Simulation study
Pedro L. Baldoni
08 April, 2025
Last updated: 2025-04-08
Checks: 7 0
Knit directory: DTU-code/analysis/
This reproducible R Markdown analysis was created with workflowr (version 1.7.1). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20240501) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 59e7c06. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: .gitignore
Ignored: .renvignore
Ignored: DTU-code.Rproj
Ignored: code/.RData
Ignored: code/mouse/data/slurm-16691239.out
Ignored: code/mouse/salmon-index/slurm-16693056.out
Ignored: code/mouse/salmon/slurm-16693640.out
Ignored: code/mouse/subread-index/.RData
Ignored: code/mouse/subread-index/buildindex.Rout
Ignored: code/mouse/subread-index/slurm-19350188.out
Ignored: code/mouse/subread/.nextflow.log
Ignored: code/mouse/subread/.nextflow/
Ignored: code/mouse/subread/log/
Ignored: code/mouse/subread/report.html
Ignored: code/mouse/subread/slurm-19356012.out
Ignored: code/mouse/subread/timeline.html
Ignored: code/mouse/subread/trace-20241127-50139746.txt
Ignored: code/pkg/.Rbuildignore
Ignored: code/pkg/.Rhistory
Ignored: code/pkg/.Rproj.user/
Ignored: code/pkg/src/.gitignore
Ignored: code/pkg/src/RcppExports.o
Ignored: code/pkg/src/pkg.so
Ignored: code/pkg/src/rcpparma_hello_world.o
Ignored: code/simulation-lenient/slurm-20927698_1.out
Ignored: code/simulation-lenient/slurm-20927698_10.out
Ignored: code/simulation-lenient/slurm-20927698_100.out
Ignored: code/simulation-lenient/slurm-20927698_101.out
Ignored: code/simulation-lenient/slurm-20927698_102.out
Ignored: code/simulation-lenient/slurm-20927698_103.out
Ignored: code/simulation-lenient/slurm-20927698_104.out
Ignored: code/simulation-lenient/slurm-20927698_105.out
Ignored: code/simulation-lenient/slurm-20927698_106.out
Ignored: code/simulation-lenient/slurm-20927698_107.out
Ignored: code/simulation-lenient/slurm-20927698_108.out
Ignored: code/simulation-lenient/slurm-20927698_109.out
Ignored: code/simulation-lenient/slurm-20927698_11.out
Ignored: code/simulation-lenient/slurm-20927698_110.out
Ignored: code/simulation-lenient/slurm-20927698_111.out
Ignored: code/simulation-lenient/slurm-20927698_112.out
Ignored: code/simulation-lenient/slurm-20927698_113.out
Ignored: code/simulation-lenient/slurm-20927698_114.out
Ignored: code/simulation-lenient/slurm-20927698_115.out
Ignored: code/simulation-lenient/slurm-20927698_116.out
Ignored: code/simulation-lenient/slurm-20927698_117.out
Ignored: code/simulation-lenient/slurm-20927698_118.out
Ignored: code/simulation-lenient/slurm-20927698_119.out
Ignored: code/simulation-lenient/slurm-20927698_12.out
Ignored: code/simulation-lenient/slurm-20927698_120.out
Ignored: code/simulation-lenient/slurm-20927698_13.out
Ignored: code/simulation-lenient/slurm-20927698_14.out
Ignored: code/simulation-lenient/slurm-20927698_15.out
Ignored: code/simulation-lenient/slurm-20927698_16.out
Ignored: code/simulation-lenient/slurm-20927698_17.out
Ignored: code/simulation-lenient/slurm-20927698_18.out
Ignored: code/simulation-lenient/slurm-20927698_19.out
Ignored: code/simulation-lenient/slurm-20927698_2.out
Ignored: code/simulation-lenient/slurm-20927698_20.out
Ignored: code/simulation-lenient/slurm-20927698_21.out
Ignored: code/simulation-lenient/slurm-20927698_22.out
Ignored: code/simulation-lenient/slurm-20927698_23.out
Ignored: code/simulation-lenient/slurm-20927698_24.out
Ignored: code/simulation-lenient/slurm-20927698_25.out
Ignored: code/simulation-lenient/slurm-20927698_26.out
Ignored: code/simulation-lenient/slurm-20927698_27.out
Ignored: code/simulation-lenient/slurm-20927698_28.out
Ignored: code/simulation-lenient/slurm-20927698_29.out
Ignored: code/simulation-lenient/slurm-20927698_3.out
Ignored: code/simulation-lenient/slurm-20927698_30.out
Ignored: code/simulation-lenient/slurm-20927698_31.out
Ignored: code/simulation-lenient/slurm-20927698_32.out
Ignored: code/simulation-lenient/slurm-20927698_33.out
Ignored: code/simulation-lenient/slurm-20927698_34.out
Ignored: code/simulation-lenient/slurm-20927698_35.out
Ignored: code/simulation-lenient/slurm-20927698_36.out
Ignored: code/simulation-lenient/slurm-20927698_37.out
Ignored: code/simulation-lenient/slurm-20927698_38.out
Ignored: code/simulation-lenient/slurm-20927698_39.out
Ignored: code/simulation-lenient/slurm-20927698_4.out
Ignored: code/simulation-lenient/slurm-20927698_40.out
Ignored: code/simulation-lenient/slurm-20927698_41.out
Ignored: code/simulation-lenient/slurm-20927698_42.out
Ignored: code/simulation-lenient/slurm-20927698_43.out
Ignored: code/simulation-lenient/slurm-20927698_44.out
Ignored: code/simulation-lenient/slurm-20927698_45.out
Ignored: code/simulation-lenient/slurm-20927698_46.out
Ignored: code/simulation-lenient/slurm-20927698_47.out
Ignored: code/simulation-lenient/slurm-20927698_48.out
Ignored: code/simulation-lenient/slurm-20927698_49.out
Ignored: code/simulation-lenient/slurm-20927698_5.out
Ignored: code/simulation-lenient/slurm-20927698_50.out
Ignored: code/simulation-lenient/slurm-20927698_51.out
Ignored: code/simulation-lenient/slurm-20927698_52.out
Ignored: code/simulation-lenient/slurm-20927698_53.out
Ignored: code/simulation-lenient/slurm-20927698_54.out
Ignored: code/simulation-lenient/slurm-20927698_55.out
Ignored: code/simulation-lenient/slurm-20927698_56.out
Ignored: code/simulation-lenient/slurm-20927698_57.out
Ignored: code/simulation-lenient/slurm-20927698_58.out
Ignored: code/simulation-lenient/slurm-20927698_59.out
Ignored: code/simulation-lenient/slurm-20927698_6.out
Ignored: code/simulation-lenient/slurm-20927698_60.out
Ignored: code/simulation-lenient/slurm-20927698_61.out
Ignored: code/simulation-lenient/slurm-20927698_62.out
Ignored: code/simulation-lenient/slurm-20927698_63.out
Ignored: code/simulation-lenient/slurm-20927698_64.out
Ignored: code/simulation-lenient/slurm-20927698_65.out
Ignored: code/simulation-lenient/slurm-20927698_66.out
Ignored: code/simulation-lenient/slurm-20927698_67.out
Ignored: code/simulation-lenient/slurm-20927698_68.out
Ignored: code/simulation-lenient/slurm-20927698_69.out
Ignored: code/simulation-lenient/slurm-20927698_7.out
Ignored: code/simulation-lenient/slurm-20927698_70.out
Ignored: code/simulation-lenient/slurm-20927698_71.out
Ignored: code/simulation-lenient/slurm-20927698_72.out
Ignored: code/simulation-lenient/slurm-20927698_73.out
Ignored: code/simulation-lenient/slurm-20927698_74.out
Ignored: code/simulation-lenient/slurm-20927698_75.out
Ignored: code/simulation-lenient/slurm-20927698_76.out
Ignored: code/simulation-lenient/slurm-20927698_77.out
Ignored: code/simulation-lenient/slurm-20927698_78.out
Ignored: code/simulation-lenient/slurm-20927698_79.out
Ignored: code/simulation-lenient/slurm-20927698_8.out
Ignored: code/simulation-lenient/slurm-20927698_80.out
Ignored: code/simulation-lenient/slurm-20927698_81.out
Ignored: code/simulation-lenient/slurm-20927698_82.out
Ignored: code/simulation-lenient/slurm-20927698_83.out
Ignored: code/simulation-lenient/slurm-20927698_84.out
Ignored: code/simulation-lenient/slurm-20927698_85.out
Ignored: code/simulation-lenient/slurm-20927698_86.out
Ignored: code/simulation-lenient/slurm-20927698_87.out
Ignored: code/simulation-lenient/slurm-20927698_88.out
Ignored: code/simulation-lenient/slurm-20927698_89.out
Ignored: code/simulation-lenient/slurm-20927698_9.out
Ignored: code/simulation-lenient/slurm-20927698_90.out
Ignored: code/simulation-lenient/slurm-20927698_91.out
Ignored: code/simulation-lenient/slurm-20927698_92.out
Ignored: code/simulation-lenient/slurm-20927698_93.out
Ignored: code/simulation-lenient/slurm-20927698_94.out
Ignored: code/simulation-lenient/slurm-20927698_95.out
Ignored: code/simulation-lenient/slurm-20927698_96.out
Ignored: code/simulation-lenient/slurm-20927698_97.out
Ignored: code/simulation-lenient/slurm-20927698_98.out
Ignored: code/simulation-lenient/slurm-20927698_99.out
Ignored: code/simulation-lenient/slurm-20927938.out
Ignored: code/simulation-lenient/summarize.Rout
Ignored: code/simulation-no-trend/
Ignored: code/simulation/.RData
Ignored: code/simulation/slurm-20927698_1.out
Ignored: code/simulation/slurm-20927698_10.out
Ignored: code/simulation/slurm-20927698_100.out
Ignored: code/simulation/slurm-20927698_101.out
Ignored: code/simulation/slurm-20927698_102.out
Ignored: code/simulation/slurm-20927698_103.out
Ignored: code/simulation/slurm-20927698_104.out
Ignored: code/simulation/slurm-20927698_105.out
Ignored: code/simulation/slurm-20927698_106.out
Ignored: code/simulation/slurm-20927698_107.out
Ignored: code/simulation/slurm-20927698_108.out
Ignored: code/simulation/slurm-20927698_109.out
Ignored: code/simulation/slurm-20927698_11.out
Ignored: code/simulation/slurm-20927698_110.out
Ignored: code/simulation/slurm-20927698_111.out
Ignored: code/simulation/slurm-20927698_112.out
Ignored: code/simulation/slurm-20927698_113.out
Ignored: code/simulation/slurm-20927698_114.out
Ignored: code/simulation/slurm-20927698_115.out
Ignored: code/simulation/slurm-20927698_116.out
Ignored: code/simulation/slurm-20927698_117.out
Ignored: code/simulation/slurm-20927698_118.out
Ignored: code/simulation/slurm-20927698_119.out
Ignored: code/simulation/slurm-20927698_12.out
Ignored: code/simulation/slurm-20927698_120.out
Ignored: code/simulation/slurm-20927698_13.out
Ignored: code/simulation/slurm-20927698_14.out
Ignored: code/simulation/slurm-20927698_15.out
Ignored: code/simulation/slurm-20927698_16.out
Ignored: code/simulation/slurm-20927698_17.out
Ignored: code/simulation/slurm-20927698_18.out
Ignored: code/simulation/slurm-20927698_19.out
Ignored: code/simulation/slurm-20927698_2.out
Ignored: code/simulation/slurm-20927698_20.out
Ignored: code/simulation/slurm-20927698_21.out
Ignored: code/simulation/slurm-20927698_22.out
Ignored: code/simulation/slurm-20927698_23.out
Ignored: code/simulation/slurm-20927698_24.out
Ignored: code/simulation/slurm-20927698_25.out
Ignored: code/simulation/slurm-20927698_26.out
Ignored: code/simulation/slurm-20927698_27.out
Ignored: code/simulation/slurm-20927698_28.out
Ignored: code/simulation/slurm-20927698_29.out
Ignored: code/simulation/slurm-20927698_3.out
Ignored: code/simulation/slurm-20927698_30.out
Ignored: code/simulation/slurm-20927698_31.out
Ignored: code/simulation/slurm-20927698_32.out
Ignored: code/simulation/slurm-20927698_33.out
Ignored: code/simulation/slurm-20927698_34.out
Ignored: code/simulation/slurm-20927698_35.out
Ignored: code/simulation/slurm-20927698_36.out
Ignored: code/simulation/slurm-20927698_37.out
Ignored: code/simulation/slurm-20927698_38.out
Ignored: code/simulation/slurm-20927698_39.out
Ignored: code/simulation/slurm-20927698_4.out
Ignored: code/simulation/slurm-20927698_40.out
Ignored: code/simulation/slurm-20927698_41.out
Ignored: code/simulation/slurm-20927698_42.out
Ignored: code/simulation/slurm-20927698_43.out
Ignored: code/simulation/slurm-20927698_44.out
Ignored: code/simulation/slurm-20927698_45.out
Ignored: code/simulation/slurm-20927698_46.out
Ignored: code/simulation/slurm-20927698_47.out
Ignored: code/simulation/slurm-20927698_48.out
Ignored: code/simulation/slurm-20927698_49.out
Ignored: code/simulation/slurm-20927698_5.out
Ignored: code/simulation/slurm-20927698_50.out
Ignored: code/simulation/slurm-20927698_51.out
Ignored: code/simulation/slurm-20927698_52.out
Ignored: code/simulation/slurm-20927698_53.out
Ignored: code/simulation/slurm-20927698_54.out
Ignored: code/simulation/slurm-20927698_55.out
Ignored: code/simulation/slurm-20927698_56.out
Ignored: code/simulation/slurm-20927698_57.out
Ignored: code/simulation/slurm-20927698_58.out
Ignored: code/simulation/slurm-20927698_59.out
Ignored: code/simulation/slurm-20927698_6.out
Ignored: code/simulation/slurm-20927698_60.out
Ignored: code/simulation/slurm-20927698_61.out
Ignored: code/simulation/slurm-20927698_62.out
Ignored: code/simulation/slurm-20927698_63.out
Ignored: code/simulation/slurm-20927698_64.out
Ignored: code/simulation/slurm-20927698_65.out
Ignored: code/simulation/slurm-20927698_66.out
Ignored: code/simulation/slurm-20927698_67.out
Ignored: code/simulation/slurm-20927698_68.out
Ignored: code/simulation/slurm-20927698_69.out
Ignored: code/simulation/slurm-20927698_7.out
Ignored: code/simulation/slurm-20927698_70.out
Ignored: code/simulation/slurm-20927698_71.out
Ignored: code/simulation/slurm-20927698_72.out
Ignored: code/simulation/slurm-20927698_73.out
Ignored: code/simulation/slurm-20927698_74.out
Ignored: code/simulation/slurm-20927698_75.out
Ignored: code/simulation/slurm-20927698_76.out
Ignored: code/simulation/slurm-20927698_77.out
Ignored: code/simulation/slurm-20927698_78.out
Ignored: code/simulation/slurm-20927698_79.out
Ignored: code/simulation/slurm-20927698_8.out
Ignored: code/simulation/slurm-20927698_80.out
Ignored: code/simulation/slurm-20927698_81.out
Ignored: code/simulation/slurm-20927698_82.out
Ignored: code/simulation/slurm-20927698_83.out
Ignored: code/simulation/slurm-20927698_84.out
Ignored: code/simulation/slurm-20927698_85.out
Ignored: code/simulation/slurm-20927698_86.out
Ignored: code/simulation/slurm-20927698_87.out
Ignored: code/simulation/slurm-20927698_88.out
Ignored: code/simulation/slurm-20927698_89.out
Ignored: code/simulation/slurm-20927698_9.out
Ignored: code/simulation/slurm-20927698_90.out
Ignored: code/simulation/slurm-20927698_91.out
Ignored: code/simulation/slurm-20927698_92.out
Ignored: code/simulation/slurm-20927698_93.out
Ignored: code/simulation/slurm-20927698_94.out
Ignored: code/simulation/slurm-20927698_95.out
Ignored: code/simulation/slurm-20927698_96.out
Ignored: code/simulation/slurm-20927698_97.out
Ignored: code/simulation/slurm-20927698_98.out
Ignored: code/simulation/slurm-20927698_99.out
Ignored: code/simulation/slurm-20927938.out
Ignored: code/simulation/summarize.Rout
Ignored: data/annotation/mm39/GRCm39.genome.fa.gz
Ignored: data/annotation/mm39/gencode.vM35.annotation.gtf.gz
Ignored: data/annotation/mm39/gencode.vM35.metadata.EntrezGene.gz
Ignored: data/annotation/mm39/gencode.vM35.transcripts.fa.gz
Ignored: data/mouse/fastq/
Ignored: ignore/
Ignored: output/mouse/
Ignored: output/simulation-lenient/
Ignored: output/simulation-no-trend/
Ignored: output/simulation/
Ignored: renv/
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (analysis/mouse.Rmd) and HTML
(docs/mouse.html) files. If you’ve configured a remote Git
repository (see ?wflow_git_remote), click on the hyperlinks
in the table below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 76ed87c | Pedro Baldoni | 2025-03-18 | Adding analysis reports |
Introduction
Setup
knitr::opts_chunk$set(dev = "png",
dpi = 300,
dev.args = list(type = "cairo-png"),
root.dir = '.',
autodep = TRUE)
options(knitr.kable.NA = "-")library(edgeR)
library(rtracklayer)
library(Rsubread)
library(GenomicRanges)
library(GenomicFeatures)
library(data.table)
library(Gviz)
library(TxDb.Mmusculus.UCSC.mm39.refGene)
library(csaw)
library(BiocParallel)
library(ggplot2)
library(stringr)
library(png)
library(patchwork)path.misc <- '../misc/mouse.Rmd'
path.quant <- list.dirs('../output/mouse/salmon',recursive = FALSE)
path.target <- '../data/mouse/misc/targets.txt'
path.entrez <- "../data/annotation/mm39/gencode.vM35.metadata.EntrezGene.gz"
path.fasta <- '../data/annotation/mm39/gencode.vM35.transcripts.fa.gz'
path.gtf <- '../data/annotation/mm39/gencode.vM35.annotation.gtf.gz'
path.bam <- "../output/mouse/subread/alignment"
path.bw <- "../output/mouse/subread/coverage/"
dir.create(path.misc,recursive = TRUE,showWarnings = FALSE)bs <- 8plotSplice2 <- function(fit, coef=ncol(fit), geneid=NULL, genecolname=NULL, rank=1L, FDR=0.05,cex.axis = 1,exonlabel = NULL,fontsize = 8,...)
