L8-ldscore-GRM
Hae Kyung Im
2020-02-06
Last updated: 2020-02-07
Checks: 5 1
Knit directory: hgen471/
This reproducible R Markdown analysis was created with workflowr (version 1.3.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown is untracked by Git. To know which version of the R Markdown file created these results, you’ll want to first commit it to the Git repo. If you’re still working on the analysis, you can ignore this warning. When you’re finished, you can run wflow_publish to commit the R Markdown file and build the HTML.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20200105) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: docs/.DS_Store
Untracked files:
Untracked: analysis/L8-GRM.Rmd
Untracked: analysis/L8-LD-score.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
There are no past versions. Publish this analysis with wflow_publish() to start tracking its development.
library(tidyverse)Registered S3 methods overwritten by 'ggplot2':
method from
[.quosures rlang
c.quosures rlang
print.quosures rlang── Attaching packages ────────────────────────────────────────────────────────────────────────────────── tidyverse 1.2.1 ──✔ ggplot2 3.1.1 ✔ purrr 0.3.2
✔ tibble 2.1.2 ✔ dplyr 0.8.1
✔ tidyr 0.8.3 ✔ stringr 1.4.0
✔ readr 1.3.1 ✔ forcats 0.4.0── Conflicts ───────────────────────────────────────────────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()work.dir ="~/Downloads/hapmap/"functions
tileplot <- function(mat)
{
mat = data.frame(mat)
mat$Var1 = factor(rownames(mat), levels=rownames(mat)) ## preserve rowname order
melted_mat <- gather(mat,key=Var2,value=value,-Var1)
melted_mat$Var2 = factor(melted_mat$Var2, levels=colnames(mat)) ## preserve colname order
rango = range(melted_mat$value)
pp <- ggplot(melted_mat,aes(x=Var1,y=Var2,fill=value)) + geom_tile() ##+scale_fill_gradientn(colours = c("#C00000", "#FF0000", "#FF8080", "#FFC0C0", "#FFFFFF", "#C0C0FF", "#8080FF", "#0000FF", "#0000C0"), limit = c(-1,1))
pp
}plot GRM to show population structure
## calculate GRM using plink, use grm.gz format
grmhead = paste0(work.dir,"output/hapmapch22")
cl_calc_grm = glue::glue("plink --bfile hapmapch22 --make-grm-gz --out {grmhead}")
