Differential Splicing
Briana Mittleman
11/11/2019
Last updated: 2019-11-18
Checks: 7 0
Knit directory: Comparative_APA/analysis/
This reproducible R Markdown analysis was created with workflowr (version 1.5.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20190902) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: code/chimp_log/
Ignored: code/human_log/
Ignored: data/RNASEQ_metadata.txt.sb-51f67ae1-HXp7Gq/
Ignored: data/metadata_HCpanel.txt.sb-a3d92a2d-b9cYoF/
Ignored: data/metadata_HCpanel.txt.sb-f4823d1e-qihGek/
Untracked files:
Untracked: ._.DS_Store
Untracked: Chimp/
Untracked: Human/
Untracked: analysis/assessReadQual.Rmd
Untracked: code/._Config_chimp.yaml
Untracked: code/._Config_human.yaml
Untracked: code/._DiffSplice.sh
Untracked: code/._LiftOrthoPAS2chimp.sh
Untracked: code/._Snakefile
Untracked: code/._SnakefilePAS
Untracked: code/._SnakefilePASfilt
Untracked: code/._bed215upbed.py
Untracked: code/._bed2SAF_gen.py
Untracked: code/._buildIndecpantro5
Untracked: code/._buildIndecpantro5.sh
Untracked: code/._buildStarIndex.sh
Untracked: code/._cleanbed2saf.py
Untracked: code/._cluster.json
Untracked: code/._cluster2bed.py
Untracked: code/._clusterLiftReverse.sh
Untracked: code/._clusterLiftprimary.sh
Untracked: code/._converBam2Junc.sh
Untracked: code/._extraSnakefiltpas
Untracked: code/._filter5percPAS.py
Untracked: code/._filterPASforMP.py
Untracked: code/._filterPostLift.py
Untracked: code/._fixExonFC.py
Untracked: code/._fixUTRexonanno.py
Untracked: code/._formathg38Anno.py
Untracked: code/._formatpantro6Anno.py
Untracked: code/._intersectLiftedPAS.sh
Untracked: code/._liftPAS19to38.sh
Untracked: code/._makeSamplyGroupsHuman_TvN.py
Untracked: code/._maphg19.sh
Untracked: code/._maphg19_subjunc.sh
Untracked: code/._mergedBam2BW.sh
Untracked: code/._nameClusters.py
Untracked: code/._overlapapaQTLPAS.sh
Untracked: code/._prepareCleanLiftedFC_5perc4LC.py
Untracked: code/._preparePAS4lift.py
Untracked: code/._primaryLift.sh
Untracked: code/._processhg38exons.py
Untracked: code/._quantJunc.sh
Untracked: code/._recLiftchim2human.sh
Untracked: code/._revLiftPAShg38to19.sh
Untracked: code/._reverseLift.sh
Untracked: code/._runChimpDiffIso.sh
Untracked: code/._runHumanDiffIso.sh
Untracked: code/._runNuclearDifffIso.sh
Untracked: code/._run_chimpverifybam.sh
Untracked: code/._run_verifyBam.sh
Untracked: code/._snakemake.batch
Untracked: code/._snakemakePAS.batch
Untracked: code/._snakemakePASchimp.batch
Untracked: code/._snakemakePAShuman.batch
Untracked: code/._snakemake_chimp.batch
Untracked: code/._snakemake_human.batch
Untracked: code/._snakemakefiltPAS.batch
Untracked: code/._snakemakefiltPAS_chimp
Untracked: code/._snakemakefiltPAS_chimp.sh
Untracked: code/._snakemakefiltPAS_human.sh
Untracked: code/._submit-snakemake-chimp.sh
Untracked: code/._submit-snakemake-human.sh
Untracked: code/._submit-snakemakePAS-chimp.sh
Untracked: code/._submit-snakemakePAS-human.sh
Untracked: code/._submit-snakemakefiltPAS-chimp.sh
Untracked: code/._submit-snakemakefiltPAS-human.sh
Untracked: code/._subset_diffisopheno_Nuclear_HvC.py
Untracked: code/._transcriptDTplotsNuclear.sh
Untracked: code/._verifyBam4973.sh
Untracked: code/._wrap_chimpverifybam.sh
Untracked: code/._wrap_verifyBam.sh
Untracked: code/.snakemake/
Untracked: code/Config_chimp.yaml
Untracked: code/Config_human.yaml
Untracked: code/DiffSplice.err
Untracked: code/DiffSplice.out
Untracked: code/DiffSplice.sh
Untracked: code/LiftClustersFirst.err
Untracked: code/LiftClustersFirst.out
Untracked: code/LiftClustersSecond.err
Untracked: code/LiftClustersSecond.out
Untracked: code/LiftOrthoPAS2chimp.sh
Untracked: code/LiftorthoPAS.err
Untracked: code/LiftorthoPASt.out
Untracked: code/Log.out
Untracked: code/Rev_liftoverPAShg19to38.err
Untracked: code/Rev_liftoverPAShg19to38.out
Untracked: code/SAF215upbed_gen.py
Untracked: code/Snakefile
Untracked: code/SnakefilePAS
Untracked: code/SnakefilePASfilt
Untracked: code/TotalTranscriptDTplot.err
Untracked: code/TotalTranscriptDTplot.out
Untracked: code/Upstream10Bases_general.py
Untracked: code/apaQTLsnake.err
Untracked: code/apaQTLsnake.out
Untracked: code/apaQTLsnakePAS.err
Untracked: code/apaQTLsnakePAS.out
Untracked: code/apaQTLsnakePAShuman.err
Untracked: code/bam2junc.err
Untracked: code/bam2junc.out
Untracked: code/bed215upbed.py
Untracked: code/bed2SAF_gen.py
Untracked: code/bed2saf.py
Untracked: code/bg_to_cov.py
Untracked: code/buildIndecpantro5
Untracked: code/buildIndecpantro5.sh
Untracked: code/buildStarIndex.sh
Untracked: code/callPeaksYL.py
Untracked: code/chooseAnno2Bed.py
