Sequencing meta data
Briana Mittleman
4/25/2019
Last updated: 2019-04-26
Checks: 6 0
Knit directory: apaQTL/analysis/
This reproducible R Markdown analysis was created with workflowr (version 1.3.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20190411) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: output/.DS_Store
Untracked files:
Untracked: .Rprofile
Untracked: ._.DS_Store
Untracked: .gitignore
Untracked: _workflowr.yml
Untracked: analysis/._PASdescriptiveplots.Rmd
Untracked: analysis/._cuttoffPercUsage.Rmd
Untracked: analysis/cuttoffPercUsage.Rmd
Untracked: apaQTL.Rproj
Untracked: code/._SnakefilePAS
Untracked: code/._SnakefilefiltPAS
Untracked: code/._apaQTLCorrectPvalMakeQQ.R
Untracked: code/._apaQTL_Nominal.sh
Untracked: code/._apaQTL_permuted.sh
Untracked: code/._bed2saf.py
Untracked: code/._callPeaksYL.py
Untracked: code/._chooseAnno2SAF.py
Untracked: code/._chooseSignalSite
Untracked: code/._chooseSignalSite.py
Untracked: code/._cluster.json
Untracked: code/._clusterPAS.json
Untracked: code/._clusterfiltPAS.json
Untracked: code/._config.yaml
Untracked: code/._config2.yaml
Untracked: code/._configOLD.yaml
Untracked: code/._convertNumeric.py
Untracked: code/._dag.pdf
Untracked: code/._filter5perc.R
Untracked: code/._filter5percPheno.py
Untracked: code/._filterpeaks.py
Untracked: code/._fixFChead.py
Untracked: code/._make5percPeakbed.py
Untracked: code/._makeFileID.py
Untracked: code/._makePheno.py
Untracked: code/._mergeAllBam.sh
Untracked: code/._mergeByFracBam.sh
Untracked: code/._mergePeaks.sh
Untracked: code/._namePeaks.py
Untracked: code/._peak2PAS.py
Untracked: code/._peakFC.sh
Untracked: code/._pheno2countonly.R
Untracked: code/._quantassign2parsedpeak.py
Untracked: code/._snakemakePAS.batch
Untracked: code/._snakemakefiltPAS.batch
Untracked: code/._submit-snakemakePAS.sh
Untracked: code/._submit-snakemakefiltPAS.sh
Untracked: code/.snakemake/
Untracked: code/APAqtl_nominal.err
Untracked: code/APAqtl_nominal.out
Untracked: code/APAqtl_permuted.err
Untracked: code/APAqtl_permuted.out
Untracked: code/BothFracDTPlotGeneRegions.err
Untracked: code/BothFracDTPlotGeneRegions.out
Untracked: code/DistPAS2Sig.py
Untracked: code/README.md
Untracked: code/Rplots.pdf
Untracked: code/Upstream100Bases_general.py
Untracked: code/bam2bw.err
Untracked: code/bam2bw.out
Untracked: code/dag.pdf
Untracked: code/dagPAS.pdf
Untracked: code/dagfiltPAS.pdf
Untracked: code/findbuginpeaks.R
Untracked: code/get100upPAS.py
Untracked: code/getSeq100up.sh
Untracked: code/getseq100up.err
Untracked: code/getseq100up.out
Untracked: code/log/
Untracked: code/run_DistPAS2Sig.err
Untracked: code/run_DistPAS2Sig.out
Untracked: code/run_distPAS2Sig.sh
Untracked: code/snakePASlog.out
Untracked: code/snakefiltPASlog.out
Untracked: data/DTmatrix/
Untracked: data/PAS/
Untracked: data/README.md
Untracked: data/SignalSiteFiles/
Untracked: data/apaQTLNominal/
Untracked: data/apaQTLPermuted/
Untracked: data/assignedPeaks/
Untracked: data/bam/
Untracked: data/bam_clean/
Untracked: data/bam_waspfilt/
Untracked: data/bed_10up/
Untracked: data/bed_clean/
Untracked: data/bed_clean_sort/
Untracked: data/bed_waspfilter/
Untracked: data/bedsort_waspfilter/
Untracked: data/fastq/
Untracked: data/filterPeaks/
Untracked: data/inclusivePeaks/
Untracked: data/inclusivePeaks_FC/
Untracked: data/mergedBG/
Untracked: data/mergedBW_byfrac/
Untracked: data/mergedBam/
Untracked: data/mergedbyFracBam/
Untracked: data/nuc_10up/
Untracked: data/nuc_10upclean/
Untracked: data/peakCoverage/
Untracked: data/peaks_5perc/
Untracked: data/phenotype/
Untracked: data/phenotype_5perc/
Untracked: data/sort/
Untracked: data/sort_clean/
Untracked: data/sort_waspfilter/
Untracked: nohup.out
Untracked: output/._.DS_Store
Untracked: output/dtPlots/
Untracked: output/fastqc/
Unstaged changes:
Modified: analysis/PASusageQC.Rmd
Modified: analysis/corrbetweenind.Rmd
Deleted: code/Upstream10Bases_general.py
Modified: code/bed2saf.py
Deleted: code/test.txt
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the R Markdown and HTML files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view them.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 111979f | brimittleman | 2019-04-26 | add seq correlations |
| html | 3557545 | brimittleman | 2019-04-25 | Build site. |
| html | 0f7ad72 | brimittleman | 2019-04-25 | Build site. |
| Rmd | 48b2ec1 | brimittleman | 2019-04-25 | add map befroe mp filter |
| html | be227e7 | brimittleman | 2019-04-25 | Build site. |
| Rmd | 6cb0a99 | brimittleman | 2019-04-25 | add seq meta pltos |
In this analysis I want to compare the sequencing depth between batches.
