lane14_comparisons
Haider Inam
2022-10-24
Last updated: 2022-10-29
Checks: 6 1
Knit directory: duplex_sequencing_screen/
This reproducible R Markdown analysis was created with workflowr (version 1.6.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of
the R Markdown file created these results, you’ll want to first commit
it to the Git repo. If you’re still working on the analysis, you can
ignore this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20200402) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 1777490. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: data/Consensus_Data/.Rhistory
Ignored: data/Consensus_Data/Novogene_lane13/sample7/variant_caller_outputs/
Ignored: data/Consensus_Data/Novogene_lane13/sample8/variant_caller_outputs/
Ignored: data/Consensus_Data/Novogene_lane14/sample13/
Ignored: data/Consensus_Data/Novogene_lane14/sample14b/
Ignored: data/Consensus_Data/Novogene_lane14/sample1_combined/
Ignored: data/Consensus_Data/Novogene_lane14/sample8/variant_caller_outputs/
Ignored: data/Consensus_Data/Novogene_lane2/abl_ref/kraken2-master/newdir/
Ignored: data/Consensus_Data/Novogene_lane4/.ipynb_checkpoints/
Ignored: data/Consensus_Data/Novogene_lane4/GalaxyData/initialanalysis/
Ignored: data/Consensus_Data/Novogene_lane4/output/
Ignored: data/Consensus_Data/Novogene_lane6/
Ignored: data/Consensus_Data/Novogene_lane7/
Ignored: data/Consensus_Data/Ranomics_Pooled/RP4/
Ignored: data/Consensus_Data/Ranomics_Pooled/RP5/
Untracked files:
Untracked: analysis/figure/
Untracked: code/merge_samples.R
Untracked: data/Consensus_Data/Novogene_lane11/sample1/.DS_Store
Untracked: data/Consensus_Data/Novogene_lane11/sample1/sscs_sorted_filtered.tsv
Untracked: data/Consensus_Data/Novogene_lane11/sample1/variant_caller_outputs/
Untracked: data/Consensus_Data/Novogene_lane11/sample2/archive/
Untracked: data/Consensus_Data/Novogene_lane11/sample2/sscs_sorted_filtered.tsv
Untracked: data/Consensus_Data/Novogene_lane11/sample2/variant_caller_outputs/
Untracked: data/Consensus_Data/Novogene_lane11/sample3/.DS_Store
Untracked: data/Consensus_Data/Novogene_lane11/sample3/sscs_sorted_filtered.tsv
Untracked: data/Consensus_Data/Novogene_lane11/sample3/variant_caller_outputs/
Untracked: data/Consensus_Data/Novogene_lane11/sample4/.DS_Store
Untracked: data/Consensus_Data/Novogene_lane11/sample4/sscs_sorted_filtered.tsv
Untracked: data/Consensus_Data/Novogene_lane11/sample4/variant_caller_outputs/
Untracked: data/Consensus_Data/Novogene_lane11/sample7/.DS_Store
Untracked: data/Consensus_Data/Novogene_lane11/sample7/archive/
Untracked: data/Consensus_Data/Novogene_lane11/sample7/sscs_sorted_filtered.tsv
Untracked: data/Consensus_Data/Novogene_lane11/sample7/variant_caller_outputs/
Untracked: data/Consensus_Data/Novogene_lane13/sample12/.DS_Store
Untracked: data/Consensus_Data/Novogene_lane14/sample10_combined/variant_caller_outputs/variants_unique_ann.csv
Untracked: data/Consensus_Data/Novogene_lane14/sample11/sscs_sorted_filtered.tsv
Untracked: data/Consensus_Data/Novogene_lane14/sample11/variant_caller_outputs/variants_unique_ann.csv
Untracked: data/Consensus_Data/Novogene_lane14/sample12/sscs_sorted_filtered.tsv
Untracked: data/Consensus_Data/Novogene_lane14/sample12/variant_caller_outputs/variants_unique_ann.csv
Untracked: data/Consensus_Data/Novogene_lane14/sample14_combined/variant_caller_outputs/variants_unique_ann.csv
Untracked: data/Consensus_Data/Novogene_lane14/sample15/variant_caller_outputs/variants_unique_ann.csv
Untracked: data/Consensus_Data/Novogene_lane14/sample16/variant_caller_outputs/variants_unique_ann.csv
Untracked: data/Consensus_Data/Novogene_lane14/sample17/variant_caller_outputs/variants_unique_ann.csv
Untracked: data/Consensus_Data/Novogene_lane14/sample18/.DS_Store
Untracked: data/Consensus_Data/Novogene_lane14/sample18/variant_caller_outputs/variants_unique_ann.csv
Untracked: data/Consensus_Data/Novogene_lane14/sample6/.DS_Store
Untracked: data/Consensus_Data/Novogene_lane14/sample7/.DS_Store
Untracked: data/Consensus_Data/Novogene_lane14/sample9/sscs_sorted_filtered.tsv
Untracked: data/Consensus_Data/Novogene_lane14/sample9/variant_caller_outputs/
Untracked: data/QC_Data_Cloning/
Unstaged changes:
