asthma_prelim_results
Last updated: 2021-10-26
Checks: 7 0
Knit directory: funcFinemapping/
This reproducible R Markdown analysis was created with workflowr (version 1.6.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it's best to always run the code in an empty environment.
The command set.seed(20210404) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version db3c6f9. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .ipynb_checkpoints/
Ignored: analysis/ldsc_results.nb.html
Ignored: analysis/results.nb.html
Ignored: analysis/snp_finemapping_results.nb.html
Ignored: analysis/splicing.nb.html
Untracked files:
Untracked: SNPs_categories,png
Untracked: SNPs_categories.png
Untracked: analysis/asthma_prelim_results_archive.Rmd
Untracked: analysis/asthma_results_cp.Rmd
Untracked: analysis/enhancer_gene_feature.Rmd
Untracked: analysis/feedback.Rmd
Untracked: analysis/gene_finemapping_results.Rmd
Untracked: analysis/learn_susie.Rmd
Untracked: analysis/mtsplice_finemapping_results.Rmd
Untracked: analysis/notes.Rmd
Untracked: analysis/snp_finemapping_results.Rmd
Untracked: analysis/splicing.Rmd
Untracked: code/.ipynb_checkpoints/
Untracked: code/ldsc_regression.sh
Untracked: code/make_plots.R
Untracked: code/run_ldsc.sh
Untracked: code/run_ldsc_with_bed.sh
Untracked: code/run_susie.R
Untracked: code/run_torus.sh
Untracked: code/split_vcf.sh
Untracked: data/num_overlaps_finemapped_SNPs_and_ctcf.txt
Untracked: data/scz_2018
Untracked: data/torus_enrichment_novel_annot.est
Untracked: data/torus_joint_enrichment.est
Untracked: data/torus_joint_refined_enrichment.est
Untracked: enhancer_gene_feature.rmd
Untracked: fig1_panels.pdf
Untracked: fig2.pdf
Untracked: fig_panel2.pdf
Untracked: gene_mapping.pdf
Untracked: output/background_SNPs_annotated_percent.txt
Untracked: panel_figure2.pdf
Untracked: test.txt
Unstaged changes:
Modified: analysis/asthma_results.Rmd
Modified: analysis/enrichment_analysis.Rmd
Deleted: output/asthma/Caldero2019_diffDA_annot_percent.txt
Deleted: output/asthma/Caldero2019_stimuDA_annot_percent.txt
Deleted: output/asthma/celltype_specific_adult_lungs_torus.est
Deleted: output/asthma/diffe_adult_blood_torus.est
Deleted: output/asthma/joint_lung_vs_blood_immune_diff_torus.est
Deleted: output/asthma/joint_lung_vs_blood_immune_stimu_torus.est
Deleted: output/asthma/lung_clusters_dict.txt
Deleted: output/asthma/lung_clusters_info.txt
Deleted: output/asthma/stimu_adult_blood_torus.est
Deleted: output/asthma/zhang2021_annot_percent.txt
Deleted: output/asthma/zhang2021_cell_type_overlaps.txt
Deleted: output/asthma/zhang2021_peaks_per_celltype.txt
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were made to the R Markdown (analysis/asthma_prelim_results.Rmd) and HTML (docs/asthma_prelim_results.html) files. If you've configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | db3c6f9 | Jing Gu | 2021-10-26 | test enrichment for asthma risk variants |
cell-type specific annotations
- Caldero et al. 2019 - ATAC-Seq data for FACS-sorted cells from whole blood I extracted ATAC-Seq peaks specific to each immune cell type from the count table that consists of a union set of peaks across cells based on the provided cutoffs.