library(tidyverse)── Attaching packages ──────────────────────────────────────────────────────────────── tidyverse 1.2.1 ──✔ ggplot2 3.1.0 ✔ purrr 0.3.2
✔ tibble 2.1.1 ✔ dplyr 0.8.0.1
✔ tidyr 0.8.3 ✔ stringr 1.3.1
✔ readr 1.3.1 ✔ forcats 0.3.0 ── Conflicts ─────────────────────────────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()Load metadata:
metadata=read.table("../data/MetaDataSequencing.txt",header = T)
meta_T=metadata %>% filter(grepl("T", Sample_ID)) %>% mutate(samp=paste("X", Sample_ID, sep=""))
meta_N=metadata %>% filter(grepl("N", Sample_ID)) %>% mutate(samp=paste("X", Sample_ID, sep=""))Read count
metadata$batch=as.factor(metadata$batch)
ggplot(metadata, aes(x=batch, group=batch, y=reads, fill=batch)) + geom_boxplot() + geom_jitter() + facet_grid(~fraction) + labs(title="Read count by batch")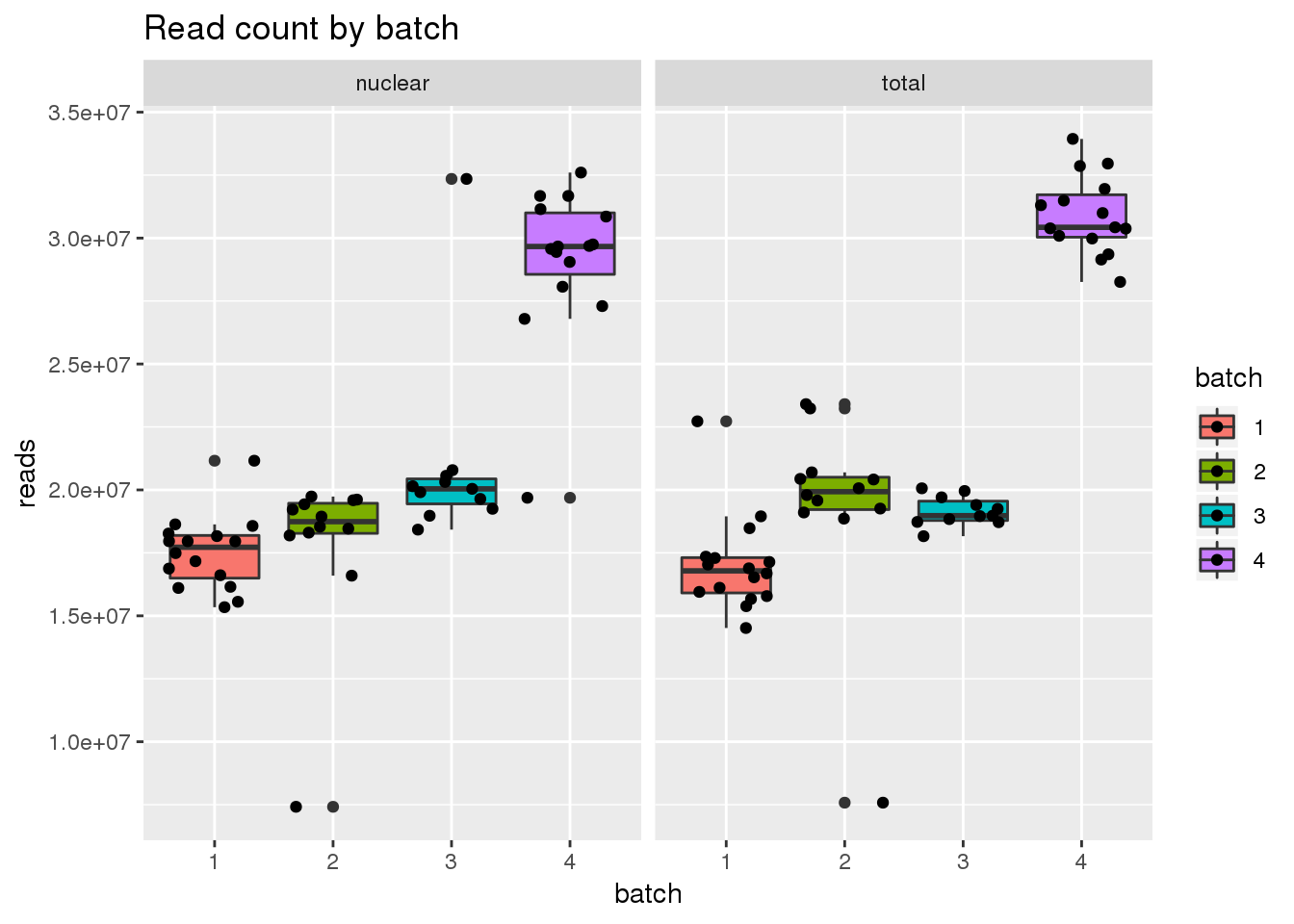
| Version | Author | Date |
|---|---|---|
| be227e7 | brimittleman | 2019-04-25 |
Mapped reads
ggplot(metadata, aes(x=batch, group=batch, y=Mapped_noMP, fill=batch)) + geom_boxplot() + geom_jitter() + facet_grid(~fraction) + labs(title="Mapped reads by batch")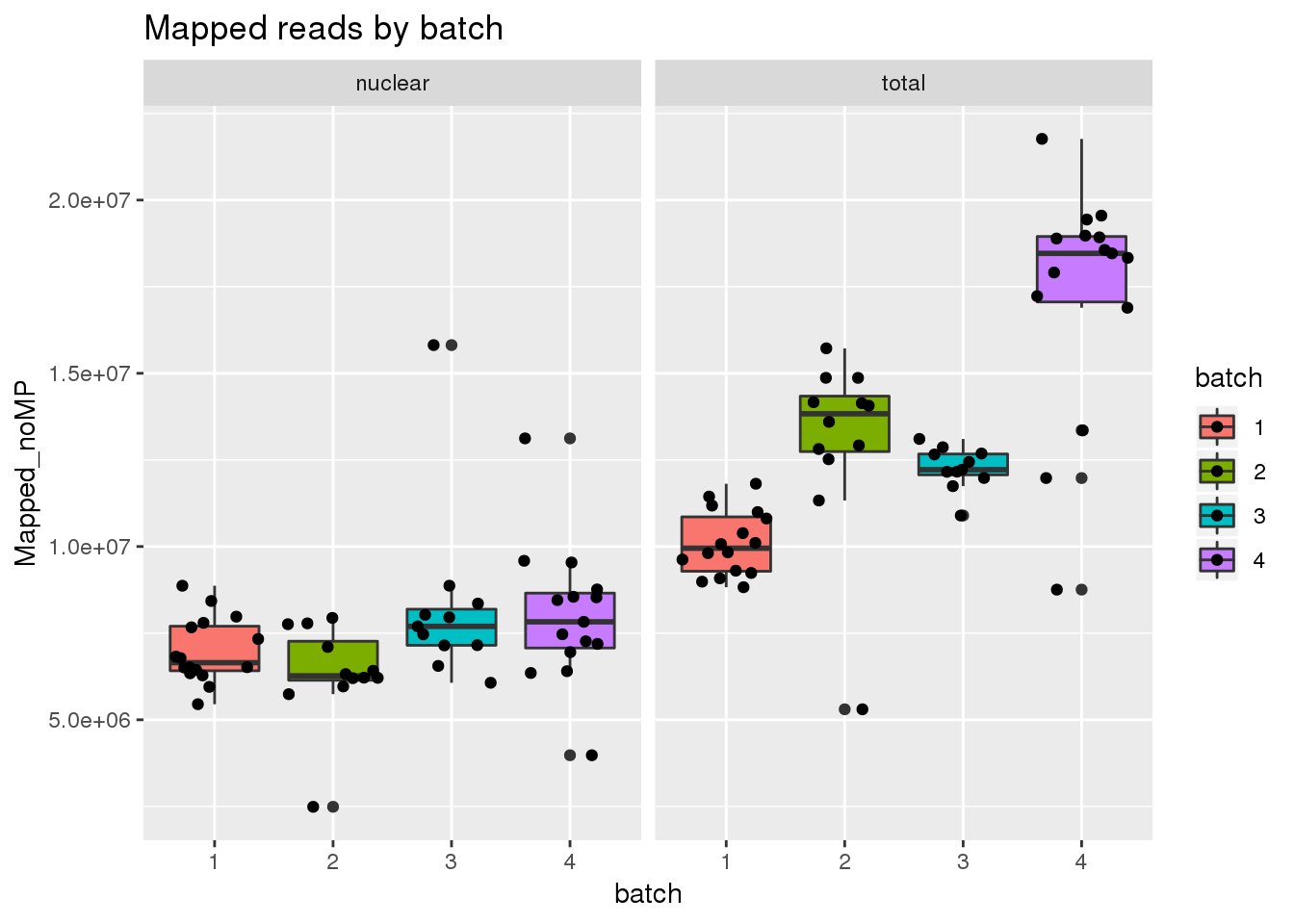
Map prop
ggplot(metadata, aes(x=batch, group=batch, y=prop_MappedwithoutMP, fill=batch)) + geom_boxplot() + geom_jitter() + facet_grid(~fraction) + labs(title="Proportion Mapped reads by batch")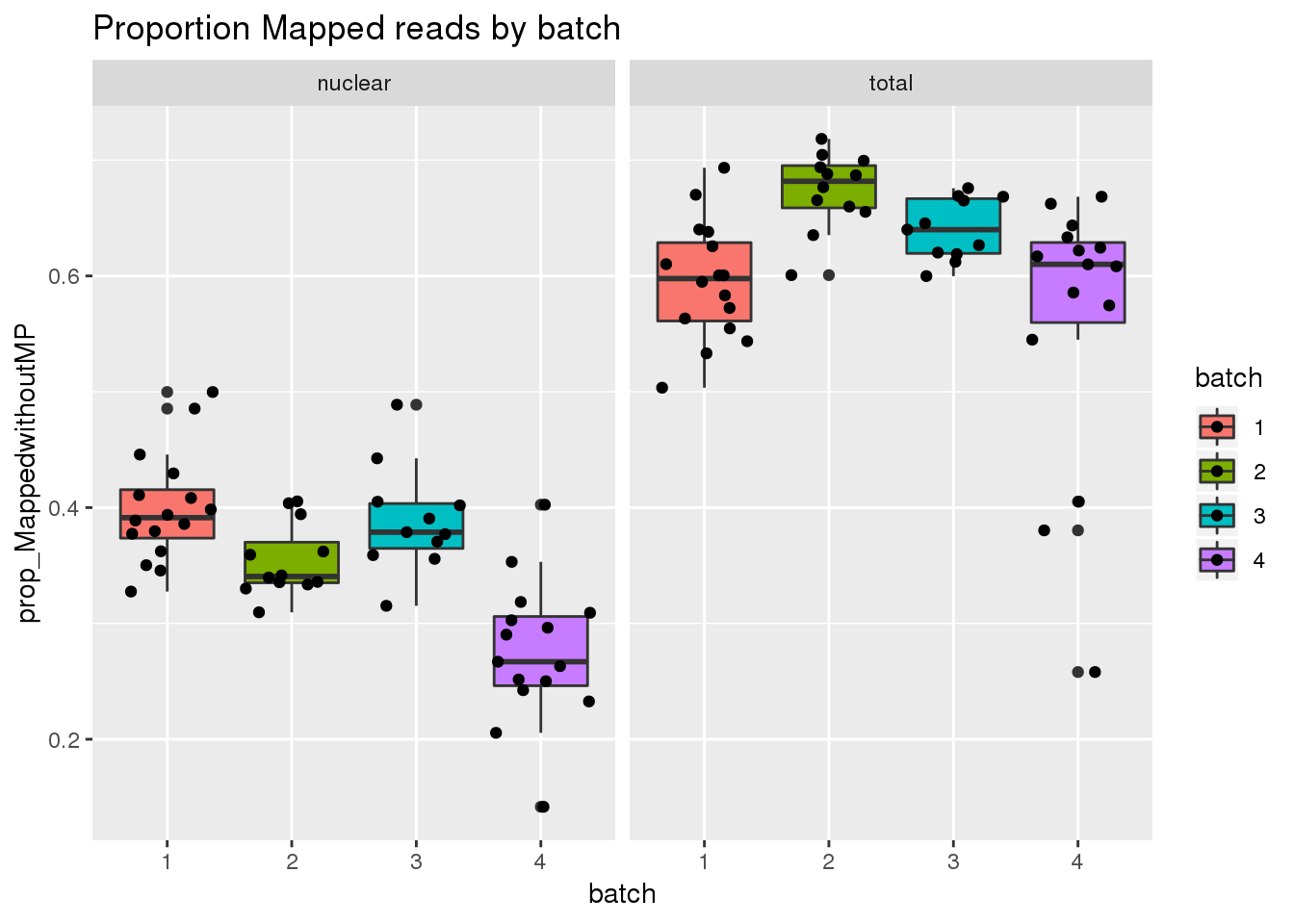
| Version | Author | Date |
|---|---|---|