# This function has been adapted from limma::plotSplice so that axis labels
# uses "transcript" and not "exon"
{
if(is.null(genecolname))
genecolname <- fit$genecolname
else
genecolname <- as.character(genecolname)
if(is.null(geneid)) {
# Find gene from specified rank
if(rank==1L)
i <- which.min(fit$gene.F.p.value[,coef])
else
i <- order(fit$gene.F.p.value[,coef])[rank]
geneid <- paste(fit$gene.genes[i,genecolname], collapse=".")
} else {
# Find gene from specified name
geneid <- as.character(geneid)
i <- which(fit$gene.genes[,genecolname]==geneid)[1]
if(!length(i)) stop(paste("geneid",geneid,"not found"))
}
# Row numbers containing exons
j <- fit$gene.firstexon[i]:fit$gene.lastexon[i]
exoncolname <- fit$exoncolname
if(!is.null(exonlabel)) exoncolname <- exonlabel
# Get strand if possible
strcol <- grepl("strand", colnames(fit$gene.genes), ignore.case=TRUE)
if(any(strcol)) geneid <- paste0(geneid, " (", as.character(fit$gene.genes[i, strcol])[1], ")")
if(is.null(exoncolname)) {
plot(fit$coefficients[j, coef], xlab="Exon", ylab="logFC (this transcript vs rest)", main=geneid, type="b")
} else {
exon.id <- fit$genes[j, exoncolname]
# xlab <- paste("Exon", exoncolname, sep=" ")
plot(fit$coefficients[j, coef], xlab="", ylab="logFC (this transcript vs rest)", main=geneid, type="b", xaxt="n",cex.lab = fontsize/12,cex.main = fontsize/12,cex.axis=fontsize/12,...)
axis(1, at=1:length(j), labels=exon.id, las=2, cex.axis=fontsize/12)
# mtext(xlab, side=1, padj=5.2)
}
# Mark the topSpliced exons
top <- topSplice(fit, coef=coef, number=Inf, test="t", sort.by="none")
m <- which(top[, genecolname] %in% fit$gene.genes[i, genecolname])
fdr <- top$FDR[m]
sig <- fdr < FDR
if(any(sig)){
fdr.sig <- fdr[sig]
if(length(unique(fdr.sig))==1)
cex <- 1.5
else {
abs.fdr <- abs(log10(fdr.sig))
from <- range(abs.fdr)
to <- c(1,2)
cex <- (abs.fdr - from[1])/diff(from) * diff(to) + to[1]
}
points((1:length(j))[sig], fit$coefficients[j[sig], coef], col="red", pch=16, cex=cex)
}
abline(h=0,lty=2)
invisible()
}
plot.barplot <- function(x,top = 10,fontsize = 8){
txt <- x$Term[1:top]
txt <- str_to_sentence(txt)
pval <- x$P.DE[1:top]
df <- data.frame(Term = txt,Value = -log10(pval))
df$Term <- factor(df$Term,levels = txt[order(pval,decreasing = TRUE)])
ggplot(data = df,aes(y = Term,x = Value)) +
geom_col(fill = 'gray',col = 'black') +
theme_bw(base_size = fontsize,base_family = 'sans') +
theme(panel.grid = element_blank(),
axis.text = element_text(colour = 'black',size = fontsize),
axis.title = element_text(colour = 'black',size = fontsize)) +
labs(x = "-log10(P-value)",y = NULL)
}
getGeneRanges <- function(entrezid,type = 'gene',flank = TRUE){
out <-
genes(TxDb.Mmusculus.UCSC.mm39.refGene, filter = list(gene_id = entrezid))
if (type == 'promoter') {
out <- promoters(out)
}
if(flank == TRUE & type == 'gene'){
flank <- ceiling(0.25*width(out))
out <- GRanges(seqnames = as.character(seqnames(out)),IRanges(start = start(out)-flank,end = end(out) + flank))
}
return(out)
}
getCoverage <- function(bam, lib.sizes, gr, param, labels = NULL, group.by = NULL, ...) {
reads <- bplapply(seq_along(bam),function(i){
reads <- extractReads(bam.file = bam[i], gr, param = param)
},...)
if (!is.null(group.by)) {
groups <- unique(group.by)
reads <- bplapply(seq_along(groups),function(i){
subreads <- reads[group.by == groups[i]]
do.call('c',subreads)
},...)
names(reads) <- groups
lib.sizes <- bplapply(seq_along(groups),function(i){
sum(lib.sizes[group.by == groups[i]])
},...)
lib.sizes <- unlist(lib.sizes)
} else{
if (is.null(labels)) {
names(reads) <- bam
} else{
names(reads) <- labels
}
}
cov <- bplapply(seq_along(reads),function(i){
as(coverage(reads[[i]])/(lib.sizes[i]/1e6), "GRanges")
},...)
names(cov) <- names(reads)
return(cov)
}
getTrack <- function(cov, fill = NULL, ylim = NULL,yTicksAt = NULL,isBAM = NULL, ...) {
if (is.null(fill)) {
fill <- rep('gray',length(cov))
}
if (is.null(ylim) & isBAM) {
max.y <- max(unlist(lapply(cov,function(i){max(i$score)})))
ylim <- c(0,round(max.y + 5, -1))
}
out <- bplapply(seq_along(cov),function(i){
DataTrack(cov[[i]],
type = "histogram",
baseline = 0,
col.baseline = 'black',
lwd.baseline = 0.25,
ylim = ylim,
yTicksAt = yTicksAt,
name = names(cov[i]),
fill = fill[i],
col.title = 'black',
background.title = "white",
col.axis = 'black',
col.histogram = NA)
},...)
return(out)
}
plotCoverage <- function(x,lib.sizes,param,anno,fontsize = 8,isBAM = TRUE,
gr = NULL,entrezid = NULL,fill = NULL, labels = NULL, group.by = NULL, flank = TRUE, main = NULL,ylim = NULL, yTicksAt = NULL, ...){
if(!is.null(entrezid)){
gr <- getGeneRanges(entrezid,flank = TRUE)
}
if(isBAM){
cov <- getCoverage(bam = x,lib.sizes = lib.sizes,gr = gr,param = param,group.by = group.by,labels = labels,...)
} else{
cov <- x
names(cov) <- labels
}
if (!is.null(fill) & !is.null(group.by)) {
groups <- unique(group.by)
fill <- lapply(seq_along(groups),function(i){
subfill <- unique(fill[group.by == groups[i]])
if(length(subfill) == 1){
return(subfill)
} else{
stop('fill argument must be unique within groups')
}
})
fill <- unlist(fill)
}
track <- getTrack(cov = cov,fill = fill,ylim = ylim,yTicksAt = yTicksAt,isBAM = isBAM)
itrack <- IdeogramTrack(genome = 'mm39',
chromosome = as.character(seqnames(gr)),
fontcolor = 'black',
cex = fontsize/12)
axisTrack <- GenomeAxisTrack(cex = fontsize/12)
plotTracks(c(itrack,track, anno),
chromosome = as.character(seqnames(gr)),
cex.axis = fontsize/12,
cex.title = fontsize/12,
cex.group = fontsize/12,
sizes = c(0.1,rep(0.4,9), 0.75),
from = start(gr),
to = end(gr))
}
plot.voom <- function(fit,fontsize = 8,...){
plot(x = fit$voom.xy$x,
y = fit$voom.xy$y,
xlab = fit$voom.xy$xlab,
ylab = fit$voom.xy$ylab,
pch = fit$voom.xy$pch,
cex = fit$voom.xy$cex,
cex.lab = fontsize/12,
cex.axis = fontsize/12,
cex.main = fontsize/12,...)
