filter(abs(deltaPAU) > .2)
nrow(effectsize_deltaPAU)[1] 1153effectSize_highdiffGenes=effectsize_deltaPAU %>% separate(intron, into=c("chrom", "start", "end", "GeneName"), sep=":") %>% dplyr::select(GeneName) %>% unique()effectsize_deltaPAU_Genes= effectsize_deltaPAU %>% separate(intron, into=c("chrom", "start", "end","gene"),sep=":") %>% group_by(gene) %>% summarise(nperGene=n())
nrow(effectsize_deltaPAU_Genes)[1] 976Pull in the PAS to see where these PAS are located.
PAS=read.table("../data/Peaks_5perc/Peaks_5perc_either_bothUsage.txt", stringsAsFactors = F, header = T) %>% filter(loc !="008559")
PAS$start=as.character(PAS$start)
PAS$end=as.character(PAS$end)Increased usage in nuclear:
effectsize_deltaPAU_nuclear= effectsize_deltaPAU %>% filter(deltaPAU < -0.2) %>% separate(intron,into=c("chr", "start", "end", "gene"), sep=":" )%>% inner_join(PAS, by=c("gene", "chr","start", "end"))ggplot(effectsize_deltaPAU_nuclear,aes(x=loc,fill=loc)) + geom_histogram(stat="count") + scale_fill_brewer(palette = "Dark2") +labs(title="Chimp: Location of PAS used more in Nuclear")Warning: Ignoring unknown parameters: binwidth, bins, pad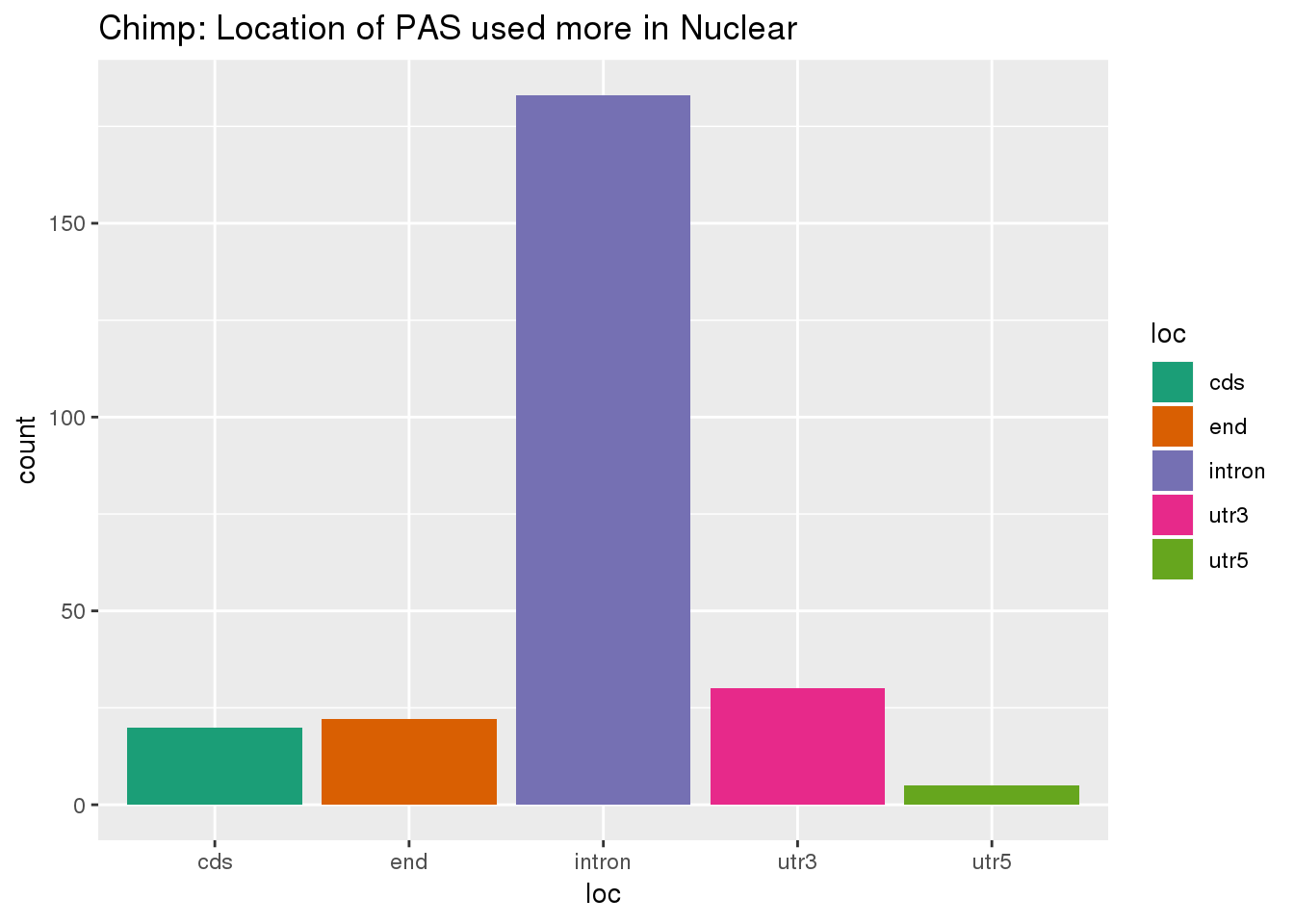
Increased usage in total:
effectsize_deltaPAU_total= effectsize_deltaPAU %>% filter(deltaPAU > 0.2) %>% separate(intron,into=c("chr", "start", "end", "gene"), sep=":" )%>% inner_join(PAS, by=c("gene", "chr","start", "end"))ggplot(effectsize_deltaPAU_total,aes(x=loc,fill=loc)) + geom_histogram(stat="count") + scale_fill_brewer(palette = "Dark2") +labs(title="Chimp: Location of PAS used more in Total")Warning: Ignoring unknown parameters: binwidth, bins, pad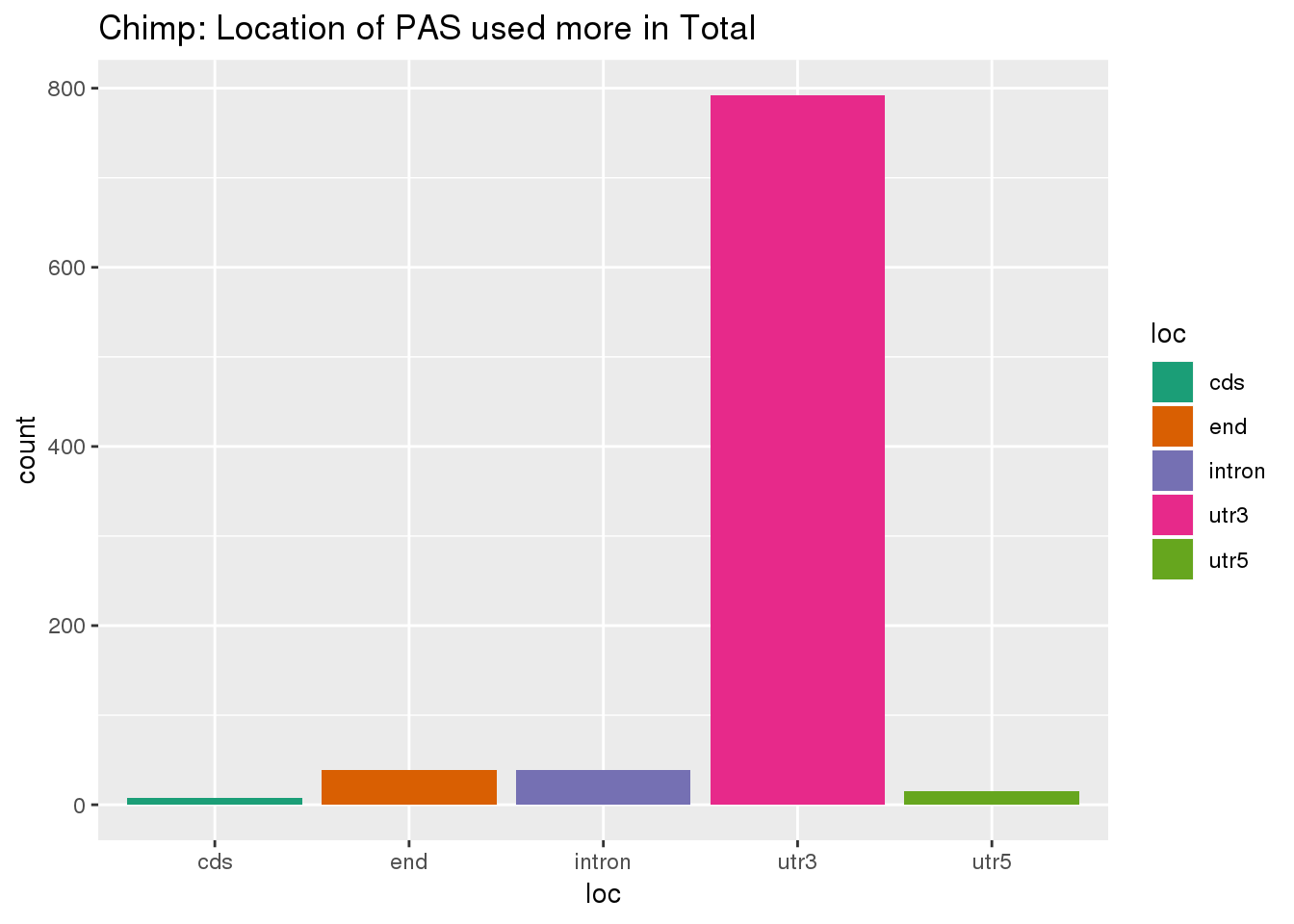
This is in line with expectation. Now I can compare this to the number of discovered PAS by plotting the proportion of PAS used more in each fraction by location.