| be227e7 | brimittleman | 2019-04-25 |
Library concentration
ggplot(metadata, aes(x=batch, group=batch, y=library_conc, fill=batch)) + geom_boxplot() + geom_jitter() + facet_grid(~fraction) + labs(title="Library concentrations by batch")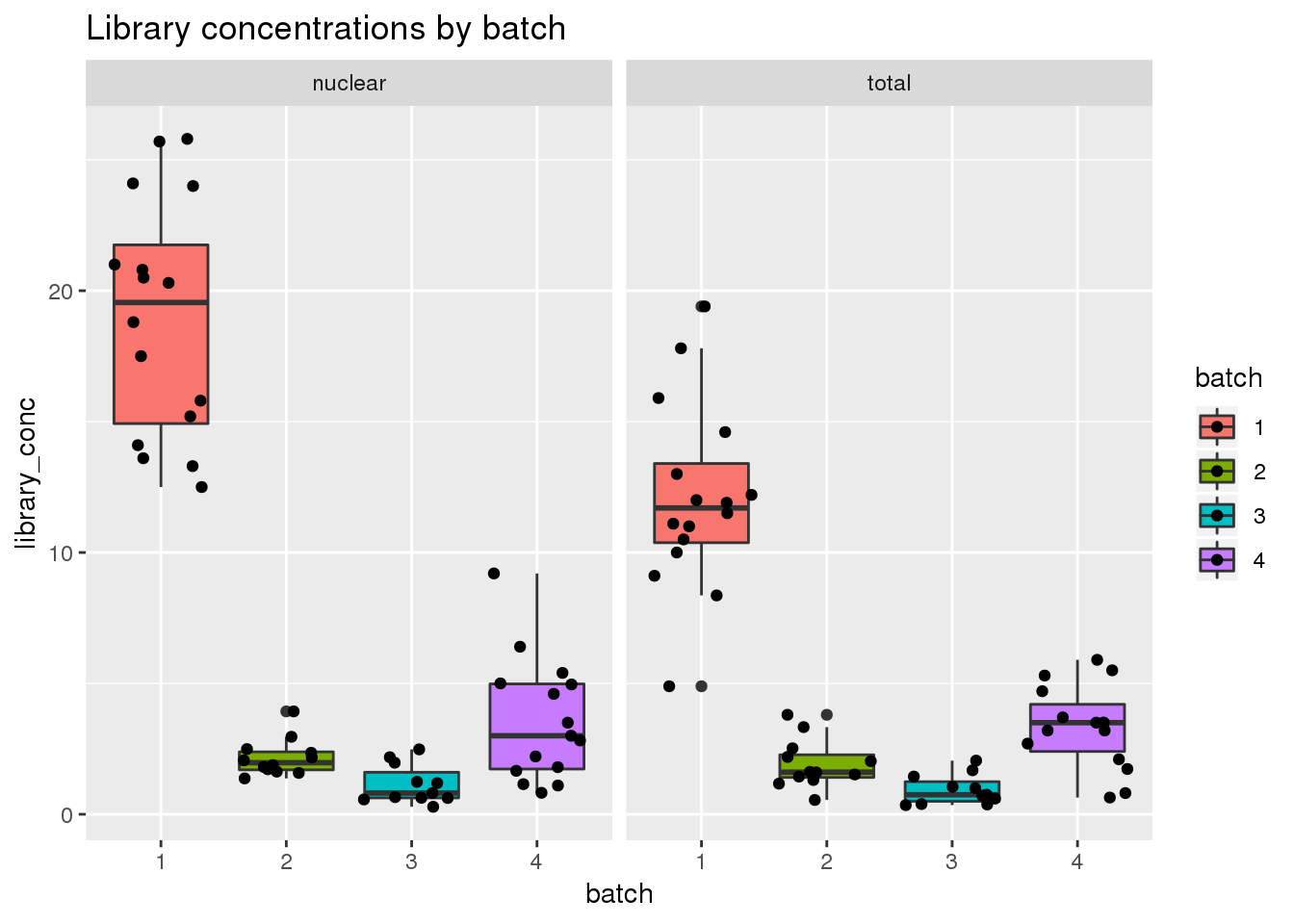
| Version | Author | Date |
|---|---|---|
| be227e7 | brimittleman | 2019-04-25 |
before mp
ggplot(metadata, aes(x=batch, group=batch, y=mapped, fill=batch)) + geom_boxplot() + geom_jitter() + facet_grid(~fraction) + labs(title="Mapped reads before MP filter by batch")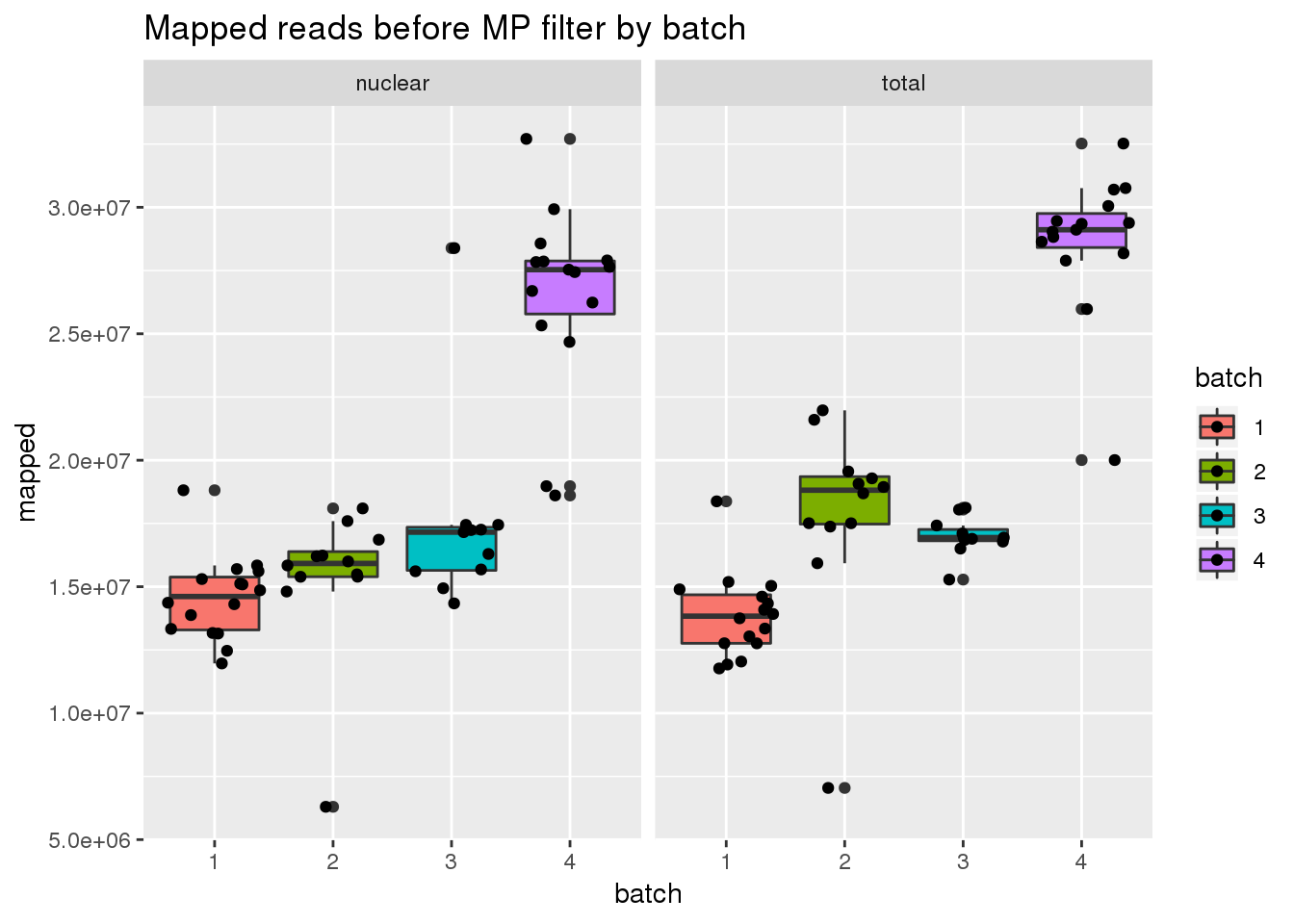
| Version | Author | Date |
|---|---|---|
| 0f7ad72 | brimittleman | 2019-04-25 |
alive perc
