Broullon DIC/TA climatology
Pasqualina Vonlanthen & Jens Daniel Müller
29 April, 2022
Last updated: 2022-04-29
Checks: 7 0
Knit directory: bgc_argo_r_argodata/
This reproducible R Markdown analysis was created with workflowr (version 1.7.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20211008) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 2d44f8a. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .RData
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: output/
Untracked files:
Untracked: code/OceanSODA_argo_extremes.R
Untracked: code/creating_dataframe.R
Untracked: code/creating_map.R
Untracked: code/merging_oceanSODA_Argo.R
Untracked: code/pH_data_timeseries.R
Unstaged changes:
Modified: analysis/_site.yml
Deleted: analysis/load_OceanSODA_biomes.Rmd
Modified: code/Workflowr_project_managment.R
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were made to the R Markdown (analysis/broullon_DIC_TA_clim.Rmd) and HTML (docs/broullon_DIC_TA_clim.html) files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| html | 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 | Build site. |
| Rmd | 8b582f0 | pasqualina-vonlanthendinenna | 2022-04-29 | added broullon climatology page, argo locations |
Task
Explore the Broullón et al. (2020) DIC / TA climatology
library(tidyverse)
library(lubridate)
library(stars)path_argo <- '/nfs/kryo/work/updata/bgc_argo_r_argodata'
path_argo_preprocessed <- paste0(path_argo, "/preprocessed_bgc_data")
path_emlr_utilities <- "/nfs/kryo/work/jenmueller/emlr_cant/utilities/files/"
path_updata <- '/nfs/kryo/work/updata'
path_broullon_clim <- paste0(path_updata, "/broullon_co2_monthly_climatology")theme_set(theme_bw())map <- map <-
