Characterize Total ApaQTLs
Briana Mittleman
10/11/2018
Last updated: 2018-10-11
workflowr checks: (Click a bullet for more information)-
✔ R Markdown file: up-to-date
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
-
✔ Environment: empty
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
-
✔ Seed:
set.seed(12345)The command
set.seed(12345)was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible. -
✔ Session information: recorded
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
-
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.✔ Repository version: f819c8e
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can usewflow_publishorwflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.Ignored files: Ignored: .DS_Store Ignored: .Rhistory Ignored: .Rproj.user/ Ignored: output/.DS_Store Untracked files: Untracked: KalistoAbundance18486.txt Untracked: analysis/genometrack_figs.Rmd Untracked: analysis/ncbiRefSeq_sm.sort.mRNA.bed Untracked: analysis/snake.config.notes.Rmd Untracked: analysis/verifyBAM.Rmd Untracked: data/18486.genecov.txt Untracked: data/APApeaksYL.total.inbrain.bed Untracked: data/NuclearApaQTLs.txt Untracked: data/PeaksUsed/ Untracked: data/RNAkalisto/ Untracked: data/TotalApaQTLs.txt Untracked: data/Totalpeaks_filtered_clean.bed Untracked: data/YL-SP-18486-T-combined-genecov.txt Untracked: data/YL-SP-18486-T_S9_R1_001-genecov.txt Untracked: data/apaExamp/ Untracked: data/bedgraph_peaks/ Untracked: data/bin200.5.T.nuccov.bed Untracked: data/bin200.Anuccov.bed Untracked: data/bin200.nuccov.bed Untracked: data/clean_peaks/ Untracked: data/comb_map_stats.csv Untracked: data/comb_map_stats.xlsx Untracked: data/comb_map_stats_39ind.csv Untracked: data/combined_reads_mapped_three_prime_seq.csv Untracked: data/ensemble_to_genename.txt Untracked: data/filtered_APApeaks_merged_allchrom_refseqTrans.closest2End.bed Untracked: data/filtered_APApeaks_merged_allchrom_refseqTrans.closest2End.noties.bed Untracked: data/first50lines_closest.txt Untracked: data/gencov.test.csv Untracked: data/gencov.test.txt Untracked: data/gencov_zero.test.csv Untracked: data/gencov_zero.test.txt Untracked: data/gene_cov/ Untracked: data/joined Untracked: data/leafcutter/ Untracked: data/merged_combined_YL-SP-threeprimeseq.bg Untracked: data/mol_overlap/ Untracked: data/nom_QTL/ Untracked: data/nom_QTL_opp/ Untracked: data/nom_QTL_trans/ Untracked: data/nuc6up/ Untracked: data/other_qtls/ Untracked: data/peakPerRefSeqGene/ Untracked: data/perm_QTL/ Untracked: data/perm_QTL_opp/ Untracked: data/perm_QTL_trans/ Untracked: data/reads_mapped_three_prime_seq.csv Untracked: data/smash.cov.results.bed Untracked: data/smash.cov.results.csv Untracked: data/smash.cov.results.txt Untracked: data/smash_testregion/ Untracked: data/ssFC200.cov.bed Untracked: data/temp.file1 Untracked: data/temp.file2 Untracked: data/temp.gencov.test.txt Untracked: data/temp.gencov_zero.test.txt Untracked: output/picard/ Untracked: output/plots/ Untracked: output/qual.fig2.pdf Unstaged changes: Modified: analysis/28ind.peak.explore.Rmd Modified: analysis/39indQC.Rmd Modified: analysis/PeakToGeneAssignment.Rmd Modified: analysis/cleanupdtseq.internalpriming.Rmd Modified: analysis/dif.iso.usage.leafcutter.Rmd Modified: analysis/diff_iso_pipeline.Rmd Modified: analysis/explore.filters.Rmd Modified: analysis/overlapMolQTL.Rmd Modified: analysis/overlap_qtls.Rmd Modified: analysis/peakOverlap_oppstrand.Rmd Modified: analysis/pheno.leaf.comb.Rmd Modified: analysis/test.max2.Rmd Modified: code/Snakefile
Expand here to see past versions:
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | f819c8e | Briana Mittleman | 2018-10-11 | add distance metric analsis total apaQTL |