PAS_loc =PAS%>% group_by(loc) %>% summarise(nloc=n())effectsize_deltaPAU_nuclear_loc=effectsize_deltaPAU_nuclear %>% group_by(loc) %>% summarise(nlocNuclear=n())
effectsize_deltaPAU_total_loc=effectsize_deltaPAU_total %>% group_by(loc) %>% summarise(nloctotal=n())
effectsize_deltaPAUProp_tot=effectsize_deltaPAU_total_loc %>% inner_join(PAS_loc, by="loc") %>% mutate(Proportion_tot=nloctotal/nloc)
effectsize_deltaPAUProp_nuc=effectsize_deltaPAU_nuclear_loc %>% inner_join(PAS_loc, by="loc") %>% mutate(Proportion_nuc=nlocNuclear/nloc)
effectsize_deltaPAUProp_both= effectsize_deltaPAUProp_nuc %>% inner_join(effectsize_deltaPAUProp_tot, by=c("loc","nloc")) %>% dplyr::rename(Nuclear=Proportion_nuc, Total=Proportion_tot) %>% select(loc, Nuclear, Total)
effectsize_deltaPAUProp_both_melt= effectsize_deltaPAUProp_both %>% melt(id.vars="loc", variable.name="Fraction", value.name = "Proportion")
effectsize_deltaPAUProp_both_melt$Fraction=as.character(effectsize_deltaPAUProp_both_melt$Fraction)Plot this:
ggplot(effectsize_deltaPAUProp_both_melt, aes(x=loc, y=Proportion, by=Fraction, fill=Fraction)) + geom_bar(stat="identity", position="dodge") + scale_fill_brewer(palette = "Dark2")+ labs(title="Chimp: Proportion of PAS differential used by location",x="") +scale_x_discrete(labels = c('Coding','5kb downstream','Intronic',"3' UTR", "5' UTR")) 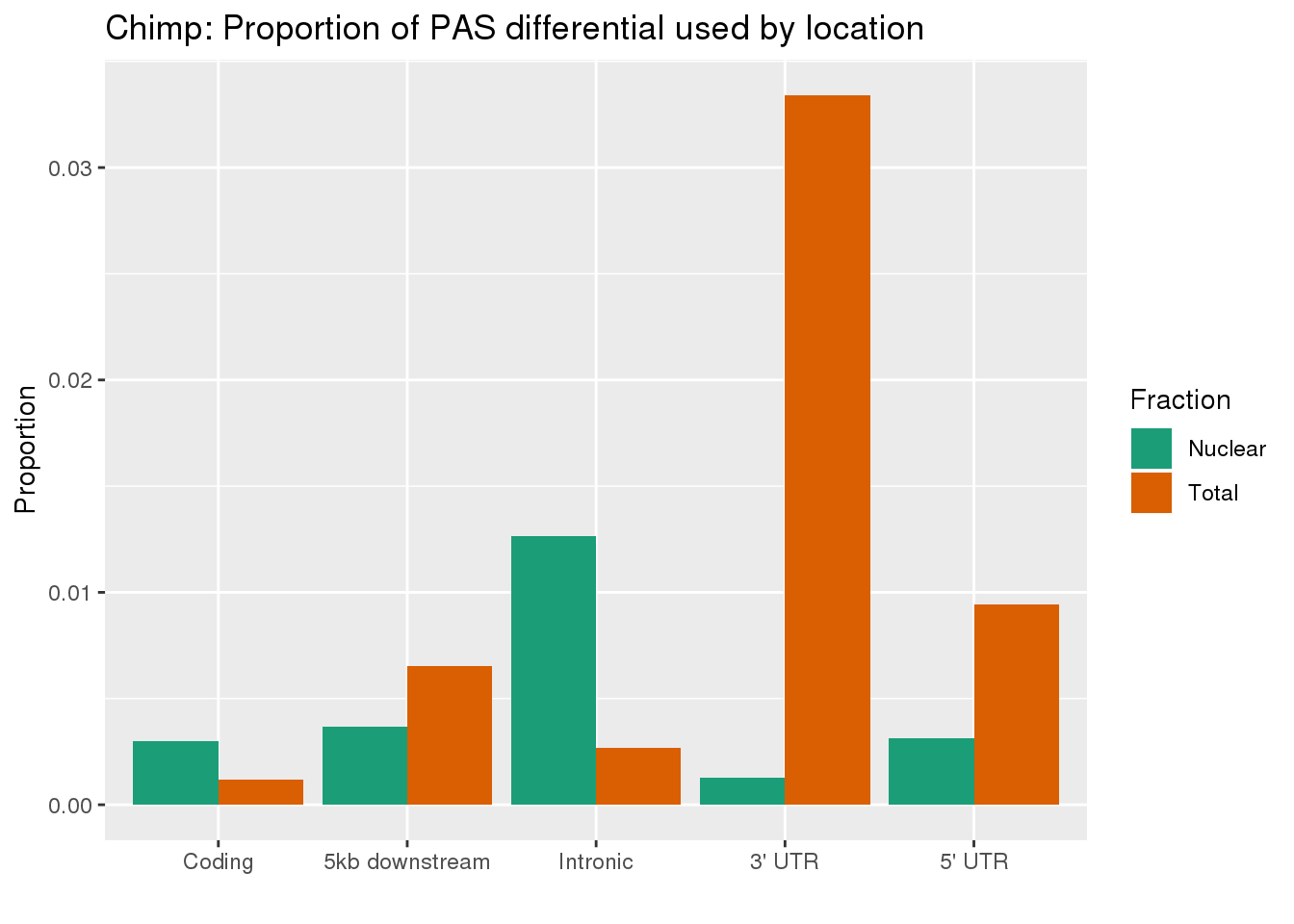