Modified: .DS_Store
Modified: analysis/DupSeq_QC_Lane11.Rmd
Modified: analysis/lane14_comparisons.Rmd
Modified: analysis/variant_caller_2022.Rmd
Modified: code/.DS_Store
Modified: data/.DS_Store
Modified: data/Consensus_Data/.DS_Store
Modified: data/Consensus_Data/Novogene_lane11/.DS_Store
Modified: data/Consensus_Data/Novogene_lane11/sample2/.DS_Store
Deleted: data/Consensus_Data/Novogene_lane11/sample2/Sample_2.1.consensus.bai
Deleted: data/Consensus_Data/Novogene_lane11/sample2/Sample_2.1.consensus.bam
Deleted: data/Consensus_Data/Novogene_lane11/sample2/Sample_2.1.consensus.variant-calls.mut
Deleted: data/Consensus_Data/Novogene_lane11/sample2/duplex_aligned.bam
Deleted: data/Consensus_Data/Novogene_lane11/sample2/duplex_aligned_filtered.tsv
Deleted: data/Consensus_Data/Novogene_lane11/sample2/duplex_reads_ann.csv
Deleted: data/Consensus_Data/Novogene_lane11/sample2/duplex_sum_ann.csv
Deleted: data/Consensus_Data/Novogene_lane11/sample2/sample2_translated.csv
Deleted: data/Consensus_Data/Novogene_lane11/sample2/sscs_aligned.bam
Deleted: data/Consensus_Data/Novogene_lane11/sample2/sscs_aligned_filtered.tsv.gz
Deleted: data/Consensus_Data/Novogene_lane11/sample2/variants_ann_sample2.csv
Deleted: data/Consensus_Data/Novogene_lane11/sample2/variants_unique_ann_sample2.csv
Modified: data/Consensus_Data/Novogene_lane11/sample5/.DS_Store
Modified: data/Consensus_Data/Novogene_lane11/sample6/.DS_Store
Deleted: data/Consensus_Data/Novogene_lane11/sample7/Sample_7.1.consensus.bai
Deleted: data/Consensus_Data/Novogene_lane11/sample7/Sample_7.1.consensus.bam
Deleted: data/Consensus_Data/Novogene_lane11/sample7/Sample_7.1.consensus.variant-calls.mut.tsv
Deleted: data/Consensus_Data/Novogene_lane11/sample7/duplex_aligned_filtered.tsv
Deleted: data/Consensus_Data/Novogene_lane11/sample7/duplex_reads_ann.csv
Deleted: data/Consensus_Data/Novogene_lane11/sample7/duplex_sum_ann.csv
Deleted: data/Consensus_Data/Novogene_lane11/sample7/sscs_aligned_filtered.tsv
Deleted: data/Consensus_Data/Novogene_lane11/sample7/sscs_reads_ann.csv
Deleted: data/Consensus_Data/Novogene_lane11/sample7/sscs_sum_ann.csv
Modified: data/Consensus_Data/Novogene_lane12/.DS_Store
Modified: data/Consensus_Data/Novogene_lane12/sample1/.DS_Store
Modified: data/Consensus_Data/Novogene_lane12/sample3/.DS_Store
Modified: data/Consensus_Data/Novogene_lane13/.DS_Store
Modified: data/Consensus_Data/Novogene_lane13/sample10/.DS_Store
Modified: data/Consensus_Data/Novogene_lane13/sample11/.DS_Store
Modified: data/Consensus_Data/Novogene_lane13/sample9/.DS_Store
Modified: data/Consensus_Data/Novogene_lane14/.DS_Store
Modified: data/Consensus_Data/Novogene_lane14/sample10_combined/.DS_Store
Deleted: data/Consensus_Data/Novogene_lane14/sample10_combined/variant_caller_outputs/variants_unique_ann.csv.gz
Modified: data/Consensus_Data/Novogene_lane14/sample11/.DS_Store
Deleted: data/Consensus_Data/Novogene_lane14/sample11/sscs_sorted_filtered.tsv.gz
Deleted: data/Consensus_Data/Novogene_lane14/sample11/variant_caller_outputs/variants_unique_ann.csv.gz
Modified: data/Consensus_Data/Novogene_lane14/sample12/.DS_Store
Deleted: data/Consensus_Data/Novogene_lane14/sample12/sscs_sorted_filtered.tsv.gz
Deleted: data/Consensus_Data/Novogene_lane14/sample12/variant_caller_outputs/variants_unique_ann.csv.gz
Modified: data/Consensus_Data/Novogene_lane14/sample14_combined/.DS_Store
Deleted: data/Consensus_Data/Novogene_lane14/sample14_combined/variant_caller_outputs/variants_unique_ann.csv.gz
Modified: data/Consensus_Data/Novogene_lane14/sample15/.DS_Store
Deleted: data/Consensus_Data/Novogene_lane14/sample15/variant_caller_outputs/variants_unique_ann.csv.gz
Modified: data/Consensus_Data/Novogene_lane14/sample16/.DS_Store
Deleted: data/Consensus_Data/Novogene_lane14/sample16/variant_caller_outputs/variants_unique_ann.csv.gz
Modified: data/Consensus_Data/Novogene_lane14/sample17/.DS_Store
Deleted: data/Consensus_Data/Novogene_lane14/sample17/variant_caller_outputs/variants_unique_ann.csv.gz
Deleted: data/Consensus_Data/Novogene_lane14/sample18/variant_caller_outputs/variants_unique_ann.csv.gz
Modified: data/Consensus_Data/Novogene_lane14/sample2_combined/.DS_Store
Modified: data/Consensus_Data/Novogene_lane14/sample3/.DS_Store