This analysis will be used to characterize the total ApaQTLs. I will run the analysis on the total APAqtls in this analysis and will then run a similar analysis on the nuclear APAqtls in another analysis. I would like to study:
- Distance metrics:
- distance from snp to TSS of gene
- Distance from snp to peak
- distance from snp to TSS of gene
- Expression metrics:
- expression of genes with significant QTLs vs other genes (by rna seq)
- expression of genes with significant QTLs vs other genes (peak coverage)
- Chrom HMM metrics:
- look at the chrom HMM interval for the significant QTLs
Upload Libraries and Data:
Library
library(workflowr)This is workflowr version 1.1.1
Run ?workflowr for help getting startedlibrary(reshape2)
library(tidyverse)── Attaching packages ──────────────────────────────────────────────────────────── tidyverse 1.2.1 ──✔ ggplot2 3.0.0 ✔ purrr 0.2.5
✔ tibble 1.4.2 ✔ dplyr 0.7.6
✔ tidyr 0.8.1 ✔ stringr 1.3.1
✔ readr 1.1.1 ✔ forcats 0.3.0── Conflicts ─────────────────────────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()library(VennDiagram)Loading required package: gridLoading required package: futile.loggerlibrary(data.table)
Attaching package: 'data.table'The following objects are masked from 'package:dplyr':
between, first, lastThe following object is masked from 'package:purrr':
transposeThe following objects are masked from 'package:reshape2':
dcast, meltlibrary(cowplot)
Attaching package: 'cowplot'The following object is masked from 'package:ggplot2':
ggsavePermuted Results from APA:
I will add a column to this dataframe that will tell me if the association is significant at 10% FDR. This will help me plot based on significance later in the analysis. I am also going to seperate the PID into relevant pieces.
totalAPA=read.table("../data/perm_QTL_trans/filtered_APApeaks_merged_allchrom_refseqGenes_pheno_Total_transcript_permResBH.txt", stringsAsFactors = F, header=T) %>% mutate(sig=ifelse(-log10(bh)>=1, 1,0 )) %>% separate(pid, sep = ":", into=c("chr", "start", "end", "id")) %>% separate(id, sep = "_", into=c("gene", "strand", "peak"))
totalAPA$sig=as.factor(totalAPA$sig)
print(names(totalAPA)) [1] "chr" "start" "end" "gene" "strand" "peak" "nvar"
[8] "shape1" "shape2" "dummy" "sid" "dist" "npval" "slope"
[15] "ppval" "bpval" "bh" "sig" Distance Metrics
Distance from snp to TSS
I ran the QTL analysis based on the starting position of the gene.
ggplot(totalAPA, aes(x=dist, fill=sig, by=sig)) + geom_density(alpha=.5) + labs(title="Distance from snp to TSS", x="Base Pairs") + scale_fill_discrete(guide = guide_legend(title = "Significant QTL"))
It looks like most of the signifcant values are 100,000 bases. This makes sense. I can zoom in on this portion.
ggplot(totalAPA, aes(x=dist, fill=sig, by=sig)) + geom_density(alpha=.5)+coord_cartesian(xlim = c(-150000, 150000))
Distance from snp to peak
To perform this analysis I need to recover the peak positions.
The peak file I used for the QTL analysis is: /project2/gilad/briana/threeprimeseq/data/mergedPeaks_comb/filtered_APApeaks_merged_allchrom_refseqTrans.noties_sm.fixed.bed
peaks=read.table("../data/PeaksUsed/filtered_APApeaks_merged_allchrom_refseqTrans.noties_sm.fixed.bed", col.names = c("chr", "peakStart", "peakEnd", "PeakNum", "PeakScore", "Strand", "Gene")) %>% mutate(peak=paste("peak", PeakNum,sep="")) %>% mutate(PeakCenter=peakStart+ (peakEnd- peakStart))I want to join the peak start to the totalAPA file but the peak column. I will then create a column that is snppos-peakcenter.
totalAPA_peakdist= totalAPA %>% inner_join(peaks, by="peak") %>% separate(sid, into=c("snpCHR", "snpLOC"), by=":")
totalAPA_peakdist$snpLOC= as.numeric(totalAPA_peakdist$snpLOC)
totalAPA_peakdist= totalAPA_peakdist %>% mutate(DisttoPeak= snpLOC-PeakCenter)Plot this by significance.