ggplot(metadata, aes(y=Mapped_noMP, col=batch, x=alive_avg)) + geom_point()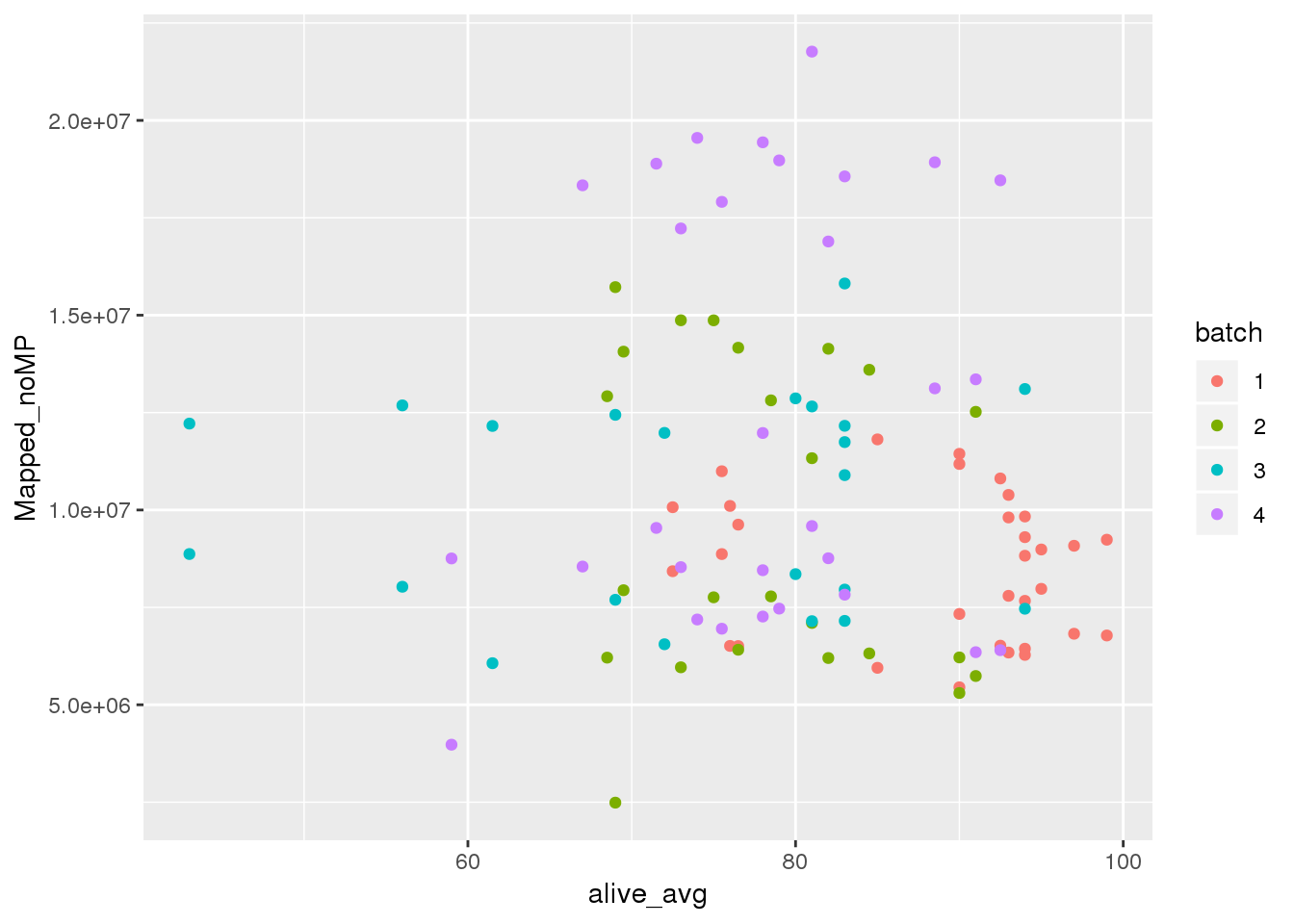 #Cq
#Cq
ggplot(metadata, aes(y=Mapped_noMP, col=batch, x=library_conc)) + geom_point() 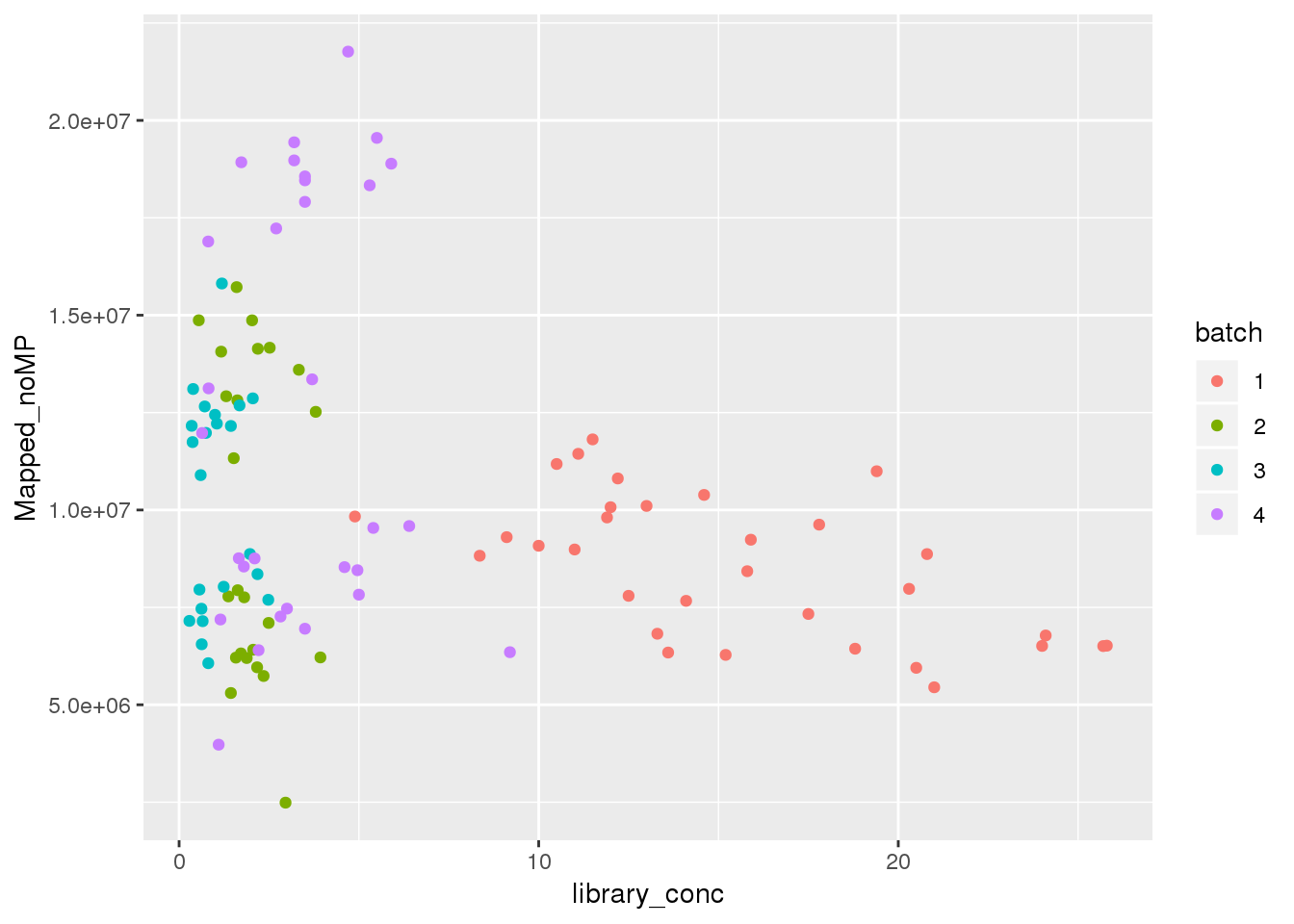
Mapped v concentration
ggplot(metadata, aes(y=Mapped_noMP, col=batch, x=Conentration)) + geom_point() + facet_grid(~fraction)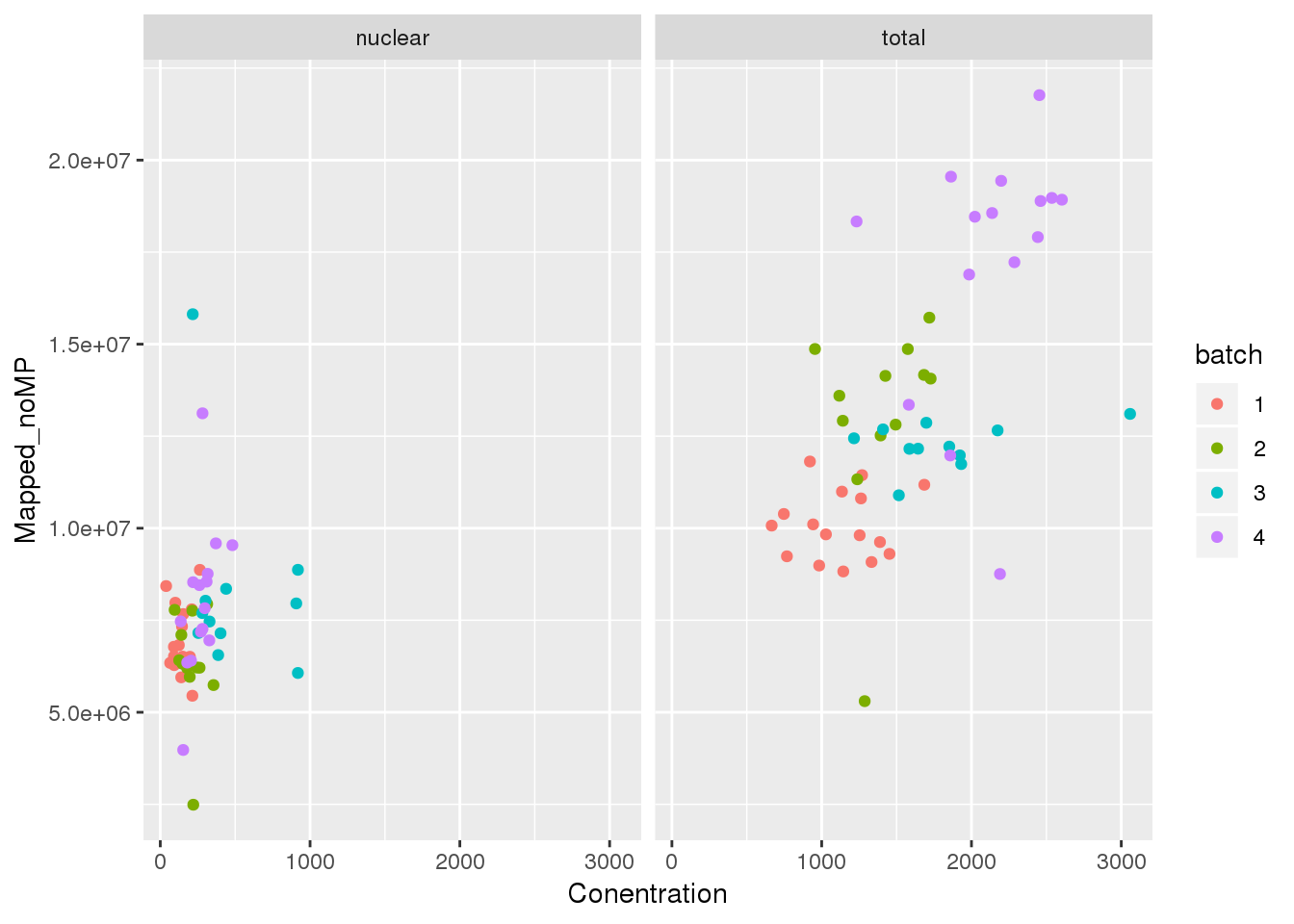
metadata_T=metadata %>% filter(fraction=="total")
summary(lm(data=metadata_T, Mapped_noMP ~ Conentration))
Call:
lm(formula = Mapped_noMP ~ Conentration, data = metadata_T)
Residuals:
Min 1Q Median 3Q Max
-6858140 -1918197 151332 1688547 6691716
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 6525031.1 1219768.8 5.349 2.01e-06 ***