read_rds(paste(path_emlr_utilities,
"map_landmask_WOA18.rds",
sep = ""))
nm_biomes <- read_rds(file = paste0(path_argo_preprocessed, "/nm_biomes.rds"))
# WOA 18 basin mask
basinmask <-
read_csv(
paste(path_emlr_utilities,
"basin_mask_WOA18.csv",
sep = ""),
col_types = cols("MLR_basins" = col_character())
)
basinmask <- basinmask %>%
filter(MLR_basins == unique(basinmask$MLR_basins)[1]) %>%
select(-c(MLR_basins, basin))Load Data
DIC
DIC_clim <- tidync::hyper_tibble(paste0(path_broullon_clim, "/TCO2_NNGv2LDEO_climatology.nc"))
nc_depth <- read_ncdf(paste0(path_broullon_clim, "/TCO2_NNGv2LDEO_climatology.nc"),
var = c('depth'))
nc_depth <- as_tibble(nc_depth)
nc_depth <- nc_depth %>%
mutate(depth_level = depth_level+0.5)
DIC_clim <- full_join(DIC_clim, nc_depth)
DIC_clim <- DIC_clim %>%
rename(DIC = TCO2_NNGv2LDEO,
month = time)
rm(nc_depth)
# table(unique(DIC_clim$latitude))
# table(unique(DIC_clim$longitude))
# table(unique(DIC_clim$time))
# table(unique(DIC_clim$depth))
# text <- read_file(paste0(path_broullon_clim, "/README_global_monthly_2020.txt"))
# Depth goes down to 5500 m, but below 1500 m DIC is an annual climatological value, rather than a monthly climatological value pCO2
pco2_clim <- tidync::hyper_tibble(paste0(path_broullon_clim, "/pCO2_NNGv2LDEO_climatology.nc"))
pco2_clim <- pco2_clim %>%
mutate(depth_level = 1) %>%
rename(pco2 = pCO2_NNGv2LDEO,
month = time)TA
TA_clim <- tidync::hyper_tibble(paste0(path_broullon_clim, "/AT_NNGv2_climatology.nc"))
nc_depth <- read_ncdf(paste0(path_broullon_clim, "/AT_NNGv2_climatology.nc"),
var = c('depth'))
nc_depth <- as_tibble(nc_depth)
nc_depth <- nc_depth %>%
mutate(depth_level = depth_level+0.5)
TA_clim <- full_join(TA_clim, nc_depth)