Untracked: code/chooseAnno2SAF.py
Untracked: code/cleanbed2saf.py
Untracked: code/cluster.json
Untracked: code/cluster2bed.py
Untracked: code/clusterLiftReverse.sh
Untracked: code/clusterLiftprimary.sh
Untracked: code/clusterPAS.json
Untracked: code/clusterfiltPAS.json
Untracked: code/converBam2Junc.sh
Untracked: code/convertNumeric.py
Untracked: code/environment.yaml
Untracked: code/extraSnakefiltpas
Untracked: code/filter5perc.R
Untracked: code/filter5percPAS.py
Untracked: code/filter5percPheno.py
Untracked: code/filterBamforMP.pysam2_gen.py
Untracked: code/filterMissprimingInNuc10_gen.py
Untracked: code/filterPASforMP.py
Untracked: code/filterPostLift.py
Untracked: code/filterSAFforMP_gen.py
Untracked: code/filterSortBedbyCleanedBed_gen.R
Untracked: code/filterpeaks.py
Untracked: code/fixExonFC.py
Untracked: code/fixFChead.py
Untracked: code/fixFChead_bothfrac.py
Untracked: code/fixUTRexonanno.py
Untracked: code/formathg38Anno.py
Untracked: code/generateStarIndex.err
Untracked: code/generateStarIndex.out
Untracked: code/generateStarIndexHuman.err
Untracked: code/generateStarIndexHuman.out
Untracked: code/intersectAnno.err
Untracked: code/intersectAnno.out
Untracked: code/intersectLiftedPAS.sh
Untracked: code/liftPAS19to38.sh
Untracked: code/liftoverPAShg19to38.err
Untracked: code/liftoverPAShg19to38.out
Untracked: code/log/
Untracked: code/make5percPeakbed.py
Untracked: code/makeFileID.py
Untracked: code/makePheno.py
Untracked: code/makeSamplyGroupsChimp_TvN.py
Untracked: code/makeSamplyGroupsHuman_TvN.py
Untracked: code/maphg19.err
Untracked: code/maphg19.out
Untracked: code/maphg19.sh
Untracked: code/maphg19_sub.err
Untracked: code/maphg19_sub.out
Untracked: code/maphg19_subjunc.sh
Untracked: code/mergedBam2BW.sh
Untracked: code/mergedbam2bw.err
Untracked: code/mergedbam2bw.out
Untracked: code/nameClusters.py
Untracked: code/namePeaks.py
Untracked: code/nuclearTranscriptDTplot.err
Untracked: code/nuclearTranscriptDTplot.out
Untracked: code/overlapPAS.err
Untracked: code/overlapPAS.out
Untracked: code/overlapapaQTLPAS.sh
Untracked: code/peak2PAS.py
Untracked: code/pheno2countonly.R
Untracked: code/prepareCleanLiftedFC_5perc4LC.py
Untracked: code/preparePAS4lift.py
Untracked: code/prepare_phenotype_table.py
Untracked: code/primaryLift.err
Untracked: code/primaryLift.out
Untracked: code/primaryLift.sh
Untracked: code/processhg38exons.py
Untracked: code/quantJunc.sh
Untracked: code/quantLiftedPAS.err
Untracked: code/quantLiftedPAS.out
Untracked: code/quantLiftedPAS.sh
Untracked: code/quatJunc.err
Untracked: code/quatJunc.out
Untracked: code/recChimpback2Human.err
Untracked: code/recChimpback2Human.out
Untracked: code/recLiftchim2human.sh
Untracked: code/revLift.err
Untracked: code/revLift.out
Untracked: code/revLiftPAShg38to19.sh
Untracked: code/reverseLift.sh
Untracked: code/runChimpDiffIso.sh
Untracked: code/runHumanDiffIso.sh
Untracked: code/runNuclearDifffIso.sh
Untracked: code/run_Chimpleafcutter_ds.err
Untracked: code/run_Chimpleafcutter_ds.out
Untracked: code/run_Chimpverifybam.err
Untracked: code/run_Chimpverifybam.out
Untracked: code/run_Humanleafcutter_ds.err
Untracked: code/run_Humanleafcutter_ds.out
Untracked: code/run_Nuclearleafcutter_ds.err
Untracked: code/run_Nuclearleafcutter_ds.out
Untracked: code/run_chimpverifybam.sh
Untracked: code/run_verifyBam.sh
Untracked: code/run_verifybam.err
Untracked: code/run_verifybam.out
Untracked: code/slurm-62824013.out
Untracked: code/slurm-62825841.out
Untracked: code/slurm-62826116.out
Untracked: code/snakePASChimp.err
Untracked: code/snakePASChimp.out
Untracked: code/snakePAShuman.out
Untracked: code/snakemake.batch
Untracked: code/snakemakeChimp.err
Untracked: code/snakemakeChimp.out
Untracked: code/snakemakeHuman.err
Untracked: code/snakemakeHuman.out
Untracked: code/snakemakePAS.batch
Untracked: code/snakemakePASFiltChimp.err
Untracked: code/snakemakePASFiltChimp.out
Untracked: code/snakemakePASFiltHuman.err
Untracked: code/snakemakePASFiltHuman.out
Untracked: code/snakemakePASchimp.batch
Untracked: code/snakemakePAShuman.batch
Untracked: code/snakemake_chimp.batch
Untracked: code/snakemake_human.batch
Untracked: code/snakemakefiltPAS.batch
Untracked: code/snakemakefiltPAS_chimp.sh
Untracked: code/snakemakefiltPAS_human.sh
Untracked: code/submit-snakemake-chimp.sh
Untracked: code/submit-snakemake-human.sh
Untracked: code/submit-snakemakePAS-chimp.sh
Untracked: code/submit-snakemakePAS-human.sh
Untracked: code/submit-snakemakefiltPAS-chimp.sh
Untracked: code/submit-snakemakefiltPAS-human.sh
Untracked: code/subset_diffisopheno.py