Modified: data/Consensus_Data/Novogene_lane14/sample4/.DS_Store
Modified: data/Consensus_Data/Novogene_lane14/sample5/.DS_Store
Modified: data/Consensus_Data/Novogene_lane14/sample9/.DS_Store
Modified: data/Consensus_Data/Novogene_lane2/.DS_Store
Modified: data/Consensus_Data/Novogene_lane2/abl_ref/.DS_Store
Modified: data/Consensus_Data/R01Figure/.DS_Store
Modified: data/Consensus_Data/archive/.DS_Store
Modified: data/Refs/.DS_Store
Modified: data/Refs/EGFR/.DS_Store
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (analysis/lane14_comparisons.Rmd)
and HTML (docs/lane14_comparisons.html) files. If you’ve
configured a remote Git repository (see ?wflow_git_remote),
click on the hyperlinks in the table below to view the files as they
were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 157f17b | haiderinam | 2022-10-24 | Lane 14 Data Added |
10.24.22 Analysis of Novogene Lane 14 Sequencing Data The sequencing here was done on EGFR il3 indep and ABL il3 indep and imat, asc treated libraries
sample14=read.csv(file = "data/Consensus_Data/Novogene_lane14/sample14_combined/variant_caller_outputs/variants_unique_ann.csv",header=T,stringsAsFactors = F)
sample14=sample14%>%
rowwise()%>%
mutate(ref_aa=strsplit(amino_acids,"/")[[1]][1],
alt_aa=strsplit(amino_acids,"/")[[1]][2])
sample14=sample14%>%mutate(maf=ct/depth)
sample14_simple=sample14%>%dplyr::select(alt_start_pos,protein_start,ref,alt,ref_aa,alt_aa,consequence_terms,ct,depth,maf)
sample15=read.csv(file = "data/Consensus_Data/Novogene_lane14/sample15/variant_caller_outputs/variants_unique_ann.csv",header=T,stringsAsFactors = F)
sample15=sample15%>%
rowwise()%>%
mutate(ref_aa=strsplit(amino_acids,"/")[[1]][1],
alt_aa=strsplit(amino_acids,"/")[[1]][2])
sample15=sample15%>%mutate(maf=ct/depth)
sample15_simple=sample15%>%dplyr::select(alt_start_pos,protein_start,ref,alt,ref_aa,alt_aa,consequence_terms,ct,depth,maf)
sample16=read.csv(file = "data/Consensus_Data/Novogene_lane14/sample16/variant_caller_outputs/variants_unique_ann.csv",header=T,stringsAsFactors = F)
sample16=sample16%>%
rowwise()%>%
mutate(ref_aa=strsplit(amino_acids,"/")[[1]][1],
alt_aa=strsplit(amino_acids,"/")[[1]][2])
sample16=sample16%>%mutate(maf=ct/depth)
sample16_simple=sample16%>%dplyr::select(alt_start_pos,protein_start,ref,alt,ref_aa,alt_aa,consequence_terms,ct,depth,maf)
sample17=read.csv(file = "data/Consensus_Data/Novogene_lane14/sample18/variant_caller_outputs/variants_unique_ann.csv",header=T,stringsAsFactors = F)
sample17=sample17%>%
rowwise()%>%
mutate(ref_aa=strsplit(amino_acids,"/")[[1]][1],
alt_aa=strsplit(amino_acids,"/")[[1]][2])
sample17=sample17%>%mutate(maf=ct/depth)
sample17_simple=sample17%>%dplyr::select(alt_start_pos,protein_start,ref,alt,ref_aa,alt_aa,consequence_terms,ct,depth,maf)
sample18=read.csv(file = "data/Consensus_Data/Novogene_lane14/sample18/variant_caller_outputs/variants_unique_ann.csv",header=T,stringsAsFactors = F)
sample18=sample18%>%
rowwise()%>%
mutate(ref_aa=strsplit(amino_acids,"/")[[1]][1],
alt_aa=strsplit(amino_acids,"/")[[1]][2])
sample18=sample18%>%mutate(maf=ct/depth)
sample18_simple=sample18%>%dplyr::select(alt_start_pos,protein_start,ref,alt,ref_aa,alt_aa,consequence_terms,ct,depth,maf)
samples_14.16=merge(sample14_simple%>%filter(consequence_terms%in%"missense_variant"),sample16_simple%>%filter(consequence_terms%in%"missense_variant"),by=c("ref_aa","protein_start","alt_aa","ref","alt","alt_start_pos","consequence_terms"))
samples_14.16=samples_14.16%>%mutate(score=log2(maf.y/maf.x),score2=log2(ct.y/ct.x))
samples_14.16_simple=samples_14.16%>%dplyr::select(ref_aa,protein_start,alt_aa,consequence_terms,ct.x,score)
samples_14.18=merge(sample14_simple%>%filter(consequence_terms%in%"missense_variant"),sample18_simple%>%filter(consequence_terms%in%"missense_variant"),by=c("ref_aa","protein_start","alt_aa","ref","alt","alt_start_pos","consequence_terms"))
samples_14.18=samples_14.18%>%mutate(score=log2(maf.y/maf.x),score2=log2(ct.y/ct.x))
samples_14.18_simple=samples_14.18%>%dplyr::select(ref_aa,protein_start,alt_aa,consequence_terms,ct.x,score)
asc_scores=merge(samples_14.16_simple,samples_14.18_simple,by=c("ref_aa","protein_start","alt_aa","consequence_terms"))