lines(fit$voom.line,col="red",lty=1)
}Data loading
targets <- read.delim(path.target,sep = ';')
targets$pool <- factor(targets$pool,levels = c('1','2','3'))cs <- catchSalmon(path.quant)Reading ../output/mouse/salmon/GSM7106766, 147556 transcripts, 100 gibbs samples
Reading ../output/mouse/salmon/GSM7106767, 147556 transcripts, 100 gibbs samples
Reading ../output/mouse/salmon/GSM7106768, 147556 transcripts, 100 gibbs samples
Reading ../output/mouse/salmon/GSM7106769, 147556 transcripts, 100 gibbs samples
Reading ../output/mouse/salmon/GSM7106770, 147556 transcripts, 100 gibbs samples
Reading ../output/mouse/salmon/GSM7106771, 147556 transcripts, 100 gibbs samples
Reading ../output/mouse/salmon/GSM7106772, 147556 transcripts, 100 gibbs samples
Reading ../output/mouse/salmon/GSM7106773, 147556 transcripts, 100 gibbs samples
Reading ../output/mouse/salmon/GSM7106774, 147556 transcripts, 100 gibbs samplestx.name <- strsplit2(rownames(cs$annotation),"\\|")
rownames(cs$annotation) <- rownames(cs$counts) <- tx.name[,1]
colnames(cs$counts) <- basename(colnames(cs$counts))entrez <- fread(path.entrez,col.names = c('TranscriptID','EntrezID'))
df.fa <- as.data.table(scanFasta(path.fasta))Warning: 2438 duplicate sequences were found in the input. Please check the summary table.TranscriptID.fa <- strsplit2(df.fa$TranscriptID,"\\|")[,1]
GeneID.fa <- strsplit2(df.fa$TranscriptID,"\\|")[,2]
TranscriptBioType.fa <- strsplit2(df.fa$TranscriptID,"\\|")[,8]
df.fa$TranscriptID <- TranscriptID.fa
df.fa$GeneID <- GeneID.fa
df.fa$TranscriptBioType <- TranscriptBioType.fa
df.fa[,allUnique := all(Unique),by = 'GeneID']
df.fa <- df.fa[order(GeneID,TranscriptID),]
gtf <- import(path.gtf)
gtf.tx <- gtf[gtf$type == 'transcript']
gtf.tx$EntrezID <- entrez$EntrezID[match(gtf.tx$transcript_id,entrez$TranscriptID)]
gtf.tx$keep.gene <- gtf.tx$gene_type %in% c('protein_coding','lncRNA') & #. (1)
as.character(seqnames(gtf.tx)) %in% paste0('chr',c(1:19,'X','Y')) & #. (2)
gtf.tx$gene_id %in% df.fa[allUnique == TRUE,unique(GeneID)] & #. (3)
!is.na(gtf.tx$EntrezID)
gtf.tx$keep.tx <- grepl("protein_coding|lncRNA",gtf.tx$transcript_type)Differential Transcript Usage
dge <- DGEList(counts = cs$counts/cs$annotation$Overdispersion,
samples = targets,
genes = cs$annotation)
dge.raw <- DGEList(counts = cs$counts,
samples = targets,
genes = cs$annotation)
dge$genes[,c('TranscriptID','GeneID')] <-
dge.raw$genes[,c('TranscriptID','GeneID')] <-
df.fa[match(rownames(dge),df.fa$TranscriptID),c('TranscriptID','GeneID')]
dge$genes[,c('TranscriptSymbol','GeneSymbol','EntrezID')] <-
dge.raw$genes[,c('TranscriptSymbol','GeneSymbol','EntrezID')] <-
mcols(gtf.tx)[match(rownames(dge),gtf.tx$transcript_id),c('transcript_name','gene_name','EntrezID')] |>
as.data.frame()keep <- filterByExpr(dge)
keep.gene <- gtf.tx$keep.gene[match(rownames(dge),gtf.tx$transcript_id)]
keep.tx <- gtf.tx$keep.tx[match(rownames(dge),gtf.tx$transcript_id)]
dge.filtr <- dge[keep & keep.gene & keep.tx,,keep.lib.sizes = FALSE]
dge.filtr <- normLibSizes(dge.filtr)
dge.raw.filtr <- dge.raw[keep & keep.gene & keep.tx,,keep.lib.sizes = FALSE]
dge.raw.filtr <- normLibSizes(dge.raw.filtr)par(mfrow = c(2,2))
y <- cpm(dge.filtr,log = TRUE)
plotMDS(y,col = dge.filtr$samples$color,
xlim = c(-6,3),ylim = c(-4,3))
plotMDS(y,labels = dge.filtr$samples$pool,col = dge.filtr$samples$color,
xlim = c(-6,3),ylim = c(-4,3))
y.nobatch <-
removeBatchEffect(x = y,
batch = dge.filtr$samples$pool,
group = dge.filtr$samples$group)
plotMDS(y.nobatch,col = dge.filtr$samples$color,
xlim = c(-6,3),ylim = c(-4,3))
plotMDS(y.nobatch,labels = dge.filtr$samples$pool,col = dge.filtr$samples$color,
xlim = c(-6,3),ylim = c(-4,3))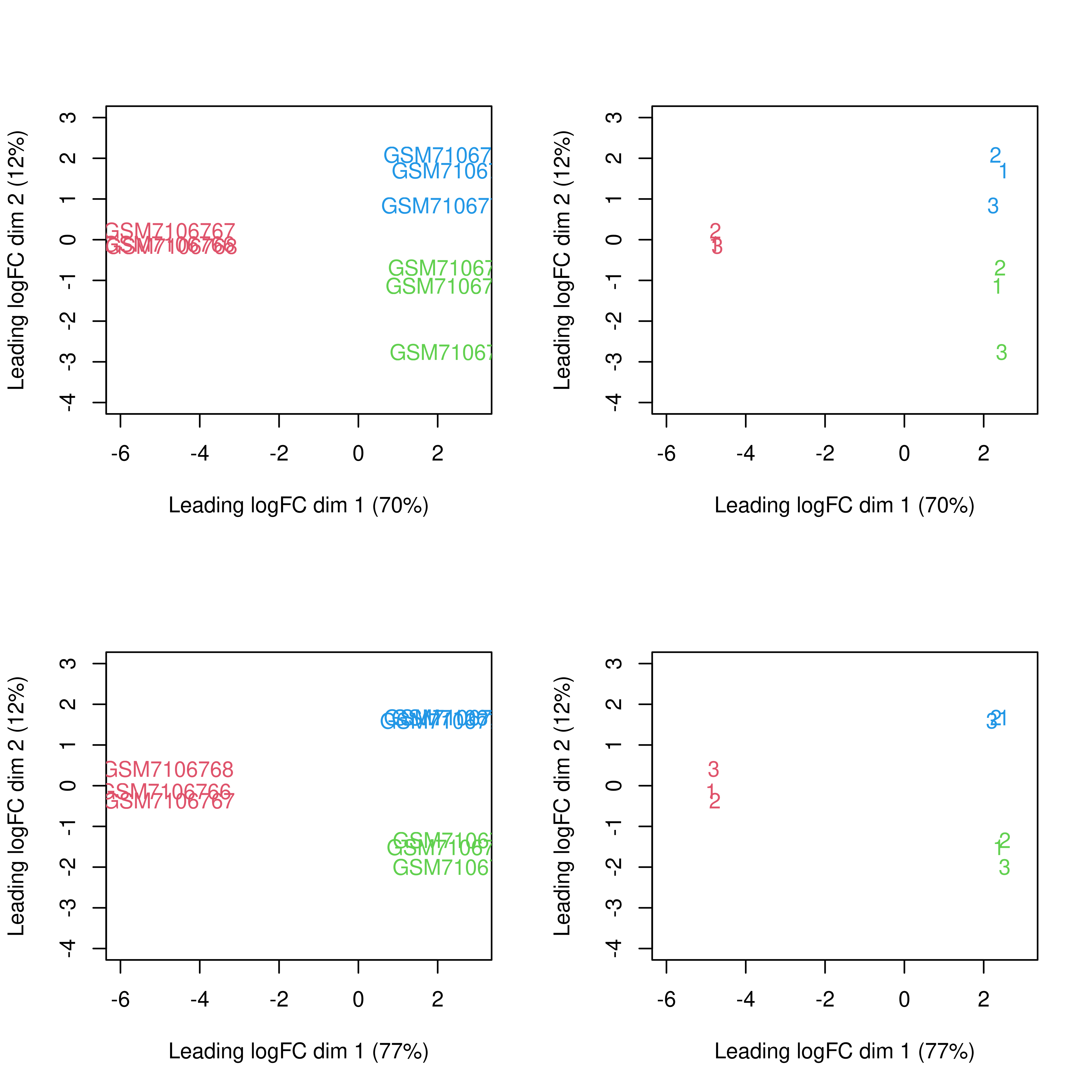
dev.off()null device
1 design <- model.matrix(~0+group,data = dge.filtr$samples)
colnames(design) <- sub("group","",colnames(design))
contr <- makeContrasts(LPvsBa = LP - Basal,
MLvsBa = ML - Basal,
MLvsLP = ML - LP,
LvsBa = 0.5*(LP+ML) - Basal,
levels = design)voom
v.raw <- voomLmFit(counts = dge.raw.filtr,
design = design,
block = dge.raw.filtr$samples$pool,
plot = TRUE,save.plot = TRUE)First intra-block correlation 0.1476588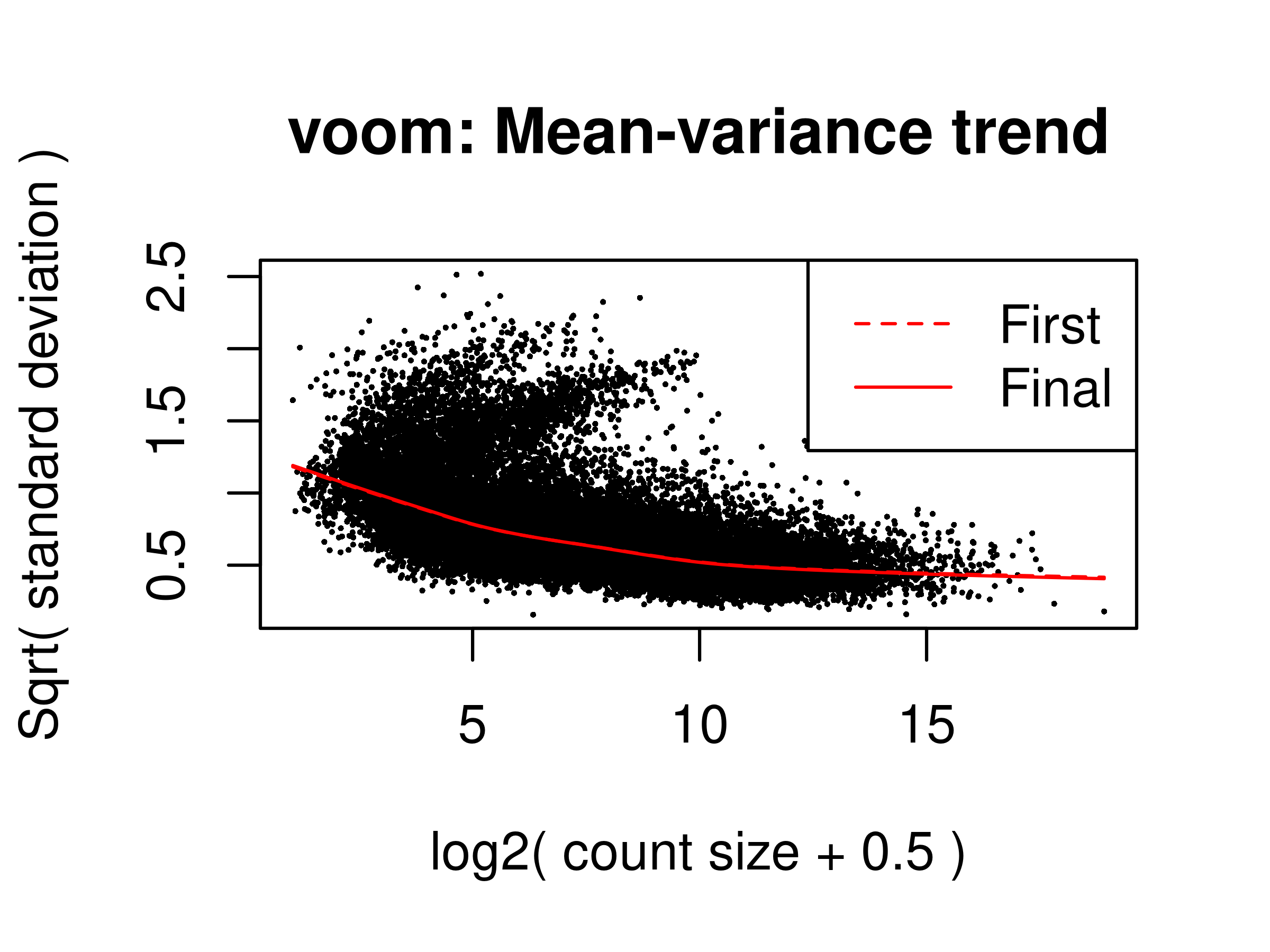
Final intra-block correlation 0.1477514v <- voomLmFit(counts = dge.filtr,
design = design,
block = dge.filtr$samples$pool,
plot = TRUE,save.plot = TRUE)First intra-block correlation 0.1501895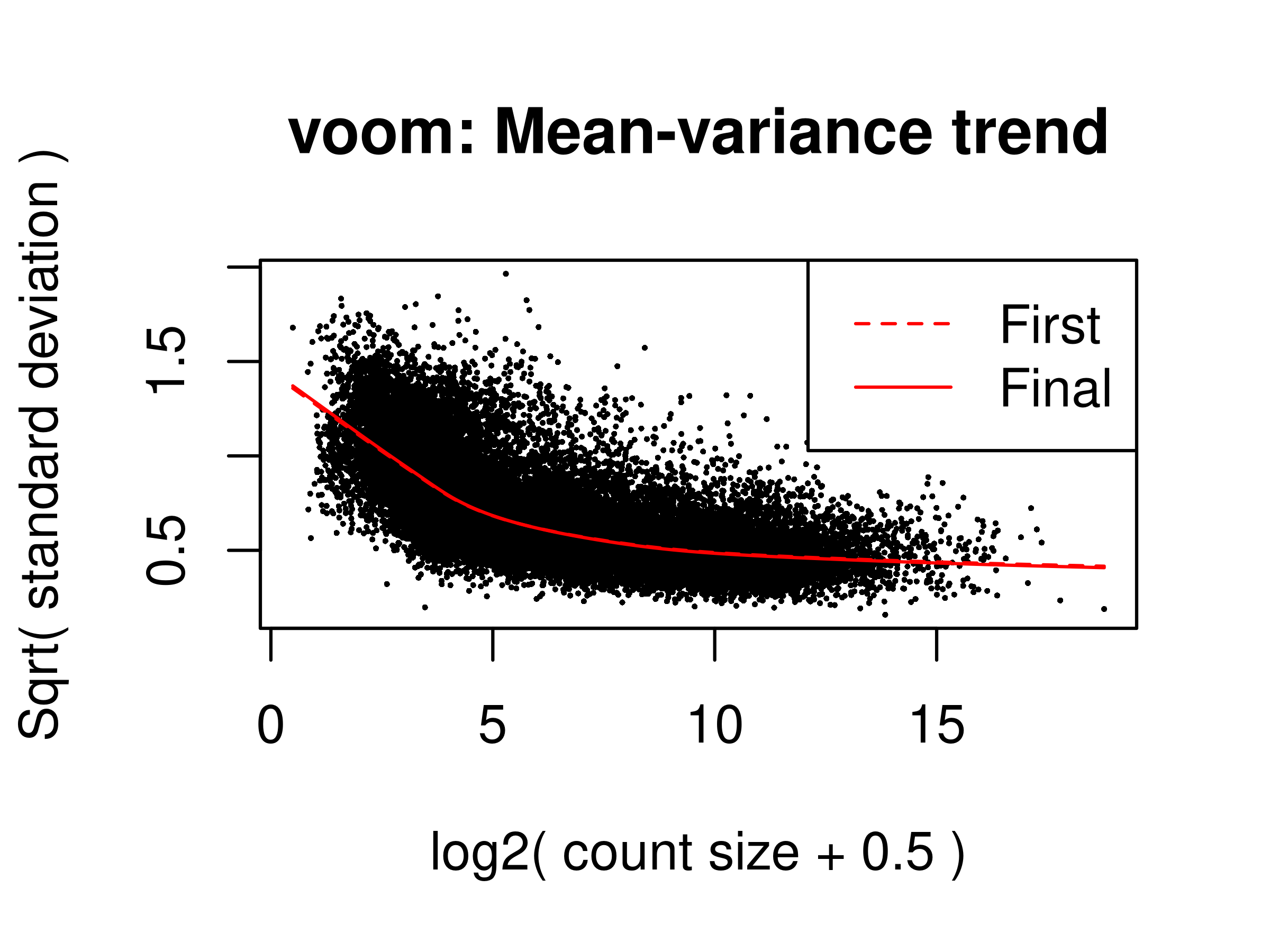
Final intra-block correlation 0.1502238fit.raw <- contrasts.fit(fit = v.raw,contrasts = contr)
fit <- contrasts.fit(fit = v,contrasts = contr)diffSplice
ds.raw <- diffSplice(fit.raw, geneid = "GeneID", exonid = "TranscriptID",robust = TRUE)Total number of exons: 29954
Total number of genes: 14632
Number of genes with 1 exon: 7067
Mean number of exons in a gene: 2
Max number of exons in a gene: 17 ds <- diffSplice(fit, geneid = "GeneID", exonid = "TranscriptID",robust = TRUE)Total number of exons: 29954
Total number of genes: 14632
Number of genes with 1 exon: 7067
Mean number of exons in a gene: 2
Max number of exons in a gene: 17 out.transcript.LPvsBa <- topSplice(ds, test = "t",number = Inf,coef = "LPvsBa")
out.gene.LPvsBa <- topSplice(ds, test = "F", number = Inf,coef = "LPvsBa")
out.transcript.MLvsBa <- topSplice(ds, test = "t",number = Inf,coef = "MLvsBa")
out.gene.MLvsBa <- topSplice(ds, test = "F", number = Inf,coef = "MLvsBa")
out.transcript.MLvsLP <- topSplice(ds, test = "t",number = Inf,coef = "MLvsLP")
out.gene.MLvsLP <- topSplice(ds, test = "F", number = Inf,coef = "MLvsLP")
out.transcript.LvsBa <- topSplice(ds, test = "t",number = Inf,coef = "LvsBa")
out.gene.LvsBa <- topSplice(ds, test = "F", number = Inf,coef = "LvsBa")summary(ds.raw$gene.df.prior) Min. 1st Qu. Median Mean 3rd Qu. Max.
3.445 3.445 3.445 3.445 3.445 3.445 summary(ds$gene.df.prior) Min. 1st Qu. Median Mean 3rd Qu. Max.