Conentration 4149.8 722.4 5.745 4.86e-07 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 2805000 on 52 degrees of freedom
Multiple R-squared: 0.3882, Adjusted R-squared: 0.3765
F-statistic: 33 on 1 and 52 DF, p-value: 4.86e-07metadata_N=metadata %>% filter(fraction=="nuclear")
summary(lm(data=metadata_N, Mapped_noMP~ Conentration))
Call:
lm(formula = Mapped_noMP ~ Conentration, data = metadata_N)
Residuals:
Min 1Q Median 3Q Max
-4831820 -875807 -215509 558782 8499292
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 6972552 446484 15.617 <2e-16 ***
Conentration 1577 1389 1.136 0.261
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 1895000 on 52 degrees of freedom
Multiple R-squared: 0.02422, Adjusted R-squared: 0.005452
F-statistic: 1.291 on 1 and 52 DF, p-value: 0.2612RNA quality
ggplot(metadata, aes(y=Mapped_noMP, col=batch, x=ratio260_280)) + geom_point() + facet_grid(~fraction)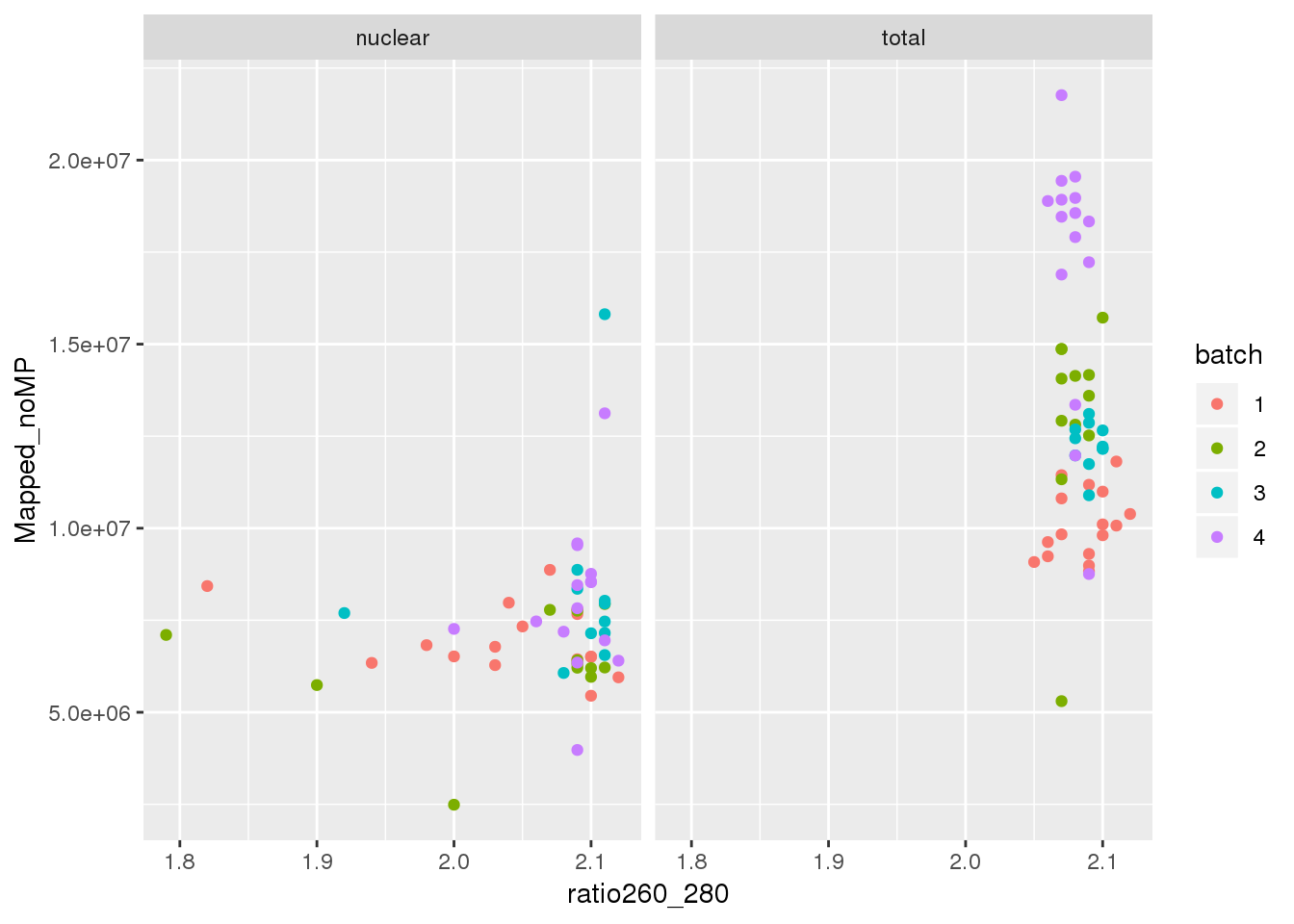
sessionInfo()R version 3.5.1 (2018-07-02)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Scientific Linux 7.4 (Nitrogen)
Matrix products: default
BLAS/LAPACK: /software/openblas-0.2.19-el7-x86_64/lib/libopenblas_haswellp-r0.2.19.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] forcats_0.3.0 stringr_1.3.1 dplyr_0.8.0.1 purrr_0.3.2
[5] readr_1.3.1 tidyr_0.8.3 tibble_2.1.1 ggplot2_3.1.0
[9] tidyverse_1.2.1
loaded via a namespace (and not attached):