samples_14.17=merge(sample14_simple%>%filter(consequence_terms%in%"missense_variant"),sample17_simple%>%filter(consequence_terms%in%"missense_variant"),by=c("ref_aa","protein_start","alt_aa","ref","alt","alt_start_pos","consequence_terms"))
samples_14.17=samples_14.17%>%mutate(score=log2(maf.y/maf.x),score2=log2(ct.y/ct.x))
samples_14.17_simple=samples_14.17%>%dplyr::select(ref_aa,protein_start,alt_aa,consequence_terms,ct.x,score)
samples_14.15=merge(sample14_simple%>%filter(consequence_terms%in%"missense_variant"),sample15_simple%>%filter(consequence_terms%in%"missense_variant"),by=c("ref_aa","protein_start","alt_aa","ref","alt","alt_start_pos","consequence_terms"))
samples_14.15=samples_14.15%>%mutate(score=log2(maf.y/maf.x),score2=log2(ct.y/ct.x))
samples_14.15_simple=samples_14.15%>%dplyr::select(ref_aa,protein_start,alt_aa,consequence_terms,ct.x,score)
imat_scores=merge(samples_14.15_simple,samples_14.17_simple,by=c("ref_aa","protein_start","alt_aa","consequence_terms"))
ggplot(samples_14.16,aes(x=score))+geom_density()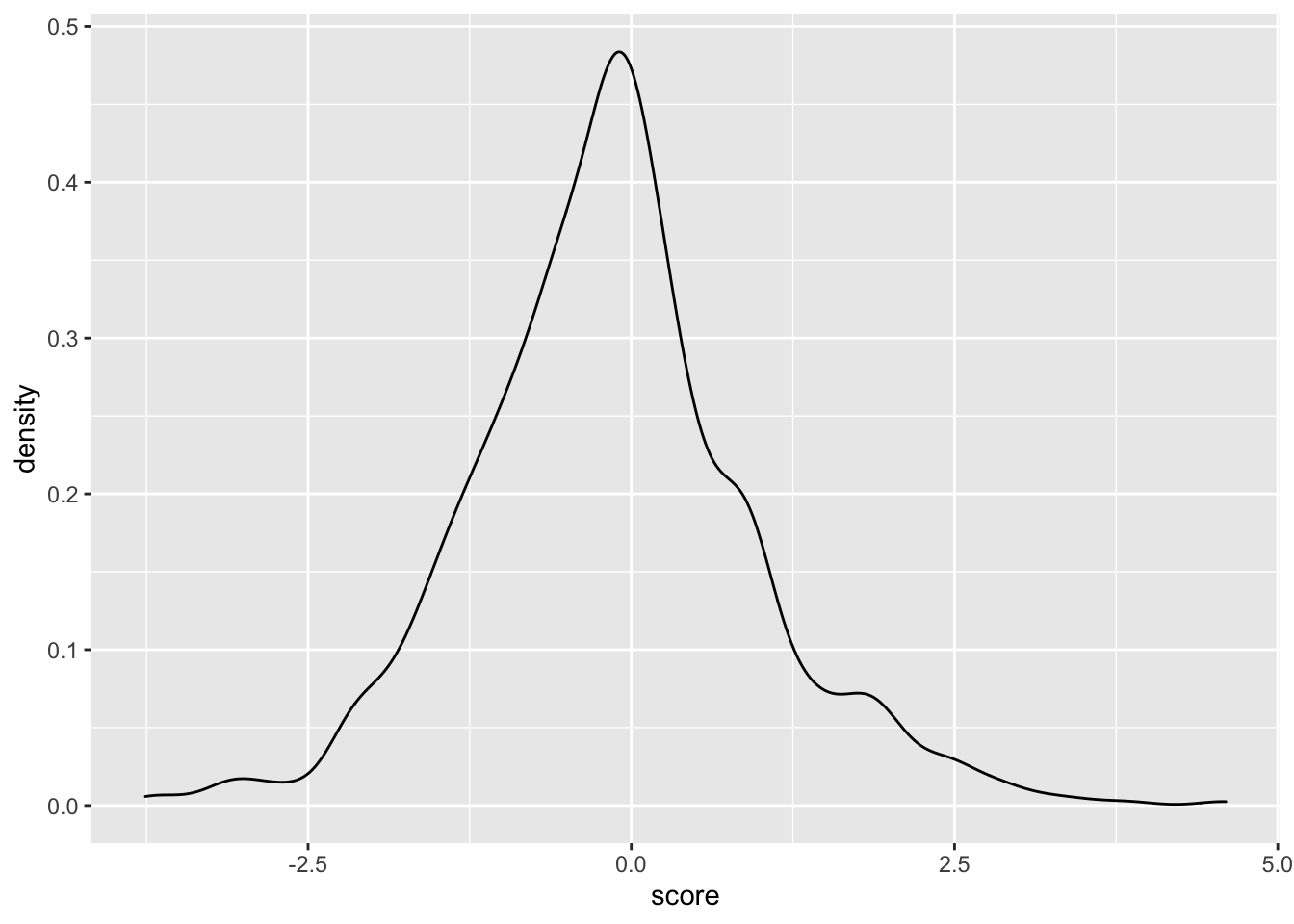
ggplot(samples_14.16,aes(x=score))+geom_histogram(bins=100)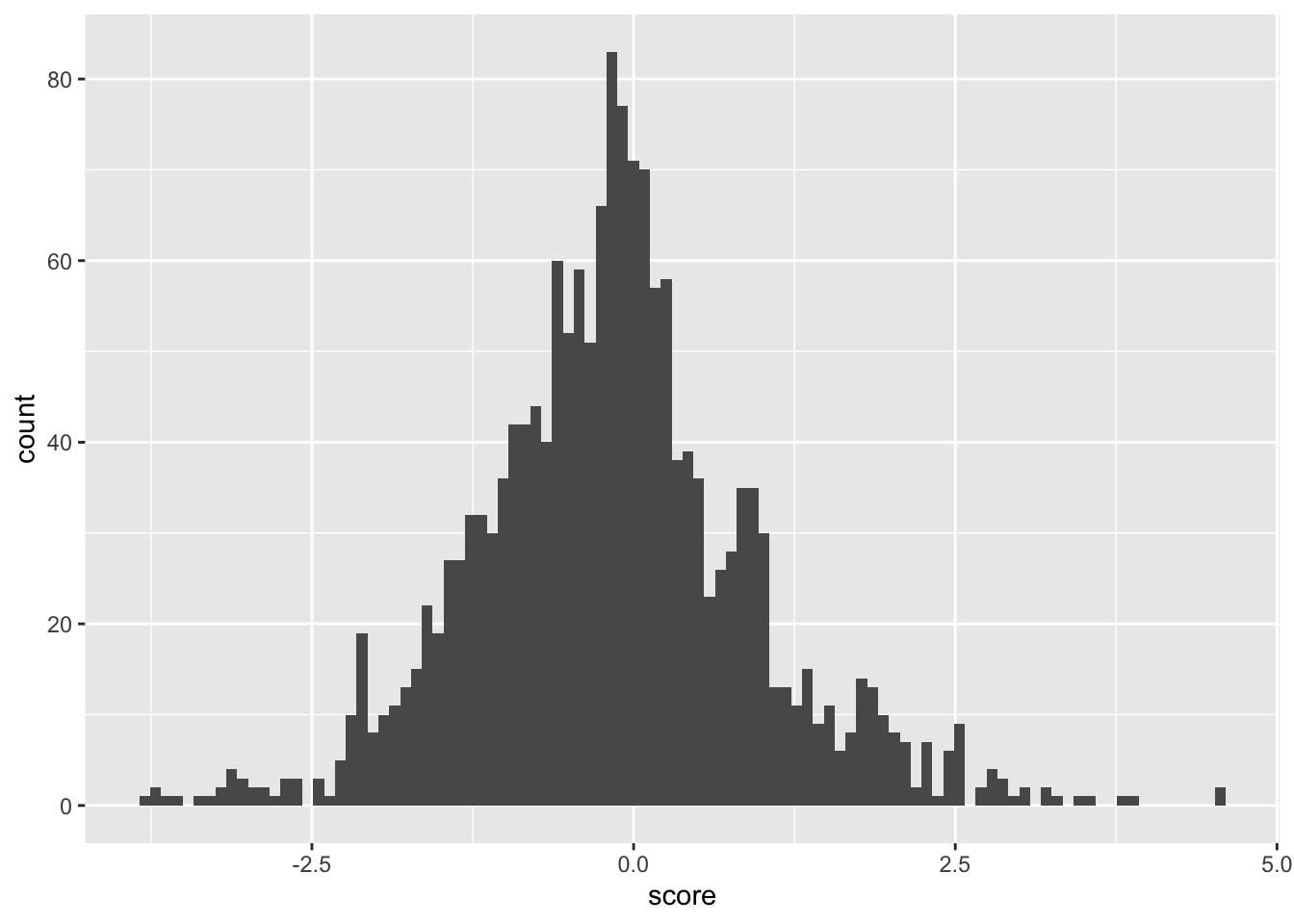
ggplot(imat_scores,aes(x=score.x,y=score.y))+geom_point()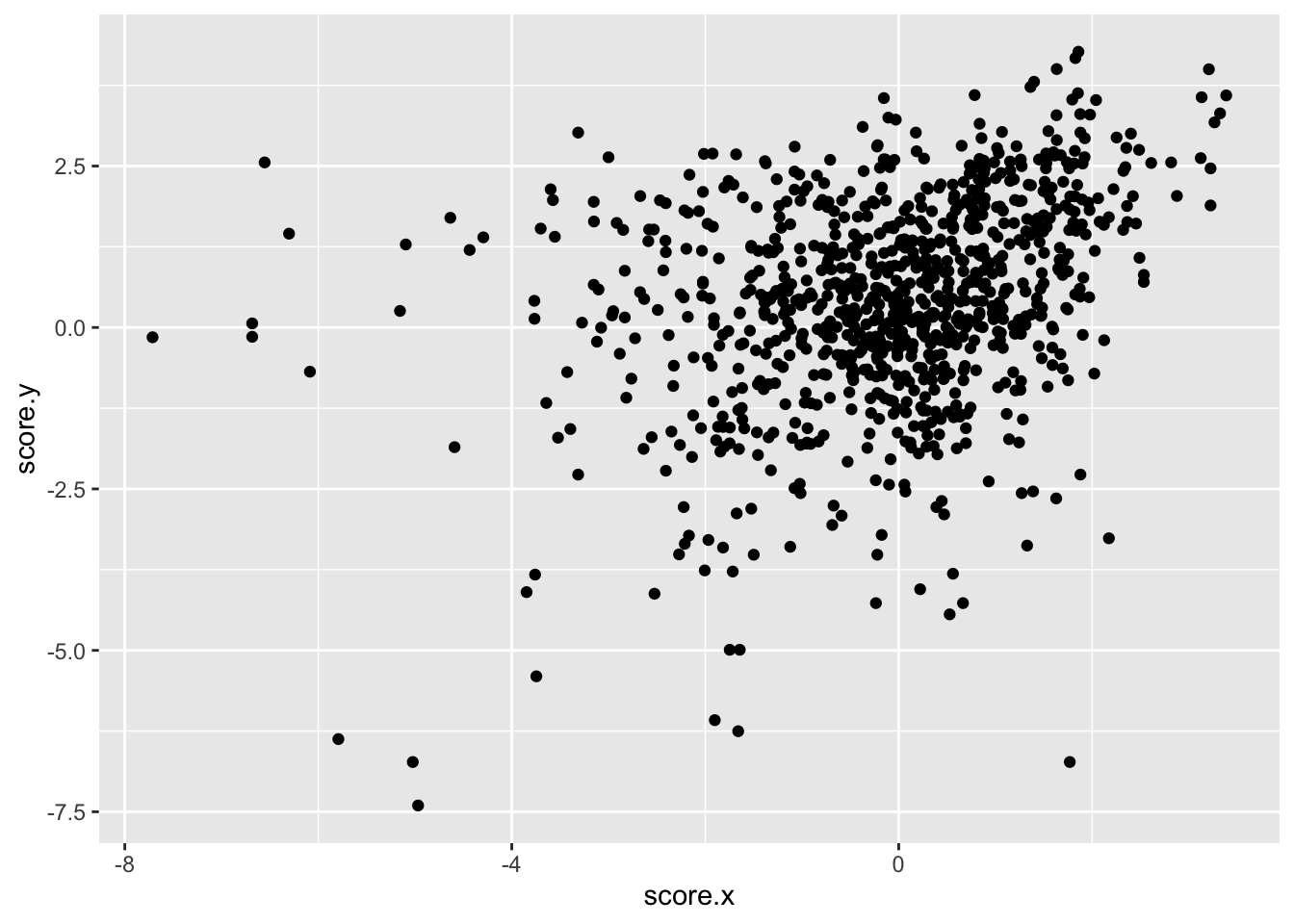
ggplot(imat_scores%>%mutate(name=paste(ref_aa,protein_start,alt_aa)),aes(x=score.x,y=score.y,label=name))+geom_text()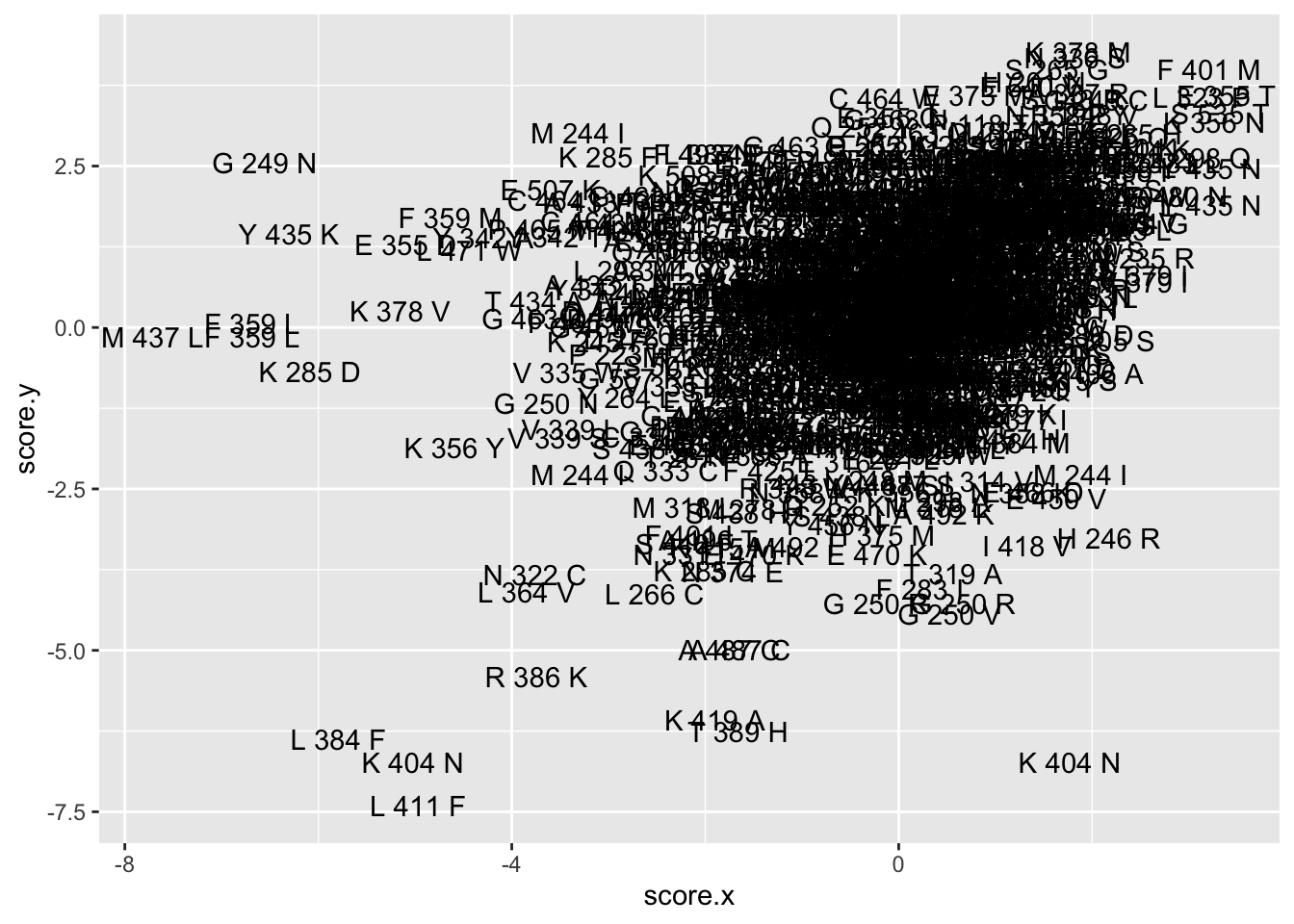
# ggplotly(plotly)
ggplot(imat_scores,aes(x=score.x,y=score.y))+geom_bin2d(bins=100)+scale_fill_continuous(type="viridis")+theme_bw()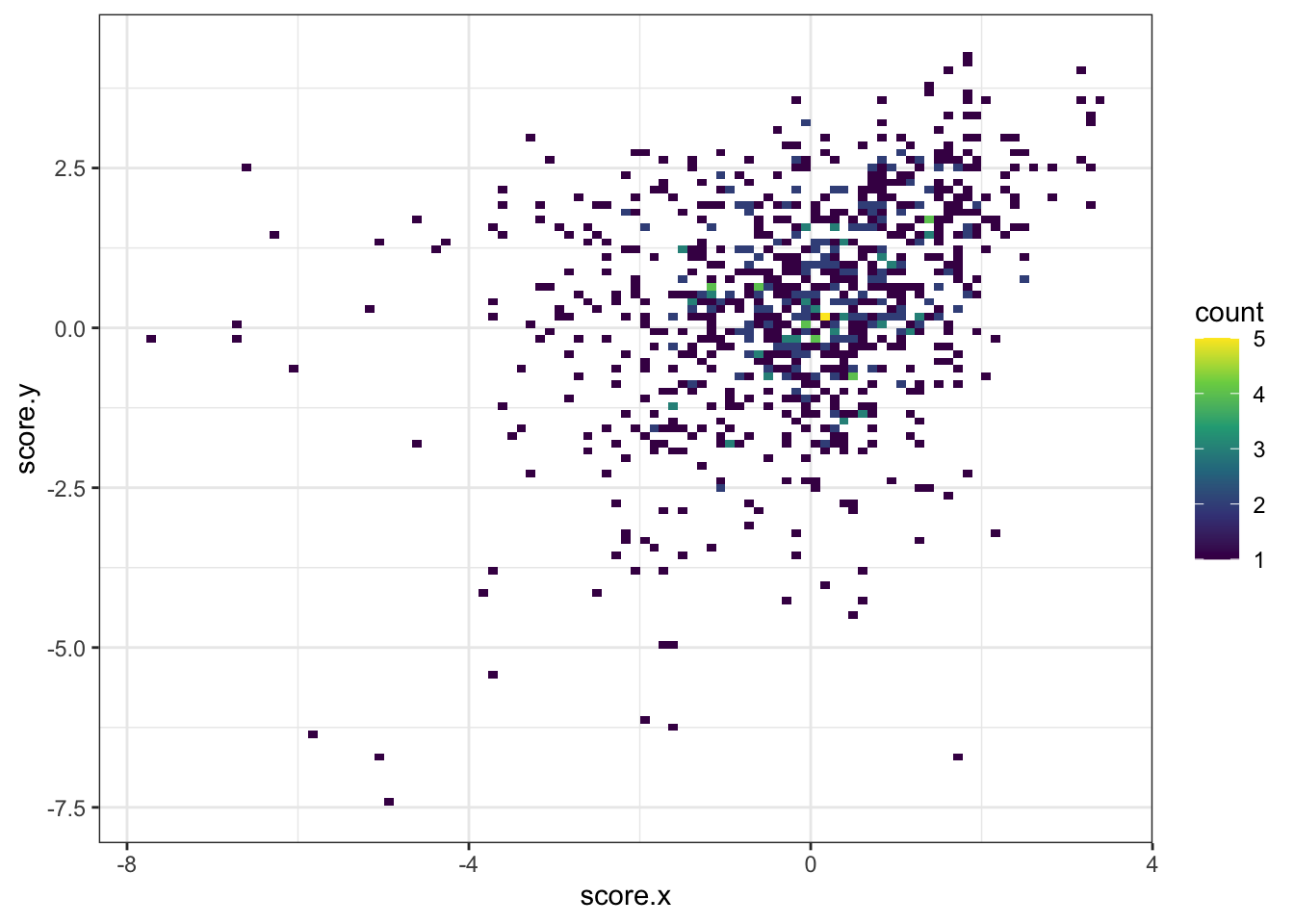
ggplot(imat_scores,aes(x=score.x,y=score.y))+stat_density_2d(aes(fill=..level..),geom = "polygon", colour="white")+scale_fill_continuous(type="viridis")+theme_bw()+xlab("Enrichment Score Imat Treatment Replicate 1")+ylab("Enrichment Score Imat Treatment Replicate 2")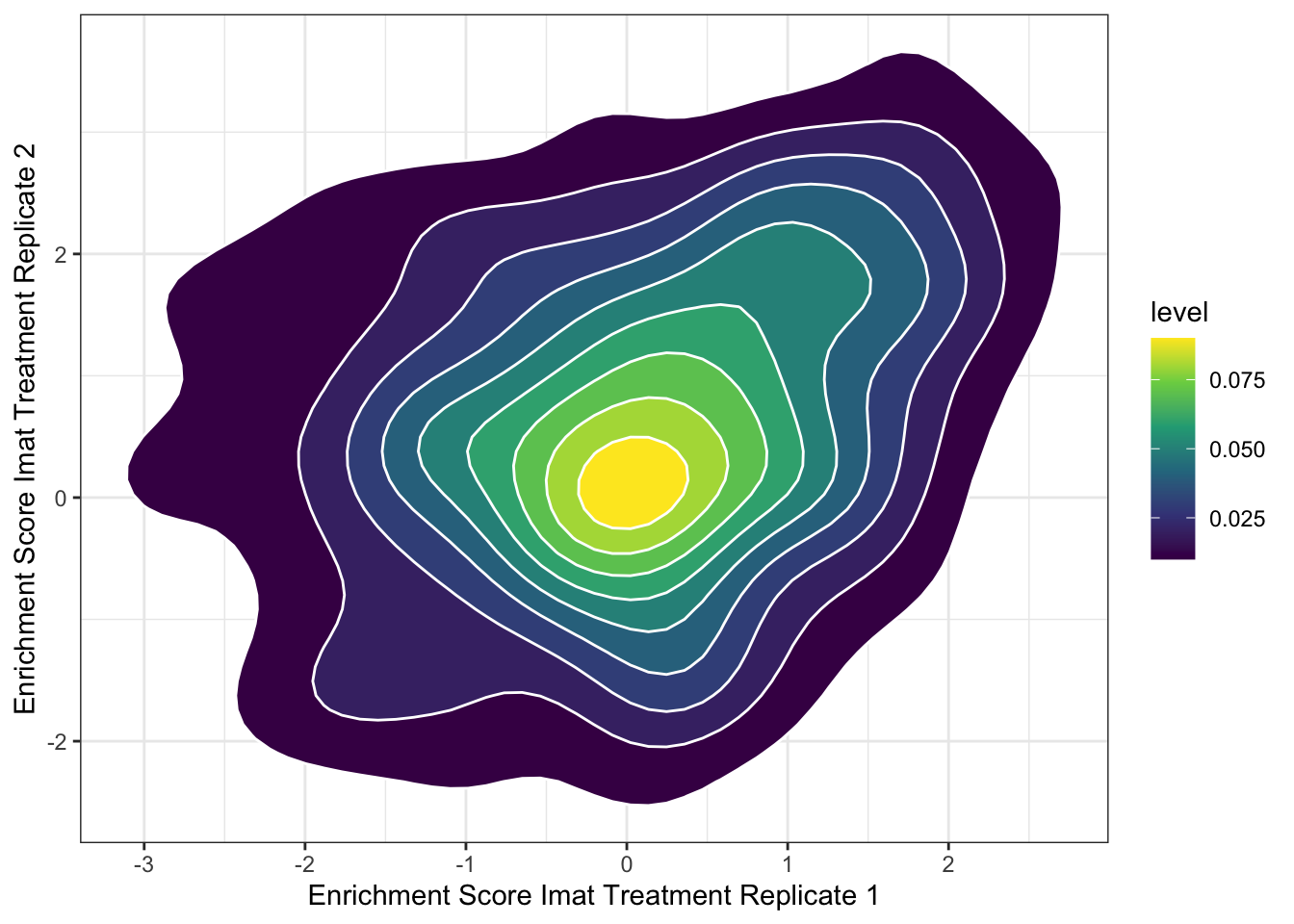
plotly=ggplot(imat_scores%>%filter(!alt_aa%in%c("PA","NY","LL"),protein_start>=242,protein_start<=492),aes(x=protein_start,y=alt_aa,fill=score.x))+geom_tile()+theme(panel.background=element_rect(fill="white", colour="white"))+scale_fill_gradient2(low ="blue",midpoint=0,mid="white", high ="red",name="Score")
ggplotly(plotly)Warning in matrix(g$fill_plotlyDomain, nrow = length(y), ncol = length(x), :
data length [4653] is not a sub-multiple or multiple of the number of rows [20]Warning in matrix(g$hovertext, nrow = length(y), ncol = length(x), byrow =
TRUE): data length [4653] is not a sub-multiple or multiple of the number of
rows [20]# cor(imat_scores$score.x,imat_scores$score.y,method="pearson")
ggplot(asc_scores,aes(x=score.x,y=score.y))+geom_point()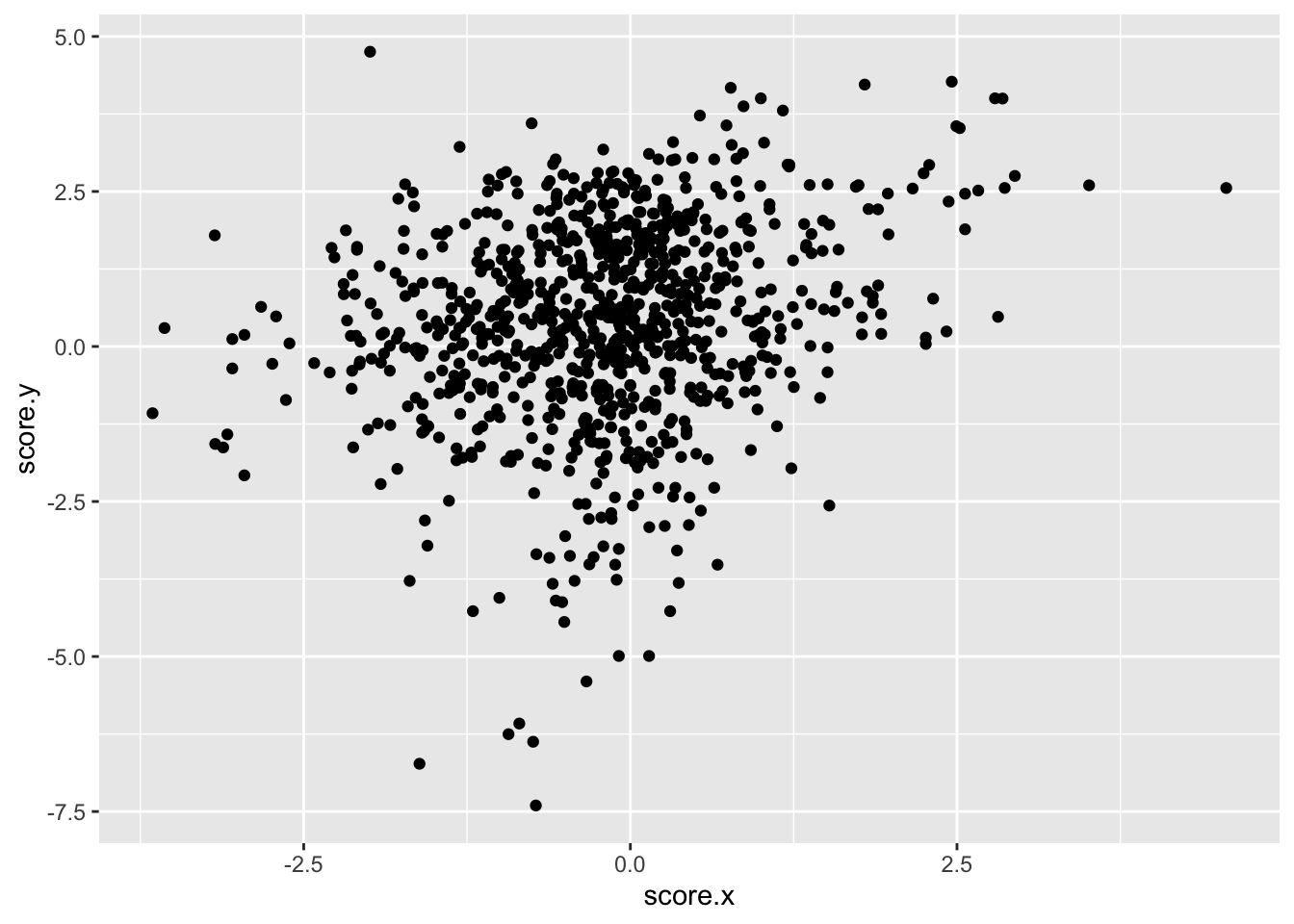
ggplot(asc_scores%>%mutate(name=paste(ref_aa,protein_start,alt_aa)),aes(x=score.x,y=score.y,label=name))+geom_text()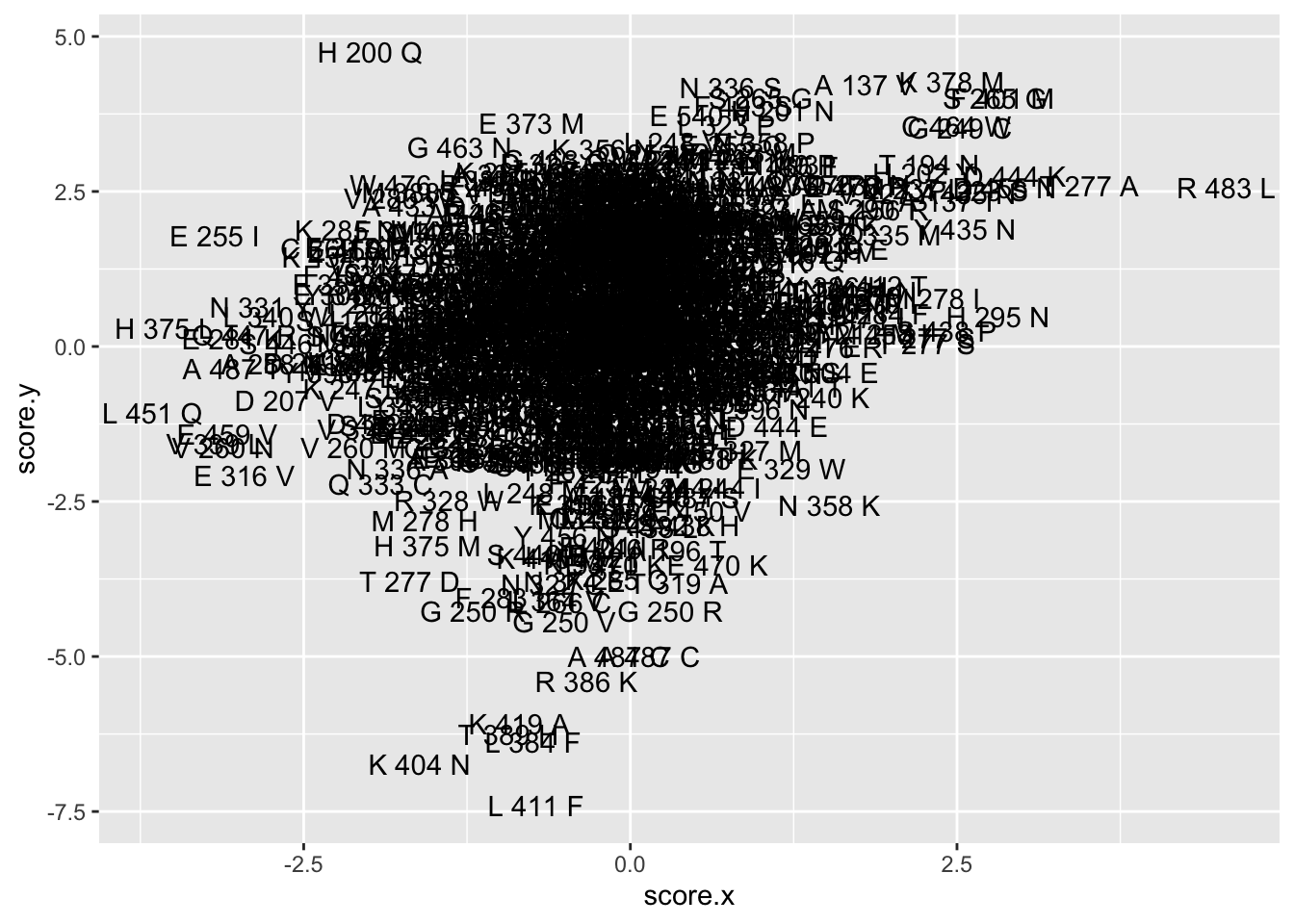
# ggplotly(plotly)
ggplot(asc_scores,aes(x=score.x,y=score.y))+geom_bin2d(bins=100)+scale_fill_continuous(type="viridis")+theme_bw()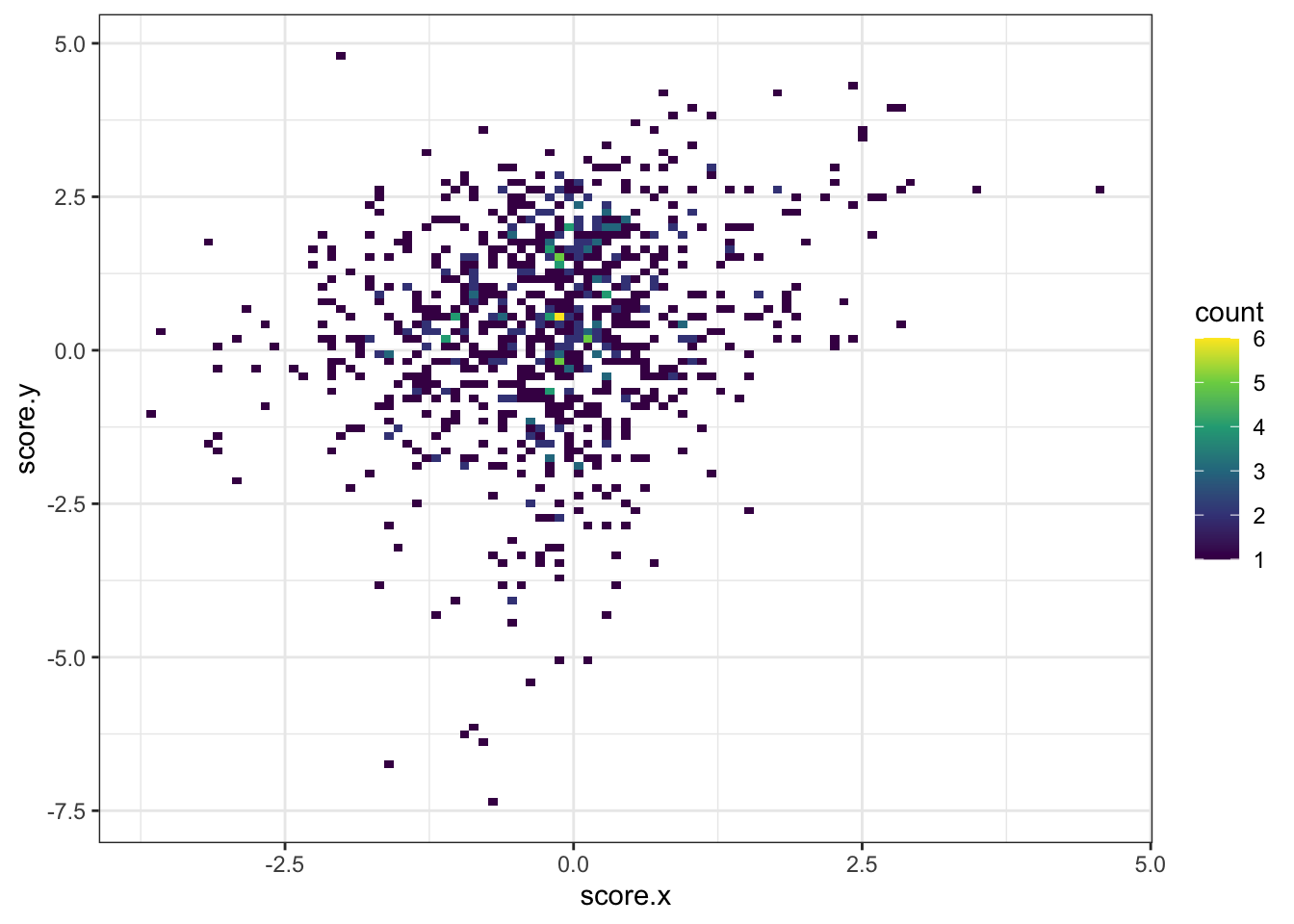
ggplot(asc_scores,aes(x=score.x,y=score.y))+stat_density_2d(aes(fill=..level..),geom = "polygon", colour="white")+scale_fill_continuous(type="viridis")+theme_bw()+xlab("Enrichment Score Asciminib Treatment Replicate 1")+ylab("Enrichment Score Asciminib Treatment Replicate 2")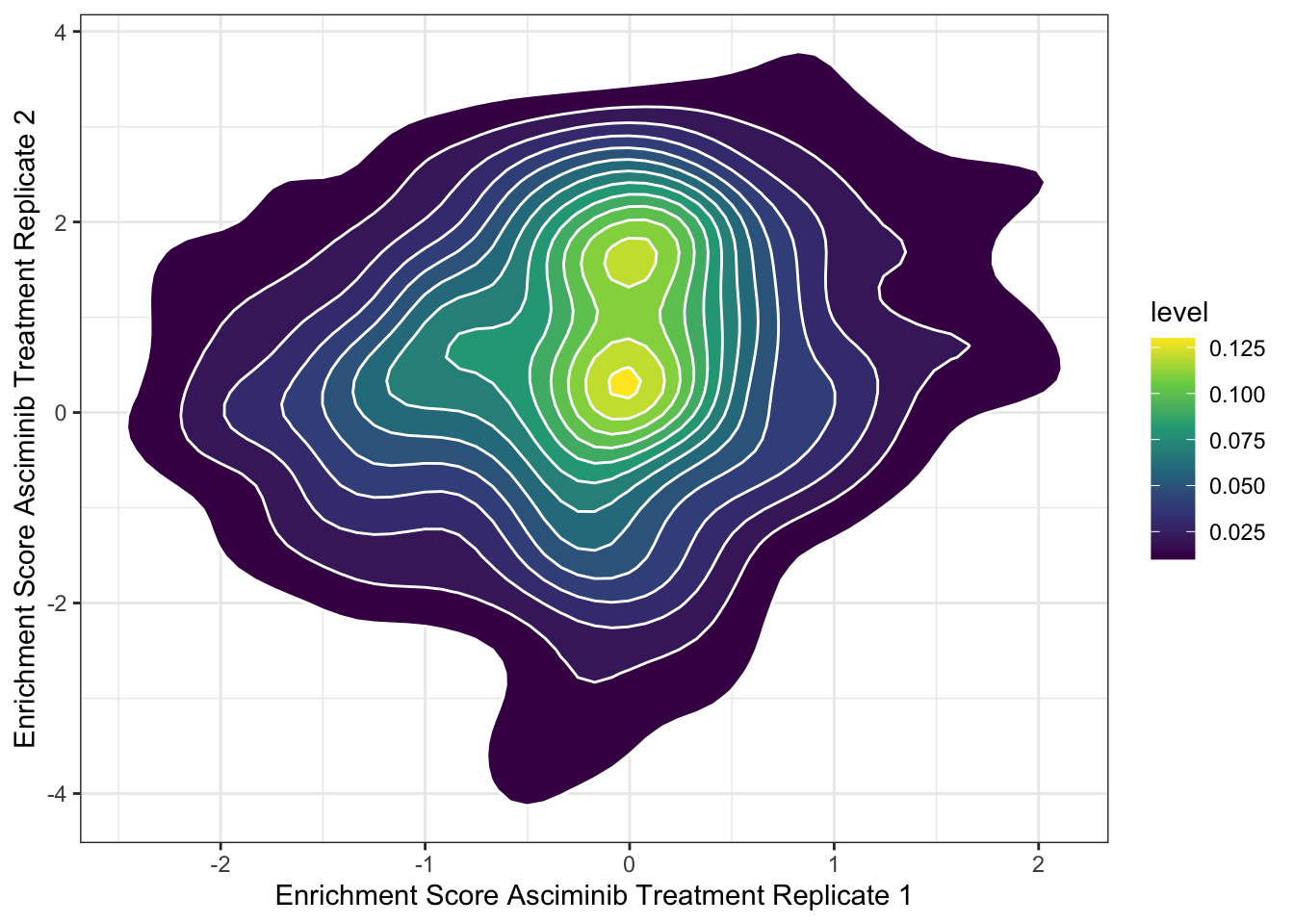
plotly=ggplot(asc_scores%>%filter(!alt_aa%in%c("PA","NY","LL"),protein_start>=242,protein_start<=492),aes(x=protein_start,y=alt_aa,fill=score.x))+geom_tile()+theme(panel.background=element_rect(fill="white", colour="white"))+scale_fill_gradient2(low ="blue",midpoint=0,mid="white", high ="red",name="Score")
ggplotly(plotly)Warning in matrix(g$fill_plotlyDomain, nrow = length(y), ncol = length(x), :
data length [4642] is not a sub-multiple or multiple of the number of rows [20]Warning in matrix(g$hovertext, nrow = length(y), ncol = length(x), byrow =
TRUE): data length [4642] is not a sub-multiple or multiple of the number of
rows [20]# a=samples_14.17%>%filter(ct.x>1)Summing counts for the two samples for Asciminib and the two samples for imatinib to see if we can improve number of mutants seen in the dataset
How uneven is the ABL library?