Rheumatoid arthritis
This analysis was done to replicate the enrichment results from figure 5b in the paper. Overall, the magnitude of enrichment estimates are much smaller than that in figure 5b. It is possible that we did not use the same GWAS dataset. They used LDSC to estimate enrichment estimates. I will check how they ran LDSC over these annotations. The stimulated immune cells tend to have slightly higher enrichment estimates, but not as significant due to overlapped CIs. However, the figure in the paper demonstrates strong enrichment of stimulated-related peaks for cells such as Bulk B, naive B, CD8+ T and Th1. 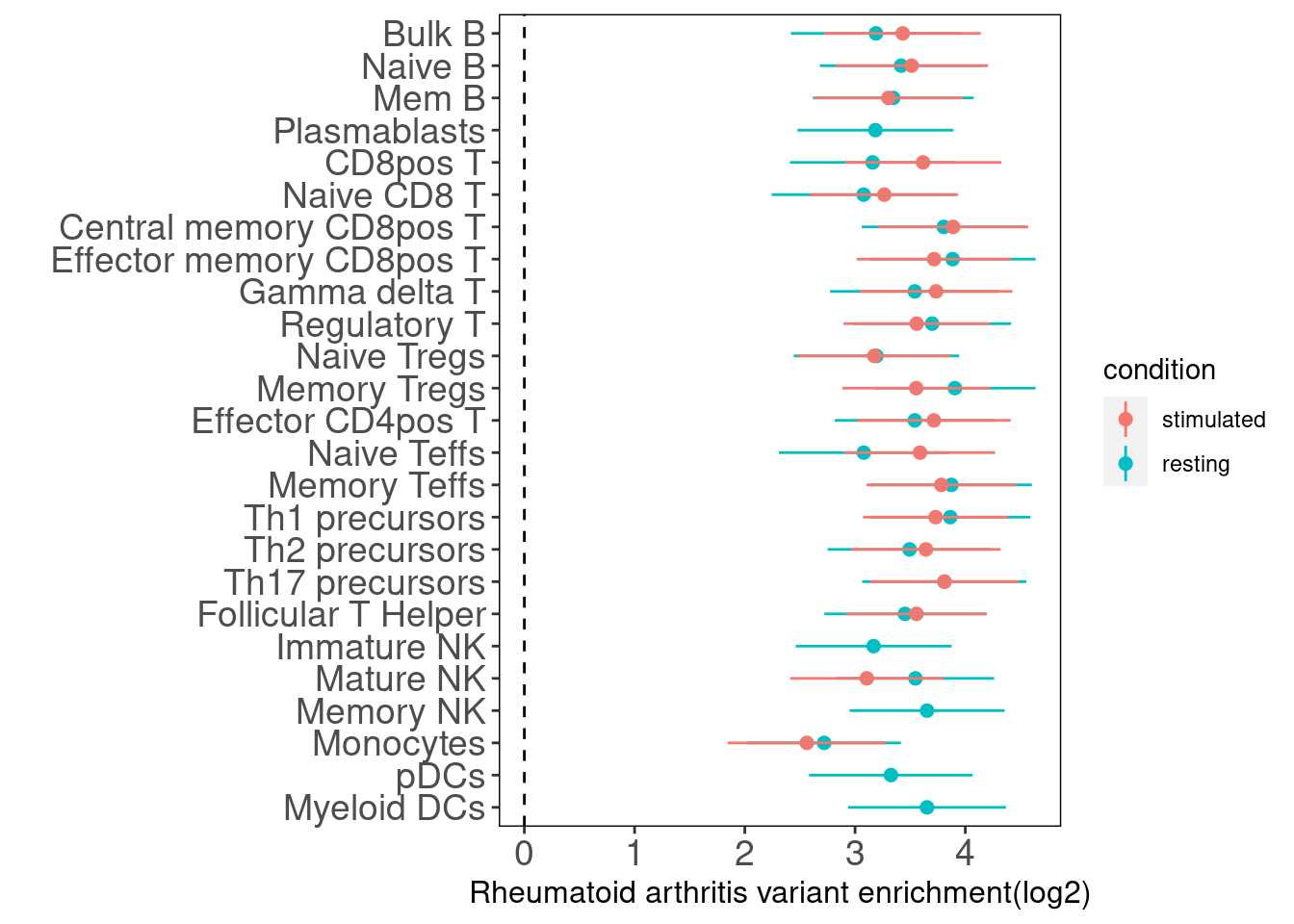
Adult-Onset Asthma
For Asthma, we did not see stronger enrichment of stimulated-related peaks compared with resting ones. But overall all immune cells whether or not stimulated (except for the stimulated mature NK) are significantly enriched for GWAS risk variants. 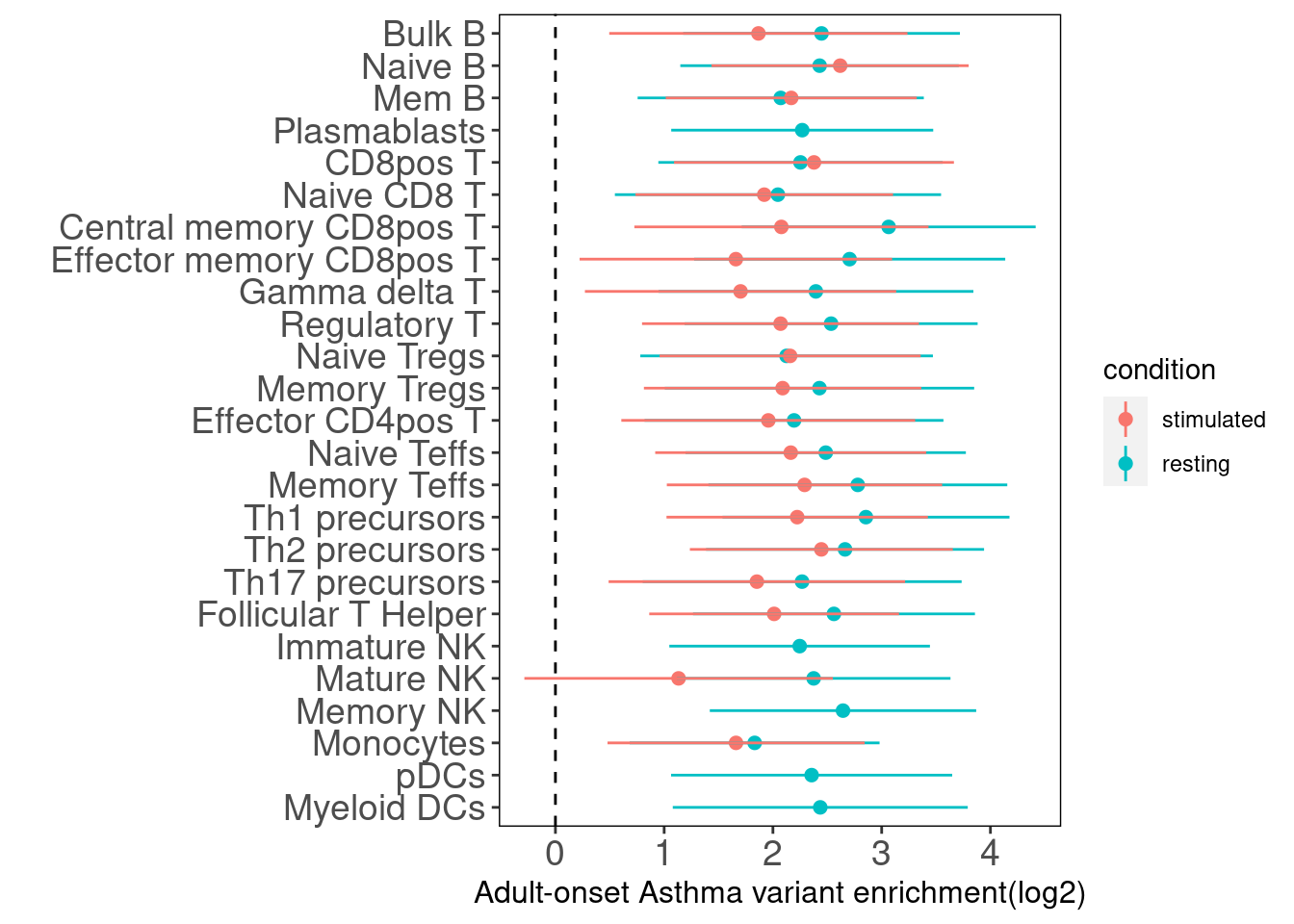
Allergy
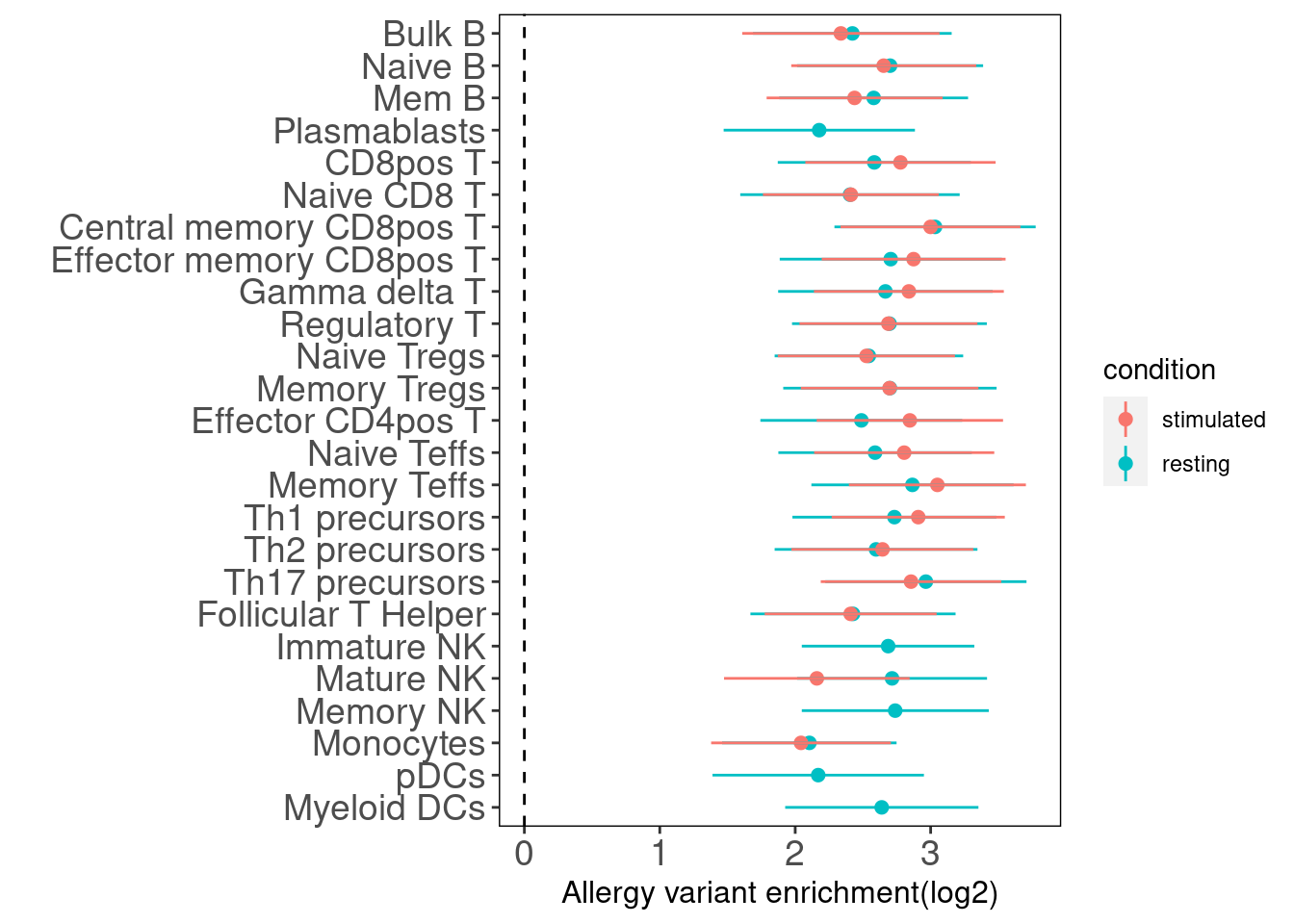
Joint Enrichment of a pair of annotations
Adult-Onset Asthma
