QC filtered sequencing reads
Evangelyn Sim
2/15/2021
Last updated: 2021-02-16
Checks: 6 1
Knit directory: Mouse_PRKO_RNAseq_bulk/
This reproducible R Markdown analysis was created with workflowr (version 1.6.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20210216) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Using absolute paths to the files within your workflowr project makes it difficult for you and others to run your code on a different machine. Change the absolute path(s) below to the suggested relative path(s) to make your code more reproducible.
| absolute | relative |
|---|---|
| /group/card2/Evangelyn_Sim/Transcriptome_chromatin_human/Sequencing_ATAC_RNA/GITHUB/Mouse_PRKO_RNAseq_bulk/output/mrna_fulllen_pe_strrev_q30.mx.PRKO.all.fix_filt | output/mrna_fulllen_pe_strrev_q30.mx.PRKO.all.fix_filt |
| /group/card2/Evangelyn_Sim/Transcriptome_chromatin_human/Sequencing_ATAC_RNA/GITHUB/Mouse_PRKO_RNAseq_bulk/output/20200211_PRKO_RNA_samplesheet.txt | output/20200211_PRKO_RNA_samplesheet.txt |
| /group/card2/Evangelyn_Sim/Transcriptome_chromatin_human/Sequencing_ATAC_RNA/GITHUB/Mouse_PRKO_RNAseq_bulk/output/mrna_fulllen_pe_strrev_q30.mx.chr | output/mrna_fulllen_pe_strrev_q30.mx.chr |
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 7f1a156. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .Rproj.user/
Untracked files:
Untracked: Mus_musculus.GRCm38.96.fulllength.saf
Untracked: analysis/00.WorkFlowR_setting.R
Untracked: header.sam
Untracked: logCPM_mrna_fulllen_pe_strrev_q30.mx.PRKO.all.fix_filt.csv
Untracked: output/20200211_PRKO_RNA_samplesheet.txt
Untracked: output/mrna_fulllen_pe_strrev_q30.mx.PRKO.all.fix_filt
Untracked: output/mrna_fulllen_pe_strrev_q30.mx.PRKO.unfiltered
Untracked: output/mrna_fulllen_pe_strrev_q30.mx.chr
Unstaged changes:
Modified: analysis/index.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were made to the R Markdown (analysis/04.MDS.Rmd) and HTML (docs/04.MDS.html) files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 7f1a156 | evangelynsim | 2021-02-16 | wflow_publish(c(“analysis/01.Generate_reference_genome.Rmd”, |
Introduction
In the GEO submission 2 processed files were uploaded.
- mrna_fulllen_pe_strrev_q30.mx.PRKO.unfiltered
- mrna_fulllen_pe_strrev_q30.mx.PRKO.all.fix_filt
They have been uploaded in the /output folder and will be used below to generate different figures.
Used libraries and functions
library(edgeR)Loading required package: limmalibrary(limma)
library(Glimma)
library(gplots)
Attaching package: 'gplots'The following object is masked from 'package:stats':
lowesslibrary(ComplexHeatmap)Loading required package: grid========================================
ComplexHeatmap version 2.0.0
Bioconductor page: http://bioconductor.org/packages/ComplexHeatmap/
Github page: https://github.com/jokergoo/ComplexHeatmap
Documentation: http://jokergoo.github.io/ComplexHeatmap-reference
If you use it in published research, please cite:
Gu, Z. Complex heatmaps reveal patterns and correlations in multidimensional
genomic data. Bioinformatics 2016.
========================================library(circlize)========================================
circlize version 0.4.10
CRAN page: https://cran.r-project.org/package=circlize
Github page: https://github.com/jokergoo/circlize
Documentation: https://jokergoo.github.io/circlize_book/book/
If you use it in published research, please cite:
Gu, Z. circlize implements and enhances circular visualization
in R. Bioinformatics 2014.