Untracked: code/subset_diffisopheno_Chimp_tvN.py
Untracked: code/subset_diffisopheno_Huma_tvN.py
Untracked: code/subset_diffisopheno_Nuclear_HvC.py
Untracked: code/transcriptDTplotsNuclear.sh
Untracked: code/transcriptDTplotsTotal.sh
Untracked: code/verifyBam4973.sh
Untracked: code/verifybam4973.err
Untracked: code/verifybam4973.out
Untracked: code/wrap_Chimpverifybam.err
Untracked: code/wrap_Chimpverifybam.out
Untracked: code/wrap_chimpverifybam.sh
Untracked: code/wrap_verifyBam.sh
Untracked: code/wrap_verifybam.err
Untracked: code/wrap_verifybam.out
Untracked: data/._RNASEQ_metadata.txt
Untracked: data/._RNASEQ_metadata.txt.sb-51f67ae1-HXp7Gq
Untracked: data/._RNASEQ_metadata.xlsx
Untracked: data/._metadata_HCpanel.txt
Untracked: data/._metadata_HCpanel.txt.sb-a3d92a2d-b9cYoF
Untracked: data/._metadata_HCpanel.txt.sb-f4823d1e-qihGek
Untracked: data/._metadata_HCpanel.xlsx
Untracked: data/._~$RNASEQ_metadata.xlsx
Untracked: data/._~$metadata_HCpanel.xlsx
Untracked: data/CompapaQTLpas/
Untracked: data/DTmatrix/
Untracked: data/DiffIso_Nuclear/
Untracked: data/DiffSplice/
Untracked: data/MapStats/
Untracked: data/NuclearHvC/
Untracked: data/Peaks_5perc/
Untracked: data/Pheno_5perc/
Untracked: data/Pheno_5perc_nuclear/
Untracked: data/Pheno_5perc_total/
Untracked: data/RNASEQ_metadata.txt
Untracked: data/RNASEQ_metadata.xlsx
Untracked: data/chainFiles/
Untracked: data/cleanPeaks_anno/
Untracked: data/cleanPeaks_byspecies/
Untracked: data/cleanPeaks_lifted/
Untracked: data/liftover_files/
Untracked: data/metadata_HCpanel.txt
Untracked: data/metadata_HCpanel.xlsx
Untracked: data/primaryLift/
Untracked: data/reverseLift/
Untracked: data/~$RNASEQ_metadata.xlsx
Untracked: data/~$metadata_HCpanel.xlsx
Untracked: output/dtPlots/
Untracked: projectNotes.Rmd
Unstaged changes:
Modified: analysis/CorrbetweenInd.Rmd
Modified: analysis/PASnumperSpecies.Rmd
Modified: analysis/annotationInfo.Rmd
Modified: analysis/verifyBAM.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the R Markdown and HTML files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view them.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | fa2c075 | brimittleman | 2019-11-18 | code for diff splice |
| html | 106f3c1 | brimittleman | 2019-11-15 | Build site. |
| Rmd | 4ad7bce | brimittleman | 2019-11-15 | look at results of cluster lift |
| html | dc91b0a | brimittleman | 2019-11-11 | Build site. |
| Rmd | b5ba82e | brimittleman | 2019-11-11 | add diff expression and diff splicing |
library(tidyverse)── Attaching packages ─────────────────────────────────────────────────────────────────────────────── tidyverse 1.2.1 ──✔ ggplot2 3.1.1 ✔ purrr 0.3.2
✔ tibble 2.1.1 ✔ dplyr 0.8.0.1
✔ tidyr 0.8.3 ✔ stringr 1.3.1
✔ readr 1.3.1 ✔ forcats 0.3.0 ── Conflicts ────────────────────────────────────────────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()library(reshape2)
Attaching package: 'reshape2'The following object is masked from 'package:tidyr':
smithsI want to use the RNA seq I collected to also perform a differential splicing analysis with leafcutter. I will follow the pipeline found at http://davidaknowles.github.io/leafcutter/articles/Usage.html. For a first pass I will use the bam files from the snakemake and differential expression analysis pipeline.
I will get clusters in both species then perform reciprocal liftover. I can use a liftover pipeline similar to the one I used for the differnetial PAS analysis.
Pipeline from example on leafcutter github.
for bamfile in `ls run/geuvadis/*chr1.bam`; do
echo Converting $bamfile to $bamfile.junc
samtools index $bamfile
regtools junctions extract -a 8 -m 50 -M 500000 $bamfile -o $bamfile.junc
echo $bamfile.junc >> test_juncfiles.txt
done
python ../clustering/leafcutter_cluster_regtools.py -j test_juncfiles.txt -m 50 -o testYRIvsEU -l 500000At this point I will be able to liftover the junctions. I can use the human corrdinates for the differential splicing step.
../scripts/leafcutter_ds.R --num_threads 4 ../example_data/testYRIvsEU_perind_numers.counts.gz example_geuvadis/groups_file.txt
I now have my RNA seq for each species. I can write a script that runs the junctions for each species.
sbatch converBam2Junc.shsbatch quantJunc.shNow I need to do reciprocal liftover with the clusters.
- /project2/gilad/briana/Comparative_APA/Human/data/RNAseq/DiffSplice/
- /project2/gilad/briana/Comparative_APA/Chimp/data/RNAseq/DiffSplice/
Chain files are in /data/chainFiles/ * panTro5ToHg38.over.chain
* hg38ToPanTro5.over.chain
I first need to make bedfiles with these clusters.
