Inverse transform sampling
Matt Bonakdarpour
University of ChicagoJanuary 5, 2025
Last updated: 2026-01-05
Checks: 6 1
Knit directory: fiveMinuteStats/analysis/
This reproducible R Markdown analysis was created with workflowr (version 1.7.1). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of
the R Markdown file created these results, you’ll want to first commit
it to the Git repo. If you’re still working on the analysis, you can
ignore this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(12345) was run prior to running the
code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 78b1165. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Unstaged changes:
Modified: analysis/Importance_sampling.Rmd
Modified: analysis/inverse_transform_sampling.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown
(analysis/inverse_transform_sampling.Rmd) and HTML
(docs/inverse_transform_sampling.html) files. If you’ve
configured a remote Git repository (see ?wflow_git_remote),
click on the hyperlinks in the table below to view the files as they
were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 67acf84 | Peter Carbonetto | 2025-12-29 | Added pdf version of inverse_transform_sampling vignette. |
| html | 67acf84 | Peter Carbonetto | 2025-12-29 | Added pdf version of inverse_transform_sampling vignette. |
| html | 5f62ee6 | Matthew Stephens | 2019-03-31 | Build site. |
| Rmd | 0cd28bd | Matthew Stephens | 2019-03-31 | workflowr::wflow_publish(all = TRUE) |
| Rmd | a86efa5 | GitHub | 2018-04-25 | fix misspelling |
| html | 34bcc51 | John Blischak | 2017-03-06 | Build site. |
| Rmd | 5fbc8b5 | John Blischak | 2017-03-06 | Update workflowr project with wflow_update (version 0.4.0). |
| Rmd | 391ba3c | John Blischak | 2017-03-06 | Remove front and end matter of non-standard templates. |
| html | 8e61683 | Marcus Davy | 2017-03-03 | rendered html using wflow_build(all=TRUE) |
| html | 5d0fa13 | Marcus Davy | 2017-03-02 | wflow_build() rendered html files |
| Rmd | d674141 | Marcus Davy | 2017-02-26 | typos, refs |
| html | c3b365a | John Blischak | 2017-01-02 | Build site. |
| Rmd | 67a8575 | John Blischak | 2017-01-02 | Use external chunk to set knitr chunk options. |
| Rmd | 5ec12c7 | John Blischak | 2017-01-02 | Use session-info chunk. |
| Rmd | dfaabc4 | mbonakda | 2016-02-02 | spin off inverse transform sampling into its own note |
See here for a PDF version of this vignette.
Prerequisites
This document assumes basic familiarity with probability theory.
Overview
Inverse transform sampling is a method for generating random numbers from any probability distribution by using its inverse cumulative distribution, \(F^{-1}(x)\). Recall that the cumulative distribution for a random variable \(X\) is \(F_X(x) = P(X \leq x)\). In what follows, we assume that our computer can, on demand, generate independent realizations of a random variable \(U\) uniformly distributed on \([0,1]\).
Algorithm
Continuous distributions
Assume we want to generate a random variable \(X\) with cumulative distribution function (CDF) \(F_X\). The inverse transform sampling algorithm is simple:
Generate \(U \sim \mathrm{Unif}(0,1)\).
Let \(X = F_X^{-1}(U)\).
Then \(X\) will follow the distribution governed by the CDF \(F_X\), which was our desired result.
Note that this algorithm works in general but is not always practical. For example, inverting \(F_X\) is easy if \(X\) is an exponential random variable, but it is harder if \(X\) is normal random variable.
Discrete distributions
Now we will consider the discrete version of the inverse transform method. Assume that \(X\) is a discrete random variable such that \(P(X = x_i) = p_i\). The algorithm proceeds as follows:
Generate \(U \sim \text{Unif}(0,1)\).
Determine the index \(k\) such that \(\sum_{j=1}^{k-1} p_j \leq U < \sum_{j=1}^k p_j\), and return \(X = x_k\).
