# Setup
# Load packages
library(pacman)
p_load(tidyverse, modeldata, skimr, janitor, tidymodels, magrittr, visdat)
# Load the credit dataset
data(credit_data)
# (Optional) Assign a new name
credit_df <- credit_data
# 'Fix' names
credit_df %<>%
clean_names() %>%
# Create a dummy variable for 'good' credit status
mutate(status_good = 1 * (status == "good")) %>%
# Drop the old status variable
select(-status)EC524: Lab 04
tidymodels Modeling, Resampling, and Prediction
2026
Lab Agenda
1. Modeling
2. Resampling and Prediction
Preprocessing
A quick catch-up of what we did:
i Loaded credit data, cleaned it up and created our outcome variable
Preprocessing
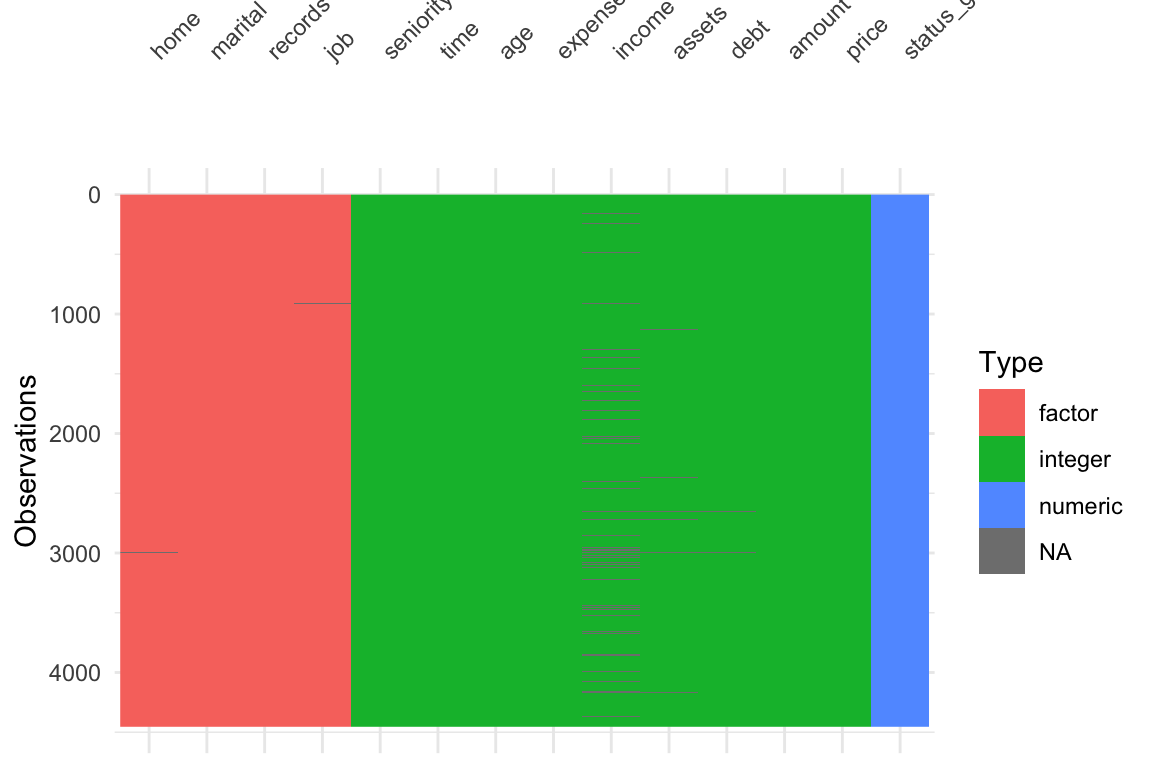
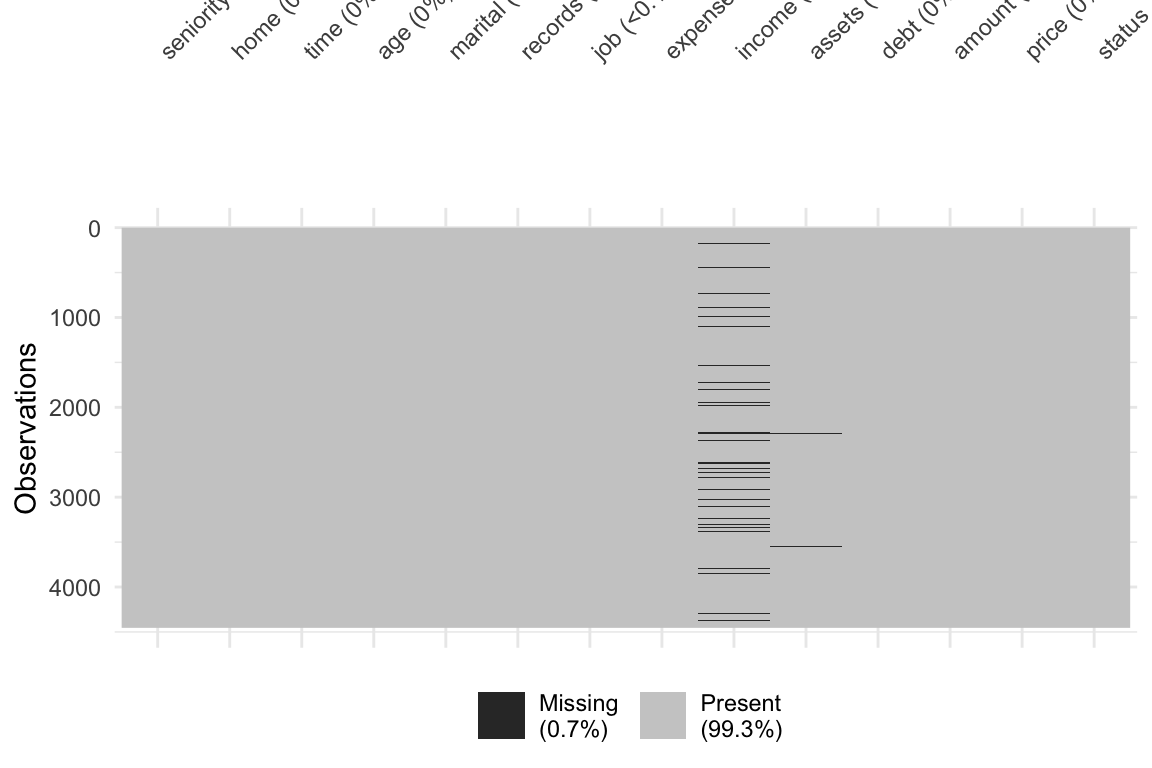
ii Visualized the data and found there is missing data
- New way:
visdat::vis_dat()andvisdat::vis_miss()
Preprocessing
iii Set up a recipe() that helped us clean the data and impute necessary missing data
Note: I am storing the preprocessing steps in an object, that I can call on later
# Define the recipe: status_good predicted by all other variables in credit_df
recipe_all = recipe(status_good ~ ., data = credit_df)
# Putting it all together
preprocessing_steps <- recipe_all %>%
# Mean imputation for numeric predictors
step_impute_mean(all_predictors() & all_numeric()) %>%
# KNN imputation for categorical predictors
step_impute_knn(all_predictors() & all_nominal(), neighbors = 5) %>%
# Create dummies for categorical variables
step_dummy(all_predictors() & all_nominal()) %>%
# Interactions
step_interact(~income:starts_with("home"))Preprocessing
iv Recall that creating a recipe() with steps are only instructions, we still need to prep() and juice() it to get a dataframe
Fitting Models
Now that we have a nice clean dataset, we can begin to fit a model
In the tidymodels universe, we need to do (at least) three things:
1. Define the Model Specification
- The Type of model we want (linear regression, KNN, or random forest)
2. Define the Engine
- The function/package that R will use to fit your model
3. Define the data and relationships to fit
- Point R to your data and define output/input
Specifying the Type of Model
Your first decision is the type of model you want to use (in tidymodels “type” = “specification”, but that may be confusing for the econometric mind)
You have many choices for the type of model you want to specify. Each comes with a different