The clusters all have _NA I dont this this is correct.
gunzip ../Human/data/RNAseq/DiffSplice/humanJunc_perind.counts.gz
gunzip ../Chimp/data/RNAseq/DiffSplice/chimpJunc_perind.counts.gz
python cluster2bed.py ../Human/data/RNAseq/DiffSplice/humanJunc_perind.counts ../Human/data/RNAseq/DiffSplice/humanJunc.bed
python cluster2bed.py ../Chimp/data/RNAseq/DiffSplice/chimpJunc_perind.counts ../Chimp/data/RNAseq/DiffSplice/chimpJunc.bed
#I need to name the clusters before I can do the lift. (this is like the naming in apa 1-n(clusters))
python nameClusters.py ../Human/data/RNAseq/DiffSplice/humanJunc.bed ../Human/data/RNAseq/DiffSplice/humanJuncNamed.bed
python nameClusters.py ../Chimp/data/RNAseq/DiffSplice/chimpJunc.bed ../Chimp/data/RNAseq/DiffSplice/chimpJuncNamed.bed
sbatch clusterLiftprimary.sh
sbatch clusterLiftReverse.shEvaluate results:
(this code is from the lift for the PAS)
unliftedH=read.table("../Human/data/RNAseq/DiffSplice/humanJunc_unlifted.bed",stringsAsFactors = F) %>% nrow()
unliftedC=read.table("../Chimp/data/RNAseq/DiffSplice/chimpJunc_unlifted.bed",stringsAsFactors = F) %>% nrow()
liftedH=read.table("../Human/data/RNAseq/DiffSplice/humanJunc_inChimp.bed",stringsAsFactors = F) %>% nrow()