[1] Rcpp_1.0.0 cellranger_1.1.0 pillar_1.3.1 compiler_3.5.1
[5] git2r_0.23.0 plyr_1.8.4 workflowr_1.3.0 tools_3.5.1
[9] digest_0.6.18 lubridate_1.7.4 jsonlite_1.6 evaluate_0.12
[13] nlme_3.1-137 gtable_0.2.0 lattice_0.20-38 pkgconfig_2.0.2
[17] rlang_0.3.1 cli_1.0.1 rstudioapi_0.10 yaml_2.2.0
[21] haven_1.1.2 withr_2.1.2 xml2_1.2.0 httr_1.3.1
[25] knitr_1.20 hms_0.4.2 generics_0.0.2 fs_1.2.6
[29] rprojroot_1.3-2 grid_3.5.1 tidyselect_0.2.5 glue_1.3.0
[33] R6_2.3.0 readxl_1.1.0 rmarkdown_1.10 reshape2_1.4.3
[37] modelr_0.1.2 magrittr_1.5 whisker_0.3-2 backports_1.1.2
[41] scales_1.0.0 htmltools_0.3.6 rvest_0.3.2 assertthat_0.2.0
[45] colorspace_1.3-2 labeling_0.3 stringi_1.2.4 lazyeval_0.2.1
[49] munsell_0.5.0 broom_0.5.1 crayon_1.3.4