parsnipfunction- Linear Regression:
linear_reg() - K-Nearest Neighbor:
nearest_neighbor() - Random Forest:
rand_forest()
- Linear Regression:
We can check what the linear regression model function
linear_reg()tells us:
Setting an Engine
Before we actually fit/train the model, we need to tell R which engine we want to use.
The engine tells R what algorithms to use, and some algos may differ greatly from others
You define the engine using the set_engine() function
You can see that we can fit a linear regression model using any of five engines
"lm""glmnet""stan"
"spark""keras"
You can use show_engines() function to see which engines are available for your model type
Setting an Engine
We will use the "lm" engine to fit a traditional linear regression model. Fancy stuff comes later.
Setting an Engine
We can also use translate() to see what is going on under the hood a bit more
Fitting (Training) Your Model
After defining the type of model and the engine, we can now begin to fit() the model
The
fit()function wants a few arguments- The
model_specobject comes from setting the type and engine (form before) - A
formulathe defines the outputs and inputs (just like usinglm()) - The
data
- The
In our case, we will have:
- Model type is
linear_reg()and our Engine is"lm" - The
formulais our linear regression wherestatus_goodis our outcome and the other vars.(.)are the predictors - Our data is our cleaned dataset
credit_clean
- Model type is
This is a simple linear regression but the tidymodels way.
tidymodels Linear Regression
We can use tidy() to see the results in a more usual way
# A tibble: 28 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 0.686 0.175 3.91 9.26e- 5
2 seniority 0.00927 0.000880 10.5 1.19e-25
3 time -0.0000822 0.000473 -0.174 8.62e- 1
4 age -0.00153 0.000720 -2.12 3.37e- 2
5 expenses -0.00212 0.000376 -5.63 1.95e- 8
6 income 0.00125 0.000981 1.27 2.03e- 1
7 assets 0.00000249 0.000000559 4.46 8.53e- 6
8 debt -0.0000175 0.00000489 -3.58 3.42e- 4
9 amount -0.000234 0.0000207 -11.3 2.21e-29
10 price 0.0000899 0.0000142 6.33 2.70e-10
# ℹ 18 more rowsWe can also use this information and use predict() as we have before.
Changing The Type of Model
Okay, so we have just gone through a complex way to run a simple regression model that you likely knew how to do in a simpler way
But the power of tidymodels is coming from its flexibility.