5.524 5.524 5.524 5.524 5.524 5.524 head(out.gene.LPvsBa,20) GeneID GeneSymbol EntrezID NExons F
ENSMUSG00000022791.20 ENSMUSG00000022791.20 Tnk2 51789 11 51.20126
ENSMUSG00000032366.16 ENSMUSG00000032366.16 Tpm1 22003 15 30.00696
ENSMUSG00000025290.18 ENSMUSG00000025290.18 Rps24 20088 9 60.87343
ENSMUSG00000026131.22 ENSMUSG00000026131.22 Dst 13518 12 19.40701
ENSMUSG00000028649.23 ENSMUSG00000028649.23 Macf1 11426 11 21.42040
ENSMUSG00000005417.18 ENSMUSG00000005417.18 Mprip 26936 8 40.43212
ENSMUSG00000026987.17 ENSMUSG00000026987.17 Baz2b 407823 11 19.50041
ENSMUSG00000022587.16 ENSMUSG00000022587.16 Ly6e 17069 10 22.12727
ENSMUSG00000035325.17 ENSMUSG00000035325.17 Sec31a 69162 7 36.49451
ENSMUSG00000021870.18 ENSMUSG00000021870.18 Slmap 83997 12 16.15625
ENSMUSG00000038143.17 ENSMUSG00000038143.17 Stox2 71069 4 95.12136
ENSMUSG00000027455.17 ENSMUSG00000027455.17 Nsfl1c 386649 4 89.52181
ENSMUSG00000017897.19 ENSMUSG00000017897.19 Eya2 14049 4 88.37603
ENSMUSG00000069601.15 ENSMUSG00000069601.15 Ank3 11735 14 12.44064
ENSMUSG00000027381.18 ENSMUSG00000027381.18 Bcl2l11 12125 7 29.38600
ENSMUSG00000025195.10 ENSMUSG00000025195.10 Dnmbp 71972 4 81.54684
ENSMUSG00000000631.21 ENSMUSG00000000631.21 Myo18a 360013 7 29.37584
ENSMUSG00000048200.13 ENSMUSG00000048200.13 Cracr2b 213573 5 53.27515
ENSMUSG00000024048.16 ENSMUSG00000024048.16 Myl12a 67268 7 27.93278
ENSMUSG00000020758.16 ENSMUSG00000020758.16 Itgb4 192897 7 35.14066
P.Value FDR
ENSMUSG00000022791.20 4.361309e-27 1.731290e-23
ENSMUSG00000032366.16 4.577104e-27 1.731290e-23
ENSMUSG00000025290.18 8.395822e-26 2.117146e-22
ENSMUSG00000026131.22 8.553399e-18 1.363001e-14
ENSMUSG00000028649.23 9.008598e-18 1.363001e-14
ENSMUSG00000005417.18 1.815237e-17 2.288711e-14
ENSMUSG00000026987.17 9.869717e-17 1.066634e-13
ENSMUSG00000022587.16 1.141236e-16 1.079181e-13
ENSMUSG00000035325.17 1.992673e-15 1.674953e-12
ENSMUSG00000021870.18 2.436839e-15 1.843468e-12
ENSMUSG00000038143.17 2.836790e-15 1.950938e-12
ENSMUSG00000027455.17 6.355105e-15 4.006364e-12
ENSMUSG00000017897.19 7.537372e-15 4.386171e-12
ENSMUSG00000069601.15 9.664430e-15 5.222244e-12
ENSMUSG00000027381.18 2.058227e-14 1.028598e-11
ENSMUSG00000025195.10 2.175488e-14 1.028598e-11
ENSMUSG00000000631.21 4.222361e-14 1.878951e-11
ENSMUSG00000048200.13 4.598223e-14 1.932531e-11
ENSMUSG00000024048.16 5.181649e-14 2.063115e-11
ENSMUSG00000020758.16 5.998492e-14 2.204800e-11head(out.gene.MLvsBa,20) GeneID GeneSymbol EntrezID NExons
ENSMUSG00000032366.16 ENSMUSG00000032366.16 Tpm1 22003 15
ENSMUSG00000025290.18 ENSMUSG00000025290.18 Rps24 20088 9
ENSMUSG00000022791.20 ENSMUSG00000022791.20 Tnk2 51789 11
ENSMUSG00000022587.16 ENSMUSG00000022587.16 Ly6e 17069 10
ENSMUSG00000026987.17 ENSMUSG00000026987.17 Baz2b 407823 11
ENSMUSG00000028649.23 ENSMUSG00000028649.23 Macf1 11426 11
ENSMUSG00000005417.18 ENSMUSG00000005417.18 Mprip 26936 8
ENSMUSG00000002985.17 ENSMUSG00000002985.17 Apoe 11816 6
ENSMUSG00000016487.16 ENSMUSG00000016487.16 Ppfibp1 67533 5
ENSMUSG00000069601.15 ENSMUSG00000069601.15 Ank3 11735 14
ENSMUSG00000026131.22 ENSMUSG00000026131.22 Dst 13518 12
ENSMUSG00000021870.18 ENSMUSG00000021870.18 Slmap 83997 12
ENSMUSG00000025195.10 ENSMUSG00000025195.10 Dnmbp 71972 4
ENSMUSG00000027455.17 ENSMUSG00000027455.17 Nsfl1c 386649 4
ENSMUSG00000034006.18 ENSMUSG00000034006.18 Slc66a2 66943 10
ENSMUSG00000017897.19 ENSMUSG00000017897.19 Eya2 14049 4
ENSMUSG00000020758.16 ENSMUSG00000020758.16 Itgb4 192897 7
ENSMUSG00000004043.15 ENSMUSG00000004043.15 Stat5a 20850 6
ENSMUSG00000035325.17 ENSMUSG00000035325.17 Sec31a 69162 7
ENSMUSG00000000631.21 ENSMUSG00000000631.21 Myo18a 360013 7
F P.Value FDR
ENSMUSG00000032366.16 45.20079 1.939648e-33 1.467344e-29
ENSMUSG00000025290.18 54.00634 1.888609e-24 7.143665e-21
ENSMUSG00000022791.20 30.99385 4.434990e-21 1.118357e-17
ENSMUSG00000022587.16 32.79022 5.998107e-21 1.134392e-17
ENSMUSG00000026987.17 26.73448 2.459973e-20 3.721939e-17
ENSMUSG00000028649.23 23.80758 5.636097e-19 7.106179e-16
ENSMUSG00000005417.18 46.60902 1.137337e-18 1.229136e-15
ENSMUSG00000002985.17 60.99870 4.792633e-18 4.532034e-15
ENSMUSG00000016487.16 86.24402 8.734216e-18 7.341594e-15
ENSMUSG00000069601.15 15.92528 1.102257e-17 8.338576e-15
ENSMUSG00000026131.22 19.02023 1.463452e-17 1.006456e-14
ENSMUSG00000021870.18 18.80178 5.613067e-17 3.538571e-14
ENSMUSG00000025195.10 123.28807 8.574236e-17 4.989546e-14
ENSMUSG00000027455.17 111.61042 3.312830e-16 1.790111e-13
ENSMUSG00000034006.18 21.02947 3.801508e-16 1.917227e-13
ENSMUSG00000017897.19 108.21631 5.028060e-16 2.377330e-13
ENSMUSG00000020758.16 44.17757 1.565734e-15 6.967517e-13
ENSMUSG00000004043.15 43.00015 2.300481e-15 9.668410e-13
ENSMUSG00000035325.17 34.81708 4.596194e-15 1.830011e-12
ENSMUSG00000000631.21 32.60327 6.530347e-15 2.470104e-12head(out.gene.MLvsLP,20) GeneID GeneSymbol EntrezID NExons
ENSMUSG00000045636.17 ENSMUSG00000045636.17 Mtus1 102103 6
ENSMUSG00000022791.20 ENSMUSG00000022791.20 Tnk2 51789 11
ENSMUSG00000001211.16 ENSMUSG00000001211.16 Agpat3 28169 5
ENSMUSG00000028337.15 ENSMUSG00000028337.15 Coro2a 107684 2
ENSMUSG00000021000.18 ENSMUSG00000021000.18 Mia2 338320 8
ENSMUSG00000026131.22 ENSMUSG00000026131.22 Dst 13518 12
ENSMUSG00000004043.15 ENSMUSG00000004043.15 Stat5a 20850 6
ENSMUSG00000002985.17 ENSMUSG00000002985.17 Apoe 11816 6
ENSMUSG00000038007.15 ENSMUSG00000038007.15 Acer2 230379 3
ENSMUSG00000023017.11 ENSMUSG00000023017.11 Asic1 11419 2
ENSMUSG00000040479.13 ENSMUSG00000040479.13 Dgkz 104418 4
ENSMUSG00000036792.13 ENSMUSG00000036792.13 Mbd5 109241 8
ENSMUSG00000024589.19 ENSMUSG00000024589.19 Nedd4l 83814 9
ENSMUSG00000026821.17 ENSMUSG00000026821.17 Ralgds 19730 6
ENSMUSG00000032000.18 ENSMUSG00000032000.18 Birc3 11796 5
ENSMUSG00000029381.17 ENSMUSG00000029381.17 Shroom3 27428 5
ENSMUSG00000023039.18 ENSMUSG00000023039.18 Krt7 110310 4
ENSMUSG00000024608.12 ENSMUSG00000024608.12 Rps14 20044 4
ENSMUSG00000090841.3 ENSMUSG00000090841.3 Myl6 17904 8
ENSMUSG00000028865.15 ENSMUSG00000028865.15 Cd164l2 69655 3
F P.Value FDR
ENSMUSG00000045636.17 21.478181 1.490869e-10 1.127842e-06
ENSMUSG00000022791.20 8.253745 1.720190e-08 6.506617e-05
ENSMUSG00000001211.16 17.314271 5.771458e-08 1.455369e-04
ENSMUSG00000028337.15 75.059913 9.558274e-08 1.807709e-04
ENSMUSG00000021000.18 9.377649 1.576217e-07 2.207502e-04
ENSMUSG00000026131.22 6.493020 1.750828e-07 2.207502e-04
ENSMUSG00000004043.15 12.095865 2.927025e-07 3.163278e-04
ENSMUSG00000002985.17 11.443989 5.565017e-07 5.262419e-04
ENSMUSG00000038007.15 24.513880 1.765148e-06 1.483705e-03
ENSMUSG00000023017.11 41.450248 5.292878e-06 4.004062e-03
ENSMUSG00000040479.13 14.129538 6.837863e-06 4.484068e-03
ENSMUSG00000036792.13 6.931654 7.112864e-06 4.484068e-03
ENSMUSG00000024589.19 6.156170 8.998482e-06 5.236424e-03
ENSMUSG00000026821.17 8.754089 9.841939e-06 5.318162e-03
ENSMUSG00000032000.18 9.752780 1.977821e-05 9.974809e-03
ENSMUSG00000029381.17 9.525990 2.433474e-05 1.150577e-02
ENSMUSG00000023039.18 11.919248 2.759647e-05 1.167563e-02
ENSMUSG00000024608.12 11.842872 2.902891e-05 1.167563e-02
ENSMUSG00000090841.3 6.055025 3.160153e-05 1.167563e-02
ENSMUSG00000028865.15 16.605212 3.184016e-05 1.167563e-02head(out.gene.LvsBa,20) GeneID GeneSymbol EntrezID NExons
ENSMUSG00000032366.16 ENSMUSG00000032366.16 Tpm1 22003 15
ENSMUSG00000025290.18 ENSMUSG00000025290.18 Rps24 20088 9
ENSMUSG00000022791.20 ENSMUSG00000022791.20 Tnk2 51789 11
ENSMUSG00000026987.17 ENSMUSG00000026987.17 Baz2b 407823 11
ENSMUSG00000028649.23 ENSMUSG00000028649.23 Macf1 11426 11
ENSMUSG00000022587.16 ENSMUSG00000022587.16 Ly6e 17069 10
ENSMUSG00000005417.18 ENSMUSG00000005417.18 Mprip 26936 8
ENSMUSG00000021870.18 ENSMUSG00000021870.18 Slmap 83997 12
ENSMUSG00000026131.22 ENSMUSG00000026131.22 Dst 13518 12
ENSMUSG00000016487.16 ENSMUSG00000016487.16 Ppfibp1 67533 5
ENSMUSG00000069601.15 ENSMUSG00000069601.15 Ank3 11735 14
ENSMUSG00000020758.16 ENSMUSG00000020758.16 Itgb4 192897 7
ENSMUSG00000034006.18 ENSMUSG00000034006.18 Slc66a2 66943 10
ENSMUSG00000027455.17 ENSMUSG00000027455.17 Nsfl1c 386649 4
ENSMUSG00000035325.17 ENSMUSG00000035325.17 Sec31a 69162 7
ENSMUSG00000017897.19 ENSMUSG00000017897.19 Eya2 14049 4
ENSMUSG00000025195.10 ENSMUSG00000025195.10 Dnmbp 71972 4
ENSMUSG00000002985.17 ENSMUSG00000002985.17 Apoe 11816 6
ENSMUSG00000038143.17 ENSMUSG00000038143.17 Stox2 71069 4
ENSMUSG00000021458.19 ENSMUSG00000021458.19 Aopep 72061 12
F P.Value FDR
ENSMUSG00000032366.16 50.17743 3.969028e-35 3.002570e-31
ENSMUSG00000025290.18 77.62153 1.317648e-28 4.984005e-25
ENSMUSG00000022791.20 51.56844 3.554692e-27 8.963749e-24
ENSMUSG00000026987.17 30.85375 4.540846e-22 8.587875e-19
ENSMUSG00000028649.23 30.53245 6.104804e-22 9.236569e-19