sample14=sample14%>%mutate(region=case_when(protein_start<250&protein_start>=242~1,
protein_start<258&protein_start>=250~2,
protein_start<266&protein_start>=258~3,
protein_start<274&protein_start>=266~4,
protein_start<282&protein_start>=274~5,
protein_start<290&protein_start>=282~6,
protein_start<298&protein_start>=290~7,
protein_start<306&protein_start>=298~8,
protein_start<314&protein_start>=306~9,
protein_start<322&protein_start>=314~10,
protein_start<330&protein_start>=322~11,
protein_start<338&protein_start>=330~12,
protein_start<346&protein_start>=338~13,
protein_start<354&protein_start>=346~14,
protein_start<362&protein_start>=354~15,
protein_start<370&protein_start>=362~16,
protein_start<378&protein_start>=370~17,
protein_start<386&protein_start>=378~18,
protein_start<394&protein_start>=386~19,
protein_start<402&protein_start>=394~20,
protein_start<410&protein_start>=402~21,
protein_start<418&protein_start>=410~22,
protein_start<426&protein_start>=418~23,
protein_start<434&protein_start>=426~24,
protein_start<442&protein_start>=434~25,
protein_start<450&protein_start>=442~26,
protein_start<458&protein_start>=450~27,
protein_start<466&protein_start>=458~28,
protein_start<474&protein_start>=466~29,
protein_start<482&protein_start>=474~30,
protein_start<490&protein_start>=482~31,
protein_start<498&protein_start>=490~32,
T~0))
calls_sum=sample14%>%filter(consequence_terms%in%"missense_variant")%>%group_by(protein_start,region)%>%summarize(unique_mutants=n(),count=sum(ct))`summarise()` has grouped output by 'protein_start'. You can override using the `.groups` argument.ggplot(calls_sum%>%filter(protein_start>=242,protein_start<=500),aes(x=protein_start,y=count))+geom_col()+cleanup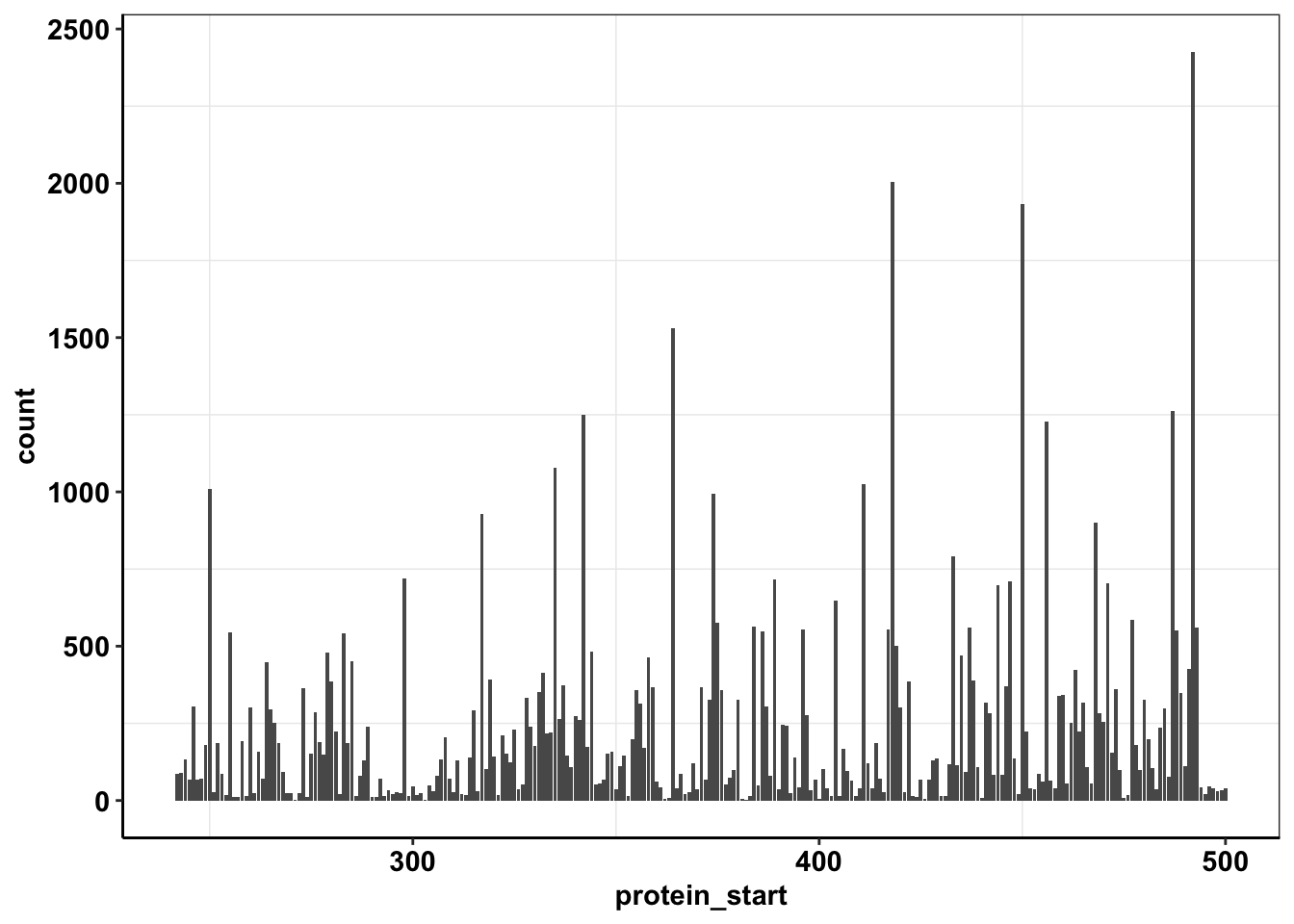
getPalette = colorRampPalette(brewer.pal(33, "Set2"))Warning in brewer.pal(33, "Set2"): n too large, allowed maximum for palette Set2 is 8
Returning the palette you asked for with that many colorsplotly=ggplot(calls_sum%>%filter(protein_start>=242,protein_start<=500),aes(x=protein_start,y=count))+geom_col(color="black",aes(fill=factor(region)))+cleanup+scale_fill_manual(values=getPalette(33))
ggplotly(plotly)sample11=read.csv(file = "data/Consensus_Data/Novogene_lane14/sample11/variant_caller_outputs/variants_unique_ann.csv",header=T,stringsAsFactors = F)
sample11=sample11%>%
rowwise()%>%
mutate(ref_aa=strsplit(amino_acids,"/")[[1]][1],
alt_aa=strsplit(amino_acids,"/")[[1]][2])
sample11=sample11%>%mutate(maf=ct/depth)
sample11=sample11%>%mutate(region=case_when(protein_start<250&protein_start>=242~1,
protein_start<258&protein_start>=250~2,
protein_start<266&protein_start>=258~3,
protein_start<274&protein_start>=266~4,
protein_start<282&protein_start>=274~5,
protein_start<290&protein_start>=282~6,
protein_start<298&protein_start>=290~7,
protein_start<306&protein_start>=298~8,
protein_start<314&protein_start>=306~9,
protein_start<322&protein_start>=314~10,
protein_start<330&protein_start>=322~11,
protein_start<338&protein_start>=330~12,
protein_start<346&protein_start>=338~13,
protein_start<354&protein_start>=346~14,
protein_start<362&protein_start>=354~15,
protein_start<370&protein_start>=362~16,
protein_start<378&protein_start>=370~17,
protein_start<386&protein_start>=378~18,
protein_start<394&protein_start>=386~19,
protein_start<402&protein_start>=394~20,
protein_start<410&protein_start>=402~21,
protein_start<418&protein_start>=410~22,
protein_start<426&protein_start>=418~23,
protein_start<434&protein_start>=426~24,
protein_start<442&protein_start>=434~25,
protein_start<450&protein_start>=442~26,
protein_start<458&protein_start>=450~27,
protein_start<466&protein_start>=458~28,
protein_start<474&protein_start>=466~29,
protein_start<482&protein_start>=474~30,
protein_start<490&protein_start>=482~31,
protein_start<498&protein_start>=490~32,
T~0))
calls_sum=sample11%>%filter(consequence_terms%in%"missense_variant")%>%group_by(protein_start,region)%>%summarize(unique_mutants=n(),count=sum(ct))`summarise()` has grouped output by 'protein_start'. You can override using the `.groups` argument.ggplot(calls_sum%>%filter(protein_start>=242,protein_start<=500),aes(x=protein_start,y=count))+geom_col()+cleanup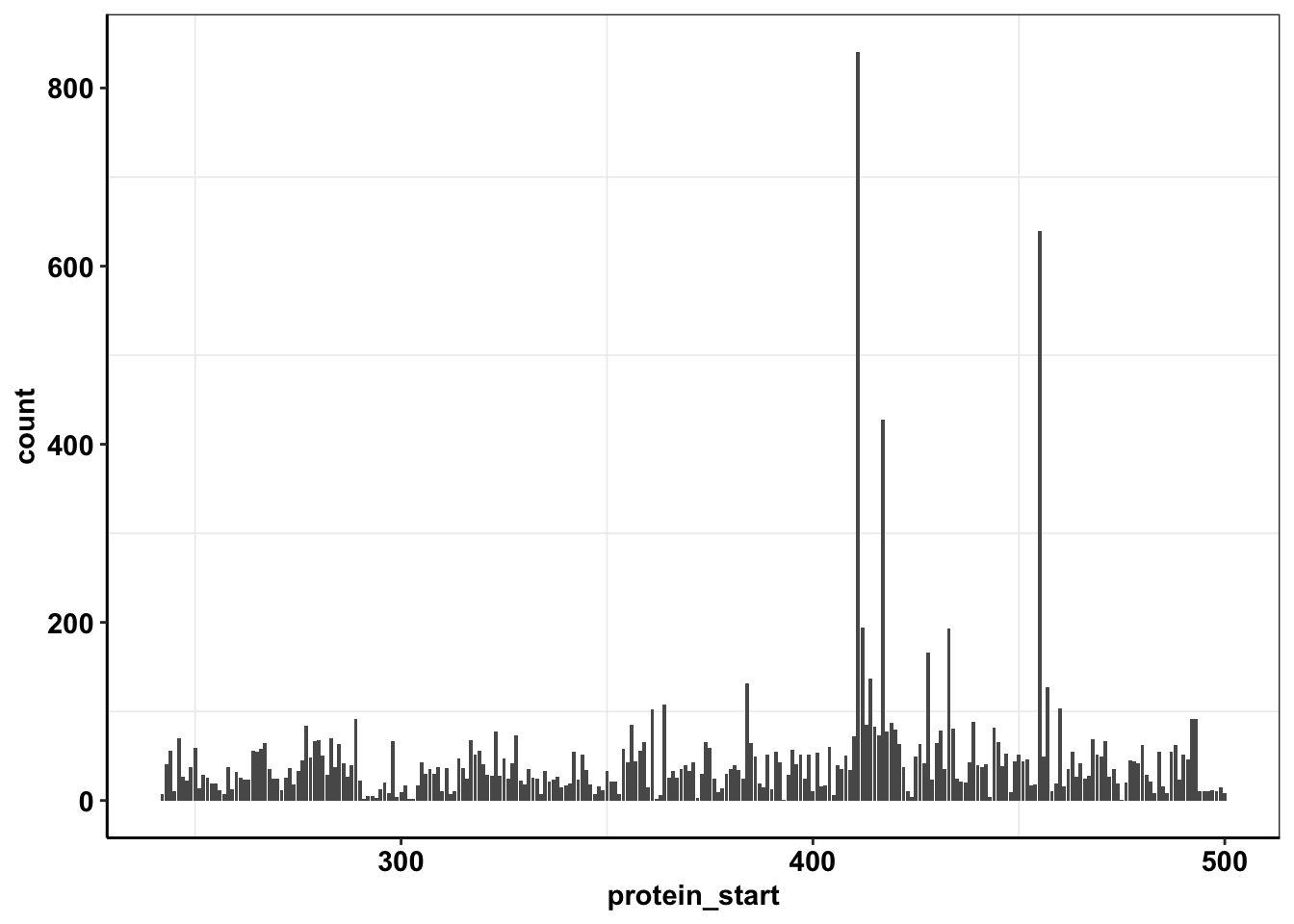