ggplot(totalAPA_peakdist, aes(x=DisttoPeak, fill=sig, by=sig)) + geom_density(alpha=.5) + labs(title="Distance from snp peak", x="log10 absolute value Distance to Peak") + scale_fill_discrete(guide = guide_legend(title = "Significant QTL"))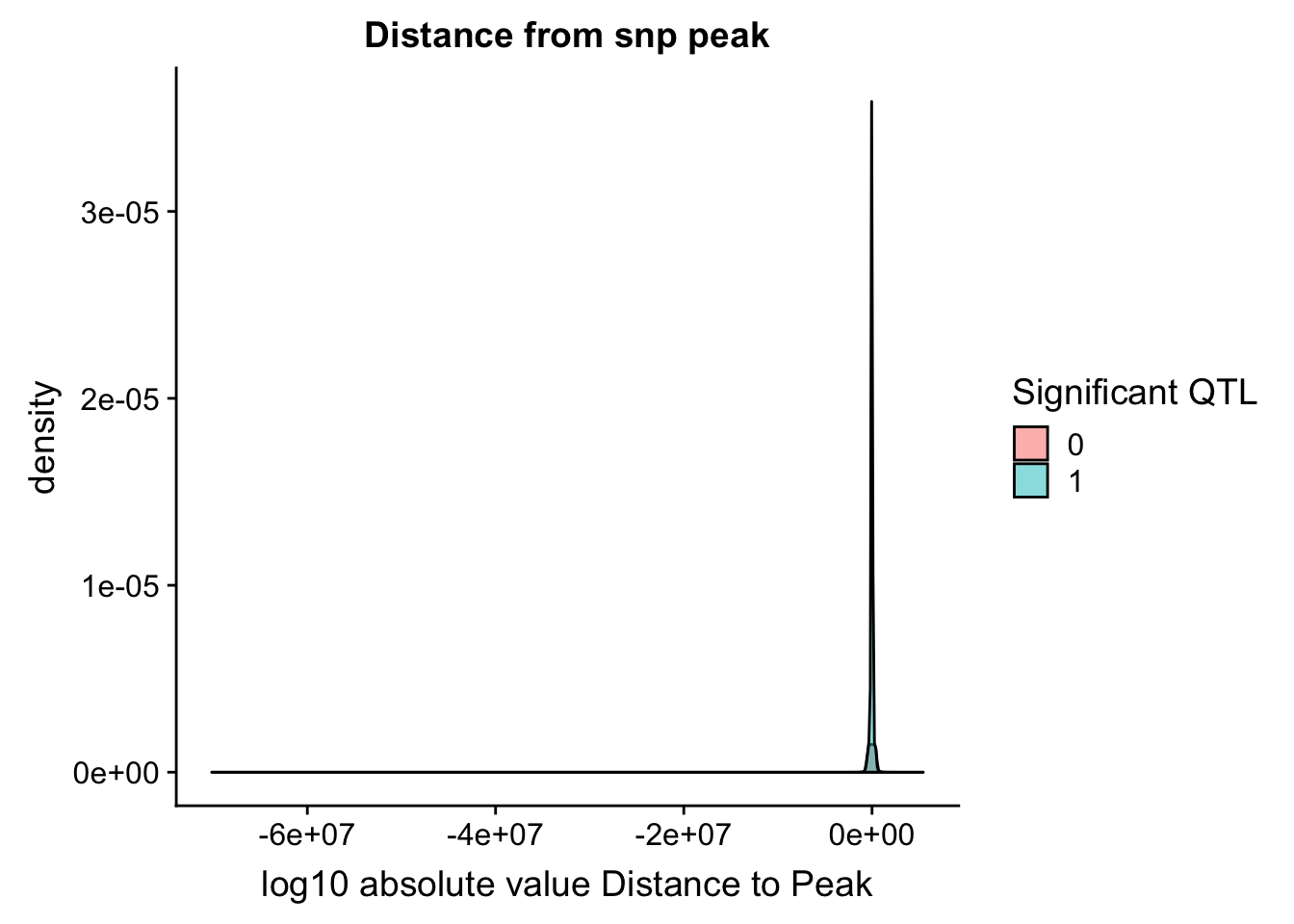
Look at the summarys based on significance:
totalAPA_peakdist_sig=totalAPA_peakdist %>% filter(sig==1)
totalAPA_peakdist_notsig=totalAPA_peakdist %>% filter(sig==0)
summary(totalAPA_peakdist_sig$DisttoPeak) Min. 1st Qu. Median Mean 3rd Qu. Max.