| Version | Author | Date |
|---|---|---|
| 62b5285 | brimittleman | 2019-10-07 |
sessionInfo()R version 3.5.1 (2018-07-02)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Scientific Linux 7.4 (Nitrogen)
Matrix products: default
BLAS/LAPACK: /software/openblas-0.2.19-el7-x86_64/lib/libopenblas_haswellp-r0.2.19.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] reshape2_1.4.3 forcats_0.3.0 stringr_1.3.1 dplyr_0.8.0.1
[5] purrr_0.3.2 readr_1.3.1 tidyr_0.8.3 tibble_2.1.1
[9] ggplot2_3.1.1 tidyverse_1.2.1
loaded via a namespace (and not attached):
[1] Rcpp_1.0.2 RColorBrewer_1.1-2 cellranger_1.1.0
[4] pillar_1.3.1 compiler_3.5.1 git2r_0.25.2
[7] plyr_1.8.4 workflowr_1.4.0 tools_3.5.1
[10] digest_0.6.18 lubridate_1.7.4 jsonlite_1.6
[13] evaluate_0.12 nlme_3.1-137 gtable_0.2.0
[16] lattice_0.20-38 pkgconfig_2.0.2 rlang_0.4.0
[19] cli_1.1.0 rstudioapi_0.10 yaml_2.2.0
[22] haven_1.1.2 withr_2.1.2 xml2_1.2.0
[25] httr_1.3.1 knitr_1.20 hms_0.4.2
[28] generics_0.0.2 fs_1.3.1 rprojroot_1.3-2
[31] grid_3.5.1 tidyselect_0.2.5 glue_1.3.0
[34] R6_2.3.0 readxl_1.1.0 rmarkdown_1.10
[37] modelr_0.1.2 magrittr_1.5 whisker_0.3-2
[40] backports_1.1.2 scales_1.0.0 htmltools_0.3.6
[43] rvest_0.3.2 assertthat_0.2.0 colorspace_1.3-2
[46] labeling_0.3 stringi_1.2.4 lazyeval_0.2.1
[49] munsell_0.5.0 broom_0.5.1 crayon_1.3.4