getPalette = colorRampPalette(brewer.pal(33, "Set2"))Warning in brewer.pal(33, "Set2"): n too large, allowed maximum for palette Set2 is 8
Returning the palette you asked for with that many colorsplotly=ggplot(calls_sum%>%filter(protein_start>=242,protein_start<=500),aes(x=protein_start,y=count))+geom_col(color="black",aes(fill=factor(region)))+cleanup+scale_fill_manual(values=getPalette(33))
ggplotly(plotly)ggplot(calls_sum%>%filter(protein_start>=242,protein_start<=500),aes(x=protein_start,y=count))+geom_col()+cleanup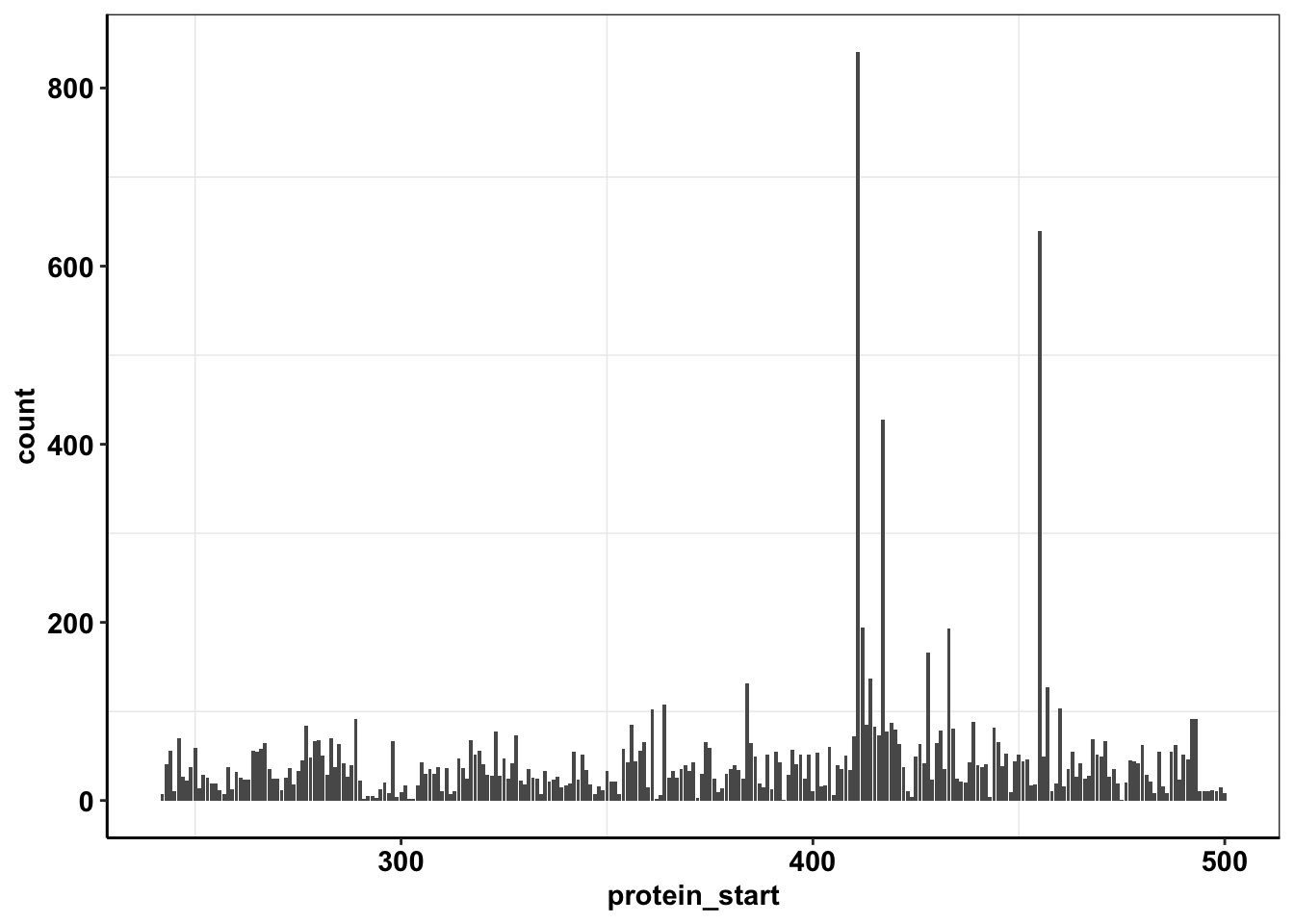
plotly=ggplot(calls_sum%>%filter(protein_start>=242,protein_start<=500,!protein_start%in%c(411,417,455)),aes(x=protein_start,y=count))+geom_col(color="black",aes(fill=factor(region)))+cleanup+scale_fill_manual(values=getPalette(33))
ggplotly(plotly)TSII Sample 11 and 12
sample11=read.csv(file = "data/Consensus_Data/Novogene_lane14/sample11/variant_caller_outputs/variants_unique_ann.csv",header=T,stringsAsFactors = F)
sample11=sample11%>%
rowwise()%>%
mutate(ref_aa=strsplit(amino_acids,"/")[[1]][1],
alt_aa=strsplit(amino_acids,"/")[[1]][2])
sample11=sample11%>%mutate(maf=ct/depth)
sample11_simple=sample11%>%dplyr::select(alt_start_pos,protein_start,ref,alt,ref_aa,alt_aa,consequence_terms,ct,depth,maf)
sample12=read.csv(file = "data/Consensus_Data/Novogene_lane14/sample12/variant_caller_outputs/variants_unique_ann.csv",header=T,stringsAsFactors = F)
sample12=sample12%>%
rowwise()%>%
mutate(ref_aa=strsplit(amino_acids,"/")[[1]][1],
alt_aa=strsplit(amino_acids,"/")[[1]][2])
sample12=sample12%>%mutate(maf=ct/depth)
sample12_simple=sample12%>%dplyr::select(alt_start_pos,protein_start,ref,alt,ref_aa,alt_aa,consequence_terms,ct,depth,maf)
samples_11.12=merge(sample11_simple%>%filter(consequence_terms%in%"missense_variant"),sample12_simple%>%filter(consequence_terms%in%"missense_variant"),by=c("ref_aa","protein_start","alt_aa","ref","alt","alt_start_pos","consequence_terms"))
samples_11.12=samples_11.12%>%mutate(score=log2(maf.y/maf.x),score2=log2(ct.y/ct.x))
samples_11.12_simple=samples_11.12%>%dplyr::select(ref_aa,protein_start,alt_aa,consequence_terms,ct.x,score)
ggplot(samples_11.12,aes(x=score))+geom_density()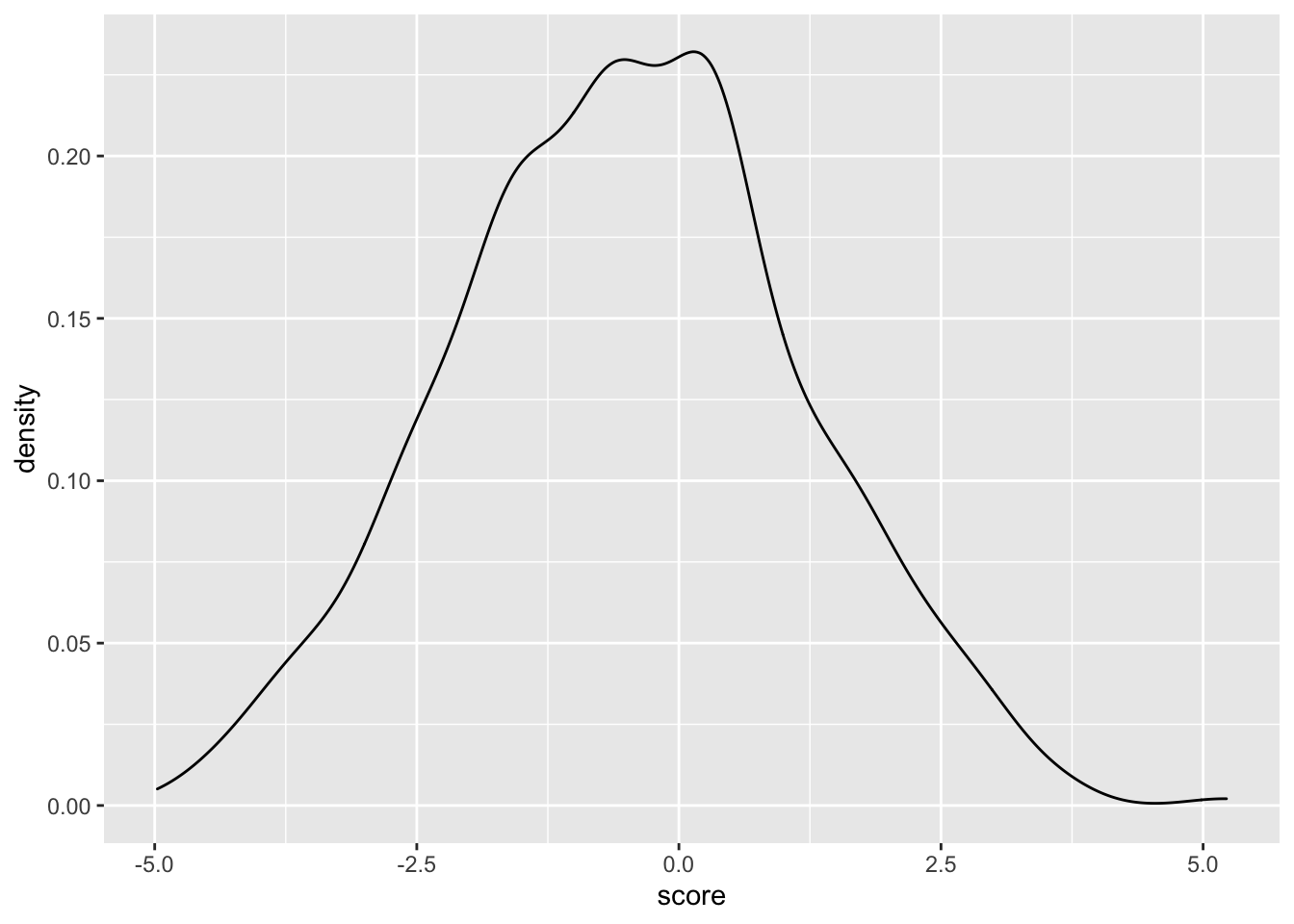
ggplot(samples_14.16,aes(x=score))+geom_histogram(bins=100)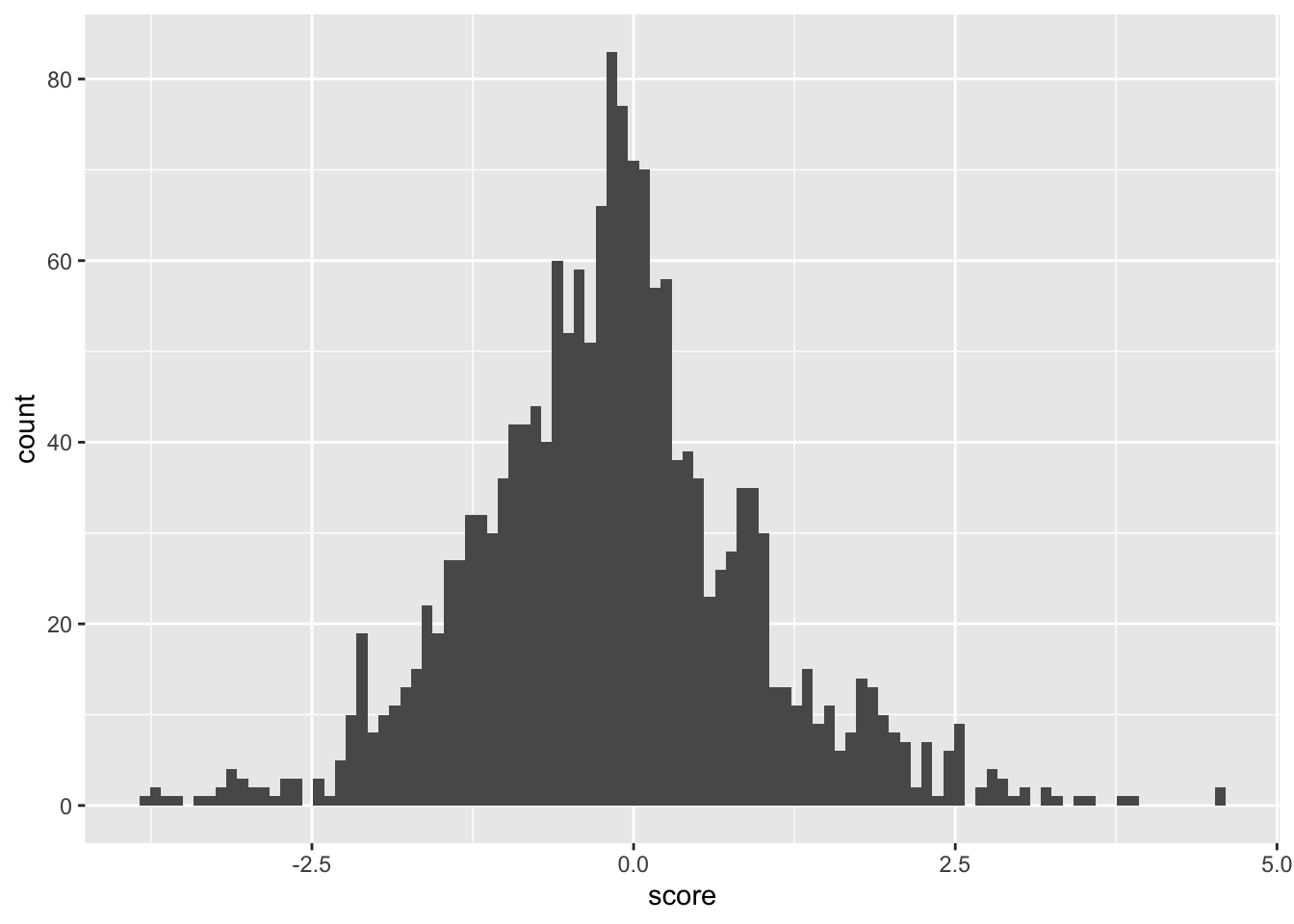
samples_11.12=samples_11.12%>%mutate(resmuts=case_when(protein_start%in%253&alt_aa%in%"H"~T,
protein_start%in%255&alt_aa%in%"V"~T,
protein_start%in%486&alt_aa%in%"S"~T,
protein_start%in%396&alt_aa%in%"P"~T,
protein_start%in%255&alt_aa%in%"K"~T,
protein_start%in%315&alt_aa%in%"I"~T,
protein_start%in%252&alt_aa%in%"H"~T,
protein_start%in%253&alt_aa%in%"F"~T,
protein_start%in%250&alt_aa%in%"E"~T,
protein_start%in%359&alt_aa%in%"C"~T,
protein_start%in%351&alt_aa%in%"T"~T,
protein_start%in%355&alt_aa%in%"G"~T,
protein_start%in%317&alt_aa%in%"L"~T,
protein_start%in%359&alt_aa%in%"I"~T,
protein_start%in%355&alt_aa%in%"A"~T,
protein_start%in%459&alt_aa%in%"K"~T,
protein_start%in%276&alt_aa%in%"G"~T,
protein_start%in%299&alt_aa%in%"L"~T,
T~F))
samples_11.12=samples_11.12%>%mutate(resresids=case_when(protein_start%in%253~T,
protein_start%in%255~T,
protein_start%in%486~T,
protein_start%in%396~T,
protein_start%in%255~T,
protein_start%in%315~T,
protein_start%in%252~T,
protein_start%in%253~T,
protein_start%in%250~T,
protein_start%in%359~T,
protein_start%in%351~T,
protein_start%in%355~T,
protein_start%in%317~T,
protein_start%in%359~T,
protein_start%in%355~T,
protein_start%in%459~T,
protein_start%in%276~T,
protein_start%in%299~T,
T~F))
highscore=samples_11.12%>%filter(!ct.x%in%1,protein_start>=242,protein_start<=494)
highscore=highscore%>%mutate(subs_name=paste(ref_aa,protein_start,alt_aa,sep = ""))
plotly=ggplot(highscore,aes(x=reorder(subs_name,-score),y=score,fill=resmuts))+geom_col()+theme(axis.text.x=element_text(angle=90, hjust=1))+scale_y_continuous(name="Enrithcment Score")+scale_x_discrete(name="Mutant")+guides(fill = guide_legend(title = "Clinically\n Observed \n Resmut"))
ggplotly(plotly)plotly=ggplot(highscore%>%filter(ct.y>=2),aes(x=reorder(subs_name,-score),y=score,fill=resresids))+geom_col()+theme(axis.text.x=element_text(angle=90, hjust=1))+scale_y_continuous(name="Enrithcment Score")+scale_x_discrete(name="Mutant")+guides(fill = guide_legend(title = "Clinically\n Observed \n Resmut"))