-634474.0 -26505.0 -3325.5 -23883.8 492.5 460051.0 summary(totalAPA_peakdist_notsig$DisttoPeak) Min. 1st Qu. Median Mean 3rd Qu. Max.
-70147526 -269853 -1269 -6620 266370 5433999 ggplot(totalAPA_peakdist, aes(y=DisttoPeak,x=sig, fill=sig, by=sig)) + geom_boxplot() + scale_fill_discrete(guide = guide_legend(title = "Significant QTL"))
Look like there are some outliers that are really far. I will remove variants greater than 1*10^6th away
totalAPA_peakdist_filt=totalAPA_peakdist %>% filter(abs(DisttoPeak) <= 1*(10^6))
ggplot(totalAPA_peakdist_filt, aes(y=DisttoPeak,x=sig, fill=sig, by=sig)) + geom_boxplot() + scale_fill_discrete(guide = guide_legend(title = "Significant QTL")) + facet_grid(.~strand)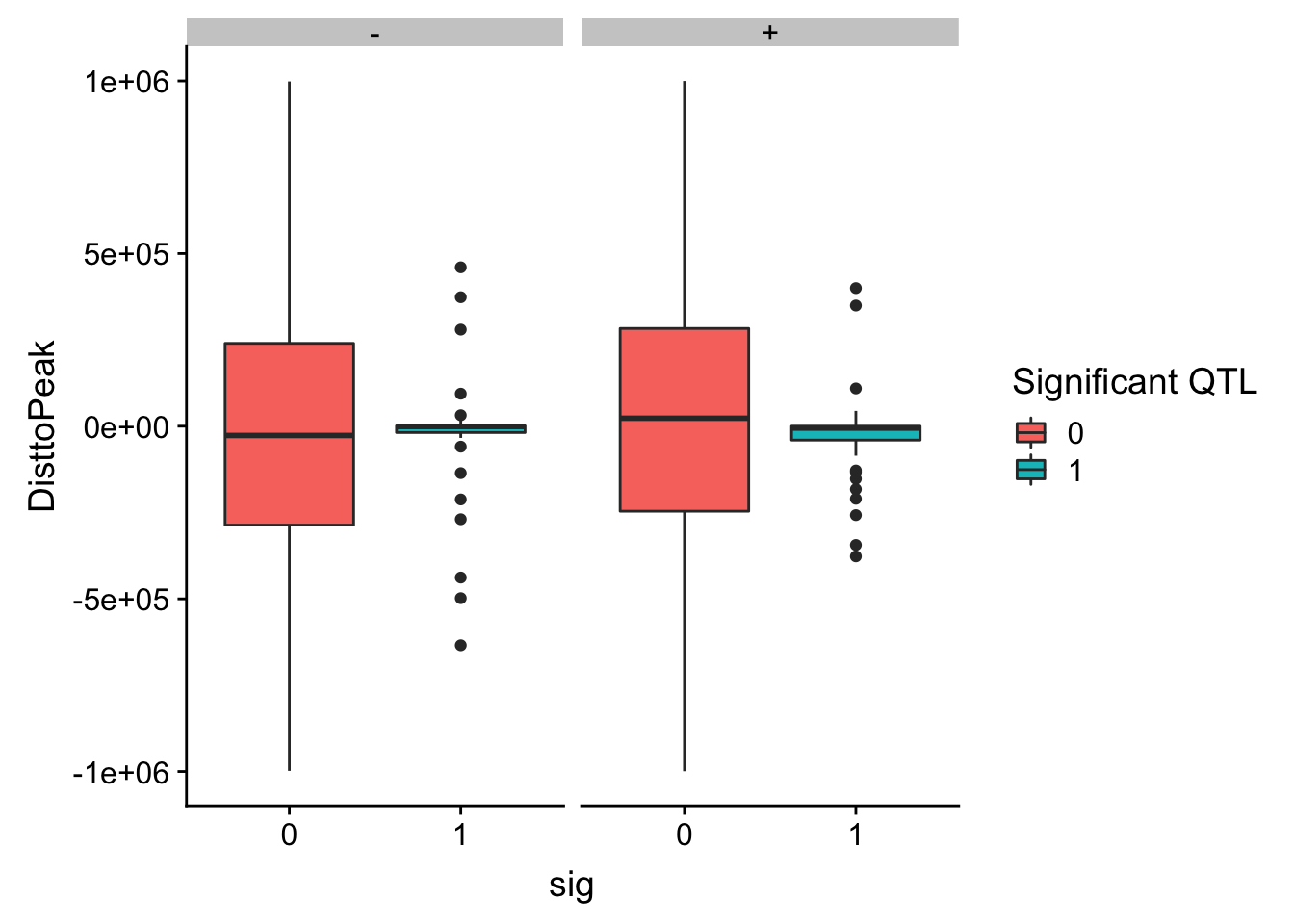
ggplot(totalAPA_peakdist_filt, aes(x=DisttoPeak, fill=sig, by=sig)) + geom_density() + scale_fill_discrete(guide = guide_legend(title = "Significant QTL")) + facet_grid(.~strand) This gives a similar distribution.