This message can be suppressed by:
suppressPackageStartupMessages(library(circlize))
========================================library(RColorBrewer)
library(mclust)Package 'mclust' version 5.4.6
Type 'citation("mclust")' for citing this R package in publications.Read files
rm1 <- read.delim("/group/card2/Evangelyn_Sim/Transcriptome_chromatin_human/Sequencing_ATAC_RNA/GITHUB/Mouse_PRKO_RNAseq_bulk/output/mrna_fulllen_pe_strrev_q30.mx.PRKO.all.fix_filt", row.names = 1)
info = read.delim("/group/card2/Evangelyn_Sim/Transcriptome_chromatin_human/Sequencing_ATAC_RNA/GITHUB/Mouse_PRKO_RNAseq_bulk/output/20200211_PRKO_RNA_samplesheet.txt", header = TRUE, sep = "\t")
info$ID = gsub("-",".",info$ID)
names(rm1) = info$ID[match(names(rm1),info$ID)]
m = match(info$ID,names(rm1))
rm2 = rm1[,m]
rm1 = rm2
sampleinfo = info
levels(factor(sampleinfo$Group))[1] "KO" "WT"levels(factor(sampleinfo$BinSex))[1] "KO_F" "KO_M" "WT_F" "WT_M"sampleinfo$colour = c("aquamarine2","mediumpurple1")[factor(sampleinfo$Group)]
table(colnames(rm2)==sampleinfo$ID)
TRUE
15 y <- DGEList(rm2)Make plot
par(mfrow=c(1,3))
plotMDS(y, pch=c(0,1)[factor(sampleinfo$Group)], col=sampleinfo$colour,dim.plot = c(1,2), cex = 2)
legend("top", legend = c("AAV_MCS","AAV_PR"), pch=c(1,0), col = c("mediumpurple1", "aquamarine2"), cex=1)