(Notice that the second step requires a search. )
Assume our random variable \(X\) can take on any one of \(K\) values with probabilities \(\{p_1, \ldots, p_K\}\). We implement the algorithm below, assuming these probabilities are stored in a vector called “pvec.”
discrete_inv_transform_sample <- function (pvec) {
K <- length(pvec)
u <- runif(1)
if (u <= pvec[1])
return(1)
for (k in seq(2,K)) {
if(sum(pvec[seq(1,k-1)]) < u && u <= sum(pvec[seq(1,k)]))
return(k)
}
}Note that this is this an inefficient implementation given here for pedagogical purposes.
Continuous example: exponential distribution
Assume \(Y\) is an exponential random variable with rate parameter \(\lambda = 2\). Recall that the probability density function is \(p(y) = 2e^{-2y}\), for \(y > 0\). First, we derive the CDF: \[ F_Y(x) = P(Y\leq x) = \int_0^x 2e^{-2y} dy = 1 - e^{-2x} \] Solving for the inverse CDF, we get that \[ F_Y^{-1}(y) = \textstyle -\frac{1}{2}\log(1-y) \]
Using our algorithm above, we first generate \(U \sim \text{Unif}(0,1)\), then set \(X = -\log(1-U)/2\). We do this in the R code below and compare the histogram of our samples with the true density of \(Y\).
num_samples <- 1000
u <- runif(num_samples)
x <- -log(1-u)/2
hist(x,n = 32,freq = FALSE,xlab = 'X',main = "")
curve(dexp(x,rate = 2),0,3,lwd = 2,add = TRUE)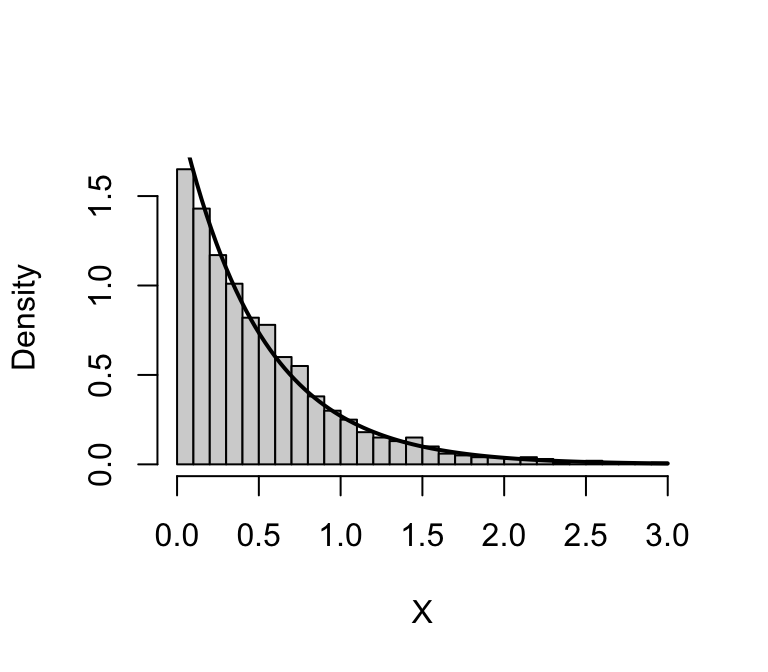
Indeed, the random draws appear are following the intended distribution.