This gives a similar distribution.
I did snp - peak. This means if the peak is downstream of the snp on the positive strand the number will be negative.
In this case the peak is downstream of the snp.
totalAPA_peakdist %>% filter(sig==1) %>% filter(strand=="+") %>% filter(DisttoPeak < 0) %>% nrow()[1] 45totalAPA_peakdist %>% filter(sig==1) %>% filter(strand=="-") %>% filter(DisttoPeak > 0) %>% nrow()[1] 16Peak is upstream of the snp.
totalAPA_peakdist %>% filter(sig==1) %>% filter(strand=="+") %>% filter(DisttoPeak > 0) %>% nrow()[1] 17totalAPA_peakdist %>% filter(sig==1) %>% filter(strand=="-") %>% filter(DisttoPeak < 0) %>% nrow()[1] 40This means there is about 50/50 distribution around the peak start.
I am going to plot a violin plot for just the significant ones.
ggplot(totalAPA_peakdist_sig, aes(x=DisttoPeak)) + geom_density()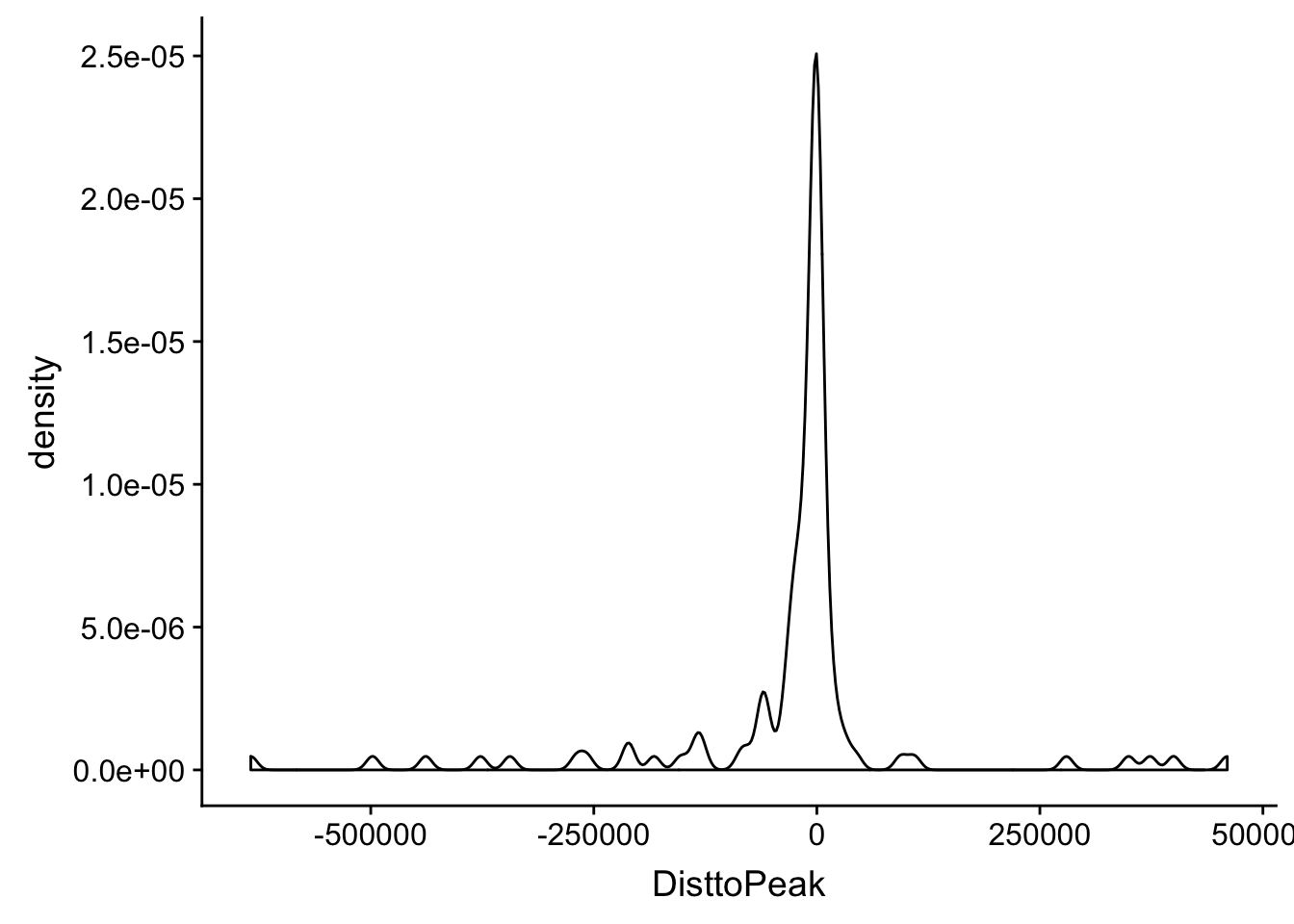 Within 1000 bases of the peak center.
Within 1000 bases of the peak center.
totalAPA_peakdist_sig %>% filter(abs(DisttoPeak) < 1000) %>% nrow()[1] 29totalAPA_peakdist_sig %>% filter(abs(DisttoPeak) < 10000) %>% nrow()[1] 57totalAPA_peakdist_sig %>% filter(abs(DisttoPeak) < 100000) %>% nrow()[1] 9829 QTLs are within 1000 bp of the peak center, 57 within 10,000bp and 98 within 100,000bp
Expression metrics
Session information
sessionInfo()R version 3.5.1 (2018-07-02)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS Sierra 10.12.6