liftedC=read.table("../Chimp/data/RNAseq/DiffSplice/chimpJunc_inHuman.bed",stringsAsFactors = F) %>% nrow()
primaryUnC=c("Chimp","Unlifted", unliftedC)
primaryUnH=c("Human","Unlifted", unliftedH)
primaryLH=c("Human","Lifted", liftedH)
primaryLC=c("Chimp","Lifted", liftedC)
header=c("species", "liftStat", "PAS")
primaryDF= as.data.frame(rbind(primaryLH,primaryLC, primaryUnH,primaryUnC))
colnames(primaryDF)=header
primaryDF$PAS=as.numeric(as.character(primaryDF$PAS))
primaryDF= primaryDF %>% group_by(species) %>% mutate(nPAS=sum(PAS)) %>% ungroup() %>% mutate(proportion=PAS/nPAS)ggplot(primaryDF,aes(x=species, y=PAS, fill=liftStat)) + geom_bar(stat="identity",position = "dodge") + scale_fill_brewer(palette = "Dark2") + labs(title="Primary Liftover Results")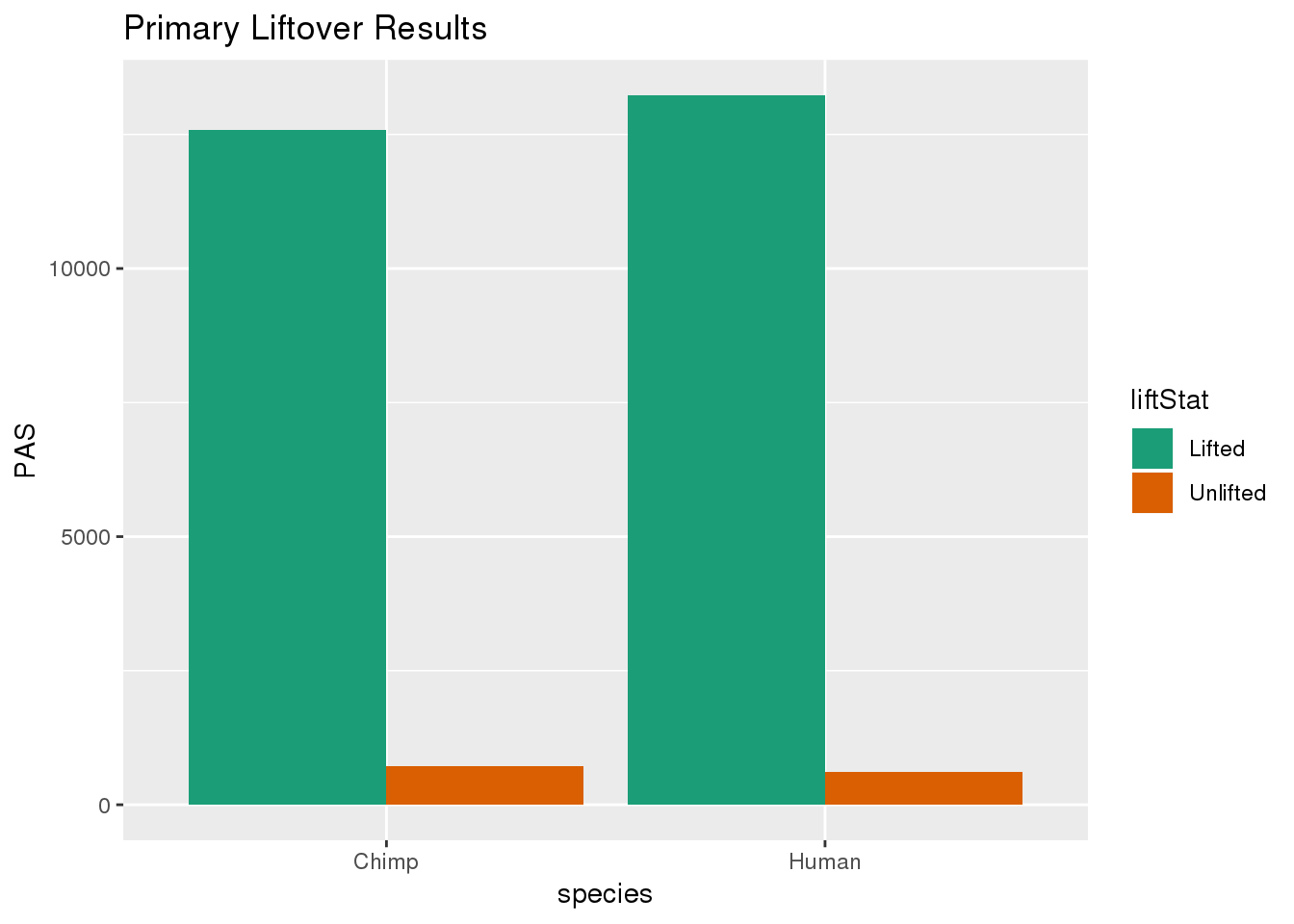
| Version | Author | Date |
|---|---|---|
| 106f3c1 | brimittleman | 2019-11-15 |
ggplot(primaryDF,aes(x=species, y=proportion, fill=liftStat)) + geom_bar(stat="identity",position = "dodge") + scale_fill_brewer(palette = "Dark2") + labs(title="Primary Liftover Results")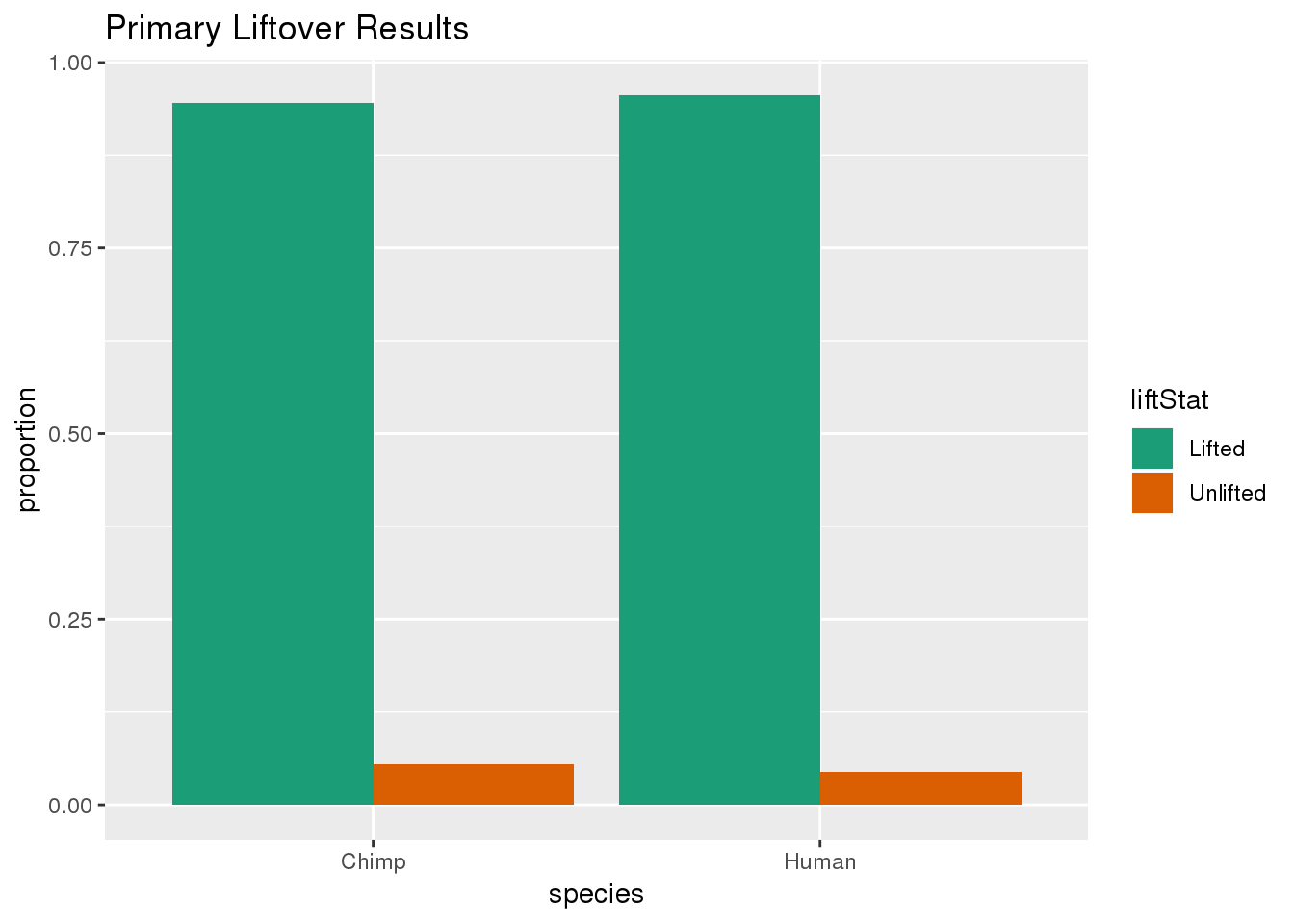
| Version | Author | Date |
|---|---|---|
| 106f3c1 | brimittleman | 2019-11-15 |
Look at the lifted:
OriginalHuman=read.table("../Human/data/RNAseq/DiffSplice/humanJunc.bed",stringsAsFactors = F)
liftedHuman=read.table("../Human/data/RNAseq/DiffSplice/humanJunc_inChimp.bed",stringsAsFactors = F)
OriginalChimp=read.table("../Chimp/data/RNAseq/DiffSplice/chimpJunc.bed",stringsAsFactors = F)
liftedChimp=read.table("../Chimp/data/RNAseq/DiffSplice/chimpJunc_inHuman.bed",stringsAsFactors = F)Reverse lift:
re_unliftedH=read.table("../Human/data/RNAseq/DiffSplice/humanJunc_inChimp_B2Human_unlifted.bed",stringsAsFactors = F) %>% nrow()
re_unliftedC=read.table("../Chimp/data/RNAseq/DiffSplice/chimpJunc_inHuman_B2Chimp_unlifted.bed",stringsAsFactors = F) %>% nrow()
re_liftedH=read.table("../Human/data/RNAseq/DiffSplice/humanJunc_inChimp_B2Human.bed",stringsAsFactors = F) %>% nrow()
re_liftedC=read.table("../Chimp/data/RNAseq/DiffSplice/chimpJunc_inHuman_B2Chimp.bed",stringsAsFactors = F) %>% nrow()
re_UnC=c("Chimp","Unlifted", re_unliftedC)
re_UnH=c("Human","Unlifted", re_unliftedH)
re_LH=c("Human","Lifted", re_liftedH)
re_LC=c("Chimp","Lifted", re_liftedC)
header=c("species", "liftStat", "PAS")
re_DF= as.data.frame(rbind(re_LH,re_LC, re_UnH,re_UnC))
colnames(re_DF)=header
re_DF$PAS=as.numeric(as.character(re_DF$PAS))