plotMDS(y, pch=c(0,15,1,16)[factor(sampleinfo$BinSex)], col=sampleinfo$colour, dim.plot = c(1,2),cex = 2)
legend("top", legend = c("AAV_MCS","AAV_PR"), pch=c(1,0), col = c("mediumpurple1", "aquamarine2"), cex=1)
legend("topleft", legend = c("F","M"), pch=c(1,16), col = c("grey"), cex=1)
plotMDS(y, cex = 0.8, dim.plot = c(1,2))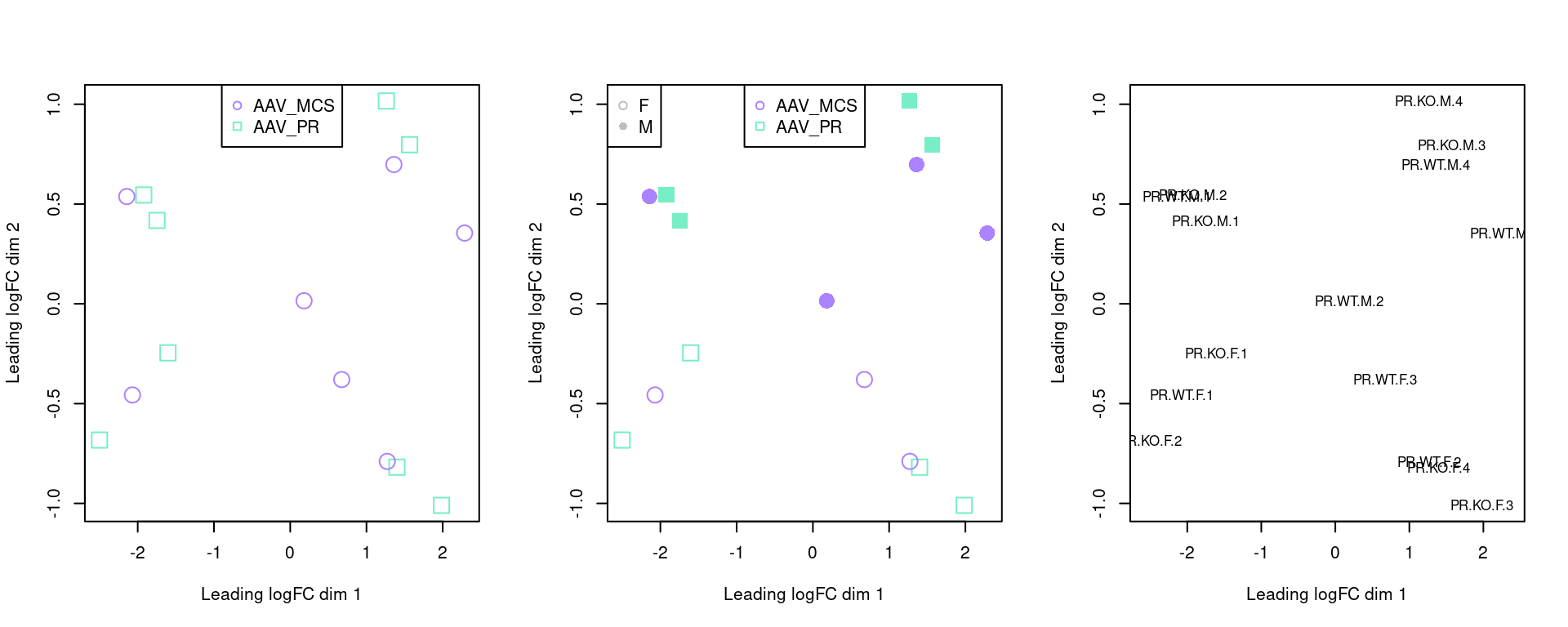
Remove Chr X & Y genes
rm3 = read.delim("/group/card2/Evangelyn_Sim/Transcriptome_chromatin_human/Sequencing_ATAC_RNA/GITHUB/Mouse_PRKO_RNAseq_bulk/output/mrna_fulllen_pe_strrev_q30.mx.chr", header = TRUE, row.names = 1)
rm3 = merge(rm3, rm2, by = "row.names")
rm3 = rm3[!grepl("Y",rm3$Chr),]
rm3 = rm3[!grepl("X",rm3$Chr),]
row.names(rm3)= rm3$Row.names
rm3 = rm3[,c(7:ncol(rm3))]
rm2 = rm3
z <- DGEList(rm3)Make MDS plot (removed Chr X & Y genes)
par(mfrow=c(1,3))
plotMDS(z, pch=c(0,1)[factor(sampleinfo$Group)], col=sampleinfo$colour,dim.plot = c(3,4), cex = 2)
legend("topright", legend = c("AAV_MCS","AAV_PR"), pch=c(1,0), col = c("purple", "springgreen3"), cex=1)
plotMDS(z, pch=c(0,15,1,16)[factor(sampleinfo$BinSex)], col=sampleinfo$colour, dim.plot = c(3,4),cex = 2)
legend("topright", legend = c("AAV_MCS","AAV_PR"), pch=c(1,0), col = c("purple", "springgreen3"), cex=1)
legend("bottomright", legend = c("F","M"), pch=c(1,16), col = c("grey"), cex=1)
plotMDS(z, cex = 0.8, dim.plot = c(3,4))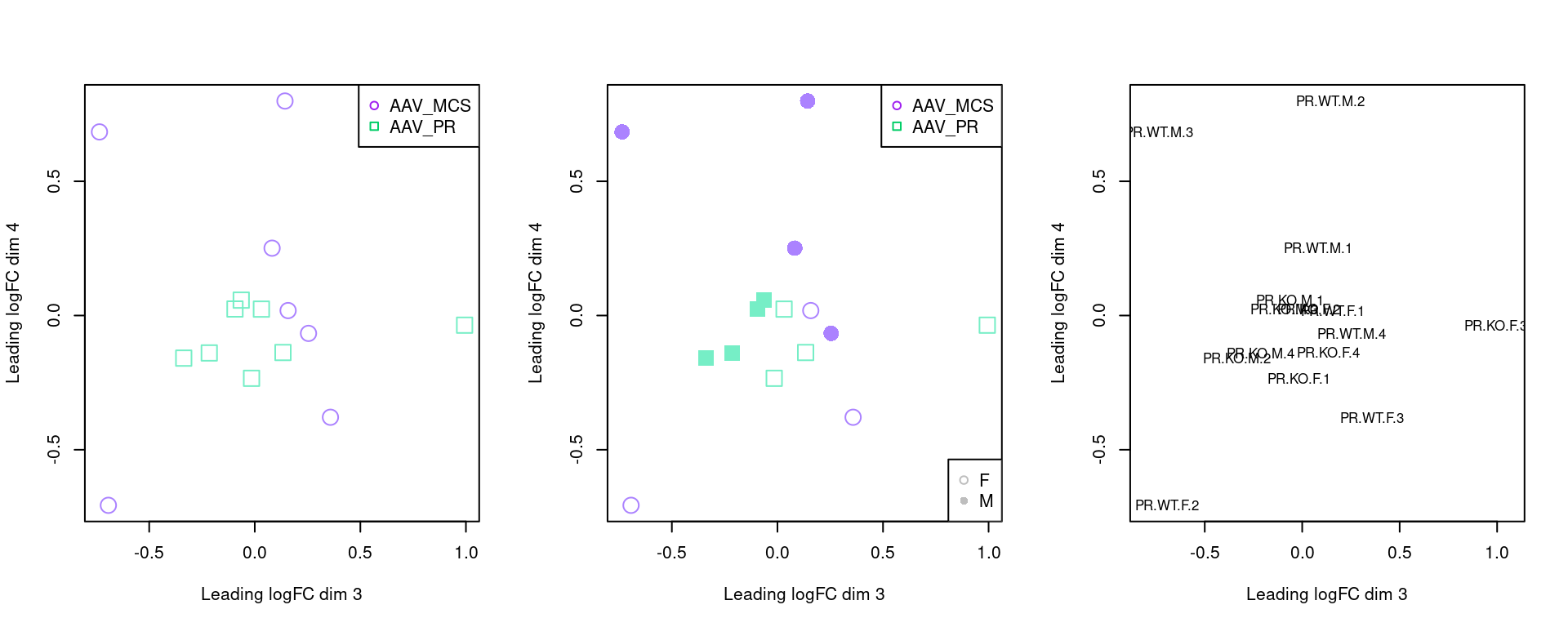