- It is very easy to change any of our decisions along the way
Let’s say instead we wanted to fit a KNN model. We just need to change the type of model and engine.
- Note Some methods like KNN can be used for either
"regression"tasks and"classification"tasks. We want to se the mode for these methods, either within thenearest_neighbor()function (nearest_neighbor(mode = "regression)) or using theset_mode()function.
We can finally choose a value for our parameters (tuning and CV later). In KNN that is just setting a value of \(k\). I’ll arbitrarily choose 6.
Resampling, Tuning, and Validating
We have omitted a big part of the ML Process: Tuning and Validating models.
As always, we begin with the setup (mostly in case this is not the same week as the other parts)
# Packages
library(pacman)
p_load(tidyverse, modeldata, skimr, janitor, tidymodels, magrittr)
# Load credit data
data(credit_data)
credit_df <- credit_data
# Clean up names
credit_df %<>% clean_names()
# Create a dummy variable for our outcome (good credit status)
credit_df %<>% mutate(status_good = 1 * (status == "good")) %>%
# drop old status variable
select(-status)Simple Sample Splitting
First simple task: Split the full credit_df dataset into a training (sub)set and a testing (sub)set
To split the sample, lets use the initial_split() function from the rsample package.
- It splits a proportion (
propargument) of your data into a training subset and leaves the rest for the testing subset.- If you wanted to sample by
stratayou could as well
- If you wanted to sample by
- It is another tool for you to use (we had used
sample_frac()previously) - The output is an
rsplitobject, not a dataframe- We will pass the object given by
initial_split()to thetraining()andtesting()functions to get the actual subsets as dataframes
- We will pass the object given by
[Note: It can be helpful to set a seed before you split the data for reproducing results]
Simple Sample Splitting
# Set the seed
set.seed(123)
# Create the split (80-20 split)
credit_split <- credit_df %>%
initial_split(prop = 0.8)
# Check the output
credit_split<Training/Testing/Total>
<3563/891/4454>Now we can impose the split by using the training() and testing() functions.
Simple Sample Splitting
After we grab the proper split data, we can check the dimensions to make sure it makes sense
Resampling
As you’ve seen in class, the goal of prediction exercises is good out-of-sample performance
To avoid overfitting, while still allowing for “optimal” flexibility, we use resampling methods.
We will use the vfold_cv() function to set up 5-fold cross validation for our model
vfold_cv() takes the following arguments:
datav, the number of foldsrepeatsthe number of times you want to repeat the cross validation (default is 1)stratais an optional variable to use for stratified sampling in the folds
Creating the folds looks like:
# set seed
set.seed(123)
# 5-fold CV on the training dataset
credit_cv <- credit_train %>% vfold_cv(v = 5)
# Check the output
credit_cv %>% tidy()# A tibble: 17,815 × 3
Row Data Fold
<int> <chr> <chr>
1 1 Analysis Fold1
2 1 Analysis Fold2
3 1 Analysis Fold4
4 1 Analysis Fold5
5 2 Analysis Fold2
6 2 Analysis Fold3
7 2 Analysis Fold4
8 2 Analysis Fold5
9 3 Analysis Fold1
10 3 Analysis Fold3
# ℹ 17,805 more rowsDefine the Data-Processing Recipe
For each split, we want to process the data (recipes) and fit the model (parsnip)
Important! We want to preprocess the training subset in each split separately from the split’s subset. This helps us avoid data leakage.
In everything we have done before, we impute using the entire data. We are supposed to be keeping them totally separate.