A pair of annotations - stimulated and resting for each immune cell type was jointly tested for their enrichment of GWAS risk variants. We observed many cell types share GWAS association signals. Conditional on resting cells,there are five stimulated cell types still show enrichment including Naive B, Memory B, CD8+ T , Naive CD8+ T and Naive Tregs cells. 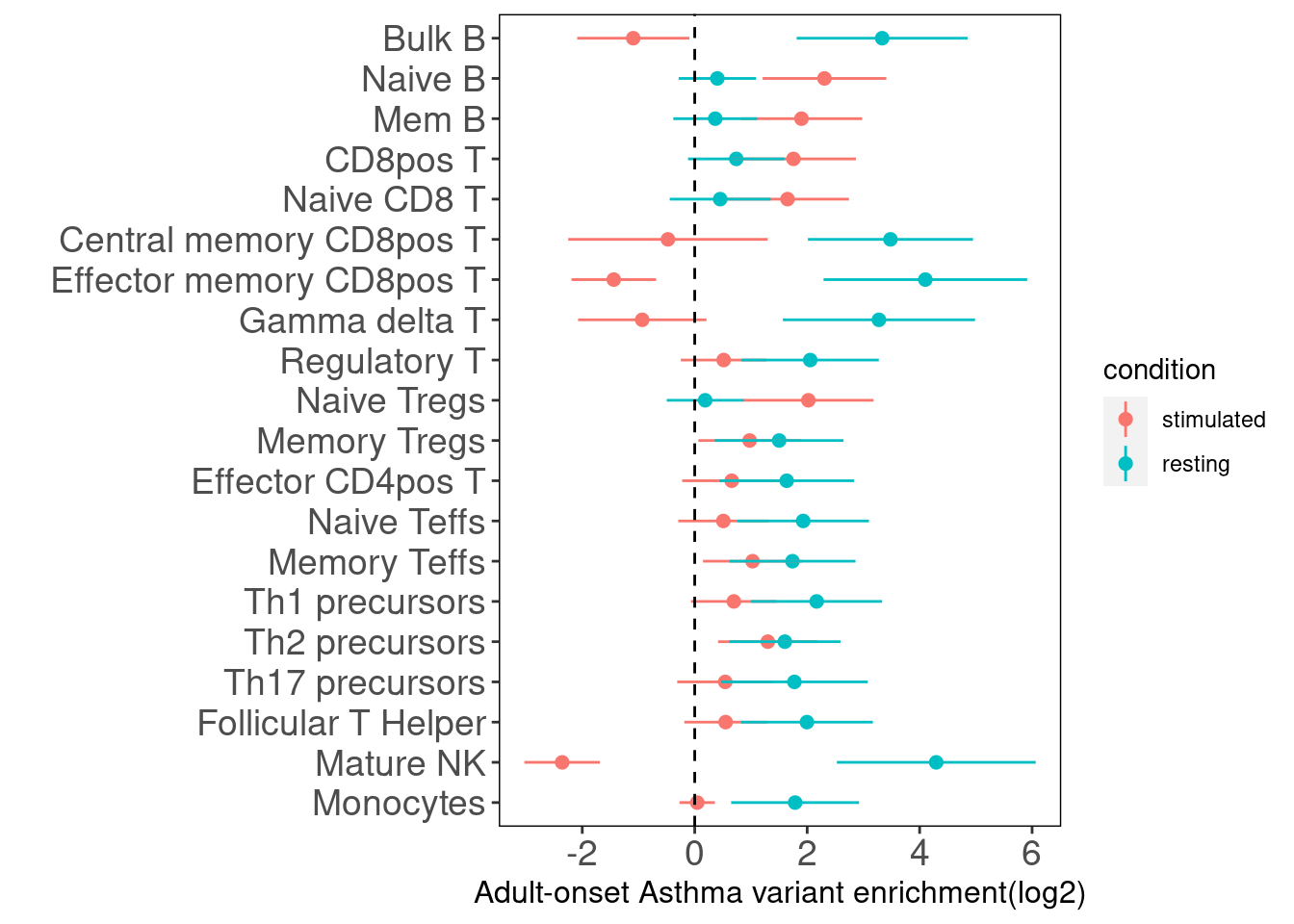
Annotations - Differential accessible peaks
Caldero et al. 2019
- Significant differentially accessible regions when compared to progenitor cells
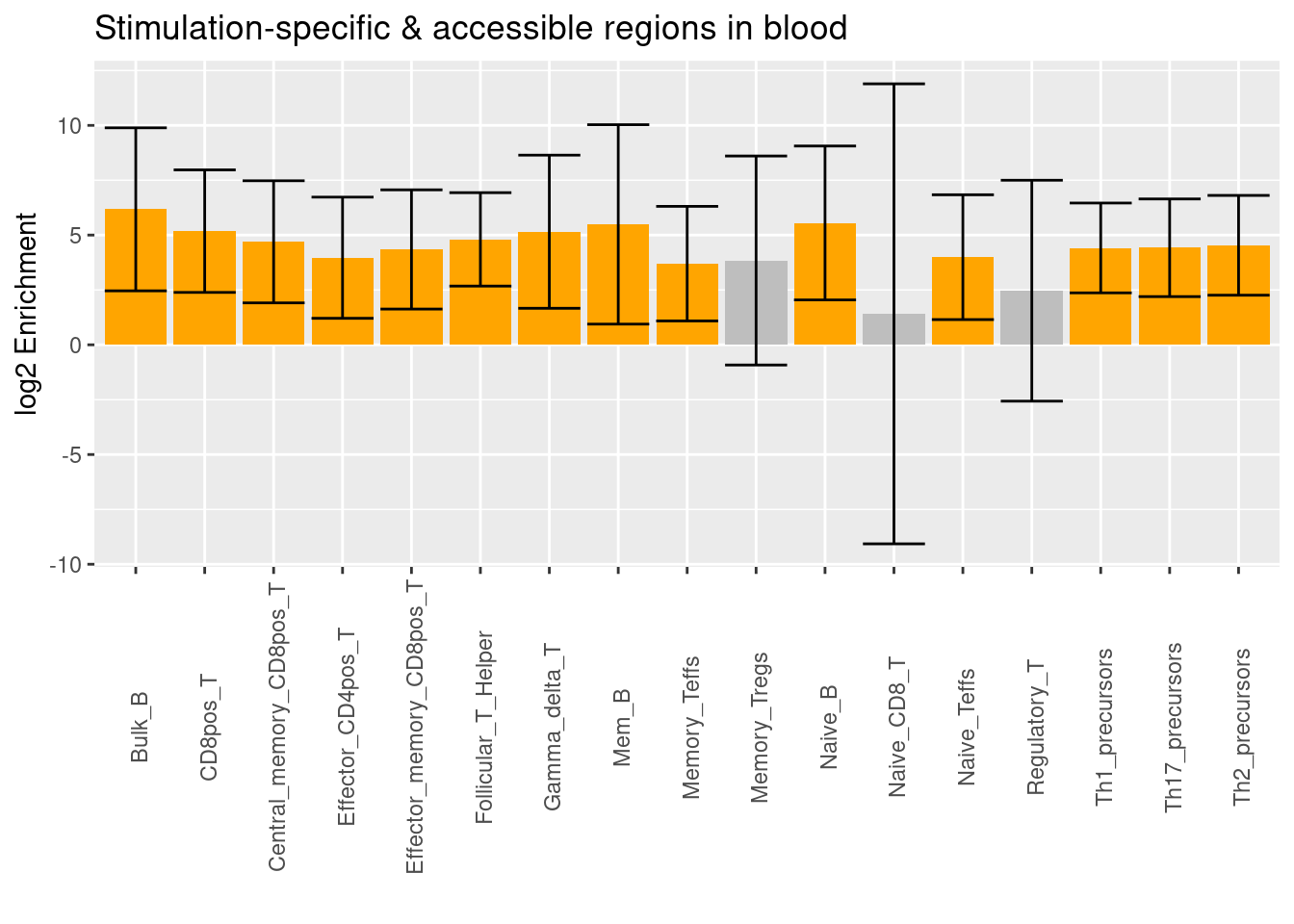