ggplotly(plotly)ggplot(samples_11.12%>%filter(!alt_aa%in%c("LL","PA"),protein_start>=242,protein_start<=494),aes(x=protein_start,y=alt_aa,fill=score))+geom_tile()+theme(panel.background=element_rect(fill="white", colour="white"))+scale_fill_gradient2(low ="blue",midpoint=0,mid="white", high ="red",name="Score")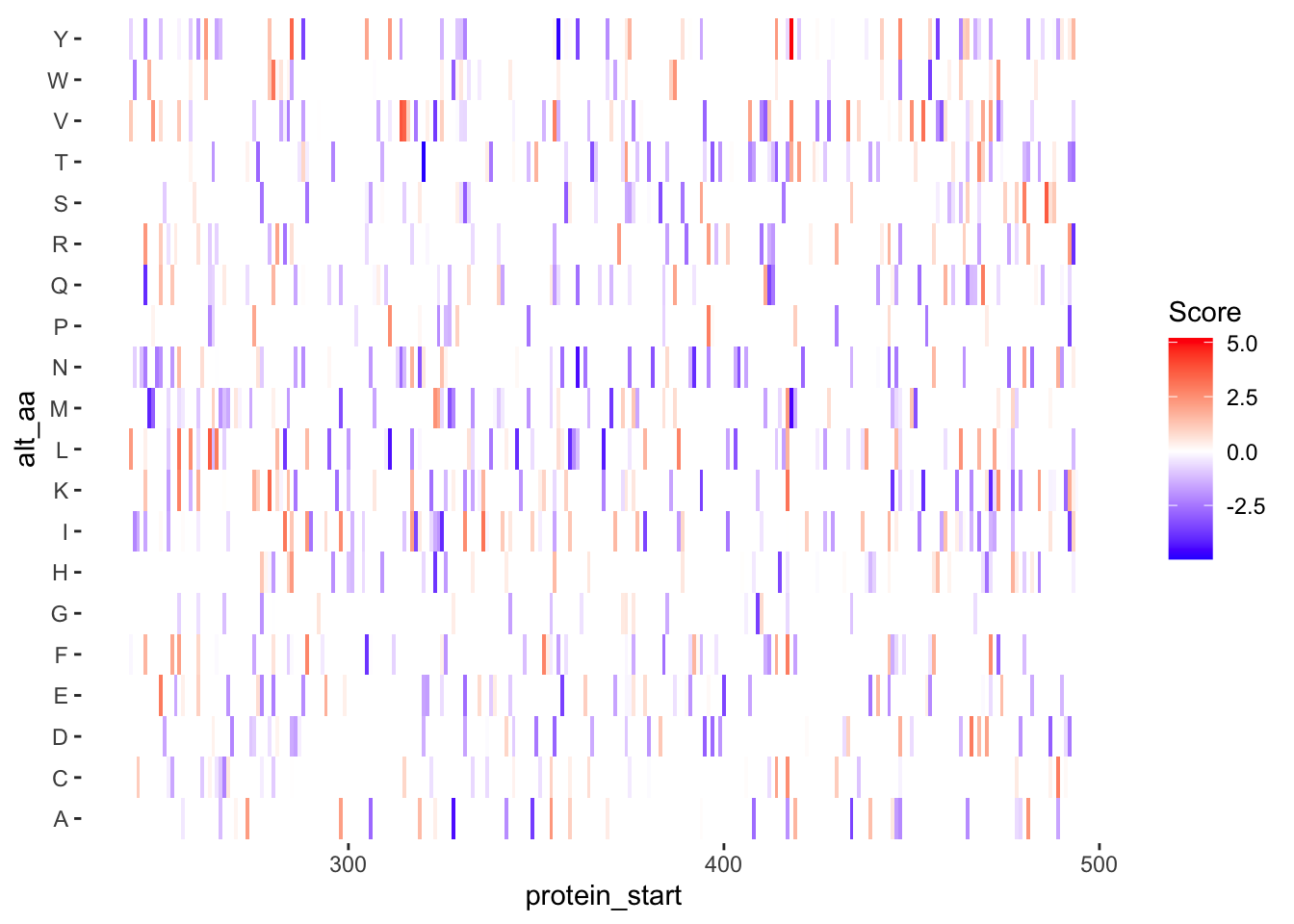
# a=highscore%>%filter(protein_start%in%"255")TSII Sample 10 and 12
source("code/merge_samples.R")
sample10=read.csv(file = "data/Consensus_Data/Novogene_lane14/sample10_combined/variant_caller_outputs/variants_unique_ann.csv",header=T,stringsAsFactors = F)
sample10=sample10%>%
rowwise()%>%
mutate(ref_aa=strsplit(amino_acids,"/")[[1]][1],
alt_aa=strsplit(amino_acids,"/")[[1]][2])
sample10=sample10%>%mutate(maf=ct/depth)
sample10_simple=sample10%>%dplyr::select(alt_start_pos,protein_start,ref,alt,ref_aa,alt_aa,consequence_terms,ct,depth,maf)
sample12=read.csv(file = "data/Consensus_Data/Novogene_lane14/sample12/variant_caller_outputs/variants_unique_ann.csv",header=T,stringsAsFactors = F)
sample12=sample12%>%
rowwise()%>%
mutate(ref_aa=strsplit(amino_acids,"/")[[1]][1],
alt_aa=strsplit(amino_acids,"/")[[1]][2])
sample12=sample12%>%mutate(maf=ct/depth)
sample12_simple=sample12%>%dplyr::select(alt_start_pos,protein_start,ref,alt,ref_aa,alt_aa,consequence_terms,ct,depth,maf)
samples_10.12=merge(sample10_simple%>%filter(consequence_terms%in%"missense_variant"),sample12_simple%>%filter(consequence_terms%in%"missense_variant"),by=c("ref_aa","protein_start","alt_aa","ref","alt","alt_start_pos","consequence_terms"))
samples_10.12=samples_10.12%>%mutate(score=log2(maf.y/maf.x),score2=log2(ct.y/ct.x))
samples_10.12_simple=samples_10.12%>%dplyr::select(ref_aa,protein_start,alt_aa,consequence_terms,ct.x,score)
ggplot(samples_10.12,aes(x=score))+geom_density()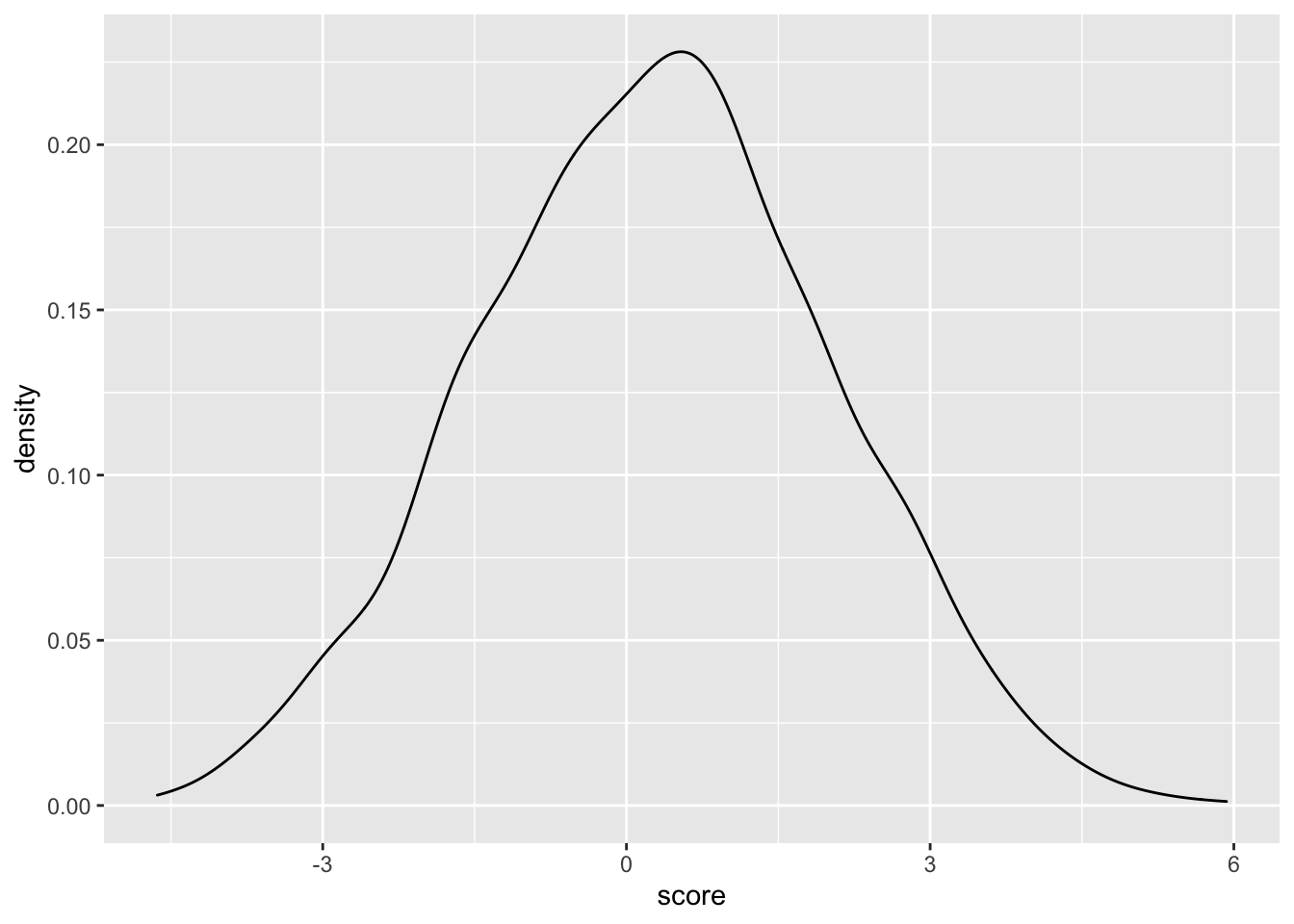
# ggplot(samples_14.16,aes(x=score))+geom_histogram(bins=100)
samples_10.12=samples_10.12%>%mutate(resmuts=case_when(protein_start%in%253&alt_aa%in%"H"~T,
protein_start%in%255&alt_aa%in%"V"~T,
protein_start%in%486&alt_aa%in%"S"~T,
protein_start%in%396&alt_aa%in%"P"~T,
protein_start%in%255&alt_aa%in%"K"~T,
protein_start%in%315&alt_aa%in%"I"~T,
protein_start%in%252&alt_aa%in%"H"~T,
protein_start%in%253&alt_aa%in%"F"~T,
protein_start%in%250&alt_aa%in%"E"~T,
protein_start%in%359&alt_aa%in%"C"~T,
protein_start%in%351&alt_aa%in%"T"~T,
protein_start%in%355&alt_aa%in%"G"~T,
protein_start%in%317&alt_aa%in%"L"~T,
protein_start%in%359&alt_aa%in%"I"~T,
protein_start%in%355&alt_aa%in%"A"~T,
protein_start%in%459&alt_aa%in%"K"~T,
protein_start%in%276&alt_aa%in%"G"~T,
protein_start%in%299&alt_aa%in%"L"~T,
T~F))
samples_10.12=samples_10.12%>%mutate(resresids=case_when(protein_start%in%253~T,
protein_start%in%255~T,
protein_start%in%486~T,
protein_start%in%396~T,
protein_start%in%255~T,
protein_start%in%315~T,
protein_start%in%252~T,
protein_start%in%253~T,
protein_start%in%250~T,
protein_start%in%359~T,
protein_start%in%351~T,
protein_start%in%355~T,
protein_start%in%317~T,
protein_start%in%359~T,
protein_start%in%355~T,
protein_start%in%459~T,
protein_start%in%276~T,
protein_start%in%299~T,
T~F))
highscore=samples_10.12%>%filter(!ct.x%in%1,protein_start>=242,protein_start<=494)
highscore=highscore%>%mutate(subs_name=paste(ref_aa,protein_start,alt_aa,sep = ""))
plotly=ggplot(highscore,aes(x=reorder(subs_name,-score),y=score,fill=resmuts))+geom_col()+theme(axis.text.x=element_text(angle=90, hjust=1))+scale_y_continuous(name="Enrithcment Score")+scale_x_discrete(name="Mutant")+guides(fill = guide_legend(title = "Clinically\n Observed \n Resmut"))
ggplotly(plotly)plotly=ggplot(highscore%>%filter(ct.y>=2),aes(x=reorder(subs_name,-score),y=score,fill=resresids))+geom_col()+theme(axis.text.x=element_text(angle=90, hjust=1))+scale_y_continuous(name="Enrithcment Score")+scale_x_discrete(name="Mutant")+guides(fill = guide_legend(title = "Clinically\n Observed \n Resmut"))