What we really want is to process the datasets independently, but using the same recipe and steps
Define the Recipe
Only the recipe, no prep(), juice(), or bake()
# Data-processing recipe
# IMPORTANT! NOTICE I AM USING THE TRAINING DATA HERE NOT THE FULL SAMPLE
credit_recipe <- recipe(status_good ~ ., data = credit_train) %>%
# Mean imputation for numeric predictors
step_impute_mean(all_predictors() & all_numeric()) %>%
# KNN imputation for categorical predictors
step_impute_knn(all_predictors() & all_nominal(), neighbors = 5) %>%
# Create dummies for categorical variables
step_dummy(all_predictors() & all_nominal()) %>%
# Interactions
step_interact(~income:starts_with("home"))
# Check the result
credit_recipe── Recipe ──────────────────────────────────────────────────────────────────────── Inputs Number of variables by roleoutcome: 1
predictor: 13── Operations • Mean imputation for: all_predictors() & all_numeric()• K-nearest neighbor imputation for: all_predictors() & all_nominal()• Dummy variables from: all_predictors() & all_nominal()• Interactions with: income:starts_with("home")Define the Model
Tell it both the type of model and the desired engine
Keeping it simple with linear_reg() and the standard "lm"
- Feel free to try whatever you want here in the code to test things out
Although not necessary for "lm", we will set the mode using set_mode(). Just good to get into the habit of things
Putting it Together in a Workflow
We have three different objects floating around
- The defined CV splits for the training dataset (
credit_cv) - The recipe for processing the data (
credit_recipe) - The model that we want to fit on the trianing data (folds)
Now we need to put them together.
- Repeating the preprocessing for each training and validation split could be annoying if you did it all by hand.
- Luckily, we will be uwing the
workflowspackage withintidymodels
How? Literally by defining a workflow with the workflow() function and then pipe in your model and then recipe
══ Workflow ════════════════════════════════════════════════════════════════════
Preprocessor: Recipe
Model: linear_reg()
── Preprocessor ────────────────────────────────────────────────────────────────
4 Recipe Steps
• step_impute_mean()
• step_impute_knn()
• step_dummy()
• step_interact()
── Model ───────────────────────────────────────────────────────────────────────
Linear Regression Model Specification (regression)
Computational engine: lm Putting it Together in a Workflow
The workflow we just wrote defines the model, the data processing, and the engine.
We will have a couple of options:
1. Fit a Single Model Without CV
2. Fit a Single Model With CV
3. Tune your model (using CV)
Putting it Together in a Workflow
The workflow we just wrote defines the model, the data processing, and the engine.
1. Fit a Single Model Without CV
If you only want to fit the model (with no CV), you can pipe the workflow to fit(data = dataframe)
- This is not what we will ultimately want as we want to cross-validate using the folds we created
# Workflow and model fit
workflow() %>%
add_model(model_lm) %>%
add_recipe(credit_recipe) %>%
fit(data = credit_df)══ Workflow [trained] ══════════════════════════════════════════════════════════
Preprocessor: Recipe
Model: linear_reg()
── Preprocessor ────────────────────────────────────────────────────────────────
4 Recipe Steps
• step_impute_mean()
• step_impute_knn()
• step_dummy()
• step_interact()
── Model ───────────────────────────────────────────────────────────────────────
Call:
stats::lm(formula = ..y ~ ., data = data)
Coefficients:
(Intercept) seniority time
6.855e-01 9.267e-03 -8.311e-05
age expenses income
-1.525e-03 -2.120e-03 1.250e-03
assets debt amount
2.493e-06 -1.750e-05 -2.344e-04
price home_other home_owner
8.991e-05 6.054e-02 2.588e-01
home_parents home_priv home_rent
2.135e-01 1.559e-01 7.157e-02
marital_married marital_separated marital_single
7.142e-02 -1.320e-01 8.053e-03
marital_widow records_yes job_freelance
1.179e-02 -2.969e-01 -1.174e-01
job_others job_partime income_x_home_other
-1.341e-01 -2.979e-01 -4.669e-04
income_x_home_owner income_x_home_parents income_x_home_priv
-5.550e-04 -4.636e-04 -5.933e-04
income_x_home_rent
-5.508e-05 Putting it Together in a Workflow
The workflow we just wrote defines the model, the data processing, and the engine.
2. Fit a Single Model With CV
If you want to estimate your model’s out-of-sample performance, then you can use fit_resamples() And we can use collect_metrics() function to assess the performance across the resamples
- By default, it will average across the folds (notice it says
n = 5, which is our folds).
fit_lm_cv <-
workflow() %>%
add_model(model_lm) %>%
add_recipe(credit_recipe) %>%
fit_resamples(credit_cv)
# Check performance
fit_lm_cv %>% collect_metrics()# A tibble: 2 × 6
.metric .estimator mean n std_err .config
<chr> <chr> <dbl> <int> <dbl> <chr>
1 rmse standard 0.390 5 0.00428 pre0_mod0_post0
2 rsq standard 0.255 5 0.0163 pre0_mod0_post0Putting it Together in a Workflow
The workflow we just wrote defines the model, the data processing, and the engine.