Discrete example
Let’s assume we want to simulate a discrete random variable \(X\) that follows the following distribution:
| \(k\) | \(\Pr(X = k)\) |
|---|---|
| 1 | 0.1 |
| 2 | 0.4 |
| 3 | 0.2 |
| 4 | 0.3 |
Below we simulate from this distribution using the “discrete_inv_transform_sample” function above, and plot both the true probability vector, and the empirical proportions from our simulation.
par(mfcol = c(1,2))
num_samples <- 1000
pvec <- c(0.1,0.4,0.2,0.3)
names(pvec) <- 1:4
samples <- rep(0,num_samples)
for(i in seq_len(num_samples)) {
samples[i] <- discrete_inv_transform_sample(pvec)
}
barplot(pvec,main = "true pdf")
barplot(table(samples),main = "empirical pdf")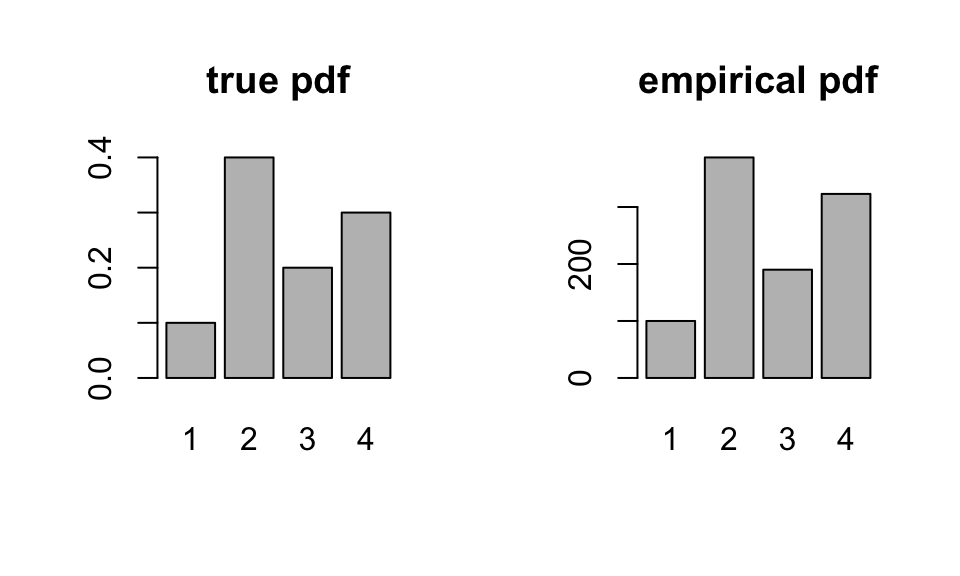
Again, the plot supports our claim that we are drawing from the correct probability distribution.
sessionInfo()
# R version 4.3.3 (2024-02-29)
# Platform: aarch64-apple-darwin20 (64-bit)
# Running under: macOS 15.7.1
#
# Matrix products: default
# BLAS: /Library/Frameworks/R.framework/Versions/4.3-arm64/Resources/lib/libRblas.0.dylib
# LAPACK: /Library/Frameworks/R.framework/Versions/4.3-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.11.0
#
# locale:
# [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
#
# time zone: America/Chicago
# tzcode source: internal
#
# attached base packages:
# [1] stats graphics grDevices utils datasets methods base
#
# loaded via a namespace (and not attached):
# [1] vctrs_0.6.5 cli_3.6.5 knitr_1.50 rlang_1.1.6
# [5] xfun_0.52 stringi_1.8.7 promises_1.3.3 jsonlite_2.0.0
# [9] workflowr_1.7.1 glue_1.8.0 rprojroot_2.0.4 git2r_0.33.0
# [13] htmltools_0.5.8.1 httpuv_1.6.14 sass_0.4.10 rmarkdown_2.29
# [17] evaluate_1.0.4 jquerylib_0.1.4 tibble_3.3.0 fastmap_1.2.0
# [21] yaml_2.3.10 lifecycle_1.0.4 whisker_0.4.1 stringr_1.5.1
# [25] compiler_4.3.3 fs_1.6.6 Rcpp_1.1.0 pkgconfig_2.0.3
# [29] later_1.4.2 digest_0.6.37 R6_2.6.1 pillar_1.11.0
# [33] magrittr_2.0.3 bslib_0.9.0 tools_4.3.3 cachem_1.1.0