re_DF= re_DF %>% group_by(species) %>% mutate(nPAS=sum(PAS)) %>% ungroup() %>% mutate(proportion=PAS/nPAS)ggplot(re_DF,aes(x=species, y=PAS, fill=liftStat)) + geom_bar(stat="identity",position = "dodge") + scale_fill_brewer(palette = "Dark2") + labs(title="Reverse Liftover Results")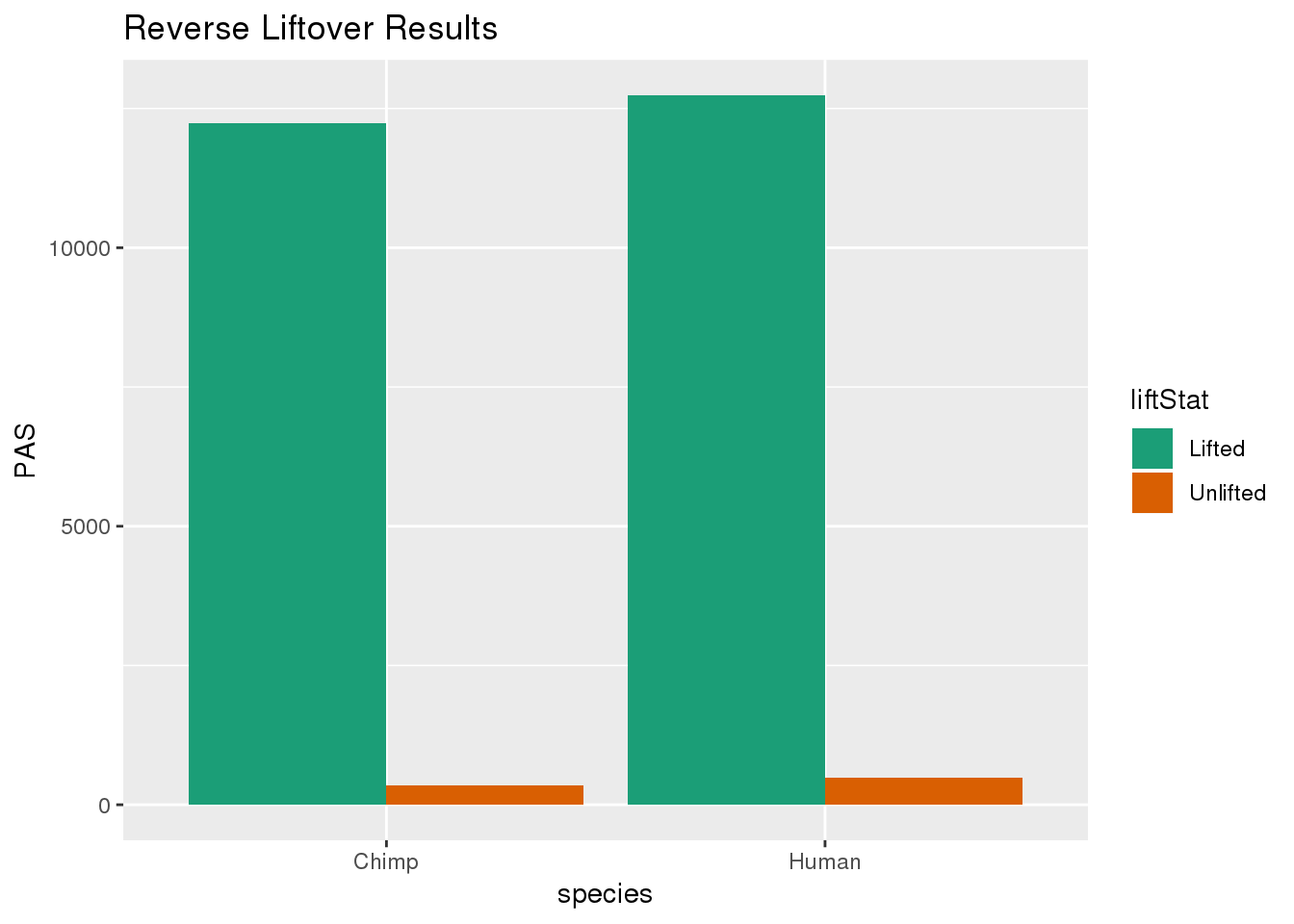
| Version | Author | Date |
|---|---|---|
| 106f3c1 | brimittleman | 2019-11-15 |
ggplot(re_DF,aes(x=species, y=proportion, fill=liftStat)) + geom_bar(stat="identity",position = "dodge") + scale_fill_brewer(palette = "Dark2")+ labs(title="Reverse Liftover Results")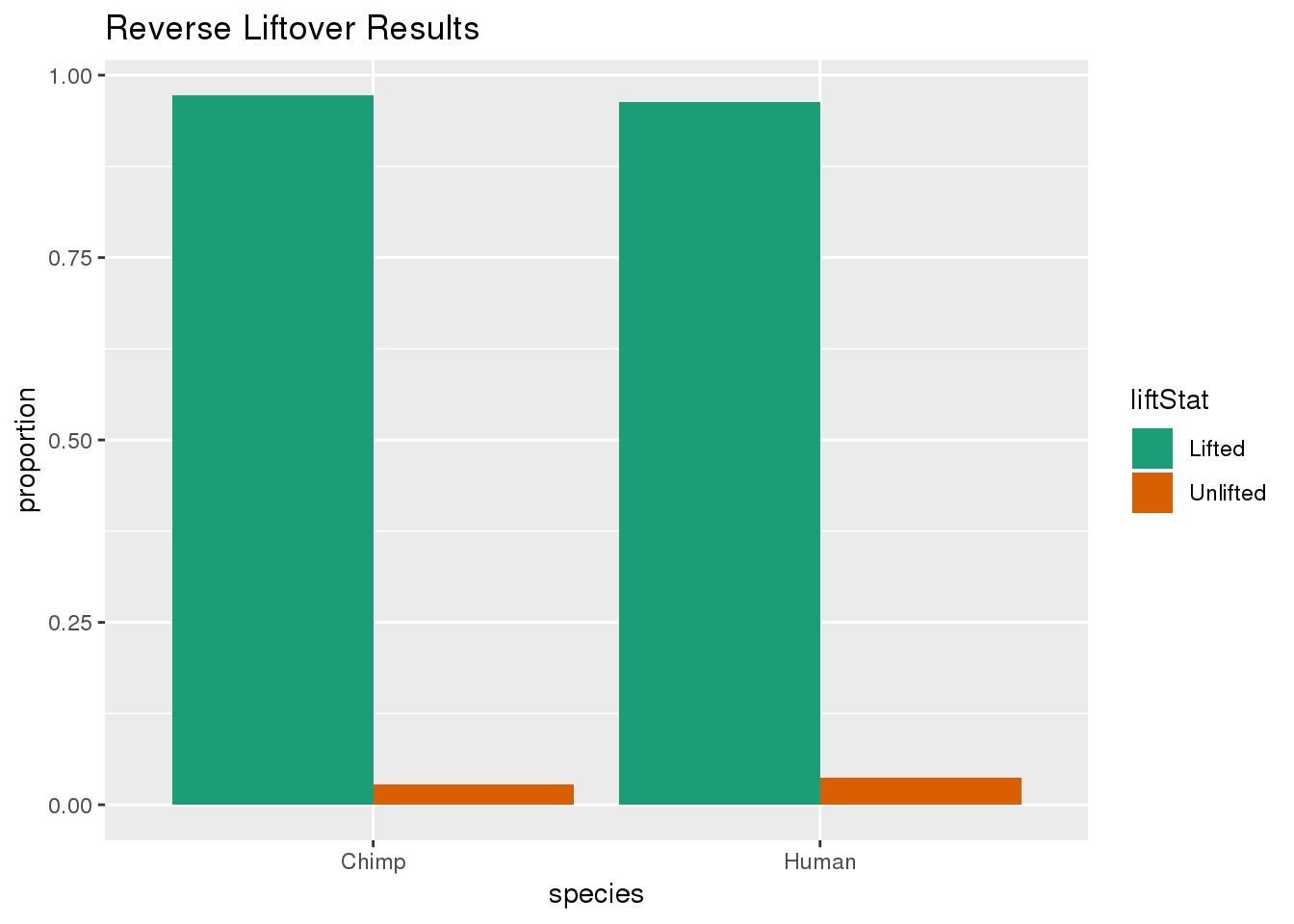
| Version | Author | Date |
|---|---|---|
| 106f3c1 | brimittleman | 2019-11-15 |
How many lifted both ways?
#human
re_liftedH/nrow(OriginalHuman)[1] 0.9200983#chimp
re_liftedC/nrow(OriginalChimp)[1] 0.9193779The next step will be to find the corresponding clusters. This is important because I will need to get the quantifications for the same introns and clusters. To do this I will need to write code that looks for the intron location from the primary lift in the reverse lift.
For now I will only look at those introns identified in both species. I need to do this because I need junctions we have quantifications for in both species.