sessionInfo()R version 3.6.1 (2019-07-05)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: CentOS Linux 7 (Core)
Matrix products: default
BLAS: /hpc/software/installed/R/3.6.1/lib64/R/lib/libRblas.so
LAPACK: /hpc/software/installed/R/3.6.1/lib64/R/lib/libRlapack.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] grid stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] mclust_5.4.6 RColorBrewer_1.1-2 circlize_0.4.10
[4] ComplexHeatmap_2.0.0 gplots_3.1.0 Glimma_1.12.0
[7] edgeR_3.26.8 limma_3.40.6 workflowr_1.6.2
loaded via a namespace (and not attached):
[1] Rcpp_1.0.5 pillar_1.4.6 compiler_3.6.1
[4] later_1.1.0.1 git2r_0.27.1 highr_0.8
[7] bitops_1.0-6 tools_3.6.1 digest_0.6.27
[10] clue_0.3-57 jsonlite_1.7.0 evaluate_0.14
[13] lifecycle_0.2.0 tibble_3.0.3 lattice_0.20-41
[16] png_0.1-7 pkgconfig_2.0.3 rlang_0.4.7
[19] rstudioapi_0.11 parallel_3.6.1 yaml_2.2.1
[22] xfun_0.18 cluster_2.1.0 stringr_1.4.0
[25] knitr_1.30 GlobalOptions_0.1.2 caTools_1.18.0
[28] gtools_3.8.2 fs_1.5.0 vctrs_0.3.2
[31] locfit_1.5-9.4 rprojroot_1.3-2 glue_1.4.2
[34] R6_2.5.0 GetoptLong_1.0.2 rmarkdown_2.5
[37] magrittr_1.5 whisker_0.4 backports_1.1.10
[40] promises_1.1.1 ellipsis_0.3.1 htmltools_0.5.0
[43] shape_1.4.4 colorspace_1.4-1 httpuv_1.5.4
[46] KernSmooth_2.23-17 stringi_1.5.3 rjson_0.2.20
[49] crayon_1.3.4