ENSMUSG00000022587.16 35.38603 8.027335e-22 1.012113e-18
ENSMUSG00000005417.18 61.33500 4.651845e-21 5.027315e-18
ENSMUSG00000021870.18 25.53692 1.644759e-20 1.555325e-17
ENSMUSG00000026131.22 22.89281 9.237884e-20 7.764955e-17
ENSMUSG00000016487.16 94.92351 1.867114e-18 1.391119e-15
ENSMUSG00000069601.15 16.88428 2.022777e-18 1.391119e-15
ENSMUSG00000020758.16 60.29860 9.167033e-18 5.779050e-15
ENSMUSG00000034006.18 23.48478 2.730988e-17 1.506684e-14
ENSMUSG00000027455.17 133.59545 2.859329e-17 1.506684e-14
ENSMUSG00000035325.17 45.96925 2.987477e-17 1.506684e-14
ENSMUSG00000017897.19 130.22635 4.057766e-17 1.918563e-14
ENSMUSG00000025195.10 129.56203 4.352040e-17 1.936658e-14
ENSMUSG00000002985.17 53.74945 4.644210e-17 1.951858e-14
ENSMUSG00000038143.17 112.48419 2.981042e-16 1.186925e-13
ENSMUSG00000021458.19 16.44419 4.155921e-16 1.571977e-13table(out.gene.LPvsBa$FDR < 0.05)
FALSE TRUE
6305 1260 out.gene.LPvsBa[out.gene.LPvsBa$GeneSymbol %in% c('Foxp1','Ezh2','Asap1'),] GeneID GeneSymbol EntrezID NExons
ENSMUSG00000030067.18 ENSMUSG00000030067.18 Foxp1 108655 11
ENSMUSG00000022377.17 ENSMUSG00000022377.17 Asap1 13196 4
ENSMUSG00000029687.17 ENSMUSG00000029687.17 Ezh2 14056 4
F P.Value FDR
ENSMUSG00000030067.18 8.903039 3.118056e-09 2.876597e-07
ENSMUSG00000022377.17 17.275861 1.146772e-06 4.689367e-05
ENSMUSG00000029687.17 6.129761 2.271890e-03 1.980052e-02head(out.transcript.LPvsBa,20) Length EffectiveLength Overdispersion
ENSMUST00000115126.9 4540 4219.417 9.377569
ENSMUST00000223999.2 525 234.370 5.287670
ENSMUST00000113696.8 1714 1393.417 6.058497
ENSMUST00000112384.10 468 194.337 6.131152
ENSMUST00000115120.9 2796 2475.417 7.562900
ENSMUST00000079195.6 10467 10146.417 7.172959
ENSMUST00000028949.16 1364 1043.417 6.052642
ENSMUST00000063433.8 2460 2139.417 4.602893
ENSMUST00000103079.4 3286 2965.417 5.194726
ENSMUST00000211882.2 9696 9375.417 3.724371
ENSMUST00000074582.7 2799 2478.417 6.911156
ENSMUST00000088132.13 2447 2126.417 5.137227
ENSMUST00000029676.12 2975 2654.417 6.743824
ENSMUST00000053670.12 1698 1377.417 1.164367
ENSMUST00000033407.13 2913 2592.417 4.063701
ENSMUST00000123765.8 2613 2292.417 3.107693
ENSMUST00000092887.11 7304 6983.417 18.089224
ENSMUST00000089140.13 1486 1165.417 6.677255
ENSMUST00000116247.3 689 373.675 2.008777
ENSMUST00000080933.13 5669 5348.417 20.189698
TranscriptID GeneID
ENSMUST00000115126.9 ENSMUST00000115126.9 ENSMUSG00000022791.20
ENSMUST00000223999.2 ENSMUST00000223999.2 ENSMUSG00000025290.18
ENSMUST00000113696.8 ENSMUST00000113696.8 ENSMUSG00000032366.16
ENSMUST00000112384.10 ENSMUST00000112384.10 ENSMUSG00000025290.18
ENSMUST00000115120.9 ENSMUST00000115120.9 ENSMUSG00000022791.20
ENSMUST00000079195.6 ENSMUST00000079195.6 ENSMUSG00000038143.17
ENSMUST00000028949.16 ENSMUST00000028949.16 ENSMUSG00000027455.17
ENSMUST00000063433.8 ENSMUST00000063433.8 ENSMUSG00000017897.19
ENSMUST00000103079.4 ENSMUST00000103079.4 ENSMUSG00000031078.16
ENSMUST00000211882.2 ENSMUST00000211882.2 ENSMUSG00000038143.17
ENSMUST00000074582.7 ENSMUST00000074582.7 ENSMUSG00000028041.18
ENSMUST00000088132.13 ENSMUST00000088132.13 ENSMUSG00000017897.19
ENSMUST00000029676.12 ENSMUST00000029676.12 ENSMUSG00000028041.18
ENSMUST00000053670.12 ENSMUST00000053670.12 ENSMUSG00000048200.13
ENSMUST00000033407.13 ENSMUST00000033407.13 ENSMUSG00000031078.16
ENSMUST00000123765.8 ENSMUST00000123765.8 ENSMUSG00000028649.23
ENSMUST00000092887.11 ENSMUST00000092887.11 ENSMUSG00000000631.21
ENSMUST00000089140.13 ENSMUST00000089140.13 ENSMUSG00000027455.17
ENSMUST00000116247.3 ENSMUST00000116247.3 ENSMUSG00000048200.13
ENSMUST00000080933.13 ENSMUST00000080933.13 ENSMUSG00000028284.14
TranscriptSymbol GeneSymbol EntrezID logFC t
ENSMUST00000115126.9 Tnk2-207 Tnk2 51789 -3.661632 -15.09738
ENSMUST00000223999.2 Rps24-205 Rps24 20088 -6.858530 -15.42431
ENSMUST00000113696.8 Tpm1-212 Tpm1 22003 7.377681 12.78083
ENSMUST00000112384.10 Rps24-201 Rps24 20088 5.793228 14.47835
ENSMUST00000115120.9 Tnk2-201 Tnk2 51789 3.586098 13.08662
ENSMUST00000079195.6 Stox2-201 Stox2 71069 -4.382013 -16.38977
ENSMUST00000028949.16 Nsfl1c-201 Nsfl1c 386649 4.997175 14.70325
ENSMUST00000063433.8 Eya2-201 Eya2 14049 6.848815 14.60487
ENSMUST00000103079.4 Cttn-202 Cttn 13043 -3.897749 -14.41250
ENSMUST00000211882.2 Stox2-209 Stox2 71069 3.430629 14.39119
ENSMUST00000074582.7 Adam15-202 Adam15 11490 -4.746046 -13.48766
ENSMUST00000088132.13 Eya2-202 Eya2 14049 -8.811995 -13.45151
ENSMUST00000029676.12 Adam15-201 Adam15 11490 4.766114 13.10433
ENSMUST00000053670.12 Cracr2b-201 Cracr2b 213573 3.521311 12.06175
ENSMUST00000033407.13 Cttn-201 Cttn 13043 3.408681 12.97064
ENSMUST00000123765.8 Macf1-204 Macf1 11426 2.279264 9.16542
ENSMUST00000092887.11 Myo18a-203 Myo18a 360013 10.490359 10.24146
ENSMUST00000089140.13 Nsfl1c-202 Nsfl1c 386649 -4.892618 -12.58070
ENSMUST00000116247.3 Cracr2b-202 Cracr2b 213573 -4.526939 -11.64256
ENSMUST00000080933.13 Map3k7-202 Map3k7 26409 -4.918089 -11.29403
P.Value FDR
ENSMUST00000115126.9 5.157769e-23 1.180459e-18
ENSMUST00000223999.2 1.840304e-22 2.105952e-18
ENSMUST00000113696.8 1.536767e-21 1.172400e-17
ENSMUST00000112384.10 3.648630e-21 2.087655e-17
ENSMUST00000115120.9 6.031321e-20 2.760777e-16
ENSMUST00000079195.6 2.288186e-16 8.728287e-13
ENSMUST00000028949.16 4.022080e-15 1.315048e-11
ENSMUST00000063433.8 4.791327e-15 1.370739e-11
ENSMUST00000103079.4 6.764080e-15 1.608709e-11
ENSMUST00000211882.2 7.028921e-15 1.608709e-11
ENSMUST00000074582.7 3.731859e-14 7.622186e-11
ENSMUST00000088132.13 3.996427e-14 7.622186e-11
ENSMUST00000029676.12 7.768998e-14 1.367762e-10
ENSMUST00000053670.12 9.629101e-14 1.536382e-10
ENSMUST00000033407.13 1.006935e-13 1.536382e-10
ENSMUST00000123765.8 1.089964e-13 1.559125e-10
ENSMUST00000092887.11 2.140131e-13 2.758112e-10
ENSMUST00000089140.13 2.169180e-13 2.758112e-10
ENSMUST00000116247.3 2.523942e-13 2.993133e-10
ENSMUST00000080933.13 2.615575e-13 2.993133e-10head(out.transcript.MLvsBa,20) Length EffectiveLength Overdispersion
ENSMUST00000223999.2 525 234.370 5.287670
ENSMUST00000113696.8 1714 1393.417 6.058497
ENSMUST00000113705.8 1759 1438.417 7.934639
ENSMUST00000112384.10 468 194.337 6.131152
ENSMUST00000153136.2 890 569.769 2.307391
ENSMUST00000111623.9 4782 4461.417 11.793842
ENSMUST00000115120.9 2796 2475.417 7.562900
ENSMUST00000063433.8 2460 2139.417 4.602893
ENSMUST00000028949.16 1364 1043.417 6.052642
ENSMUST00000212396.2 6100 5779.417 3.624965
ENSMUST00000016631.14 4851 4530.417 3.612108
ENSMUST00000152639.8 2539 2218.417 1.158605
ENSMUST00000191127.8 1818 1497.417 5.514662
ENSMUST00000115126.9 4540 4219.417 9.377569
ENSMUST00000088132.13 2447 2126.417 5.137227
ENSMUST00000108751.8 4021 3700.417 7.431071
ENSMUST00000089140.13 1486 1165.417 6.677255
ENSMUST00000080933.13 5669 5348.417 20.189698
ENSMUST00000113697.8 1742 1421.417 14.708784
ENSMUST00000105910.2 986 665.431 1.056655
TranscriptID GeneID
ENSMUST00000223999.2 ENSMUST00000223999.2 ENSMUSG00000025290.18
ENSMUST00000113696.8 ENSMUST00000113696.8 ENSMUSG00000032366.16
ENSMUST00000113705.8 ENSMUST00000113705.8 ENSMUSG00000032366.16
ENSMUST00000112384.10 ENSMUST00000112384.10 ENSMUSG00000025290.18
ENSMUST00000153136.2 ENSMUST00000153136.2 ENSMUSG00000026987.17
ENSMUST00000111623.9 ENSMUST00000111623.9 ENSMUSG00000016487.16
ENSMUST00000115120.9 ENSMUST00000115120.9 ENSMUSG00000022791.20
ENSMUST00000063433.8 ENSMUST00000063433.8 ENSMUSG00000017897.19
ENSMUST00000028949.16 ENSMUST00000028949.16 ENSMUSG00000027455.17
ENSMUST00000212396.2 ENSMUST00000212396.2 ENSMUSG00000025195.10
ENSMUST00000016631.14 ENSMUST00000016631.14 ENSMUSG00000016487.16
ENSMUST00000152639.8 ENSMUST00000152639.8 ENSMUSG00000026987.17
ENSMUST00000191127.8 ENSMUST00000191127.8 ENSMUSG00000022587.16
ENSMUST00000115126.9 ENSMUST00000115126.9 ENSMUSG00000022791.20
ENSMUST00000088132.13 ENSMUST00000088132.13 ENSMUSG00000017897.19
ENSMUST00000108751.8 ENSMUST00000108751.8 ENSMUSG00000005417.18
ENSMUST00000089140.13 ENSMUST00000089140.13 ENSMUSG00000027455.17
ENSMUST00000080933.13 ENSMUST00000080933.13 ENSMUSG00000028284.14
ENSMUST00000113697.8 ENSMUST00000113697.8 ENSMUSG00000032366.16
ENSMUST00000105910.2 ENSMUST00000105910.2 ENSMUSG00000028865.15
TranscriptSymbol GeneSymbol EntrezID logFC t
ENSMUST00000223999.2 Rps24-205 Rps24 20088 -5.687391 -15.756599
ENSMUST00000113696.8 Tpm1-212 Tpm1 22003 7.642484 13.397756
ENSMUST00000113705.8 Tpm1-215 Tpm1 22003 -8.749564 -11.520187
ENSMUST00000112384.10 Rps24-201 Rps24 20088 5.142645 12.927014
ENSMUST00000153136.2 Baz2b-213 Baz2b 407823 -3.814401 -11.952368
ENSMUST00000111623.9 Ppfibp1-202 Ppfibp1 67533 -5.169667 -15.522062
ENSMUST00000115120.9 Tnk2-201 Tnk2 51789 3.024369 11.120303
ENSMUST00000063433.8 Eya2-201 Eya2 14049 5.800124 16.704455
ENSMUST00000028949.16 Nsfl1c-201 Nsfl1c 386649 5.369472 16.470947