- Significant differentially accessible regions under stimulation
sessionInfo()R version 4.0.4 (2021-02-15)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Scientific Linux 7.4 (Nitrogen)
Matrix products: default
BLAS/LAPACK: /software/openblas-0.3.13-el7-x86_64/lib/libopenblas_haswellp-r0.3.13.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] knitr_1.31 ggplot2_3.3.3 workflowr_1.6.2
loaded via a namespace (and not attached):
[1] Rcpp_1.0.7 highr_0.8 pillar_1.5.0 compiler_4.0.4
[5] bslib_0.2.4 later_1.1.0.1 jquerylib_0.1.3 git2r_0.28.0
[9] tools_4.0.4 digest_0.6.27 jsonlite_1.7.2 evaluate_0.14
[13] lifecycle_1.0.0 tibble_3.0.6 gtable_0.3.0 pkgconfig_2.0.3
[17] rlang_0.4.11 DBI_1.1.1 yaml_2.2.1 xfun_0.21
[21] withr_2.4.2 dplyr_1.0.4 stringr_1.4.0 generics_0.1.0
[25] fs_1.5.0 vctrs_0.3.8 sass_0.3.1 tidyselect_1.1.1
[29] rprojroot_2.0.2 grid_4.0.4 glue_1.4.2 R6_2.5.1
[33] fansi_0.5.0 rmarkdown_2.7 farver_2.1.0 purrr_0.3.4
[37] magrittr_2.0.1 whisker_0.4 scales_1.1.1 promises_1.2.0.1
[41] ellipsis_0.3.2 htmltools_0.5.1.1 assertthat_0.2.1 colorspace_2.0-2
[45] httpuv_1.5.5 labeling_0.4.2 utf8_1.2.2 stringi_1.5.3
[49] munsell_0.5.0 crayon_1.4.1