2. Fit a Single Model With CV
- If you wanted to see how the model performs on each fold then you can set
summarize = Finsidecollect_metrics()
# A tibble: 10 × 5
id .metric .estimator .estimate .config
<chr> <chr> <chr> <dbl> <chr>
1 Fold1 rmse standard 0.393 pre0_mod0_post0
2 Fold1 rsq standard 0.257 pre0_mod0_post0
3 Fold2 rmse standard 0.387 pre0_mod0_post0
4 Fold2 rsq standard 0.255 pre0_mod0_post0
5 Fold3 rmse standard 0.377 pre0_mod0_post0
6 Fold3 rsq standard 0.308 pre0_mod0_post0
7 Fold4 rmse standard 0.391 pre0_mod0_post0
8 Fold4 rsq standard 0.246 pre0_mod0_post0
9 Fold5 rmse standard 0.404 pre0_mod0_post0
10 Fold5 rsq standard 0.206 pre0_mod0_post0Putting it Together in a Workflow
RMSE and \(R^{2}\) are the default metrics. We can change them using the metrics argument inside of fit_resamples()
You can pass whichever metric you want to use from the yardstick package.
- Suppose we want Mean Absolute Error (MAE) as well
# Define Workflow, then fit model on folds
fit_lm_cv <-
workflow() %>%
add_model(model_lm) %>%
add_recipe(credit_recipe) %>%
fit_resamples(credit_cv, metrics = metric_set(rmse, rsq, mae))
# Check performance
fit_lm_cv %>% collect_metrics()# A tibble: 3 × 6
.metric .estimator mean n std_err .config
<chr> <chr> <dbl> <int> <dbl> <chr>
1 mae standard 0.320 5 0.00386 pre0_mod0_post0
2 rmse standard 0.390 5 0.00428 pre0_mod0_post0
3 rsq standard 0.255 5 0.0163 pre0_mod0_post0Putting it Together in a Workflow
The workflow we just wrote defines the model, the data processing, and the engine.
3. Tune your model (using CV)
Instead of “just” estimating out-of-sample model performance for a specified model and pre-processing operations, we will often want to tune a model’s hyperparameter(s)
Suppose we are using \(k\) nearest neighbors instead of a linear regression. Then we either need to specify a \(k\) or we need to tune k.
By tune we mean estimate the out-of-sample performance for various values of \(k\) and then choose the “best” \(k\) (based on some metric that defines “best”).
To begin, let’s define a KNN model using nearest_neighbor(). We will tell it to tune by specifiying we want to tune() as our neighbors value.
Tuning KNN
Almost ready to fit the model, but we still need to let R know which values to try for the parameter we are tuning
To tune our hyperparameter(s), we will use the tune_grid() function (instead of fit() or fit_resamples() functions)
tune_grid() works similarly to fit_resamples() except it takes an additional argument: grid
- We will pass the possible values of our hyperparameter(s) to the
gridargument and it will evaluate each fold of our samples on each set of hyperparameter(s) we fed it
Tuning KNN
KNN only needs one hyperparameter: neighbors. We can give the grid argument a data.frame of all the values we would like to try, where the column’s name matches the hyperparameter we are trying to tune neighbors
- If, like me, you don’t know what may be “reasonable” values, you can see what is suggested by the function with the same name as the hyperparameter (
neighbors()in this case)
Tuning KNN
Caution
As you add more values of hyperparameters, you fit more models.
More models means more computation.
More computation means more time.
Keep in mind that you are already fitting a model on each fold (5 in our example) so if you want to test 100 values of neighbors then you need to fit \(100 \times 5\) models
Let’s try 1 neighbor (surely not the best) and then 5, and then 10 to 100 by 10s
Tuning KNN
Let’s try 1 neighbor (surely not the best) and then 5, and then 10 to 100 by 10s
# Define the workflow
workflow_knn <- workflow() %>%
add_model(model_knn) %>%
add_recipe(credit_recipe)
# Fit the workflow on our predefined folds and hyperparameters
fit_knn_cv <- workflow_knn %>%
tune_grid(
# CV data
credit_cv,
# Grid where we put our values
grid = data.frame(neighbors = c(1, 5, seq(10,100,10))),
metrics = metric_set(rmse, rsq, mae)
)
# Check performance
fit_knn_cv %>% collect_metrics()# A tibble: 36 × 7
neighbors .metric .estimator mean n std_err .config
<dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
1 1 mae standard 0.298 5 0.00884 pre0_mod01_post0
2 1 rmse standard 0.546 5 0.00808 pre0_mod01_post0
3 1 rsq standard 0.0706 5 0.00940 pre0_mod01_post0
4 5 mae standard 0.301 5 0.00667 pre0_mod02_post0
5 5 rmse standard 0.449 5 0.00632 pre0_mod02_post0
6 5 rsq standard 0.129 5 0.0130 pre0_mod02_post0
7 10 mae standard 0.304 5 0.00520 pre0_mod03_post0
8 10 rmse standard 0.424 5 0.00488 pre0_mod03_post0
9 10 rsq standard 0.160 5 0.0126 pre0_mod03_post0
10 20 mae standard 0.308 5 0.00469 pre0_mod04_post0
# ℹ 26 more rowsTuning KNN
Recognize what we just did was pretty cool: We changed very little to move from a cross-validated regression model to a CV-tuned KNN model.
The output is also cool, but overwhelming.