ENSMUST00000212396.2 Dnmbp-205 Dnmbp 71972 -4.949646 -16.313049
ENSMUST00000016631.14 Ppfibp1-201 Ppfibp1 67533 2.802251 14.052487
ENSMUST00000152639.8 Baz2b-212 Baz2b 407823 2.157634 10.099342
ENSMUST00000191127.8 Ly6e-211 Ly6e 17069 2.068564 10.315025
ENSMUST00000115126.9 Tnk2-207 Tnk2 51789 -2.350822 -10.266132
ENSMUST00000088132.13 Eya2-202 Eya2 14049 -4.659598 -14.794916
ENSMUST00000108751.8 Mprip-203 Mprip 26936 -2.371271 -11.596947
ENSMUST00000089140.13 Nsfl1c-202 Nsfl1c 386649 -5.028457 -14.203461
ENSMUST00000080933.13 Map3k7-202 Map3k7 26409 -3.073747 -12.299050
ENSMUST00000113697.8 Tpm1-213 Tpm1 22003 -6.685821 -9.171201
ENSMUST00000105910.2 Cd164l2-201 Cd164l2 69655 5.798399 15.648113
P.Value FDR
ENSMUST00000223999.2 6.615265e-23 1.177349e-18
ENSMUST00000113696.8 1.028836e-22 1.177349e-18
ENSMUST00000113705.8 4.385732e-19 3.345875e-15
ENSMUST00000112384.10 6.248789e-19 3.575401e-15
ENSMUST00000153136.2 1.021654e-18 4.676520e-15
ENSMUST00000111623.9 2.108433e-17 8.042617e-14
ENSMUST00000115120.9 9.876622e-17 3.229232e-13
ENSMUST00000063433.8 1.376041e-16 3.936681e-13
ENSMUST00000028949.16 2.005359e-16 5.099628e-13
ENSMUST00000212396.2 2.593348e-16 5.935395e-13
ENSMUST00000016631.14 4.456740e-16 9.272856e-13
ENSMUST00000152639.8 2.088132e-15 3.982590e-12
ENSMUST00000191127.8 2.329411e-15 4.101018e-12
ENSMUST00000115126.9 2.828287e-15 4.623643e-12
ENSMUST00000088132.13 3.419658e-15 4.963794e-12
ENSMUST00000108751.8 3.470123e-15 4.963794e-12
ENSMUST00000089140.13 9.878043e-15 1.329875e-11
ENSMUST00000080933.13 2.308478e-14 2.781647e-11
ENSMUST00000113697.8 2.309227e-14 2.781647e-11
ENSMUST00000105910.2 6.230321e-14 7.129668e-11head(out.transcript.MLvsLP,20) Length EffectiveLength Overdispersion
ENSMUST00000059115.13 6548 6227.417 9.176039
ENSMUST00000105389.8 3678 3357.417 6.816707
ENSMUST00000051379.14 3964 3643.417 4.172038
ENSMUST00000115126.9 4540 4219.417 9.377569
ENSMUST00000107756.4 4154 3833.417 3.221990
ENSMUST00000127304.2 642 330.598 1.000000
ENSMUST00000111303.2 4169 3848.417 4.142688
ENSMUST00000135874.2 2943 2622.417 1.184062
ENSMUST00000219140.3 4194 3873.417 3.283965
ENSMUST00000194192.3 6039 5718.417 2.210632
ENSMUST00000115672.2 2909 2588.417 1.597177
ENSMUST00000135803.8 801 481.644 1.400323
ENSMUST00000154087.2 3958 3637.417 1.094463
ENSMUST00000236652.2 367 131.390 1.373704
ENSMUST00000069430.15 2869 2548.417 18.887385
ENSMUST00000227670.2 2788 2467.417 4.264119
ENSMUST00000023758.9 4114 3793.417 2.843403
ENSMUST00000074352.11 3956 3635.417 2.850714
ENSMUST00000112745.8 6037 5716.417 1.820644
ENSMUST00000224156.2 1149 828.417 2.150502
TranscriptID GeneID
ENSMUST00000059115.13 ENSMUST00000059115.13 ENSMUSG00000045636.17
ENSMUST00000105389.8 ENSMUST00000105389.8 ENSMUSG00000001211.16
ENSMUST00000051379.14 ENSMUST00000051379.14 ENSMUSG00000045636.17
ENSMUST00000115126.9 ENSMUST00000115126.9 ENSMUSG00000022791.20
ENSMUST00000107756.4 ENSMUST00000107756.4 ENSMUSG00000028337.15
ENSMUST00000127304.2 ENSMUST00000127304.2 ENSMUSG00000028337.15
ENSMUST00000111303.2 ENSMUST00000111303.2 ENSMUSG00000040479.13
ENSMUST00000135874.2 ENSMUST00000135874.2 ENSMUSG00000038007.15
ENSMUST00000219140.3 ENSMUST00000219140.3 ENSMUSG00000021000.18
ENSMUST00000194192.3 ENSMUST00000194192.3 ENSMUSG00000026131.22
ENSMUST00000115672.2 ENSMUST00000115672.2 ENSMUSG00000032000.18
ENSMUST00000135803.8 ENSMUST00000135803.8 ENSMUSG00000026821.17
ENSMUST00000154087.2 ENSMUST00000154087.2 ENSMUSG00000004043.15
ENSMUST00000236652.2 ENSMUST00000236652.2 ENSMUSG00000024608.12
ENSMUST00000069430.15 ENSMUST00000069430.15 ENSMUSG00000021000.18
ENSMUST00000227670.2 ENSMUST00000227670.2 ENSMUSG00000023017.11
ENSMUST00000023758.9 ENSMUST00000023758.9 ENSMUSG00000023017.11
ENSMUST00000074352.11 ENSMUST00000074352.11 ENSMUSG00000044252.19
ENSMUST00000112745.8 ENSMUST00000112745.8 ENSMUSG00000036792.13
ENSMUST00000224156.2 ENSMUST00000224156.2 ENSMUSG00000021488.9
TranscriptSymbol GeneSymbol EntrezID logFC t
ENSMUST00000059115.13 Mtus1-202 Mtus1 102103 -2.0851123 -9.114762
ENSMUST00000105389.8 Agpat3-204 Agpat3 28169 -2.1726694 -7.559051
ENSMUST00000051379.14 Mtus1-201 Mtus1 102103 1.5150895 6.772110
ENSMUST00000115126.9 Tnk2-207 Tnk2 51789 1.3047327 6.155252
ENSMUST00000107756.4 Coro2a-202 Coro2a 107684 3.5810737 8.663712
ENSMUST00000127304.2 Coro2a-204 Coro2a 107684 -3.5810737 -8.663712
ENSMUST00000111303.2 Dgkz-203 Dgkz 104418 -1.6243741 -6.457184
ENSMUST00000135874.2 Acer2-204 Acer2 230379 1.8309349 6.693407
ENSMUST00000219140.3 Mia2-222 Mia2 338320 2.5820521 5.551469
ENSMUST00000194192.3 Dst-219 Dst 13518 -1.7276415 -5.308965
ENSMUST00000115672.2 Birc3-202 Birc3 11796 1.1491612 5.828626
ENSMUST00000135803.8 Ralgds-206 Ralgds 19730 1.2628102 5.629199
ENSMUST00000154087.2 Stat5a-207 Stat5a 20850 -0.8444577 -5.463278
ENSMUST00000236652.2 Rps14-210 Rps14 20044 2.4882551 5.806016
ENSMUST00000069430.15 Mia2-201 Mia2 338320 -1.7823252 -5.153312
ENSMUST00000227670.2 Asic1-203 Asic1 11419 4.8735920 6.438187
ENSMUST00000023758.9 Asic1-201 Asic1 11419 -4.8735920 -6.438187
ENSMUST00000074352.11 Osbpl1a-201 Osbpl1a 64291 1.2413215 5.534303
ENSMUST00000112745.8 Mbd5-202 Mbd5 109241 1.0326529 5.042686
ENSMUST00000224156.2 Nsd1-203 Nsd1 18193 -1.5401000 -5.202684
P.Value FDR
ENSMUST00000059115.13 1.841272e-11 4.214118e-07
ENSMUST00000105389.8 6.696371e-09 7.662992e-05
ENSMUST00000051379.14 3.254736e-08 2.483038e-04
ENSMUST00000115126.9 5.096525e-08 2.916104e-04
ENSMUST00000107756.4 9.558274e-08 3.646004e-04
ENSMUST00000127304.2 9.558274e-08 3.646004e-04
ENSMUST00000111303.2 4.215333e-07 1.378233e-03
ENSMUST00000135874.2 7.047137e-07 2.016098e-03
ENSMUST00000219140.3 9.038275e-07 2.298433e-03
ENSMUST00000194192.3 1.070058e-06 2.449042e-03
ENSMUST00000115672.2 1.233191e-06 2.565822e-03
ENSMUST00000135803.8 1.401066e-06 2.672183e-03
ENSMUST00000154087.2 2.414273e-06 4.158917e-03
ENSMUST00000236652.2 2.544013e-06 4.158917e-03
ENSMUST00000069430.15 3.784006e-06 5.773636e-03
ENSMUST00000227670.2 5.292878e-06 6.469612e-03
ENSMUST00000023758.9 5.292878e-06 6.469612e-03
ENSMUST00000074352.11 5.428534e-06 6.469612e-03
ENSMUST00000112745.8 5.604875e-06 6.469612e-03
ENSMUST00000224156.2 5.653526e-06 6.469612e-03head(out.transcript.LvsBa,20) Length EffectiveLength Overdispersion
ENSMUST00000223999.2 525 234.370 5.287670
ENSMUST00000113696.8 1714 1393.417 6.058497
ENSMUST00000112384.10 468 194.337 6.131152
ENSMUST00000115120.9 2796 2475.417 7.562900
ENSMUST00000115126.9 4540 4219.417 9.377569
ENSMUST00000153136.2 890 569.769 2.307391
ENSMUST00000079195.6 10467 10146.417 7.172959
ENSMUST00000028949.16 1364 1043.417 6.052642
ENSMUST00000113705.8 1759 1438.417 7.934639
ENSMUST00000063433.8 2460 2139.417 4.602893
ENSMUST00000111623.9 4782 4461.417 11.793842
ENSMUST00000191127.8 1818 1497.417 5.514662
ENSMUST00000080933.13 5669 5348.417 20.189698
ENSMUST00000152639.8 2539 2218.417 1.158605
ENSMUST00000088132.13 2447 2126.417 5.137227
ENSMUST00000123765.8 2613 2292.417 3.107693
ENSMUST00000016631.14 4851 4530.417 3.612108
ENSMUST00000212396.2 6100 5779.417 3.624965
ENSMUST00000089140.13 1486 1165.417 6.677255
ENSMUST00000211882.2 9696 9375.417 3.724371
TranscriptID GeneID
ENSMUST00000223999.2 ENSMUST00000223999.2 ENSMUSG00000025290.18
ENSMUST00000113696.8 ENSMUST00000113696.8 ENSMUSG00000032366.16
ENSMUST00000112384.10 ENSMUST00000112384.10 ENSMUSG00000025290.18
ENSMUST00000115120.9 ENSMUST00000115120.9 ENSMUSG00000022791.20
ENSMUST00000115126.9 ENSMUST00000115126.9 ENSMUSG00000022791.20
ENSMUST00000153136.2 ENSMUST00000153136.2 ENSMUSG00000026987.17
ENSMUST00000079195.6 ENSMUST00000079195.6 ENSMUSG00000038143.17
ENSMUST00000028949.16 ENSMUST00000028949.16 ENSMUSG00000027455.17
ENSMUST00000113705.8 ENSMUST00000113705.8 ENSMUSG00000032366.16
ENSMUST00000063433.8 ENSMUST00000063433.8 ENSMUSG00000017897.19
ENSMUST00000111623.9 ENSMUST00000111623.9 ENSMUSG00000016487.16
ENSMUST00000191127.8 ENSMUST00000191127.8 ENSMUSG00000022587.16
ENSMUST00000080933.13 ENSMUST00000080933.13 ENSMUSG00000028284.14
ENSMUST00000152639.8 ENSMUST00000152639.8 ENSMUSG00000026987.17
ENSMUST00000088132.13 ENSMUST00000088132.13 ENSMUSG00000017897.19
ENSMUST00000123765.8 ENSMUST00000123765.8 ENSMUSG00000028649.23
ENSMUST00000016631.14 ENSMUST00000016631.14 ENSMUSG00000016487.16
ENSMUST00000212396.2 ENSMUST00000212396.2 ENSMUSG00000025195.10
ENSMUST00000089140.13 ENSMUST00000089140.13 ENSMUSG00000027455.17
ENSMUST00000211882.2 ENSMUST00000211882.2 ENSMUSG00000038143.17
TranscriptSymbol GeneSymbol EntrezID logFC t
ENSMUST00000223999.2 Rps24-205 Rps24 20088 -6.159657 -18.97073
ENSMUST00000113696.8 Tpm1-212 Tpm1 22003 7.676108 13.84655
ENSMUST00000112384.10 Rps24-201 Rps24 20088 5.603892 15.11578
ENSMUST00000115120.9 Tnk2-201 Tnk2 51789 3.535542 13.76596