I can make files with the human and chimp coordintats for the clusters that lift both ways. I will have to number each cluster
Human cluser:
humanRevlift=read.table("../Human/data/RNAseq/DiffSplice/humanJunc_inChimp_B2Human.bed",stringsAsFactors = F,col.names = c("Hchr","Hstart", "Hend", "cluster", "score", "strand")) %>% select(-strand)
#number clusters
humanRevlift %>% select(cluster) %>% unique() %>% nrow()[1] 5085humanRevlift$score=as.character(humanRevlift$score)
humanRevlift= humanRevlift %>% mutate(Name=paste("Human", score, sep="_")) %>% select(-score)
humanInChimp=read.table("../Human/data/RNAseq/DiffSplice/humanJunc_inChimp.bed",stringsAsFactors = F,col.names = c("Cchr","Cstart", "Cend", "cluster", "score", "strand"))%>% select(-strand)
humanInChimp$score=as.character(humanInChimp$score)
humanInChimp= humanInChimp %>% mutate(Name=paste("Human", score, sep="_")) %>% select(-score)
humanliftedBoth=humanRevlift %>% inner_join(humanInChimp, by=c("cluster", "Name"))Chimp clusters:
chimpRevLift=read.table("../Chimp/data/RNAseq/DiffSplice/chimpJunc_inHuman_B2Chimp.bed",stringsAsFactors = F,col.names = c("Cchr","Cstart", "Cend", "cluster", "score", "strand")) %>% select(-strand)
chimpRevLift$score=as.character(chimpRevLift$score)
chimpRevLift= chimpRevLift %>% mutate(Name=paste("Chimp", score, sep="_")) %>% select(-score)
chimpRevLift %>% select(cluster) %>% unique() %>% nrow()[1] 4886chimpInHuman=read.table("../Chimp/data/RNAseq/DiffSplice/chimpJunc_inHuman.bed",stringsAsFactors = F,col.names = c("Hchr","Hstart", "Hend", "cluster", "score", "strand"))%>% select(-strand)
chimpInHuman$score=as.character(chimpInHuman$score)
chimpInHuman= chimpInHuman %>% mutate(Name=paste("Chimp", score, sep="_")) %>% select(-score)
chimpliftedBoth=chimpRevLift %>% inner_join(chimpInHuman, by=c("cluster", "Name"))Try to join these by the human and chimp coordinates
AllClusters=chimpliftedBoth %>% inner_join(humanliftedBoth, by=c("Cchr", "Cstart","Cend", "Hchr", "Hstart", "Hend")) %>% mutate(ChimpName=paste(Cchr,Cstart,Cend, cluster.x, sep=":" ),HumanName=paste(Hchr,Hstart,Hend, cluster.y, sep=":" ) )
nrow(AllClusters)[1] 7080AllClusters %>% select(cluster.x) %>% unique() %>% nrow()[1] 2967AllClusters %>% select(cluster.y) %>% unique() %>% nrow()[1] 2968This means there are ~7k isoforms from about 3k genes. This is from the ~5k clusters I had before. I can move on with these using the human names.
I will have to go back and figure out how to call clusters for more genes.
I need to reformat these back into the counts format.
#chr1:17055:17233:clu_1
AllClustersNames=AllClusters %>% select(HumanName, ChimpName)
ChimpCluster=read.table("../Chimp/data/RNAseq/DiffSplice/chimpJunc_perind_numers.counts.gz") %>% rownames_to_column(var="ChimpName")
FilteredChimpCluster= ChimpCluster %>% inner_join(AllClustersNames, by="ChimpName")
#map human onto these
HumanCluster=read.table("../Human/data/RNAseq/DiffSplice/humanJunc_perind_numers.counts.gz") %>% rownames_to_column(var="HumanName")
FilteredClusterBoth=HumanCluster %>% inner_join(FilteredChimpCluster, by="HumanName") %>% select(-ChimpName)
#create group file- this should have the name of the bams and the group
Bams=as.data.frame(colnames(FilteredClusterBoth)) %>% mutate(Species=ifelse(grepl("H",colnames(FilteredClusterBoth)), "Human", "Chimp")) %>% slice(2:n())
#mkdir ../data/DiffSplice
write.table(Bams, "../data/DiffSplice/groups_file.txt", col.names = F, row.names = F, quote = F, sep="\t" )
write.table(FilteredClusterBoth, "../data/DiffSplice/BothSpec_perind.counts", col.names = T, row.names = F, quote = F, sep="\t" )Remove the first name in header and zip the file:
(manually)
vi ../data/DiffSplice/BothSpec_perind.counts
gzip ../data/DiffSplice/BothSpec_perind.countsI will run the differential splicing analysis with the human exon file for now. Download the NCBI refseq exons from the table browser. I also downloaded the names with the common names so I can fix the exon file.
python processhg38exons.py
gzip ../data/DiffSplice/hg38_ncbiRefseq_exonsfixedRun leafcutter with python 2
sbatch DiffSplice.shError: duplicate ‘row.names’ are not allowed
sessionInfo()R version 3.5.1 (2018-07-02)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Scientific Linux 7.4 (Nitrogen)
Matrix products: default
BLAS/LAPACK: /software/openblas-0.2.19-el7-x86_64/lib/libopenblas_haswellp-r0.2.19.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] reshape2_1.4.3 forcats_0.3.0 stringr_1.3.1 dplyr_0.8.0.1
[5] purrr_0.3.2 readr_1.3.1 tidyr_0.8.3 tibble_2.1.1
[9] ggplot2_3.1.1 tidyverse_1.2.1
loaded via a namespace (and not attached):
[1] tidyselect_0.2.5 haven_1.1.2 lattice_0.20-38
[4] colorspace_1.3-2 generics_0.0.2 htmltools_0.3.6
[7] yaml_2.2.0 rlang_0.4.0 later_0.7.5
[10] pillar_1.3.1 glue_1.3.0 withr_2.1.2
[13] RColorBrewer_1.1-2 modelr_0.1.2 readxl_1.1.0
[16] plyr_1.8.4 munsell_0.5.0 gtable_0.2.0
[19] workflowr_1.5.0 cellranger_1.1.0 rvest_0.3.2
[22] evaluate_0.12 labeling_0.3 knitr_1.20
[25] httpuv_1.4.5 broom_0.5.1 Rcpp_1.0.2
[28] promises_1.0.1 scales_1.0.0 backports_1.1.2
[31] jsonlite_1.6 fs_1.3.1 hms_0.4.2
[34] digest_0.6.18 stringi_1.2.4 grid_3.5.1
[37] rprojroot_1.3-2 cli_1.1.0 tools_3.5.1
[40] magrittr_1.5 lazyeval_0.2.1 crayon_1.3.4
[43] whisker_0.3-2 pkgconfig_2.0.2 xml2_1.2.0
[46] lubridate_1.7.4 assertthat_0.2.0 rmarkdown_1.10
[49] httr_1.3.1 rstudioapi_0.10 R6_2.3.0
[52] nlme_3.1-137 git2r_0.26.1 compiler_3.5.1