We can use the show_best() function to have it tell us which value(s) of the hyperparameters performed best according to the metrics we asked for. We either need to give show_best() the name of the metric to use, or it will choose one for us.
show_best()also defaults to showing the top 5 models but you can change that with thenargumentselect_best()allows us to select the best model (based on the metrics)
# A tibble: 5 × 7
neighbors .metric .estimator mean n std_err .config
<dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
1 60 rmse standard 0.404 5 0.00261 pre0_mod08_post0
2 70 rmse standard 0.404 5 0.00245 pre0_mod09_post0
3 50 rmse standard 0.404 5 0.00283 pre0_mod07_post0
4 80 rmse standard 0.404 5 0.00232 pre0_mod10_post0
5 40 rmse standard 0.404 5 0.00315 pre0_mod06_post0# A tibble: 3 × 7
neighbors .metric .estimator mean n std_err .config
<dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
1 100 rsq standard 0.210 5 0.0105 pre0_mod12_post0
2 90 rsq standard 0.210 5 0.0105 pre0_mod11_post0
3 80 rsq standard 0.209 5 0.0105 pre0_mod10_post0We Can Visualize It
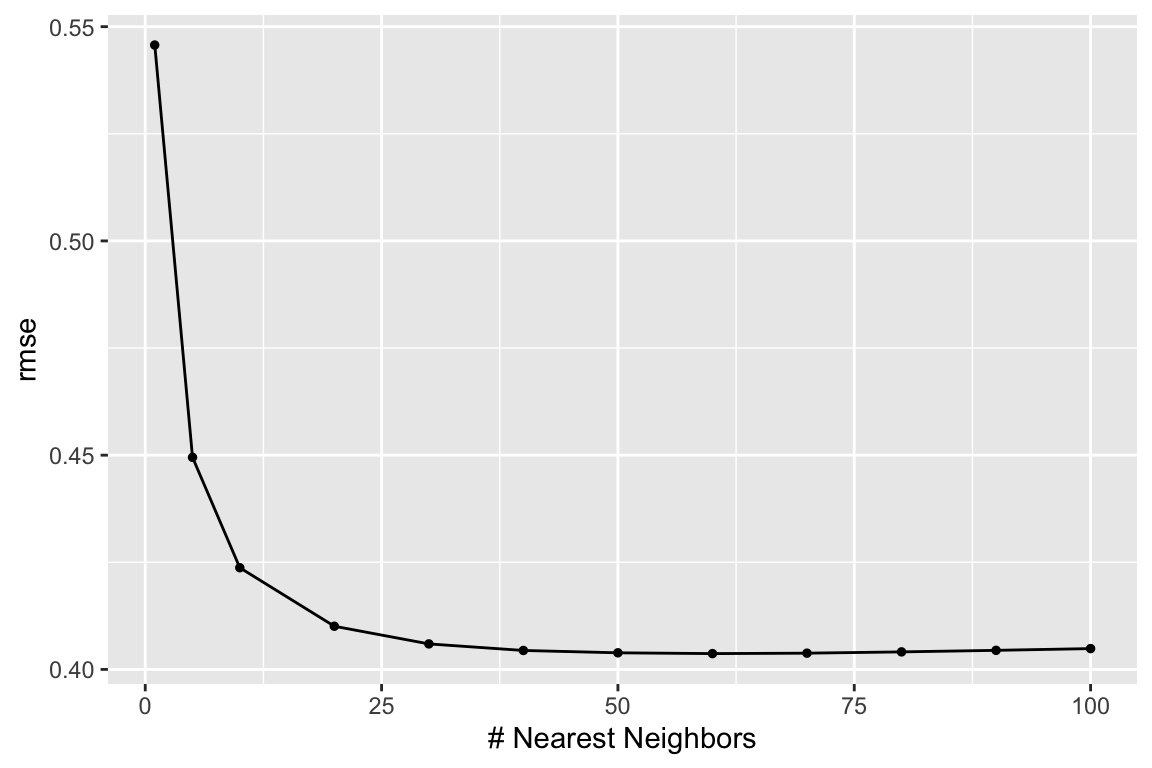
We can use the autoplot() function for a nice and quick picture (we can improve on it too).
We just need to tell it what metric we want to map.