ENSMUST00000115126.9 Tnk2-207 Tnk2 51789 -2.791865 -13.32287
ENSMUST00000153136.2 Baz2b-213 Baz2b 407823 -4.532428 -12.50956
ENSMUST00000079195.6 Stox2-201 Stox2 71069 -3.617704 -17.83454
ENSMUST00000028949.16 Nsfl1c-201 Nsfl1c 386649 5.316455 17.78489
ENSMUST00000113705.8 Tpm1-215 Tpm1 22003 -9.120773 -10.63324
ENSMUST00000063433.8 Eya2-201 Eya2 14049 6.346188 17.75111
ENSMUST00000111623.9 Ppfibp1-202 Ppfibp1 67533 -6.280208 -15.37448
ENSMUST00000191127.8 Ly6e-211 Ly6e 17069 1.958496 10.99907
ENSMUST00000080933.13 Map3k7-202 Map3k7 26409 -3.994988 -14.40998
ENSMUST00000152639.8 Baz2b-212 Baz2b 407823 2.082520 10.64372
ENSMUST00000088132.13 Eya2-202 Eya2 14049 -6.485333 -16.24813
ENSMUST00000123765.8 Macf1-204 Macf1 11426 2.268261 10.55133
ENSMUST00000016631.14 Ppfibp1-201 Ppfibp1 67533 2.702292 14.12929
ENSMUST00000212396.2 Dnmbp-205 Dnmbp 71972 -4.120276 -15.78702
ENSMUST00000089140.13 Nsfl1c-202 Nsfl1c 386649 -4.769588 -15.77816
ENSMUST00000211882.2 Stox2-209 Stox2 71069 2.957028 15.65141
P.Value FDR
ENSMUST00000223999.2 6.429734e-27 1.471573e-22
ENSMUST00000113696.8 1.480320e-23 1.694004e-19
ENSMUST00000112384.10 4.816206e-22 3.674284e-18
ENSMUST00000115120.9 5.233455e-21 2.994452e-17
ENSMUST00000115126.9 2.560395e-20 1.171995e-16
ENSMUST00000153136.2 1.111277e-19 4.238967e-16
ENSMUST00000079195.6 2.358351e-17 5.901374e-14
ENSMUST00000028949.16 2.543375e-17 5.901374e-14
ENSMUST00000113705.8 2.558412e-17 5.901374e-14
ENSMUST00000063433.8 2.677715e-17 5.901374e-14
ENSMUST00000111623.9 2.836332e-17 5.901374e-14
ENSMUST00000191127.8 1.582381e-16 3.017997e-13
ENSMUST00000080933.13 2.078587e-16 3.510075e-13
ENSMUST00000152639.8 2.147116e-16 3.510075e-13
ENSMUST00000088132.13 2.884188e-16 4.400694e-13
ENSMUST00000123765.8 3.152938e-16 4.510081e-13
ENSMUST00000016631.14 3.778748e-16 5.087307e-13
ENSMUST00000212396.2 6.198230e-16 7.578050e-13
ENSMUST00000089140.13 6.291036e-16 7.578050e-13
ENSMUST00000211882.2 7.788334e-16 8.912581e-13table(out.transcript.LPvsBa$FDR < 0.05)
FALSE TRUE
20398 2489 out.transcript.LPvsBa[out.transcript.LPvsBa$GeneSymbol %in% c('Foxp1','Ezh2','Asap1'),] Length EffectiveLength Overdispersion
ENSMUST00000175838.2 420 163.758 1.159129
ENSMUST00000177083.8 4415 4094.417 13.839170
ENSMUST00000177229.8 1848 1527.417 16.866677
ENSMUST00000110115.9 4535 4214.417 18.135542
ENSMUST00000177371.8 6164 5843.417 3.344945
ENSMUST00000081721.13 2787 2466.417 6.653742
ENSMUST00000092648.13 2115 1794.417 4.728582
ENSMUST00000154163.9 853 533.020 1.566080
ENSMUST00000124058.8 1213 892.417 3.563683
ENSMUST00000176565.8 3727 3406.417 8.946068
ENSMUST00000177437.8 2468 2147.417 5.693538
ENSMUST00000113322.9 7177 6856.417 5.676970
ENSMUST00000177307.8 2121 1800.417 6.298976
ENSMUST00000114618.8 2775 2454.417 9.490129
ENSMUST00000177410.2 618 309.424 1.435156
ENSMUST00000177227.8 967 646.467 1.373630
ENSMUST00000113321.7 1176 855.417 1.235459
ENSMUST00000170327.3 834 514.204 1.613701
ENSMUST00000175799.8 4237 3916.417 12.753683
TranscriptID GeneID
ENSMUST00000175838.2 ENSMUST00000175838.2 ENSMUSG00000030067.18
ENSMUST00000177083.8 ENSMUST00000177083.8 ENSMUSG00000022377.17
ENSMUST00000177229.8 ENSMUST00000177229.8 ENSMUSG00000030067.18
ENSMUST00000110115.9 ENSMUST00000110115.9 ENSMUSG00000022377.17
ENSMUST00000177371.8 ENSMUST00000177371.8 ENSMUSG00000022377.17
ENSMUST00000081721.13 ENSMUST00000081721.13 ENSMUSG00000029687.17
ENSMUST00000092648.13 ENSMUST00000092648.13 ENSMUSG00000029687.17
ENSMUST00000154163.9 ENSMUST00000154163.9 ENSMUSG00000030067.18
ENSMUST00000124058.8 ENSMUST00000124058.8 ENSMUSG00000030067.18
ENSMUST00000176565.8 ENSMUST00000176565.8 ENSMUSG00000030067.18
ENSMUST00000177437.8 ENSMUST00000177437.8 ENSMUSG00000030067.18
ENSMUST00000113322.9 ENSMUST00000113322.9 ENSMUSG00000030067.18
ENSMUST00000177307.8 ENSMUST00000177307.8 ENSMUSG00000030067.18
ENSMUST00000114618.8 ENSMUST00000114618.8 ENSMUSG00000029687.17
ENSMUST00000177410.2 ENSMUST00000177410.2 ENSMUSG00000030067.18
ENSMUST00000177227.8 ENSMUST00000177227.8 ENSMUSG00000030067.18
ENSMUST00000113321.7 ENSMUST00000113321.7 ENSMUSG00000030067.18
ENSMUST00000170327.3 ENSMUST00000170327.3 ENSMUSG00000029687.17
ENSMUST00000175799.8 ENSMUST00000175799.8 ENSMUSG00000022377.17
TranscriptSymbol GeneSymbol EntrezID logFC
ENSMUST00000175838.2 Foxp1-216 Foxp1 108655 3.7178808
ENSMUST00000177083.8 Asap1-216 Asap1 13196 2.5112479
ENSMUST00000177229.8 Foxp1-225 Foxp1 108655 -4.8272966
ENSMUST00000110115.9 Asap1-203 Asap1 13196 -3.7673263
ENSMUST00000177371.8 Asap1-217 Asap1 13196 -1.3858171
ENSMUST00000081721.13 Ezh2-201 Ezh2 14056 -1.1168666
ENSMUST00000092648.13 Ezh2-202 Ezh2 14056 0.9855436
ENSMUST00000154163.9 Foxp1-214 Foxp1 108655 1.0557781
ENSMUST00000124058.8 Foxp1-210 Foxp1 108655 -2.6334500
ENSMUST00000176565.8 Foxp1-220 Foxp1 108655 0.9961347
ENSMUST00000177437.8 Foxp1-230 Foxp1 108655 -0.4735165
ENSMUST00000113322.9 Foxp1-204 Foxp1 108655 -0.3314275
ENSMUST00000177307.8 Foxp1-228 Foxp1 108655 0.4493037
ENSMUST00000114618.8 Ezh2-204 Ezh2 14056 0.5753112
ENSMUST00000177410.2 Foxp1-229 Foxp1 108655 -0.7587415
ENSMUST00000177227.8 Foxp1-224 Foxp1 108655 -0.3202687
ENSMUST00000113321.7 Foxp1-203 Foxp1 108655 -0.4053769
ENSMUST00000170327.3 Ezh2-211 Ezh2 14056 -0.1919945
ENSMUST00000175799.8 Asap1-206 Asap1 13196 0.1526919
t P.Value FDR
ENSMUST00000175838.2 6.1216449 4.870324e-08 6.518544e-06
ENSMUST00000177083.8 6.1084573 1.099600e-06 8.170954e-05
ENSMUST00000177229.8 -4.9893592 4.292241e-06 2.307853e-04
ENSMUST00000110115.9 -4.1672062 2.466891e-04 5.519037e-03
ENSMUST00000177371.8 -3.8943742 5.204844e-04 9.529861e-03
ENSMUST00000081721.13 -3.8680119 5.591062e-04 1.002842e-02
ENSMUST00000092648.13 3.3287243 2.348844e-03 2.772491e-02
ENSMUST00000154163.9 2.7391013 7.820229e-03 6.355880e-02
ENSMUST00000124058.8 -2.6991106 8.721437e-03 6.833534e-02
ENSMUST00000176565.8 2.3972268 1.921090e-02 1.144999e-01
ENSMUST00000177437.8 -1.9049437 6.092441e-02 2.366560e-01
ENSMUST00000113322.9 -1.5396204 1.281922e-01 3.656017e-01
ENSMUST00000177307.8 1.4620755 1.482262e-01 3.982543e-01
ENSMUST00000114618.8 1.3876042 1.756426e-01 4.388094e-01
ENSMUST00000177410.2 -1.0309554 3.061349e-01 5.920661e-01
ENSMUST00000177227.8 -0.8878730 3.776705e-01 6.605338e-01
ENSMUST00000113321.7 -0.6711939 5.043210e-01 7.561680e-01
ENSMUST00000170327.3 -0.4787420 6.356488e-01 8.370302e-01
ENSMUST00000175799.8 0.3257419 7.469172e-01 8.961055e-01gene.unique <- out.gene.LPvsBa[out.gene.LPvsBa$FDR < 0.05,'GeneSymbol']
tx.unique <-
out.transcript.LPvsBa[out.transcript.LPvsBa$FDR < 0.05 &
out.transcript.LPvsBa$GeneSymbol %in% gene.unique,"GeneSymbol"]
table(table(tx.unique))
1 2 3 4 5 6 7 9 11
333 668 96 32 21 6 2 3 1 x <- as.data.table(out.transcript.LPvsBa[out.transcript.LPvsBa$FDR < 0.05 &
out.transcript.LPvsBa$GeneSymbol %in% gene.unique,])
x.sign <- x[GeneSymbol %in% names(table(tx.unique)[table(tx.unique) == 2]),][,.(min = sign(min(logFC)),max = sign(max(logFC))),by = 'GeneSymbol']
x.sign[,table(min,max)] max
min -1 1
-1 8 652
1 0 8out.gene.LPvsBa[out.gene.LPvsBa$GeneSymbol %in% c('Eya2'),] GeneID GeneSymbol EntrezID NExons F
ENSMUSG00000017897.19 ENSMUSG00000017897.19 Eya2 14049 4 88.37603
P.Value FDR
ENSMUSG00000017897.19 7.537372e-15 4.386171e-12out.transcript.LPvsBa[out.transcript.LPvsBa$GeneSymbol %in% c('Eya2'),] Length EffectiveLength Overdispersion
ENSMUST00000063433.8 2460 2139.417 4.602893
ENSMUST00000088132.13 2447 2126.417 5.137227
ENSMUST00000150638.8 679 364.332 3.585353
ENSMUST00000150669.2 959 638.483 1.635973
TranscriptID GeneID
ENSMUST00000063433.8 ENSMUST00000063433.8 ENSMUSG00000017897.19
ENSMUST00000088132.13 ENSMUST00000088132.13 ENSMUSG00000017897.19
ENSMUST00000150638.8 ENSMUST00000150638.8 ENSMUSG00000017897.19
ENSMUST00000150669.2 ENSMUST00000150669.2 ENSMUSG00000017897.19
TranscriptSymbol GeneSymbol EntrezID logFC t
ENSMUST00000063433.8 Eya2-201 Eya2 14049 6.848815 14.604867
ENSMUST00000088132.13 Eya2-202 Eya2 14049 -8.811995 -13.451515
ENSMUST00000150638.8 Eya2-205 Eya2 14049 -2.742883 -4.100140
ENSMUST00000150669.2 Eya2-206 Eya2 14049 -2.143804 -3.099637
P.Value FDR
ENSMUST00000063433.8 4.791327e-15 1.370739e-11
ENSMUST00000088132.13 3.996427e-14 7.622186e-11
ENSMUST00000150638.8 2.966532e-04 6.304086e-03
ENSMUST00000150669.2 4.232068e-03 4.224132e-02go <- goana(de = as.character(out.gene.LPvsBa[out.gene.LPvsBa$FDR < 0.05,"EntrezID"]),species = "Mm")
top.go <- topGO(go,number = Inf,ontology = "BP")
head(top.go,20) Term Ont N DE
GO:0009987 cellular process BP 17623 1106
GO:0071840 cellular component organization or biogenesis BP 6717 578
GO:0016043 cellular component organization BP 6506 565
GO:0065007 biological regulation BP 12967 850
GO:0008152 metabolic process BP 11602 791