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.5/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.5/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] grid stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] bindrcpp_0.2.2 cowplot_0.9.3 data.table_1.11.8
[4] VennDiagram_1.6.20 futile.logger_1.4.3 forcats_0.3.0
[7] stringr_1.3.1 dplyr_0.7.6 purrr_0.2.5
[10] readr_1.1.1 tidyr_0.8.1 tibble_1.4.2
[13] ggplot2_3.0.0 tidyverse_1.2.1 reshape2_1.4.3
[16] workflowr_1.1.1
loaded via a namespace (and not attached):
[1] tidyselect_0.2.4 haven_1.1.2 lattice_0.20-35
[4] colorspace_1.3-2 htmltools_0.3.6 yaml_2.2.0
[7] rlang_0.2.2 R.oo_1.22.0 pillar_1.3.0
[10] glue_1.3.0 withr_2.1.2 R.utils_2.7.0
[13] lambda.r_1.2.3 modelr_0.1.2 readxl_1.1.0
[16] bindr_0.1.1 plyr_1.8.4 munsell_0.5.0
[19] gtable_0.2.0 cellranger_1.1.0 rvest_0.3.2
[22] R.methodsS3_1.7.1 evaluate_0.11 labeling_0.3
[25] knitr_1.20 broom_0.5.0 Rcpp_0.12.19
[28] formatR_1.5 backports_1.1.2 scales_1.0.0
[31] jsonlite_1.5 hms_0.4.2 digest_0.6.17
[34] stringi_1.2.4 rprojroot_1.3-2 cli_1.0.1
[37] tools_3.5.1 magrittr_1.5 lazyeval_0.2.1
[40] futile.options_1.0.1 crayon_1.3.4 whisker_0.3-2
[43] pkgconfig_2.0.2 xml2_1.2.0 lubridate_1.7.4
[46] assertthat_0.2.0 rmarkdown_1.10 httr_1.3.1
[49] rstudioapi_0.8 R6_2.3.0 nlme_3.1-137
[52] git2r_0.23.0 compiler_3.5.1
This reproducible R Markdown analysis was created with workflowr 1.1.1