Visualize it Part 02
# Plot metrics' averages across values of 'k'
fit_knn_cv %>%
collect_metrics(summarize = T) %>%
ggplot(aes(x = neighbors, y = mean)) +
geom_line(size = 0.7, alpha = 0.6) +
geom_point(size = 2.5) +
facet_wrap(~ toupper(.metric), scales = "free", nrow = 1) +
scale_x_continuous("Neighbors (k)", labels = scales::label_number()) +
scale_y_continuous("Estimate") +
scale_color_viridis_d("CV Folds:") +
theme_minimal() +
theme(legend.position = "bottom")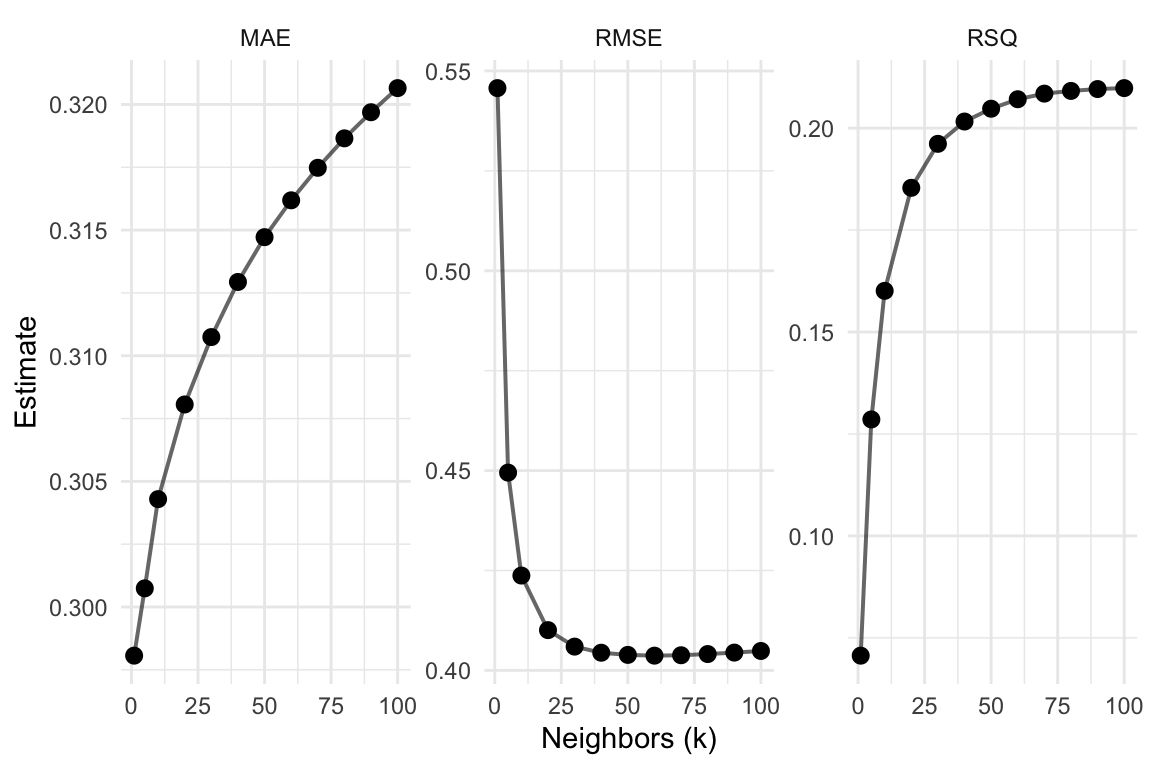
Visualize it Part 03
# Show each fold's metrics across values of 'k'
fit_knn_cv %>%
collect_metrics(summarize = F) %>%
ggplot(aes(x = neighbors, y = .estimate, color = id)) +
geom_line(size = 0.7, alpha = 0.6) +
geom_point(size = 2.5) +
facet_wrap(~ toupper(.metric), scales = "free", nrow = 1) +
scale_x_continuous("Neighbors (k)", labels = scales::label_number()) +
scale_y_continuous("Estimate") +
scale_color_viridis_d("CV Folds:") +
theme_minimal() +
theme(legend.position = "bottom")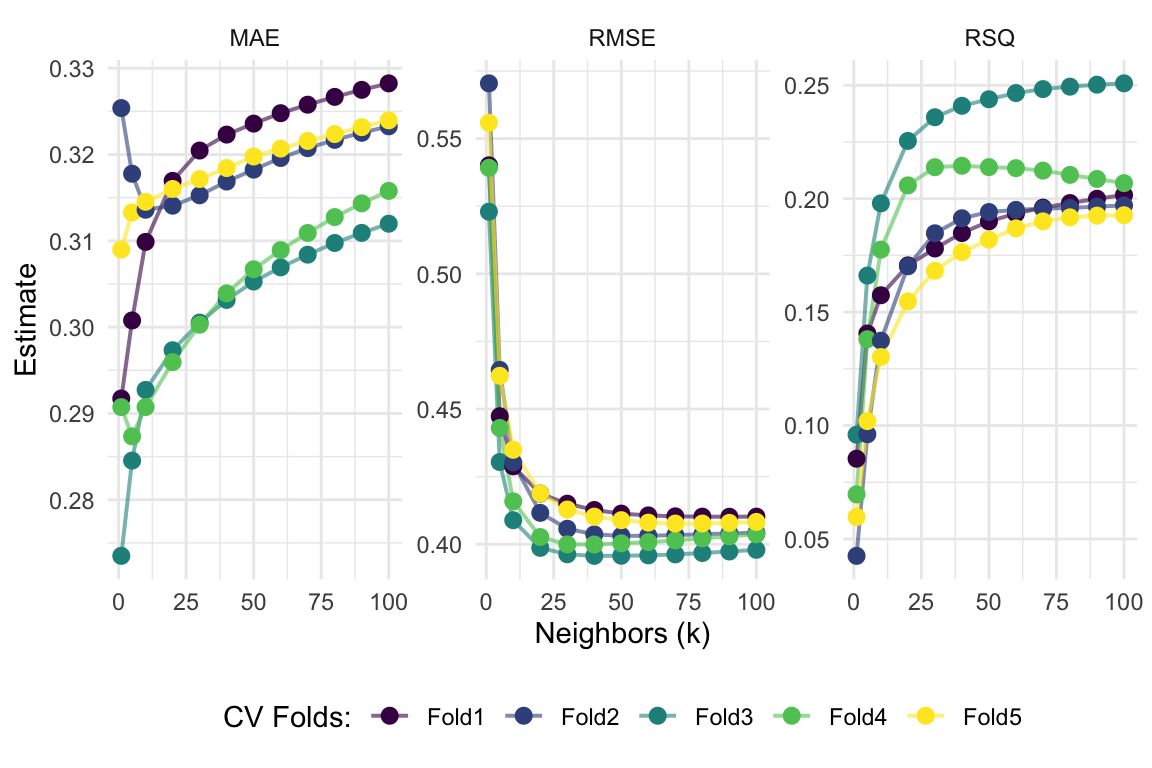
Lastly: Prediction!
We are just about finished
We will wrap it up by taking our selected model (the best model), fitting it on all of the training data, and then predict onto the test data
- Recall we pick our “best” model (using whatever
metricwe prefer) using theselect_best()function
Once we found the “best” model, we wrap up our workflow() using finalize_workflow()
- This function wants (1) your initial workflow and (2) your best model
# The final workflow for our KNN model
final_knn <-
workflow_knn %>%
finalize_workflow(select_best(fit_knn_cv, metric = "rmse"))
# Check out the final workflow object
final_knn══ Workflow ════════════════════════════════════════════════════════════════════
Preprocessor: Recipe
Model: nearest_neighbor()
── Preprocessor ────────────────────────────────────────────────────────────────
4 Recipe Steps
• step_impute_mean()
• step_impute_knn()
• step_dummy()
• step_interact()
── Model ───────────────────────────────────────────────────────────────────────
K-Nearest Neighbor Model Specification (regression)
Main Arguments:
neighbors = 60
Computational engine: kknn Prediction