print(cl_calc_grm) ## run this in the command lineplink --bfile hapmapch22 --make-grm-gz --out ~/Downloads/hapmap/output/hapmapch22## define function to read GRM (in grm-gz format) into matrix
grmgz2mat = function(grmhead)
{
## given plink like header, it reads thd grm file and returns matrix of grm
grm = read.table(paste0(grmhead,".grm.gz"),header=F)
grmid = read.table(paste0(grmhead,".grm.id"),header=F)
grmat = matrix(0,max(grm$V1),max(grm$V2))
rownames(grmat) = grmid$V2
colnames(grmat) = grmid$V2
## fill lower matrix of GRM
grmat[upper.tri(grmat,diag=TRUE)]= grm$V4
## make upper = lower, need to subtract diag
grmat + t(grmat) - diag(diag(grmat))
}
## select a 2 CEU and 2 YRI individuals
## 1345 NA07349 NA07347 NA07346 1 0 CEU
## 1353 NA12376 NA12546 NA12489 2 0 CEU
## Y004 NA18500 NA18501 NA18502 1 0 YRI missing NA18502
## Y051 NA19208 NA19207 NA19206 1 0 YRI
## Y058 NA19221 NA19223 NA19222 2 0 YRI
ind = c("NA07349","NA07347","NA07346","NA12376","NA12546","NA12489",
"NA19208","NA19207","NA19206", "NA19221","NA19223","NA19222")
## read grm calculated in plink into R matrix
grmat = grmgz2mat(grmhead)
## plot grmat to see the population structure
tileplot(grmat)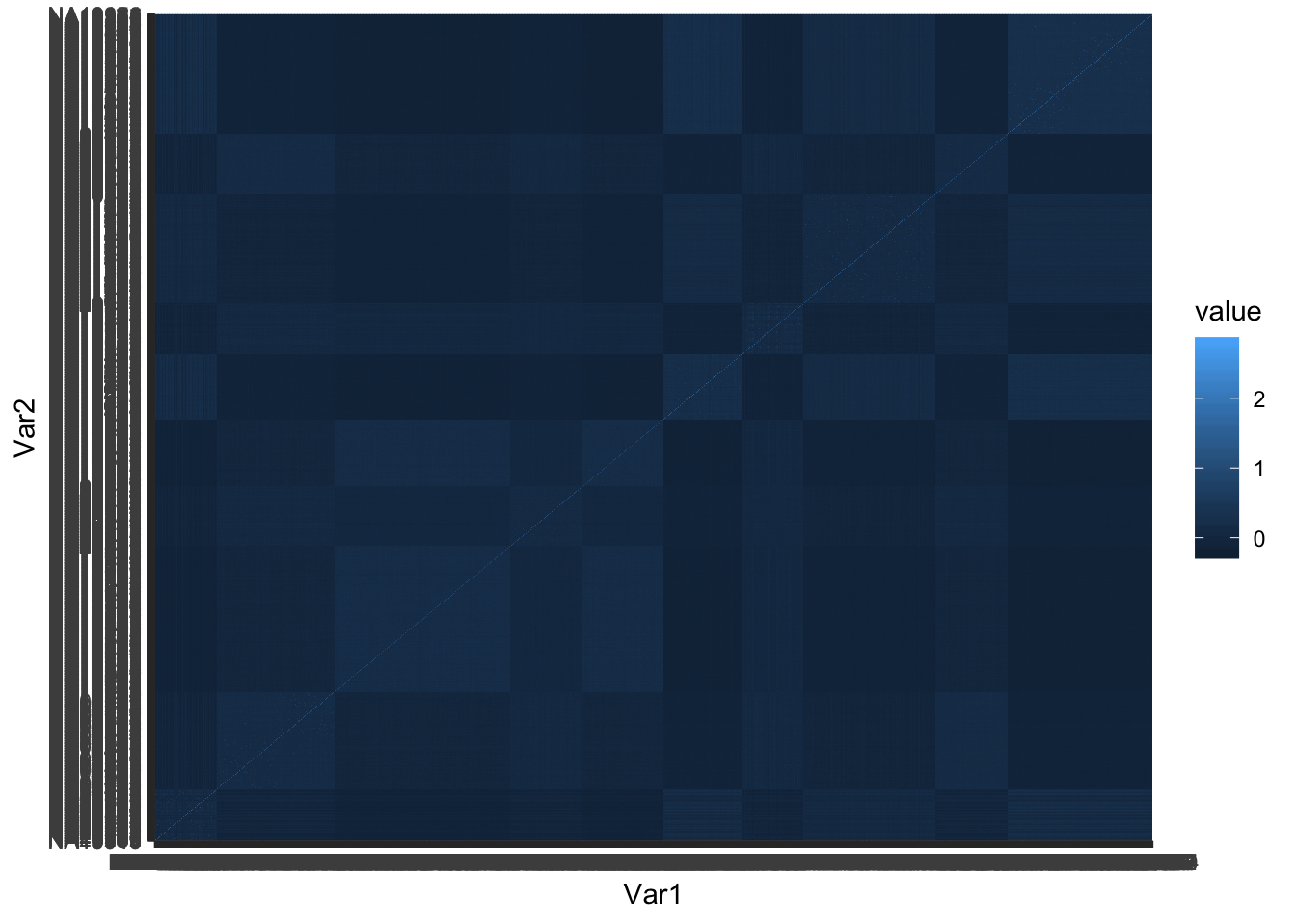
sessionInfo()R version 3.6.0 (2019-04-26)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS Mojave 10.14.6
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] forcats_0.4.0 stringr_1.4.0 dplyr_0.8.1 purrr_0.3.2
[5] readr_1.3.1 tidyr_0.8.3 tibble_2.1.2 ggplot2_3.1.1
[9] tidyverse_1.2.1
loaded via a namespace (and not attached):
[1] Rcpp_1.0.2 cellranger_1.1.0 plyr_1.8.4 pillar_1.4.1
[5] compiler_3.6.0 git2r_0.25.2 workflowr_1.3.0 tools_3.6.0
[9] digest_0.6.19 lubridate_1.7.4 jsonlite_1.6 evaluate_0.14
[13] nlme_3.1-139 gtable_0.3.0 lattice_0.20-38 pkgconfig_2.0.2
[17] rlang_0.4.1 cli_1.1.0 rstudioapi_0.10 yaml_2.2.0
[21] haven_2.1.0 xfun_0.7 withr_2.1.2 xml2_1.2.0
[25] httr_1.4.0 knitr_1.23 hms_0.4.2 generics_0.0.2
[29] fs_1.3.1 rprojroot_1.3-2 grid_3.6.0 tidyselect_0.2.5
[33] glue_1.3.1 R6_2.4.0 readxl_1.3.1 rmarkdown_1.13
[37] modelr_0.1.4 magrittr_1.5 backports_1.1.4 scales_1.0.0
[41] htmltools_0.4.0 rvest_0.3.4 assertthat_0.2.1 colorspace_1.4-1
[45] labeling_0.3 stringi_1.4.3 lazyeval_0.2.2 munsell_0.5.0
[49] broom_0.5.2 crayon_1.3.4