ggplotly(plotly)ggplot(samples_10.12%>%filter(!alt_aa%in%c("LL","PA"),protein_start>=242,protein_start<=494),aes(x=protein_start,y=alt_aa,fill=score))+geom_tile()+theme(panel.background=element_rect(fill="white", colour="white"))+scale_fill_gradient2(low ="blue",midpoint=0,mid="white", high ="red",name="Score")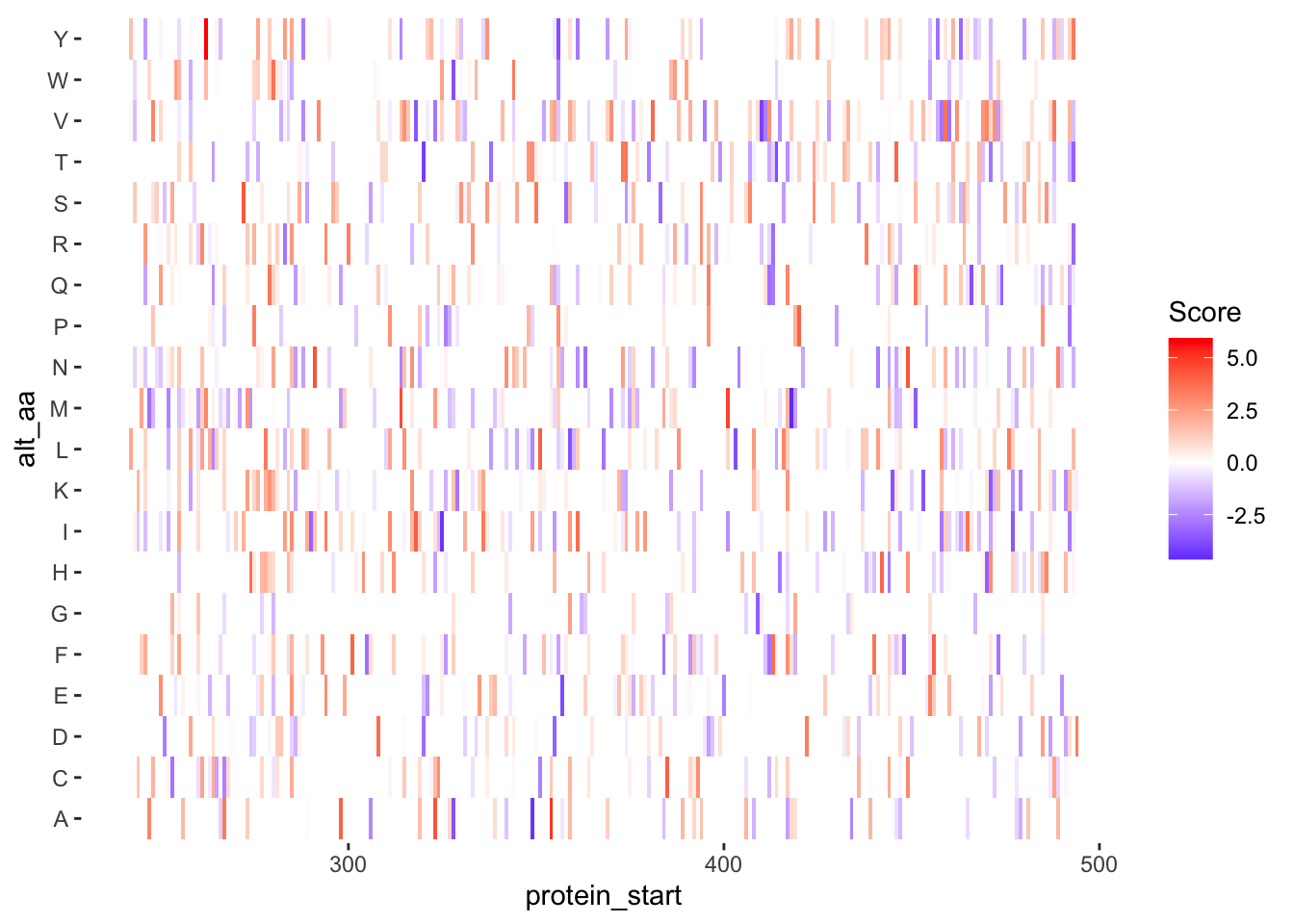
ggplot(samples_10.12%>%filter(!alt_aa%in%c("LL","PA"),protein_start>=242,protein_start<=494),aes(x=protein_start,y=alt_aa,fill=score))+geom_tile()+theme(panel.background=element_rect(fill="white", colour="white"))+scale_fill_gradient2(low ="blue",midpoint=3,mid="white", high ="red",name="Score")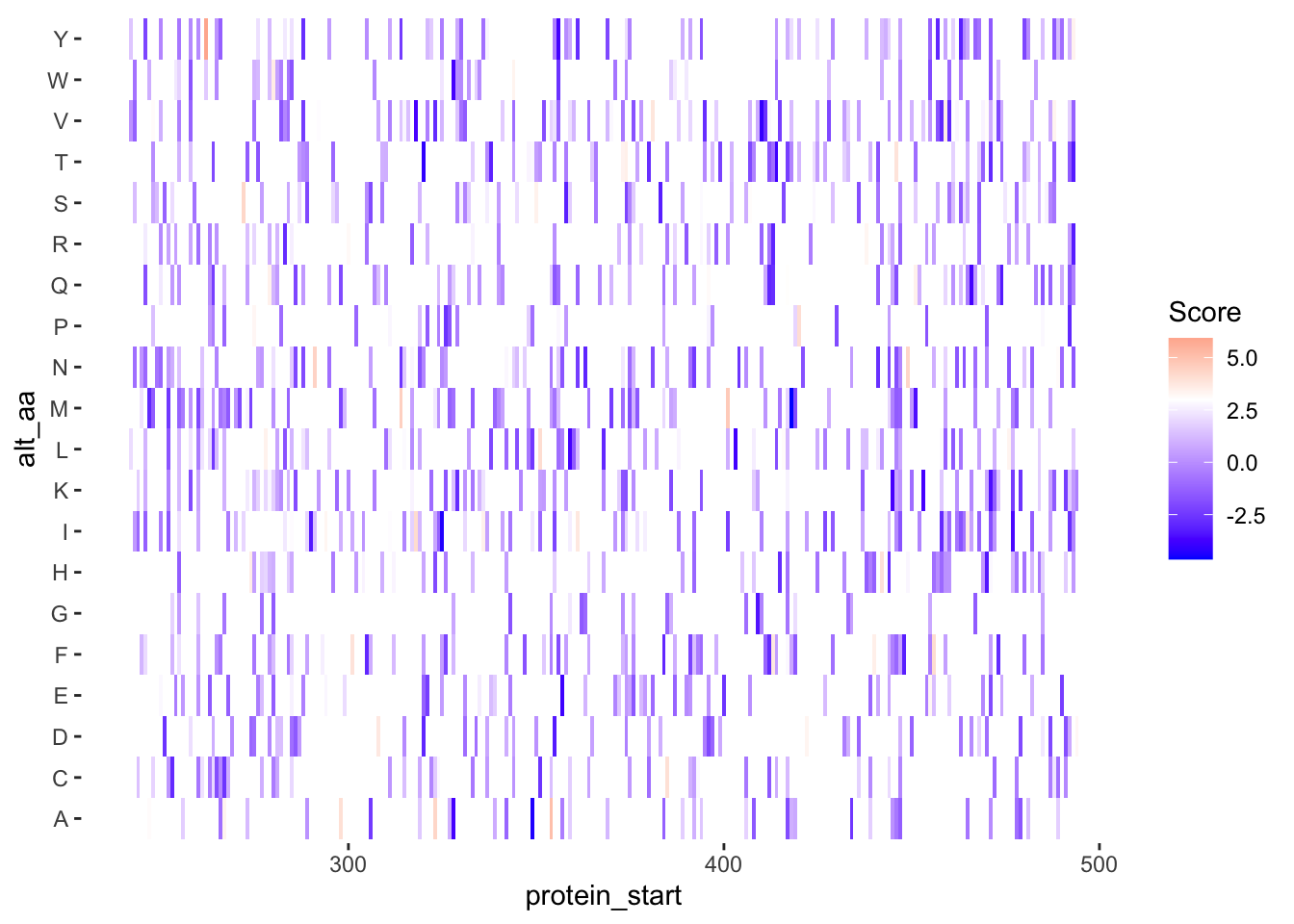
# a=highscore%>%filter(protein_start%in%"255")
# sum(highscore$ct.y)
sample10_simple=sample10_simple%>%
mutate(error_status=case_when(protein_start%in%c(1:241,495:700)~T,
T~F))
plotly=ggplot(sample10_simple%>%
filter(protein_start>=99,protein_start<=600)%>%
mutate(mutant=paste(protein_start,alt_aa)),aes(x=protein_start,y=maf,color=error_status))+
geom_col()+
scale_color_manual(values = c("red","blue"))+
theme_bw()+
theme(legend.position = "none")
ggplotly(plotly)plotly=ggplot(sample10_simple%>%
filter(protein_start>=99,protein_start<=600,ct>=3)%>%
mutate(mutant=paste(protein_start,alt_aa)),aes(x=protein_start,y=maf,fill=error_status))+
geom_col()+
# geom_col(position = "dodge")+
scale_fill_manual(values = c("blue","red"))+
theme_bw()+
theme(legend.position = "none")
# scale_y_continuous(trans="log10")
ggplotly(plotly)plotly=ggplot(sample10_simple%>%
filter(protein_start>=99,protein_start<=600,ct>=1)%>%
mutate(mutant=paste(protein_start,alt_aa)),aes(x=protein_start,y=maf))+
geom_bar(aes(fill=error_status),stat="sum")+
scale_fill_manual(values = c("blue","red"))+
theme_bw()+
theme(legend.position = "none")
ggplotly(plotly)#Things to do with the error rate plot:
#1. Look at sscs vs ngs vs dcs
#2. Combine two samples and see how that improves error rates
# setwd("../")
lane14sortedsamples=merge_samples("Novogene_lane14/sample10_combined","Novogene_lane14/sample11")
lane14sortedsamples=lane14sortedsamples%>%
rowwise()%>%
mutate(ref_aa=strsplit(amino_acids,"/")[[1]][1],
alt_aa=strsplit(amino_acids,"/")[[1]][2])
lane14sortedsamples=lane14sortedsamples%>%mutate(maf=ct/depth)
lane14sorted_simple=lane14sortedsamples%>%dplyr::select(alt_start_pos,protein_start,ref,alt,ref_aa,alt_aa,consequence_terms,ct,depth,maf)
lane14sorted_simple=lane14sorted_simple%>%
mutate(error_status=case_when(protein_start%in%c(1:241,495:700)~T,
T~F))
plotly=ggplot(lane14sorted_simple%>%
filter(protein_start>=99,protein_start<=600,ct>=3)%>%
mutate(mutant=paste(protein_start,alt_aa)),aes(x=protein_start,y=maf))+
geom_bar(aes(fill=error_status),stat="sum")+
scale_fill_manual(values = c("blue","red"))+
theme_bw()+
theme(legend.position = "none")
ggplotly(plotly)# library(ggplot2)
sessionInfo()R version 4.0.0 (2020-04-24)
Platform: x86_64-apple-darwin17.0 (64-bit)
Running under: macOS 10.16
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.0/Resources/lib/libRblas.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.0/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] parallel stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] RColorBrewer_1.1-2 doParallel_1.0.15 iterators_1.0.12 foreach_1.5.0
[5] tictoc_1.0 plotly_4.9.2.1 ggplot2_3.3.3 dplyr_1.0.6
[9] stringr_1.4.0
loaded via a namespace (and not attached):
[1] tidyselect_1.1.0 xfun_0.31 bslib_0.3.1 purrr_0.3.4
[5] colorspace_1.4-1 vctrs_0.3.8 generics_0.0.2 htmltools_0.5.2
[9] viridisLite_0.3.0 yaml_2.2.1 utf8_1.1.4 rlang_0.4.11
[13] isoband_0.2.1 jquerylib_0.1.4 later_1.0.0 pillar_1.6.1
[17] glue_1.4.1 withr_2.4.2 DBI_1.1.0 lifecycle_1.0.0
[21] munsell_0.5.0 gtable_0.3.0 workflowr_1.6.2 htmlwidgets_1.5.1
[25] codetools_0.2-16 evaluate_0.14 labeling_0.3 knitr_1.28
[29] fastmap_1.1.0 crosstalk_1.1.0.1 httpuv_1.5.2 fansi_0.4.1
[33] Rcpp_1.0.4.6 promises_1.1.0 backports_1.1.7 scales_1.1.1
[37] jsonlite_1.7.2 farver_2.0.3 fs_1.4.1 digest_0.6.25
[41] stringi_1.7.5 rprojroot_1.3-2 grid_4.0.0 tools_4.0.0
[45] magrittr_2.0.1 sass_0.4.1 lazyeval_0.2.2 tibble_3.1.2
[49] crayon_1.4.1 whisker_0.4 tidyr_1.1.3 pkgconfig_2.0.3
[53] MASS_7.3-55 ellipsis_0.3.2 data.table_1.12.8 assertthat_0.2.1
[57] rmarkdown_2.14 httr_1.4.2 R6_2.4.1 git2r_0.27.1
[61] compiler_4.0.0