Now we fit this finalized workflow to the full training dataset (credit_train).
Enter fit().
# Fitting our final workflow
final_fit_knn <- final_knn %>% fit(data = credit_train)
# Examine the final workflow
final_fit_knn══ Workflow [trained] ══════════════════════════════════════════════════════════
Preprocessor: Recipe
Model: nearest_neighbor()
── Preprocessor ────────────────────────────────────────────────────────────────
4 Recipe Steps
• step_impute_mean()
• step_impute_knn()
• step_dummy()
• step_interact()
── Model ───────────────────────────────────────────────────────────────────────
Call:
kknn::train.kknn(formula = ..y ~ ., data = data, ks = min_rows(60, data, 5))
Type of response variable: continuous
minimal mean absolute error: 0.3129085
Minimal mean squared error: 0.1606585
Best kernel: optimal
Best k: 60Prediction
Finally, we predict onto the testing data by using predict()
Prediction but Easier
Once we have a final workflow (final_knn is the workflow plus the selected best model), you can pass it to the last_fit() function
Along with the initial split (credit_split in our case) to both:
Fit your final model on your full training dataset
Make predictions onto the testing dataset (defined by the initial split object)
This last_fit() approach streamlines your work and also lets you easily collect metrics using the collect_metrics() function from before
# Write over 'final_fit_knn' with this last_fit() approach
final_fit_knn <- final_knn %>% last_fit(credit_split)
# Collect metrics on the test data!
final_fit_knn %>% collect_metrics()# A tibble: 2 × 4
.metric .estimator .estimate .config
<chr> <chr> <dbl> <chr>
1 rmse standard 0.392 pre0_mod0_post0
2 rsq standard 0.225 pre0_mod0_post0Prediction but Easier
If instead you want predictions, just use the collect_predictions() on the output from last_fit()
# A tibble: 6 × 5
.pred id status_good .row .config
<dbl> <chr> <dbl> <int> <chr>
1 0.955 train/test split 1 6 pre0_mod0_post0
2 0.940 train/test split 1 8 pre0_mod0_post0
3 0.952 train/test split 1 16 pre0_mod0_post0
4 0.290 train/test split 0 23 pre0_mod0_post0
5 0.919 train/test split 1 24 pre0_mod0_post0
6 0.957 train/test split 1 28 pre0_mod0_post0EC 524 Week 04 Lab | tidymodels cont.