rm(nc_depth)
# read_file(paste0(path_broullon_clim, "/README_Global_monthly_2019.txt"))
TA_clim <- TA_clim %>%
rename(TA = AT_NNGv2,
month = time)
# Depth goes down to 5500 m, but below 1500 m TA is an annual climatological value, rather than a monthly climatological value Join data
broullon_clim <- full_join(DIC_clim, TA_clim)
broullon_clim <- full_join(broullon_clim, pco2_clim)
rm(DIC_clim, pco2_clim, TA_clim)Harmonise data
# put longitude and latitude labels to the center of the grid (.5º)
broullon_clim <- broullon_clim %>%
rename(lon = longitude,
lat = latitude) %>%
mutate(lon = if_else(lon < 20, lon + 360, lon))
broullon_clim_SO <- broullon_clim %>%
filter(lat <= -30)Write data
broullon_clim %>%
write_rds(file = paste0(path_argo_preprocessed, "/broullon_TA_DIC_clim.rds"))
broullon_clim_SO %>%
write_rds(file = paste0(path_argo_preprocessed, "/broullon_TA_DIC_clim_SO.rds"))Plot data
# join regional separations
broullon_clim_SO <- inner_join(broullon_clim_SO, nm_biomes)
broullon_clim_SO <- inner_join(broullon_clim_SO, basinmask)DIC
broullon_clim_SO %>%
group_split(depth) %>%
map(
~map +
geom_tile(data = .x,
aes(x = lon,
y = lat,
fill = DIC))+
scale_fill_viridis_c()+
lims(y = c(-85, -28))+
facet_wrap(~month, ncol = 2)+
labs(title = paste0('Broullon et al. (2020) DIC clim, depth: ', unique(.x$depth)))
)[[1]]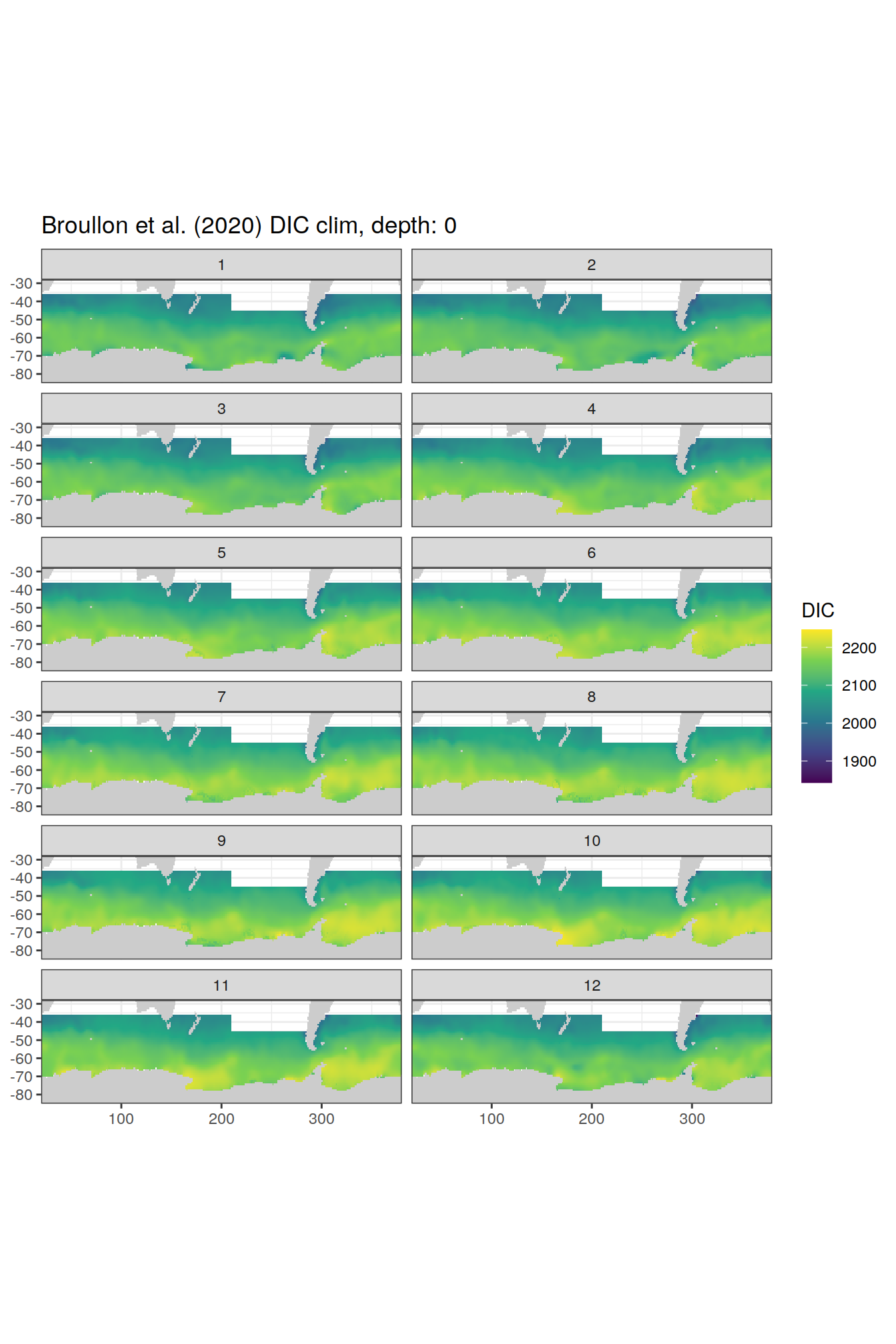
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[2]]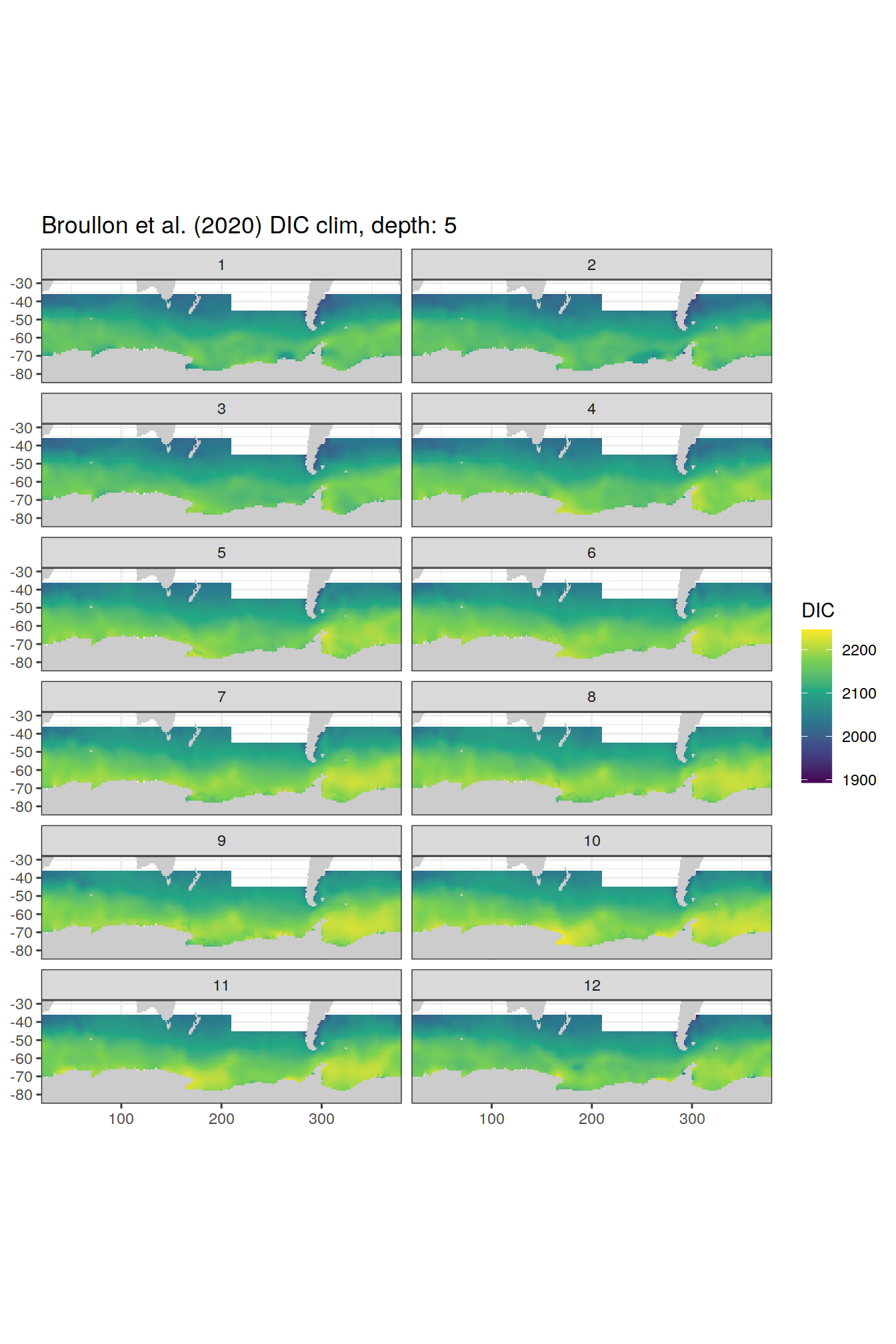
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[3]]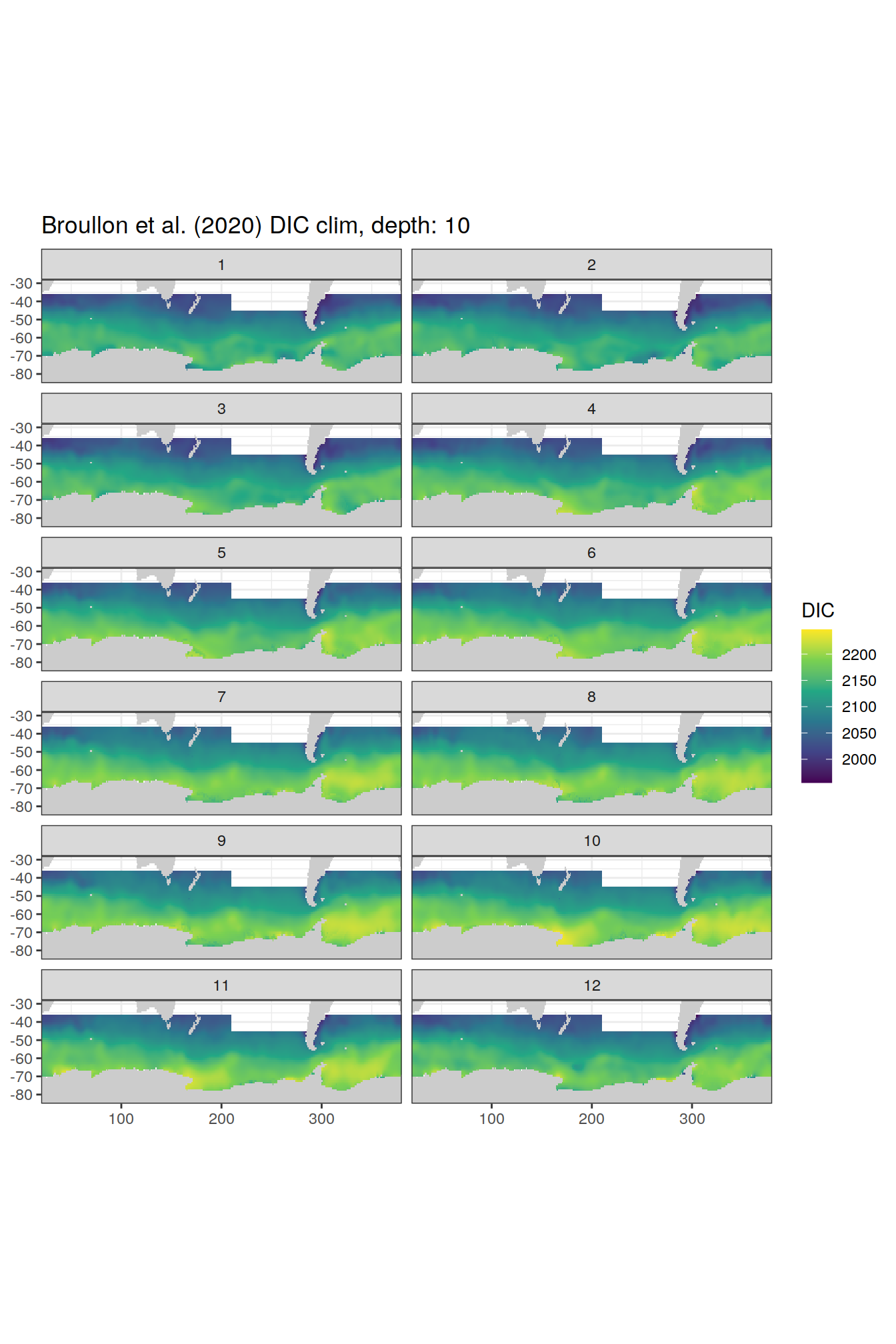
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[4]]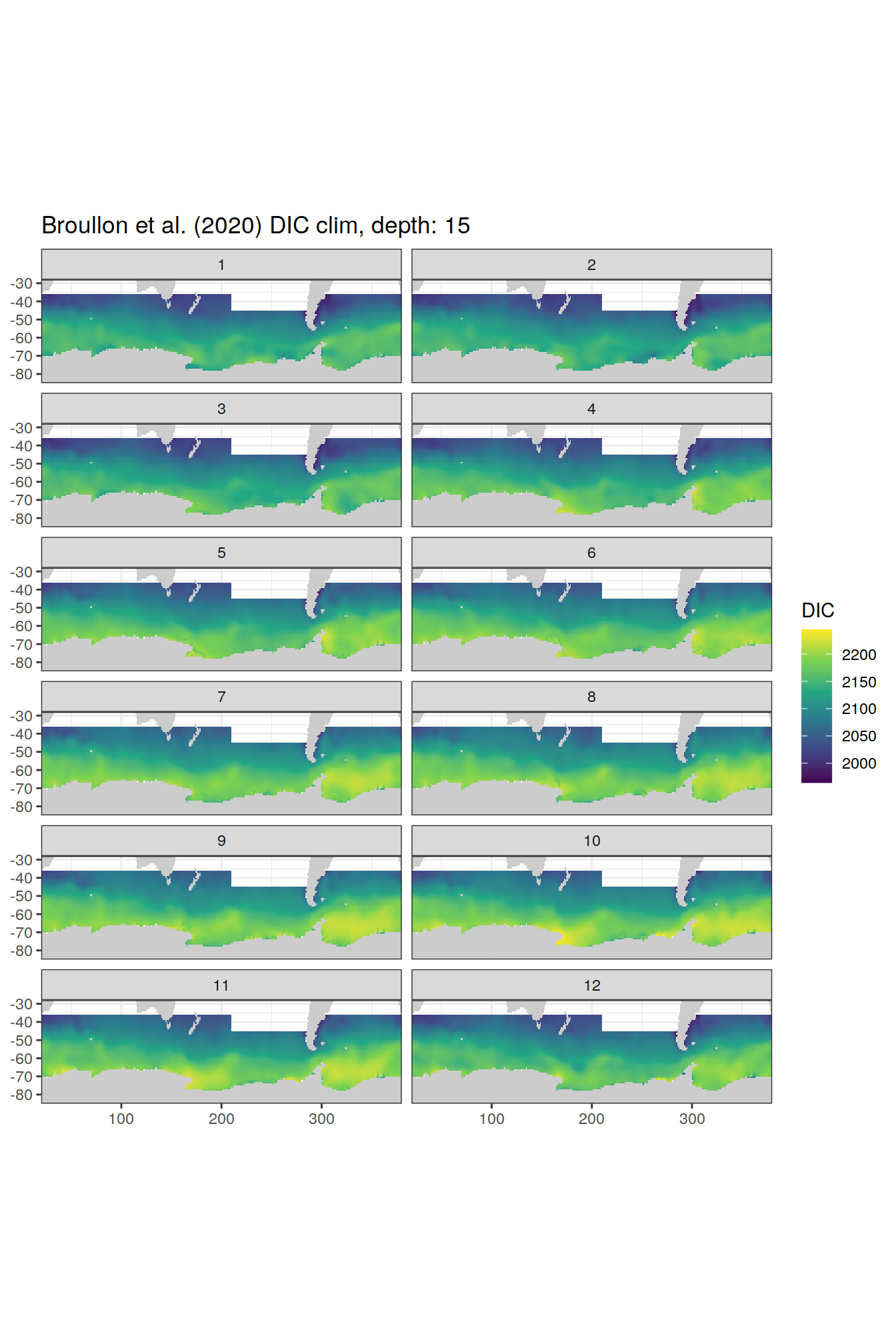