GO:0044237 cellular metabolic process BP 10118 719
GO:0044238 primary metabolic process BP 10134 717
GO:0006807 nitrogen compound metabolic process BP 9563 689
GO:0050789 regulation of biological process BP 12570 821
GO:0050794 regulation of cellular process BP 11938 792
GO:0071704 organic substance metabolic process BP 11172 755
GO:0051641 cellular localization BP 3560 356
GO:0006996 organelle organization BP 3584 357
GO:0051179 localization BP 5383 461
GO:0043170 macromolecule metabolic process BP 9533 660
GO:0048518 positive regulation of biological process BP 6679 520
GO:0048519 negative regulation of biological process BP 5817 471
GO:0048522 positive regulation of cellular process BP 6141 486
GO:0048523 negative regulation of cellular process BP 5339 442
GO:0051128 regulation of cellular component organization BP 2632 281
P.DE
GO:0009987 9.730938e-118
GO:0071840 4.186403e-79
GO:0016043 3.135819e-78
GO:0065007 5.941126e-70
GO:0008152 1.509438e-69
GO:0044237 4.701855e-67
GO:0044238 7.279417e-66
GO:0006807 3.453766e-65
GO:0050789 1.786064e-64
GO:0050794 1.291881e-63
GO:0071704 3.954474e-62
GO:0051641 1.838009e-58
GO:0006996 3.179435e-58
GO:0051179 5.084191e-58
GO:0043170 8.028239e-54
GO:0048518 9.955001e-54
GO:0048519 3.561348e-52
GO:0048522 2.811899e-51
GO:0048523 7.894808e-51
GO:0051128 2.239253e-50Plots
fig.voom <-
wrap_elements(full = ~ plot.voom(v,ylim = c(0.16,2.54)))
file.voom <- tempfile("voom",fileext = '.png')
png(file.voom,width = 4,height = 3,units = 'in',res = 300)
par(mar = c(3, 3, 2, 0.25),mgp = c(2,1,0))
fig.voom
dev.off()png
2 fig.voom <- readPNG(file.voom, native = TRUE)
plot.voom(v,ylim = c(0.16,2.54))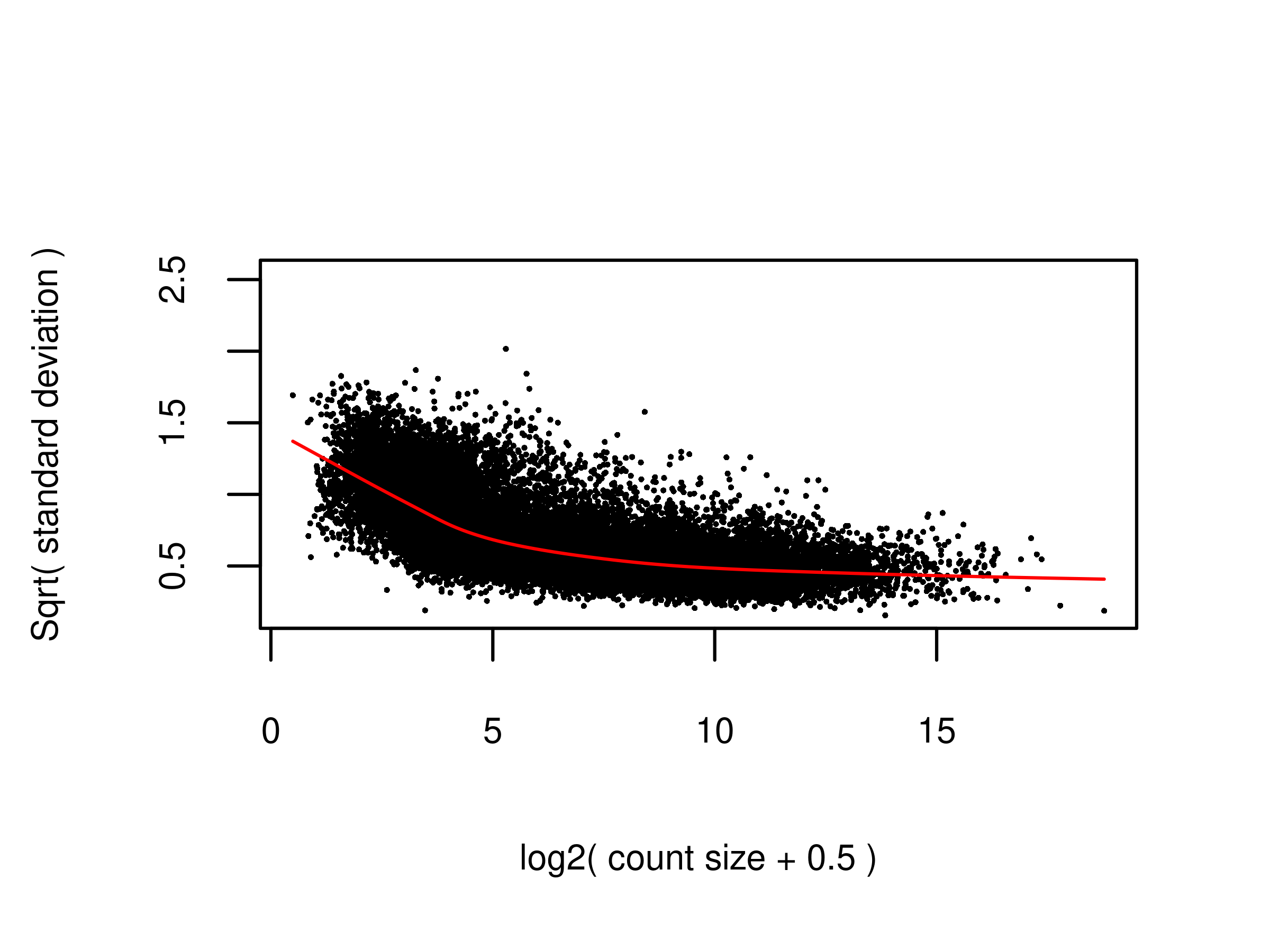
fig.voom.raw <- wrap_elements(full = ~ plot.voom(v.raw,ylim = c(0.16,2.54)))
file.voom.raw <- tempfile("voomraw",fileext = '.png')
png(file.voom.raw,width = 4,height = 3,units = 'in',res = 300)
par(mar = c(3, 3, 2, 0.25),mgp = c(2,1,0))
fig.voom.raw
dev.off()png
2 fig.voom.raw <- readPNG(file.voom.raw, native = TRUE)
plot.voom(v.raw,ylim = c(0.16,2.54))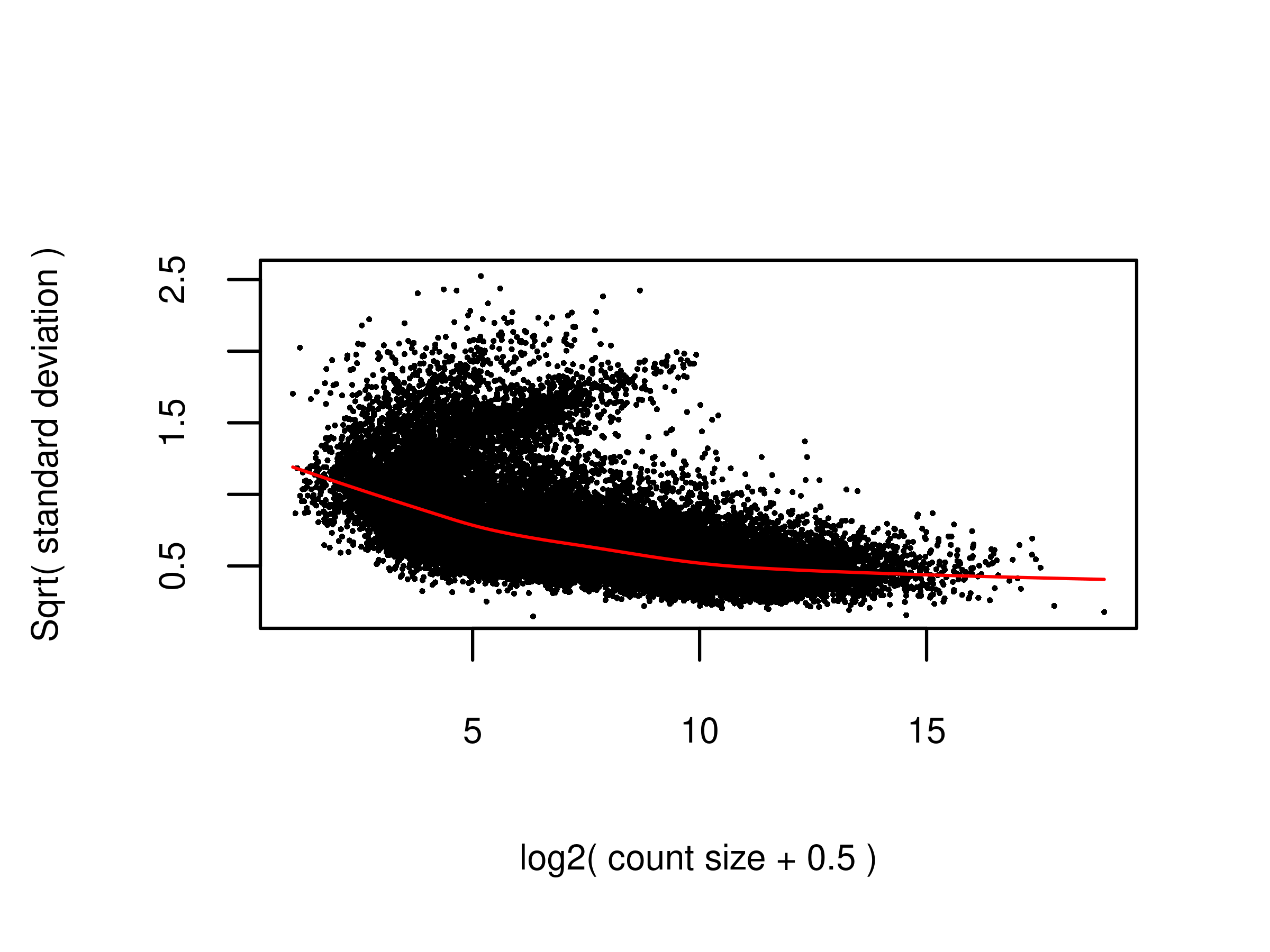
fig.splice <-
wrap_elements(full = ~ plotSplice2(fit = ds,coef = 'LPvsBa',geneid = 'Foxp1',
genecolname = 'GeneSymbol',
exonlabel = 'TranscriptSymbol',
ylim = c(-5.5,4.5)))
file.splice <- tempfile("splice",fileext = '.png')
png(file.splice,width = 4,height = 3,units = 'in',res = 300)
par(mar = c(4, 3, 2, 0.25),mgp = c(2,1,0))
fig.splice
dev.off()png
2 fig.splice <- readPNG(file.splice, native = TRUE)
plotSplice2(fit = ds,coef = 'LPvsBa',geneid = 'Foxp1',
genecolname = 'GeneSymbol',exonlabel = 'TranscriptSymbol',
ylim = c(-5.5,4.5))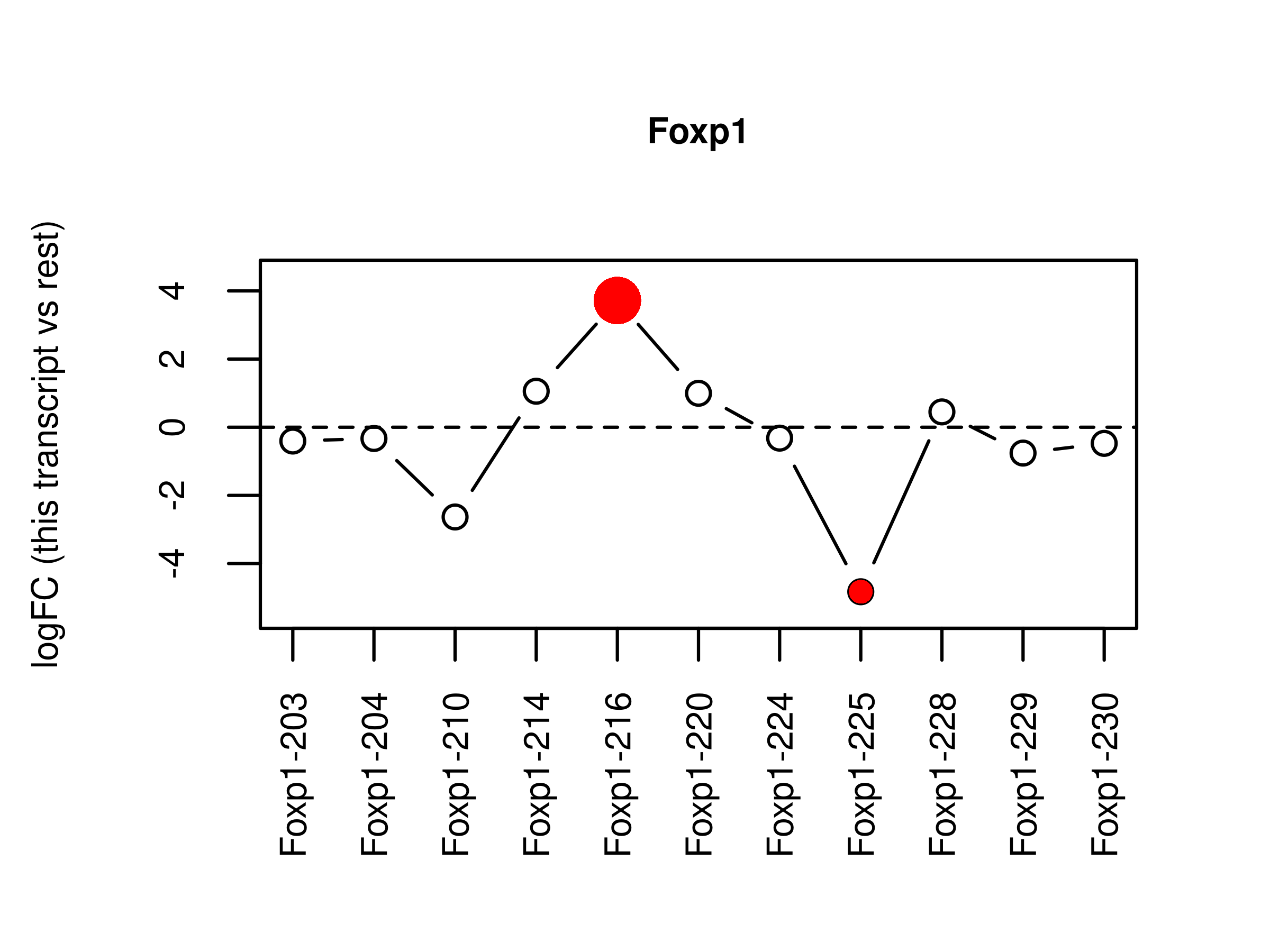
file.barplot <- tempfile("barplot",fileext = '.png')
png(file.barplot,width = 4,height = 3,units = 'in',res = 300)
plot.barplot(top.go,top = 10)
dev.off()png
2 fig.barplot <- readPNG(file.barplot, native = TRUE)
plot.barplot(top.go,top = 10)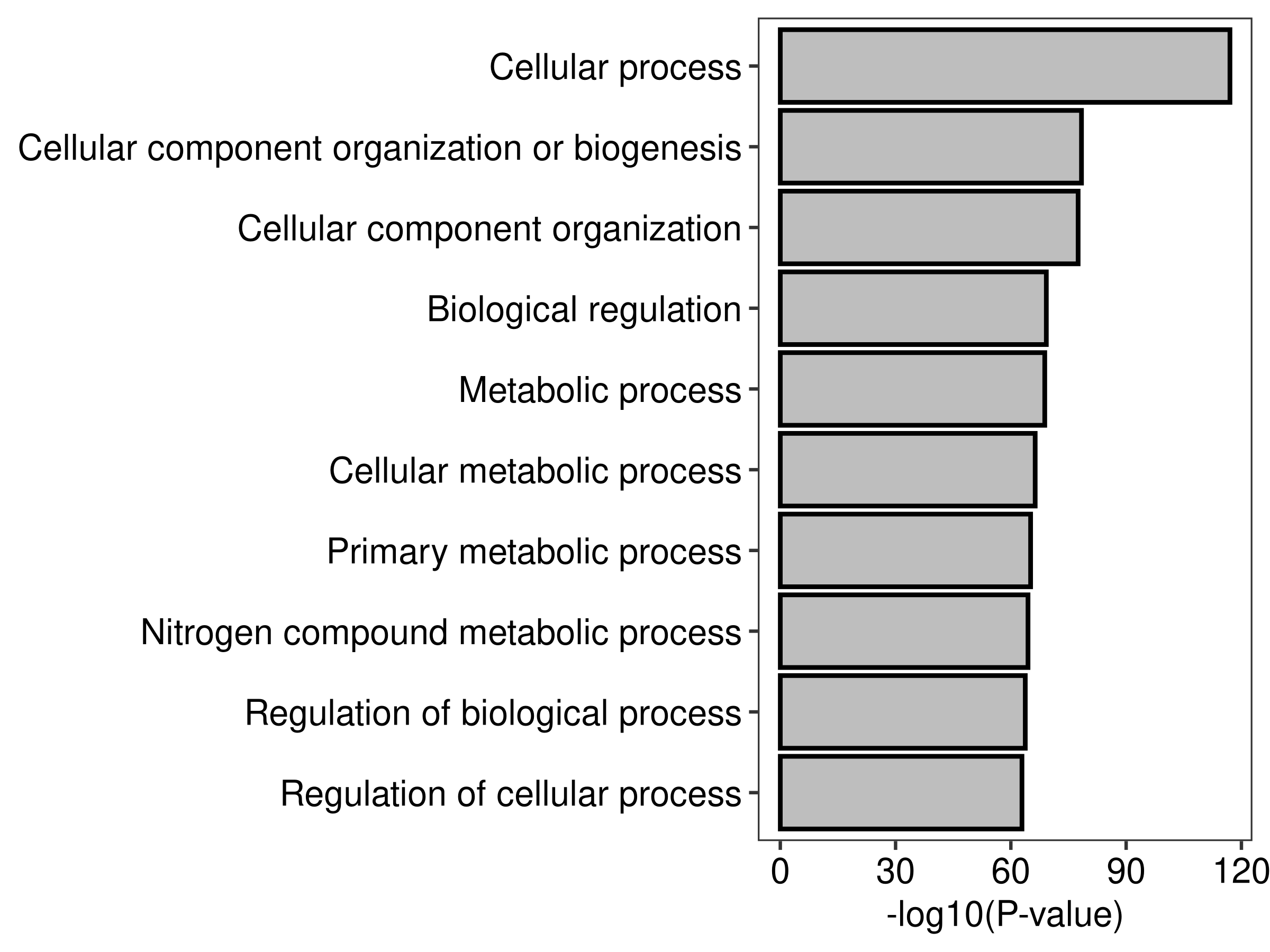
BAM <- file.path(path.bam,paste0(dge.filtr$samples$sample,'.bam'))
BW <- file.path(path.bw,paste0(dge.filtr$samples$sample,'.bw'))
txdb <- makeTxDbFromGRanges(gtf[(gtf$transcript_id %in% gtf.tx$transcript_id) &
(gtf$transcript_id %in% ds$genes$TranscriptID)])Warning in call_fun_in_txdbmaker("makeTxDbFromGRanges", ...): makeTxDbFromGRanges() has moved to the txdbmaker package. Please call
txdbmaker::makeTxDbFromGRanges() to get rid of this warning.Warning in .get_cds_IDX(mcols0$type, mcols0$phase): The "phase" metadata column contains non-NA values for features of type
stop_codon. This information was ignored.geneTrack <-
GeneRegionTrack(txdb,showId=TRUE,
just.group = 'right',
background.title = "white",
min.height=10,
fill = 'grey',
name = 'Transcripts',
col.title = 'black',
fontcolor.group = 'black')
ranges(geneTrack)$symbol <-
gtf.tx$transcript_name[match(ranges(geneTrack)$symbol,gtf.tx$transcript_id)]
param <- readParam(pe = 'both',restrict = paste0("chr", c(1:19, "X", "Y")))
geneTrack.left <- geneTrack
geneTrack.left@dp@pars$just.group <- 'left'
Eya2 <- tempfile("Eya2",fileext = 'png')
png(Eya2,width = 4,height = 6,res = 300,units = 'in')
plotCoverage(gr = GRanges('chr2',IRanges(165415686,165545256)),
x = BAM,
fontsize = 8,
lib.sizes = dge.filtr$samples$lib.size*dge.filtr$samples$norm.factors,
param = param,
anno = geneTrack.left,
fill = c(rep('#df5b6d',3),rep('#5ece5a',3),rep("#2f95e2",3)),
ylim = c(-0.75,6.75),
yTicksAt = c(0,3,6),
labels = with(dge.filtr$samples,paste(group,pool,sep = '.')))
dev.off()png
2 fig.Eya2 <- readPNG(Eya2, native = TRUE)
plotCoverage(gr = GRanges('chr2',IRanges(165415686,165545256)),
x = BAM,
fontsize = 8,
lib.sizes = dge.filtr$samples$lib.size*dge.filtr$samples$norm.factors,
param = param,
anno = geneTrack.left,
fill = c(rep('#df5b6d',3),rep('#5ece5a',3),rep("#2f95e2",3)),
ylim = c(-0.75,6.75),
yTicksAt = c(0,3,6),
labels = with(dge.filtr$samples,paste(group,pool,sep = '.')))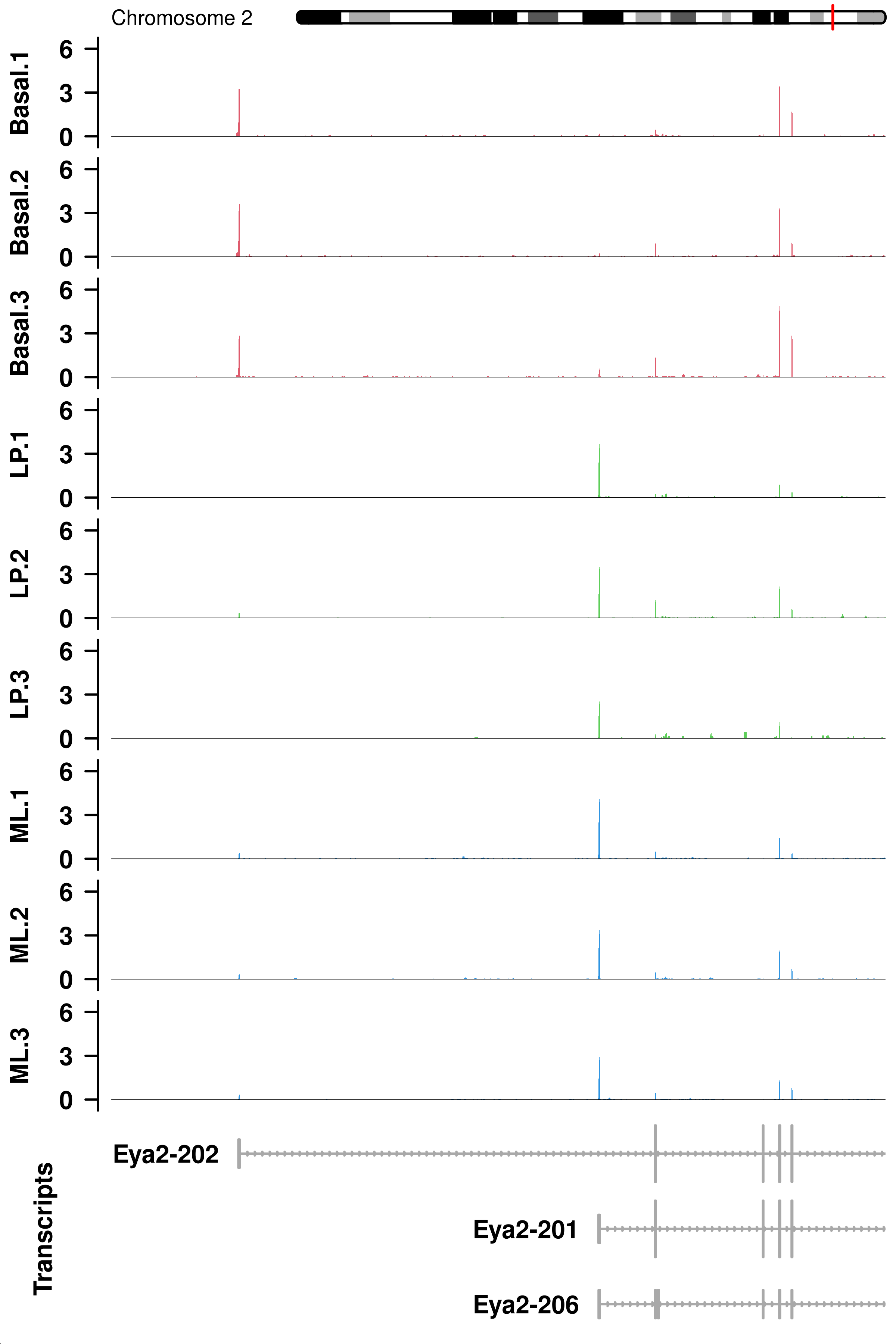
Output files
fig.ds <- wrap_plots(A = wrap_elements(fig.voom.raw),
B = wrap_elements(fig.voom),
C = wrap_elements(fig.barplot),
D = wrap_elements(fig.splice),
E = wrap_elements(fig.Eya2),
design = c(area(1,1),area(1,2),
area(2,1),area(3,1),area(2,2,3,2)),
heights = c(3,3,3)/8) +
plot_annotation(tag_levels = 'a',
theme = theme(plot.tag = element_text(size = 8)))
ggsave(plot = fig.ds,
filename = file.path(path.misc,'Figure-CaseStudy.pdf'),
device = 'pdf',width = 8,height = 9,units = 'in',dpi = 300)
sessionInfo()R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Red Hat Enterprise Linux 9.3 (Plow)
Matrix products: default
BLAS: /stornext/System/data/software/rhel/9/base/tools/R/4.4.1/lib64/R/lib/libRblas.so
LAPACK: /stornext/System/data/software/rhel/9/base/tools/R/4.4.1/lib64/R/lib/libRlapack.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
time zone: Australia/Melbourne
tzcode source: system (glibc)
attached base packages:
[1] grid stats4 stats graphics grDevices datasets utils
[8] methods base
other attached packages:
[1] patchwork_1.3.0
[2] png_0.1-8
[3] stringr_1.5.1
[4] ggplot2_3.5.1
[5] BiocParallel_1.38.0
[6] csaw_1.38.0
[7] SummarizedExperiment_1.34.0
[8] MatrixGenerics_1.16.0
[9] matrixStats_1.5.0
[10] TxDb.Mmusculus.UCSC.mm39.refGene_3.19.0
[11] Gviz_1.48.0
[12] data.table_1.17.0
[13] GenomicFeatures_1.56.0
[14] AnnotationDbi_1.66.0
[15] Biobase_2.64.0
[16] Rsubread_2.18.0
[17] rtracklayer_1.64.0
[18] GenomicRanges_1.56.2
[19] GenomeInfoDb_1.40.1
[20] IRanges_2.38.1
[21] S4Vectors_0.42.1
[22] BiocGenerics_0.50.0
[23] edgeR_4.5.9
[24] limma_3.63.9
[25] workflowr_1.7.1
loaded via a namespace (and not attached):
[1] later_1.4.1 BiocIO_1.14.0 bitops_1.0-9
[4] filelock_1.0.3 tibble_3.2.1 R.oo_1.27.0
[7] XML_3.99-0.18 rpart_4.1.23 lifecycle_1.0.4
[10] httr2_1.1.0 rprojroot_2.0.4 processx_3.8.6
[13] lattice_0.22-6 vroom_1.6.5 ensembldb_2.28.1
[16] backports_1.5.0 magrittr_2.0.3 Hmisc_5.2-2
[19] sass_0.4.9 rmarkdown_2.29 jquerylib_0.1.4
[22] yaml_2.3.10 metapod_1.12.0 httpuv_1.6.15
[25] DBI_1.2.3 RColorBrewer_1.1-3 abind_1.4-8
[28] zlibbioc_1.50.0 R.utils_2.13.0 AnnotationFilter_1.28.0
[31] biovizBase_1.52.0 RCurl_1.98-1.16 nnet_7.3-19
[34] VariantAnnotation_1.50.0 rappdirs_0.3.3 git2r_0.35.0
[37] GenomeInfoDbData_1.2.12 codetools_0.2-20 DelayedArray_0.30.1
[40] xml2_1.3.7 tidyselect_1.2.1 UCSC.utils_1.0.0
[43] farver_2.1.2 BiocFileCache_2.12.0 base64enc_0.1-3
[46] GenomicAlignments_1.40.0 jsonlite_1.9.1 Formula_1.2-5
[49] systemfonts_1.2.1 tools_4.4.1 progress_1.2.3
[52] ragg_1.3.3 Rcpp_1.0.14 glue_1.8.0
[55] gridExtra_2.3 SparseArray_1.4.8 xfun_0.51
[58] dplyr_1.1.4 withr_3.0.2 BiocManager_1.30.25
[61] fastmap_1.2.0 latticeExtra_0.6-30 callr_3.7.6
[64] digest_0.6.37 R6_2.6.1 gridGraphics_0.5-1
[67] textshaping_1.0.0 colorspace_2.1-1 GO.db_3.19.1
[70] jpeg_0.1-10 dichromat_2.0-0.1 biomaRt_2.60.1
[73] RSQLite_2.3.9 R.methodsS3_1.8.2 generics_0.1.3
[76] renv_1.1.2 prettyunits_1.2.0 httr_1.4.7
[79] htmlwidgets_1.6.4 S4Arrays_1.4.1 org.Mm.eg.db_3.19.1
[82] whisker_0.4.1 pkgconfig_2.0.3 gtable_0.3.6
[85] blob_1.2.4 XVector_0.44.0 htmltools_0.5.8.1
[88] ProtGenerics_1.36.0 scales_1.3.0 knitr_1.49
[91] rstudioapi_0.17.1 tzdb_0.4.0 rjson_0.2.23
[94] checkmate_2.3.2 curl_6.2.1 cachem_1.1.0
[97] parallel_4.4.1 foreign_0.8-86 restfulr_0.0.15
[100] pillar_1.10.1 vctrs_0.6.5 promises_1.3.2
[103] dbplyr_2.5.0 cluster_2.1.6 htmlTable_2.4.3
[106] evaluate_1.0.3 readr_2.1.5 cli_3.6.4
[109] locfit_1.5-9.12 compiler_4.4.1 Rsamtools_2.20.0
[112] rlang_1.1.5 crayon_1.5.3 labeling_0.4.3
[115] interp_1.1-6 ps_1.9.0 getPass_0.2-4
[118] fs_1.6.5 stringi_1.8.4 deldir_2.0-4
[121] txdbmaker_1.0.1 munsell_0.5.1 Biostrings_2.72.1
[124] lazyeval_0.2.2 Matrix_1.7-0 BSgenome_1.72.0
[127] hms_1.1.3 bit64_4.6.0-1 KEGGREST_1.44.1
[130] statmod_1.5.0 memoise_2.0.1 bslib_0.9.0
[133] bit_4.6.0