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[5]]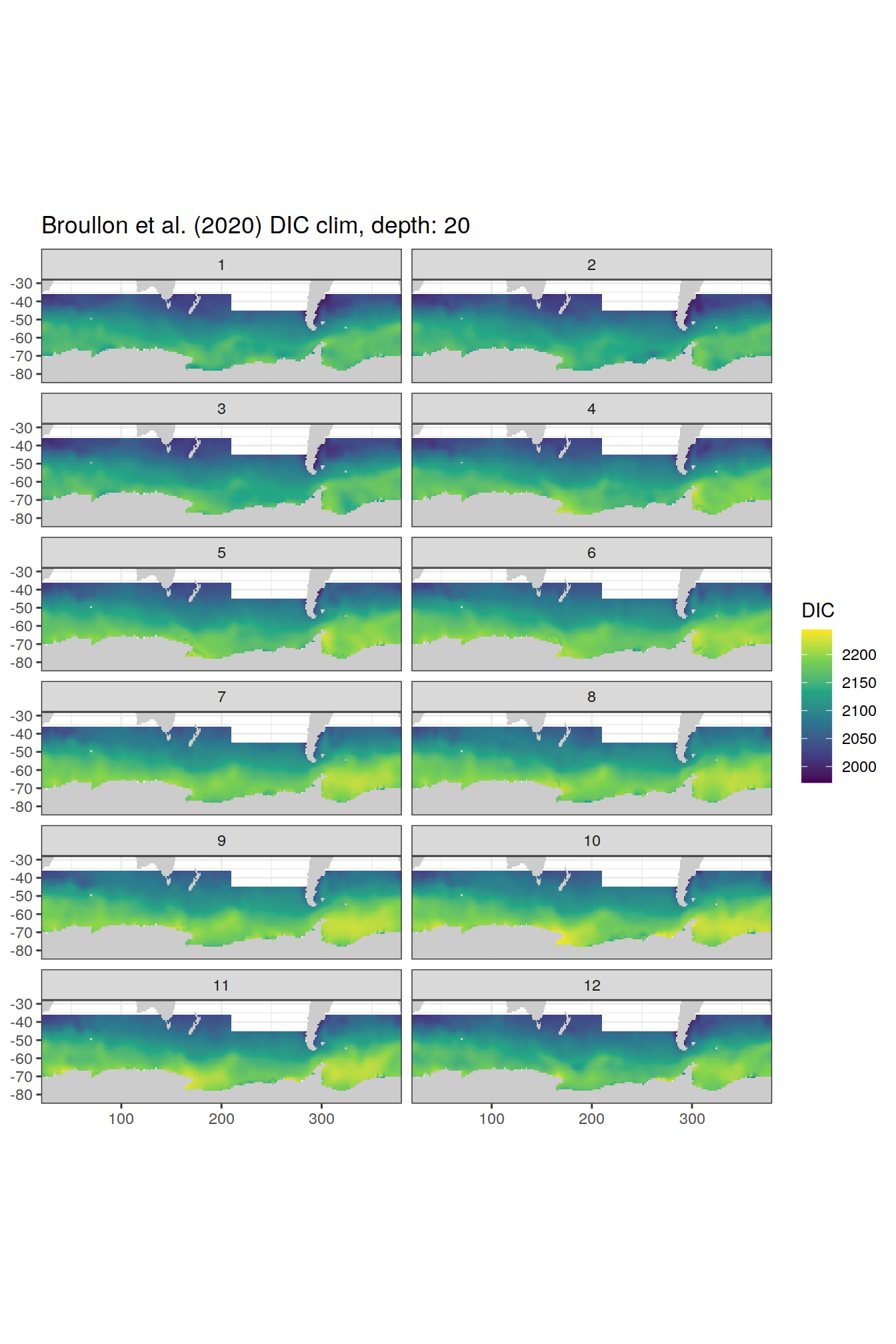
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[6]]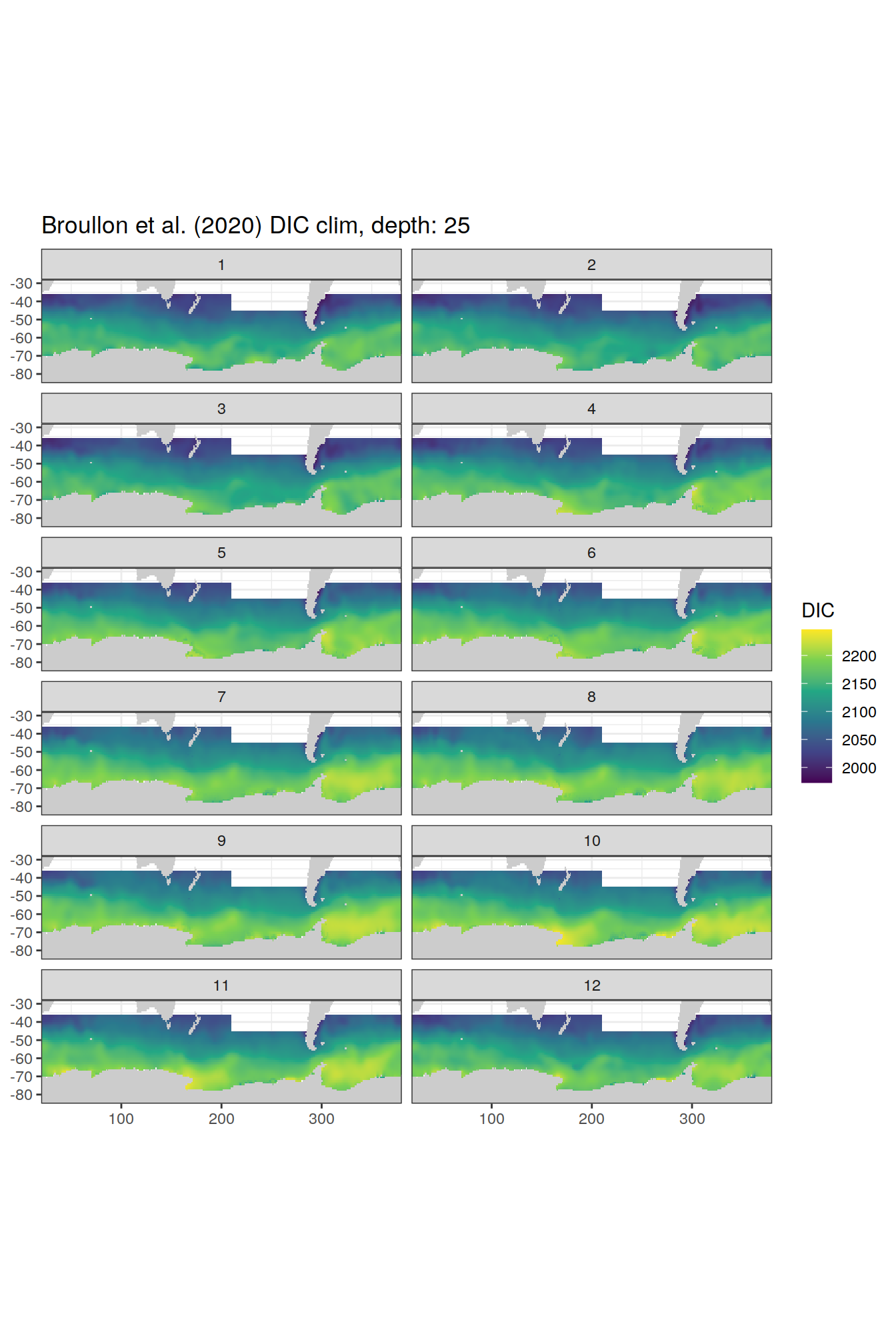
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[7]]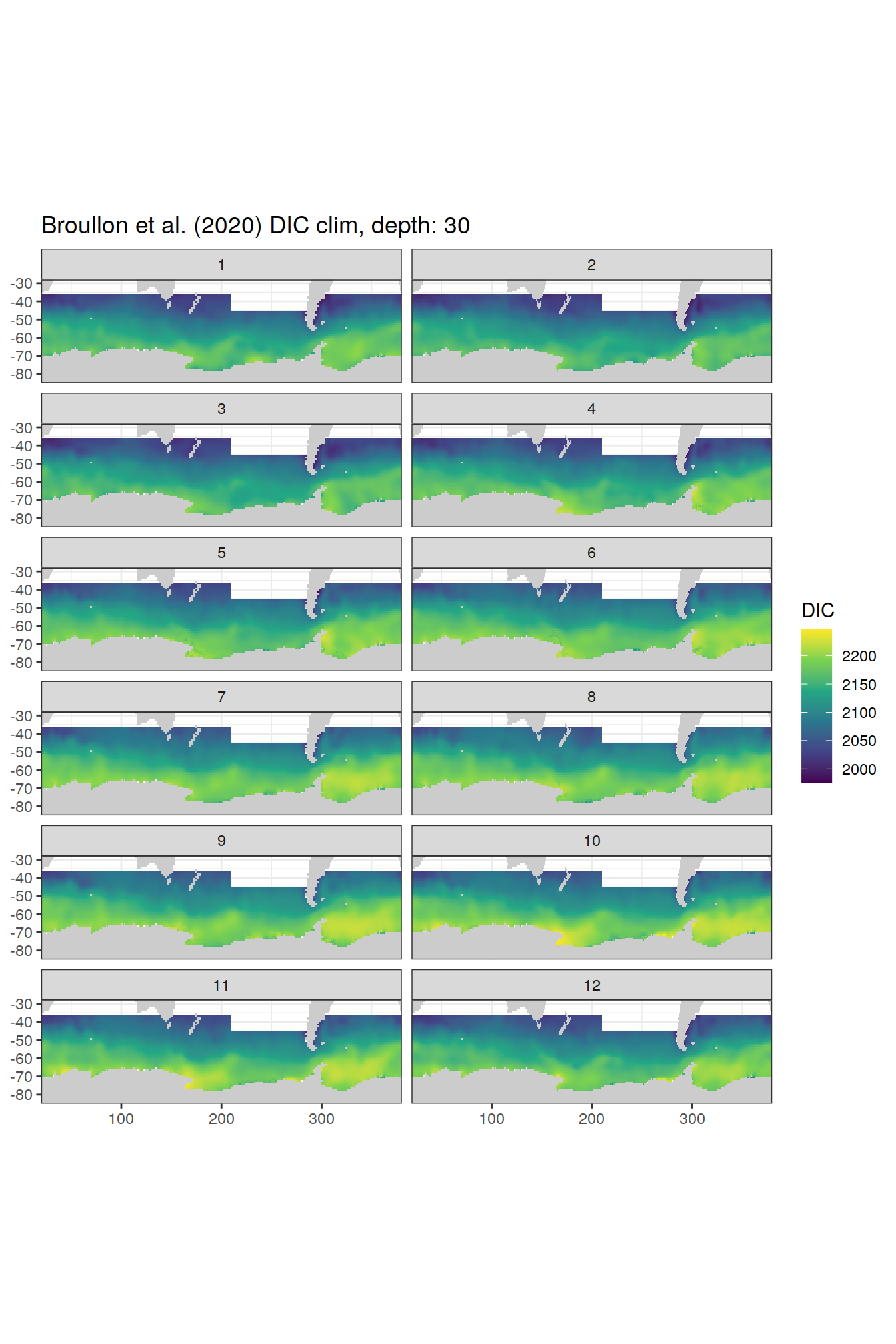
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[8]]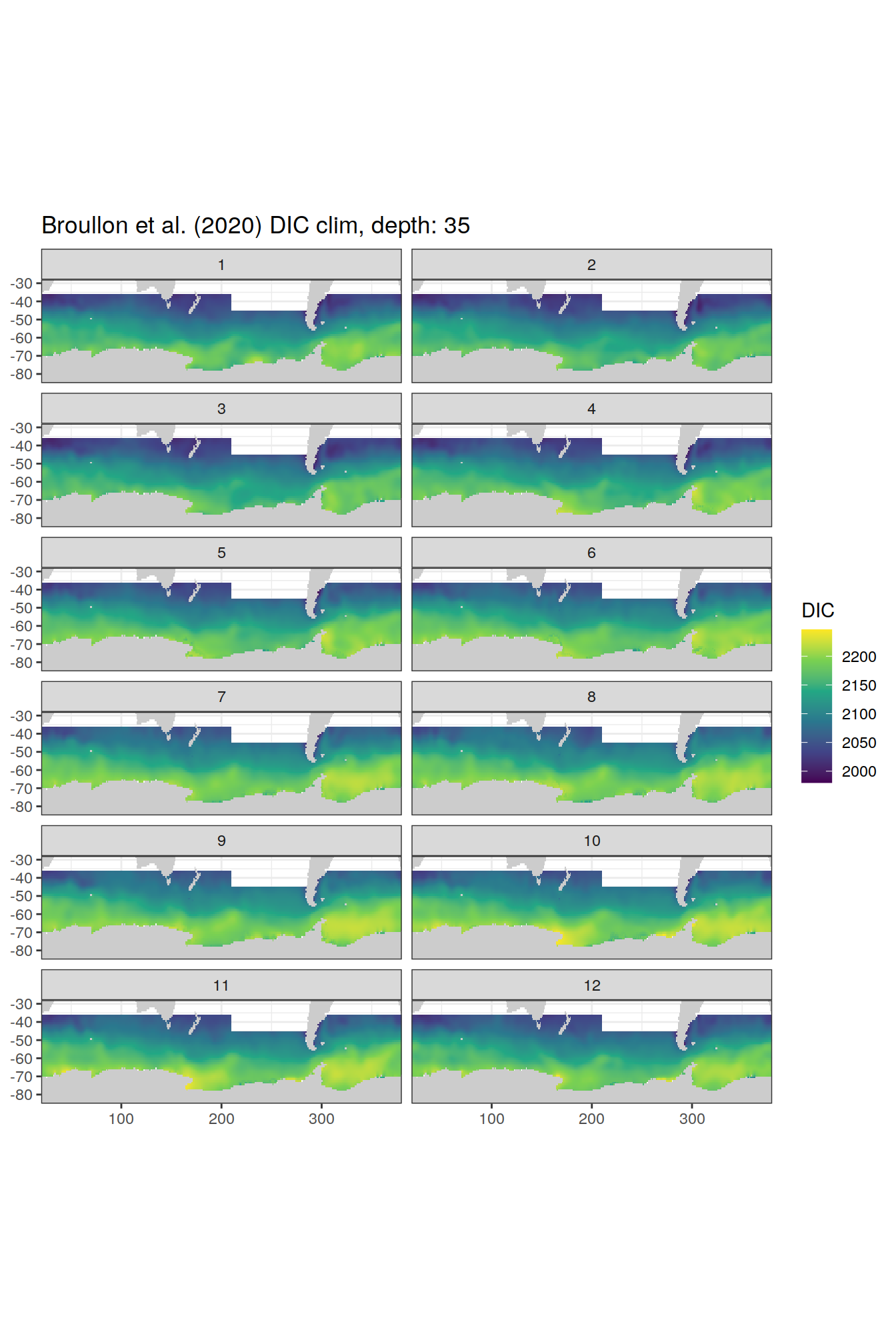
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[9]]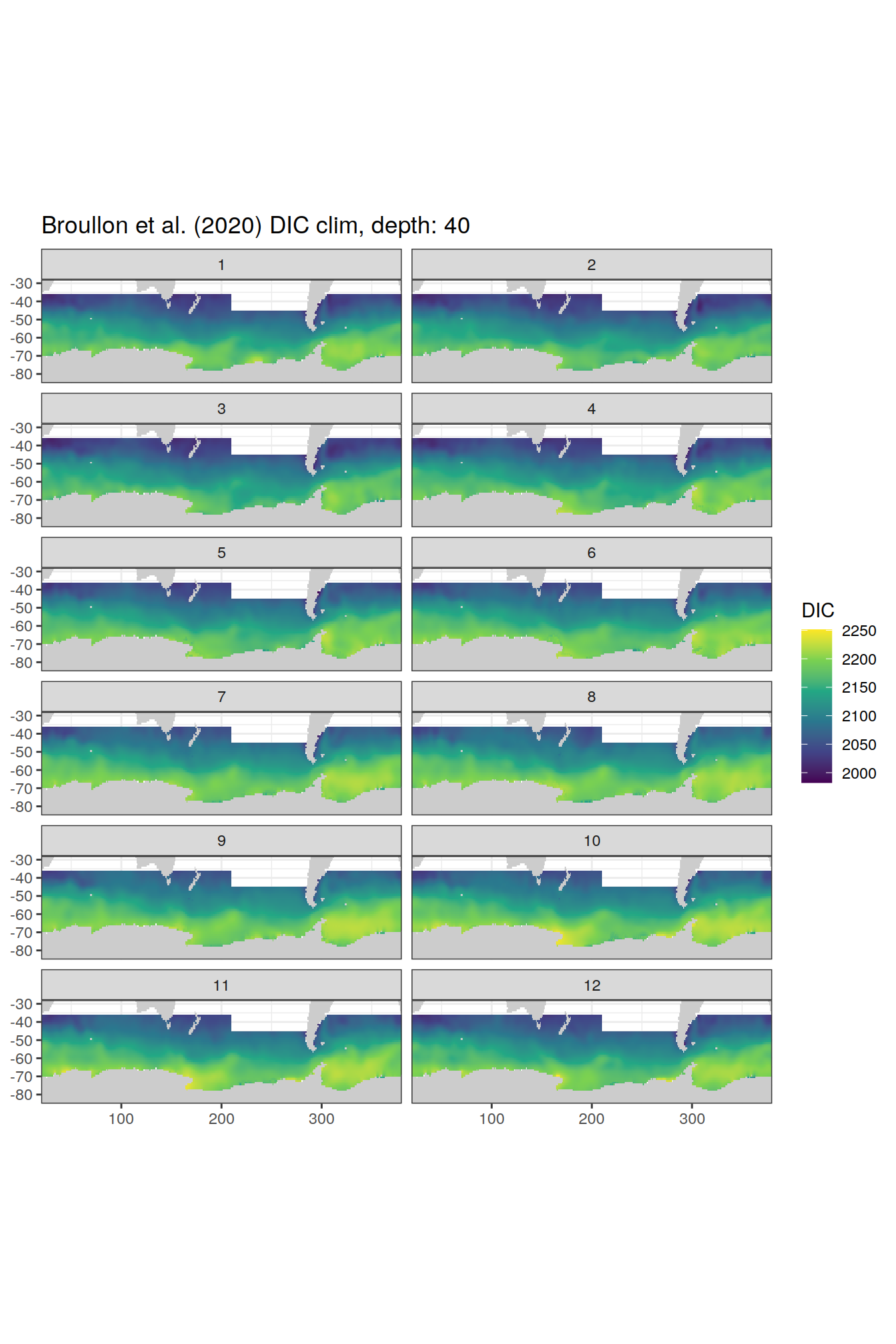
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[10]]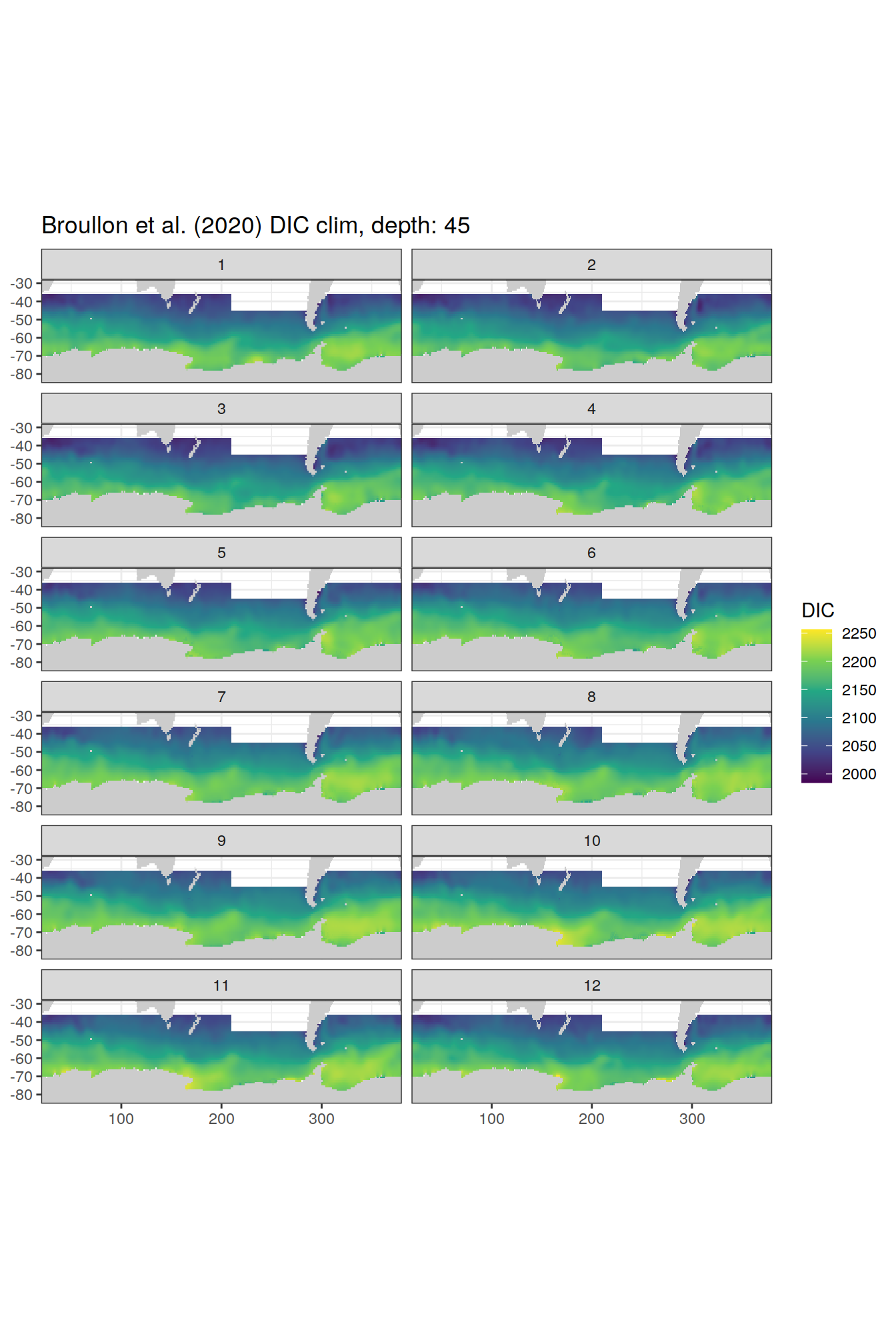
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[11]]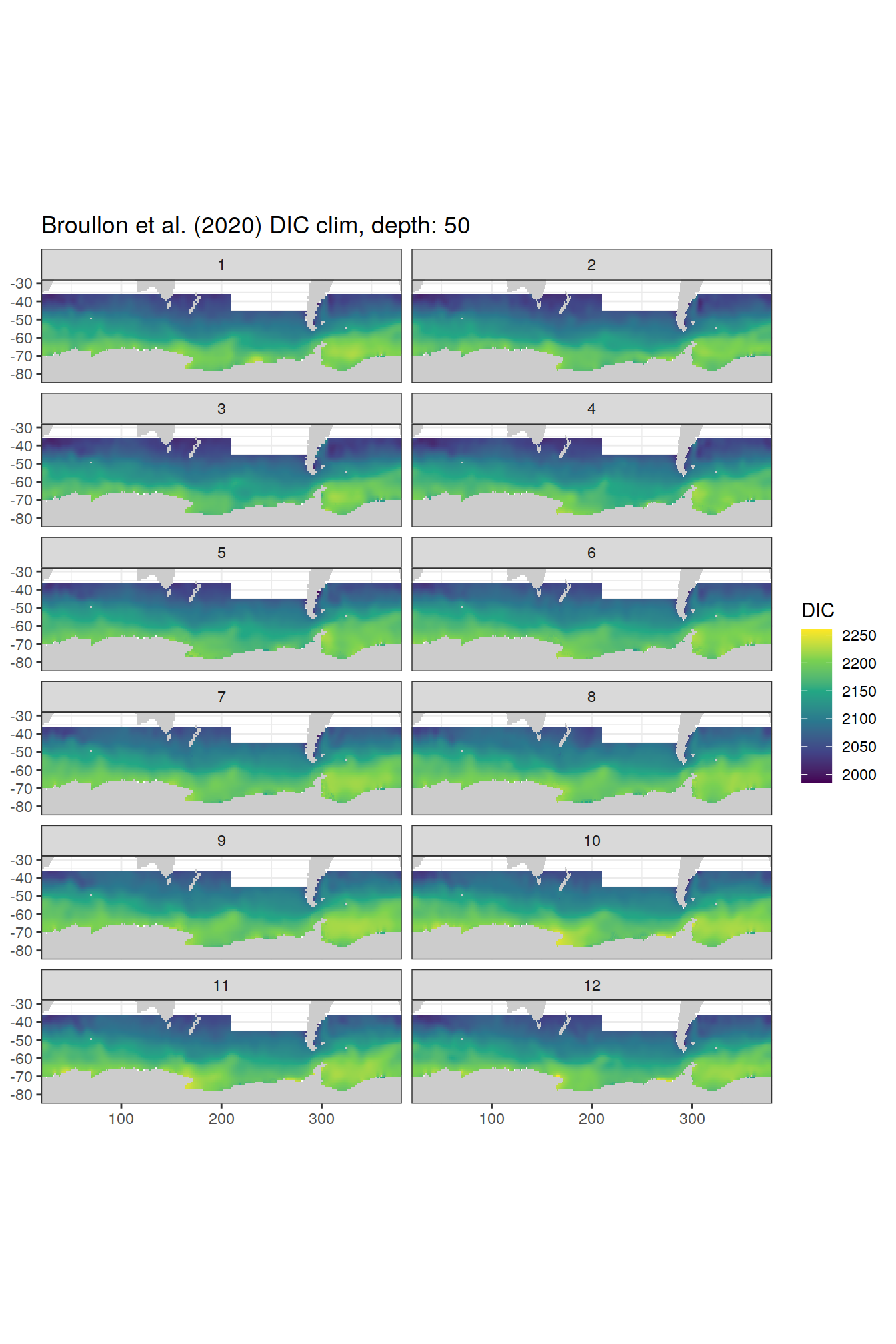
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[12]]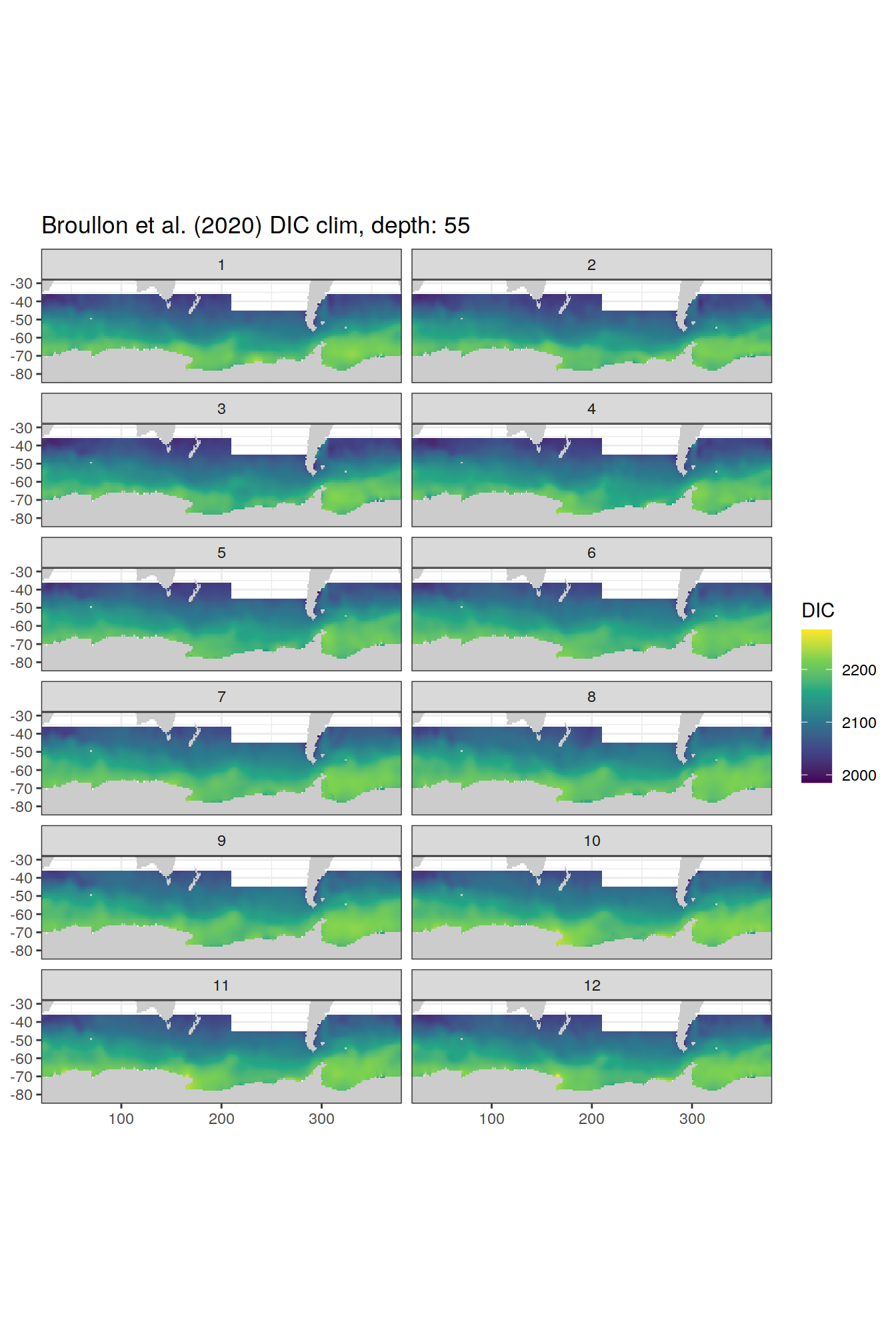
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[13]]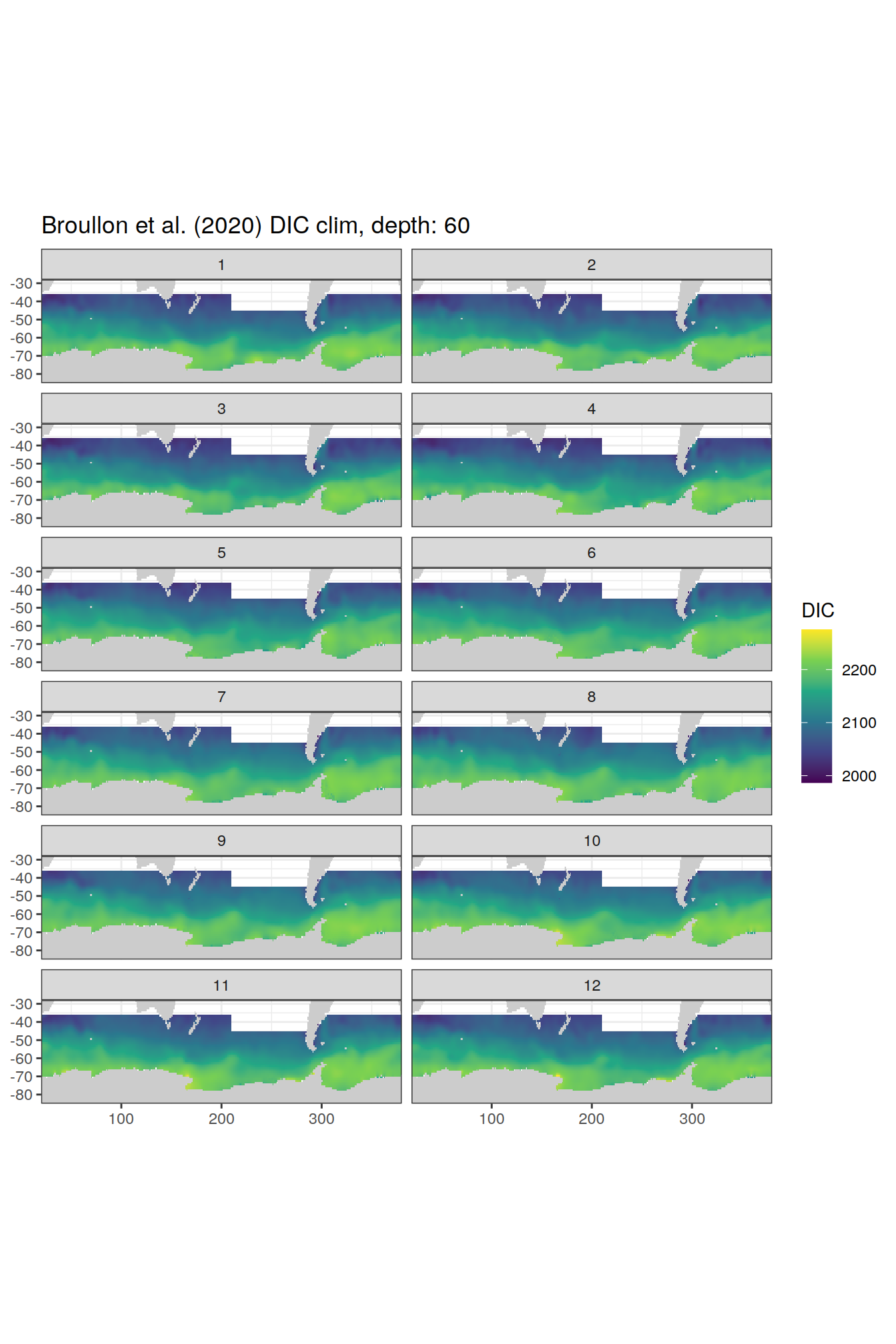
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[14]]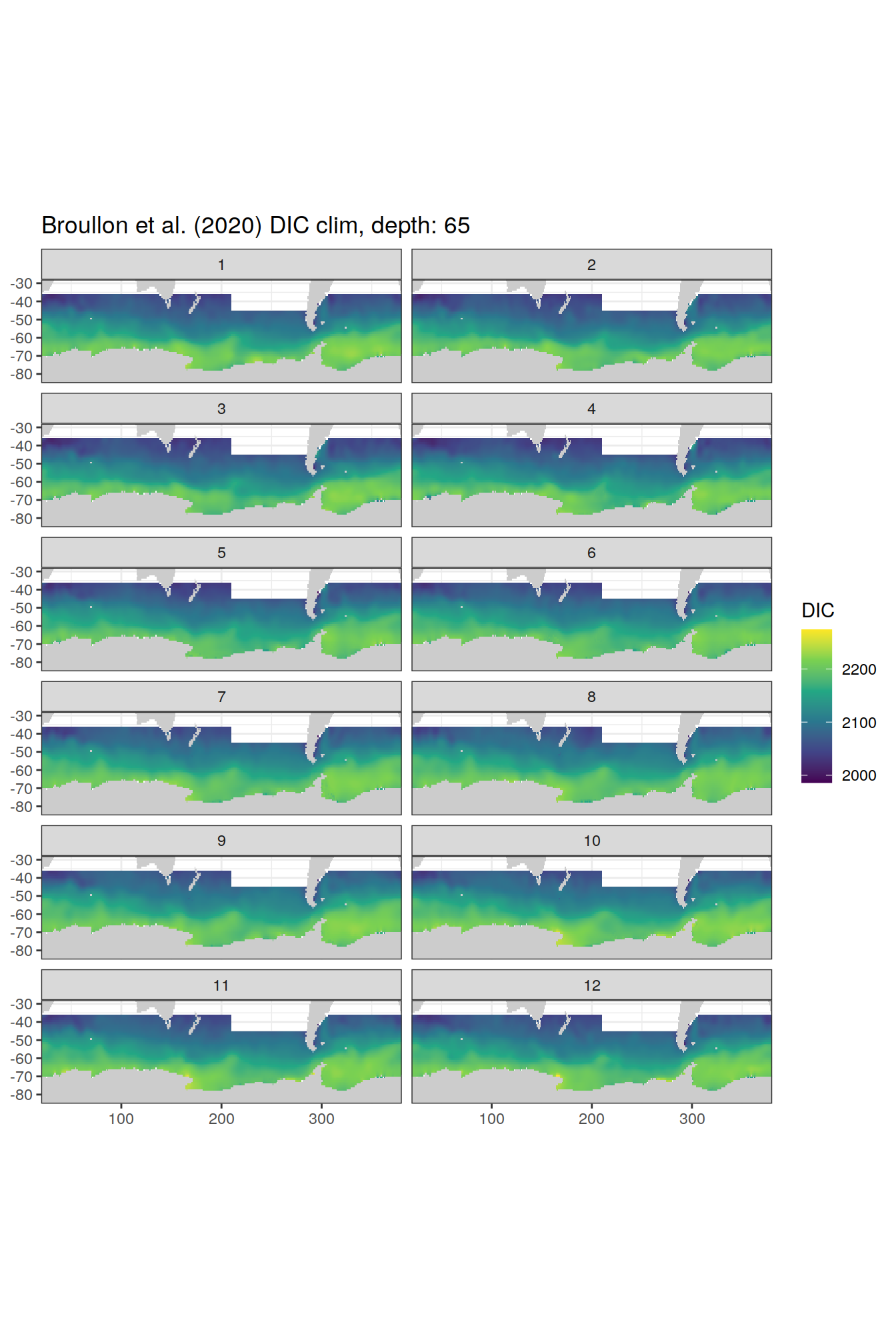
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[15]]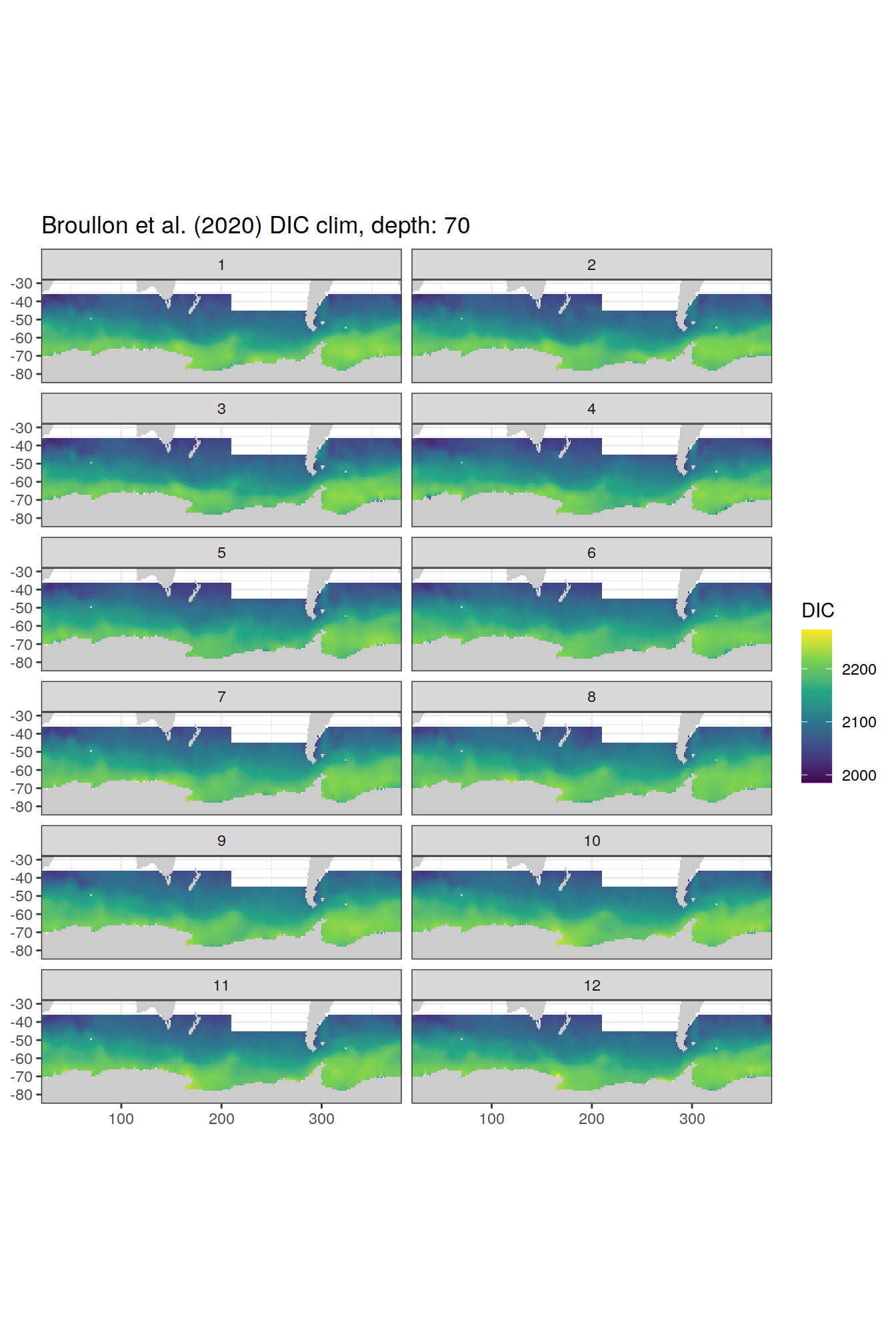
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[16]]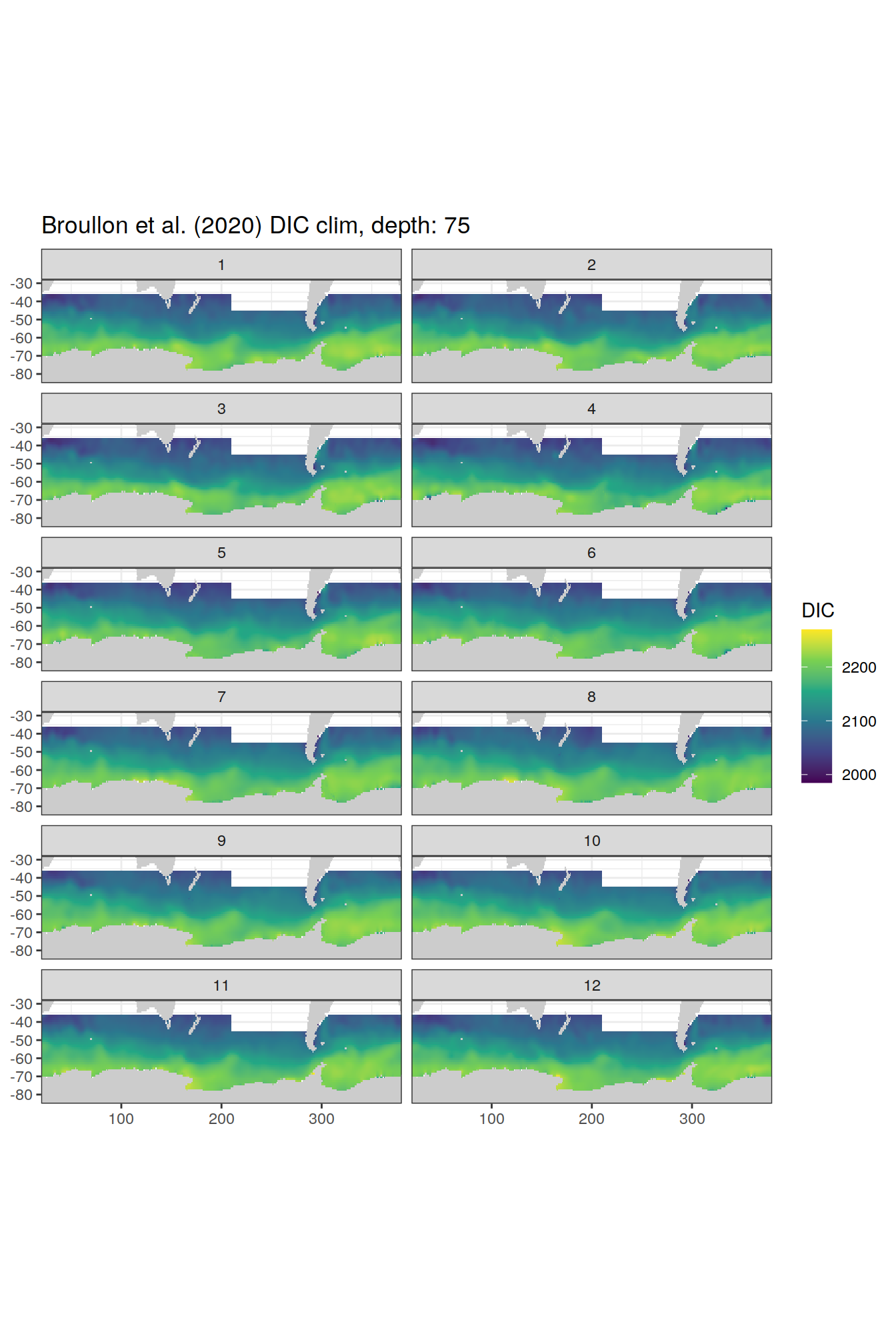
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[17]]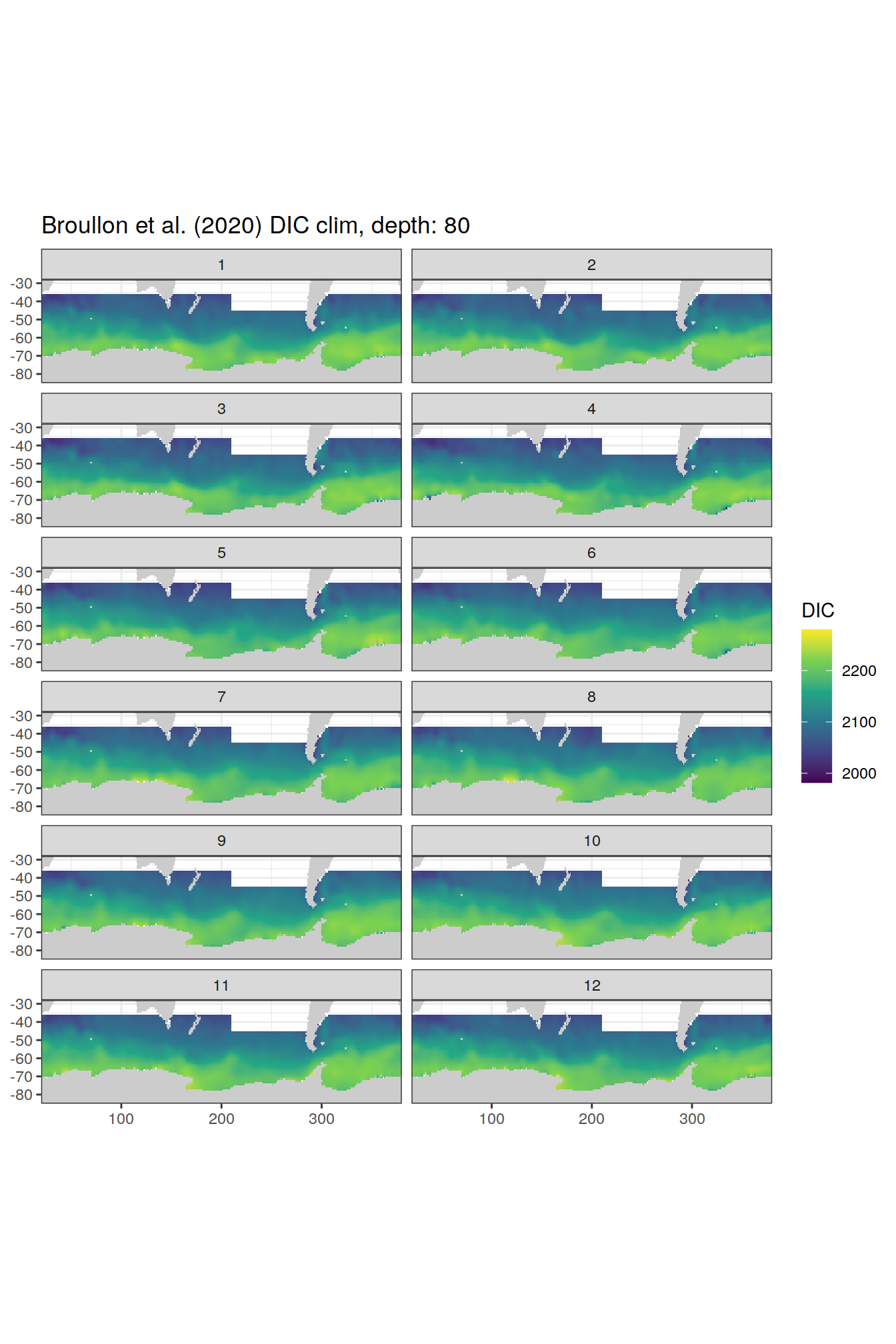
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[18]]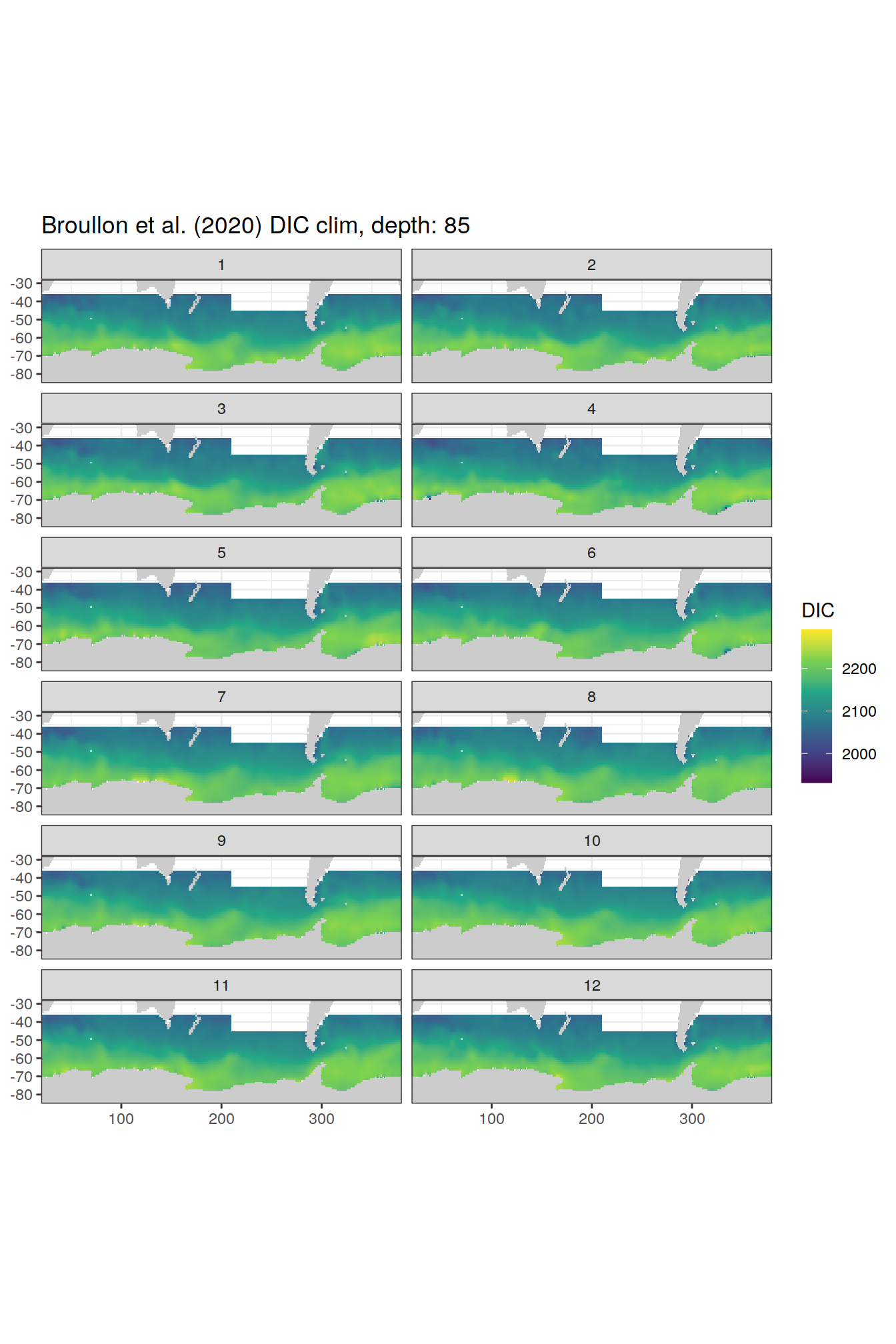
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[19]]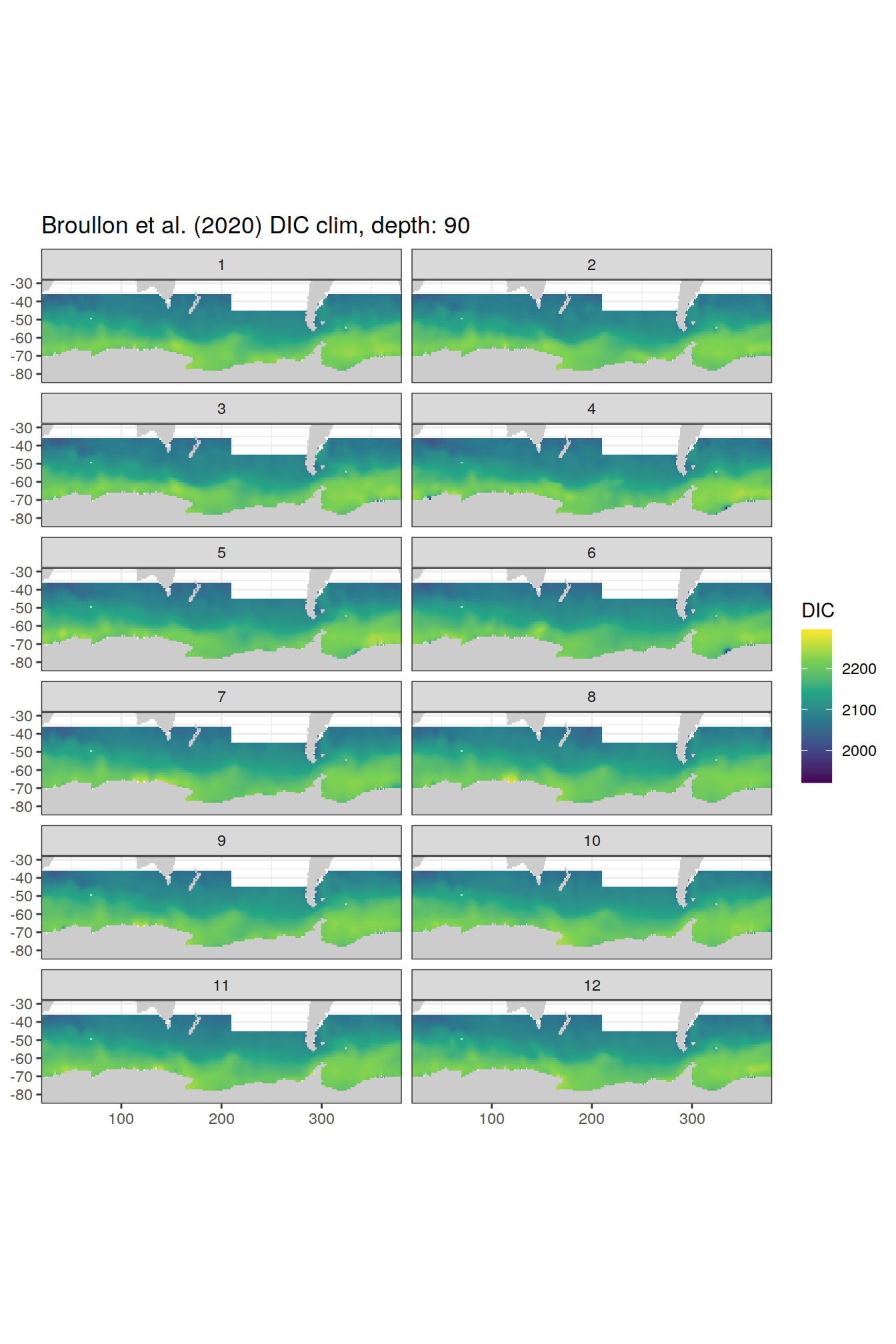
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[20]]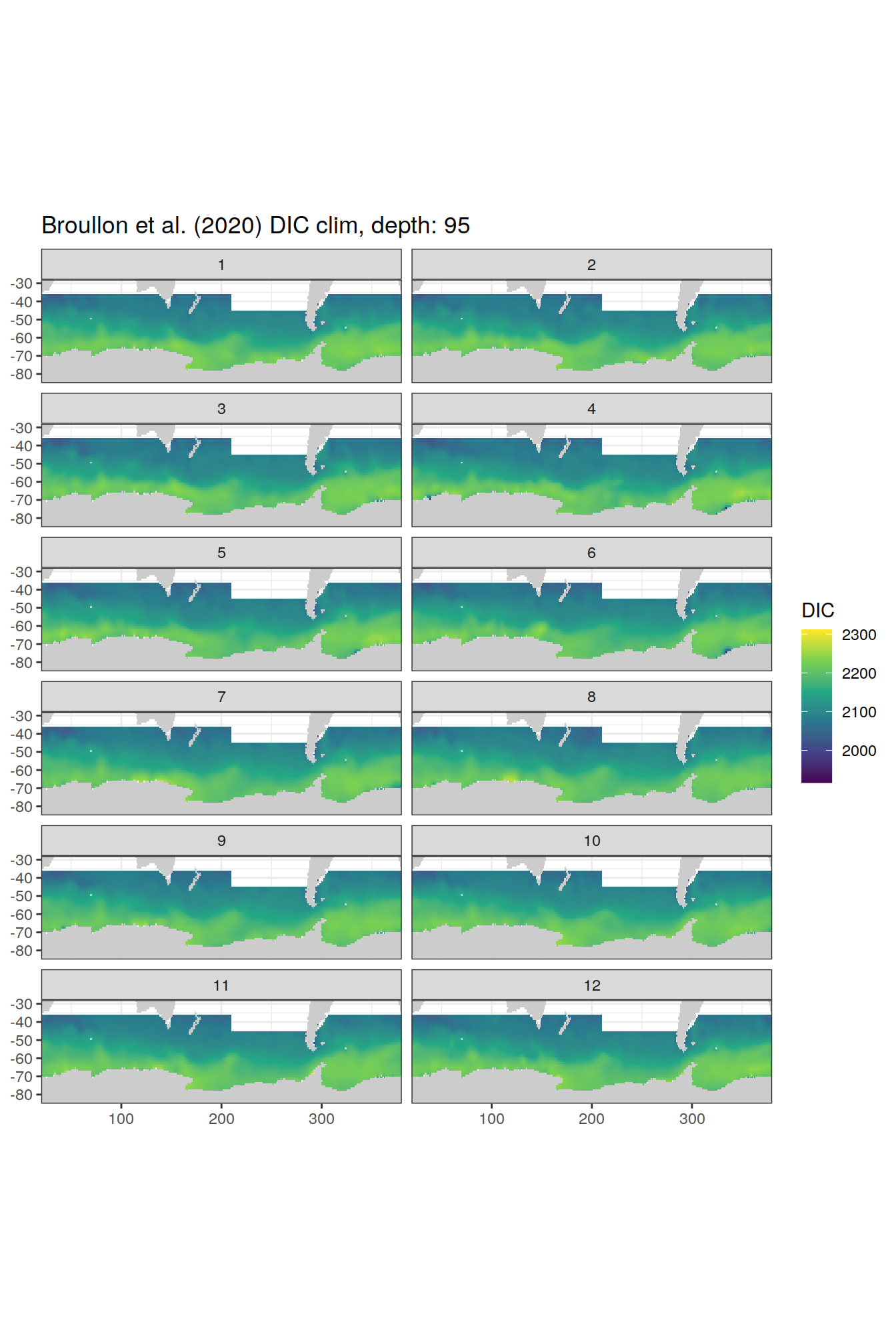
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[21]]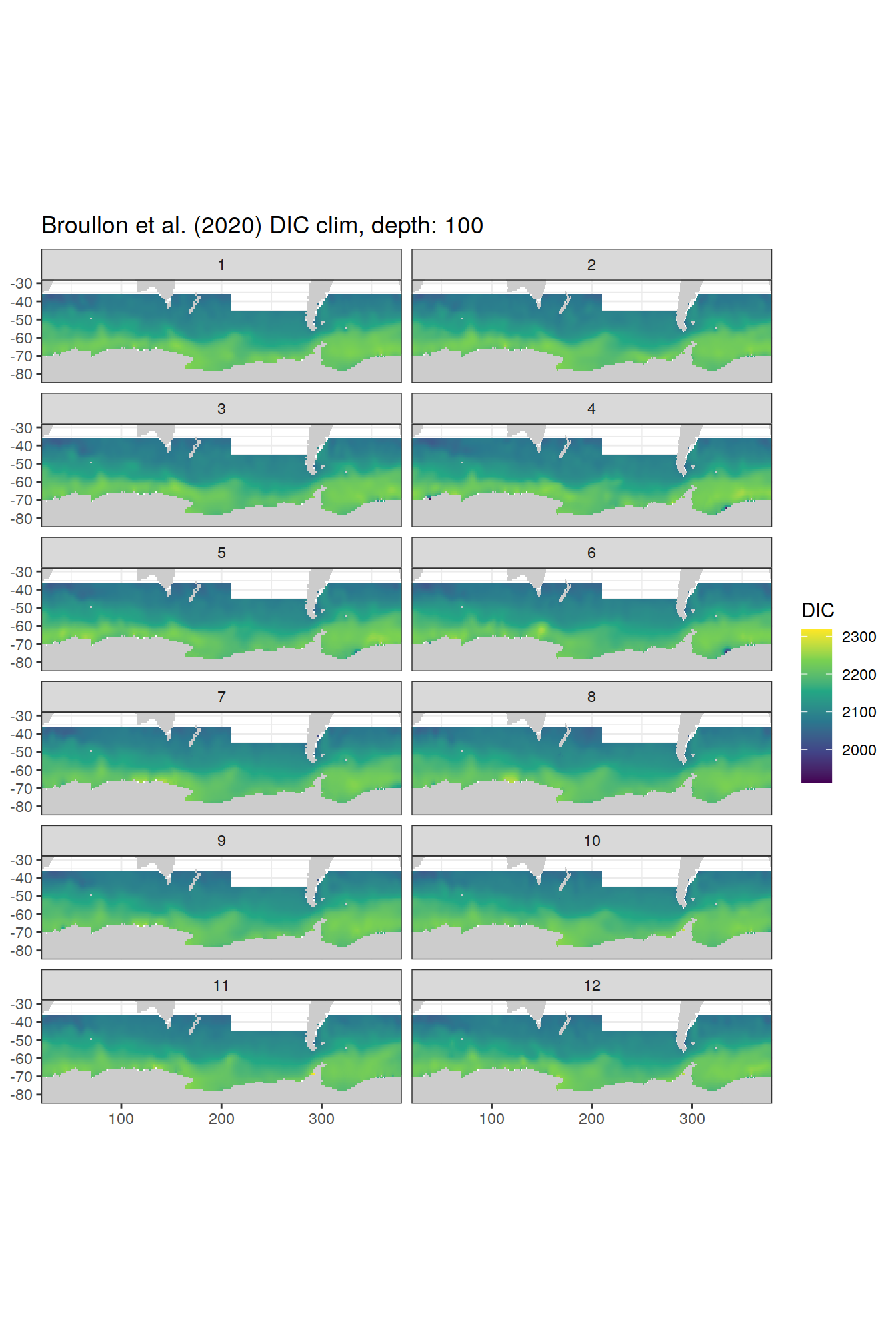
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[22]]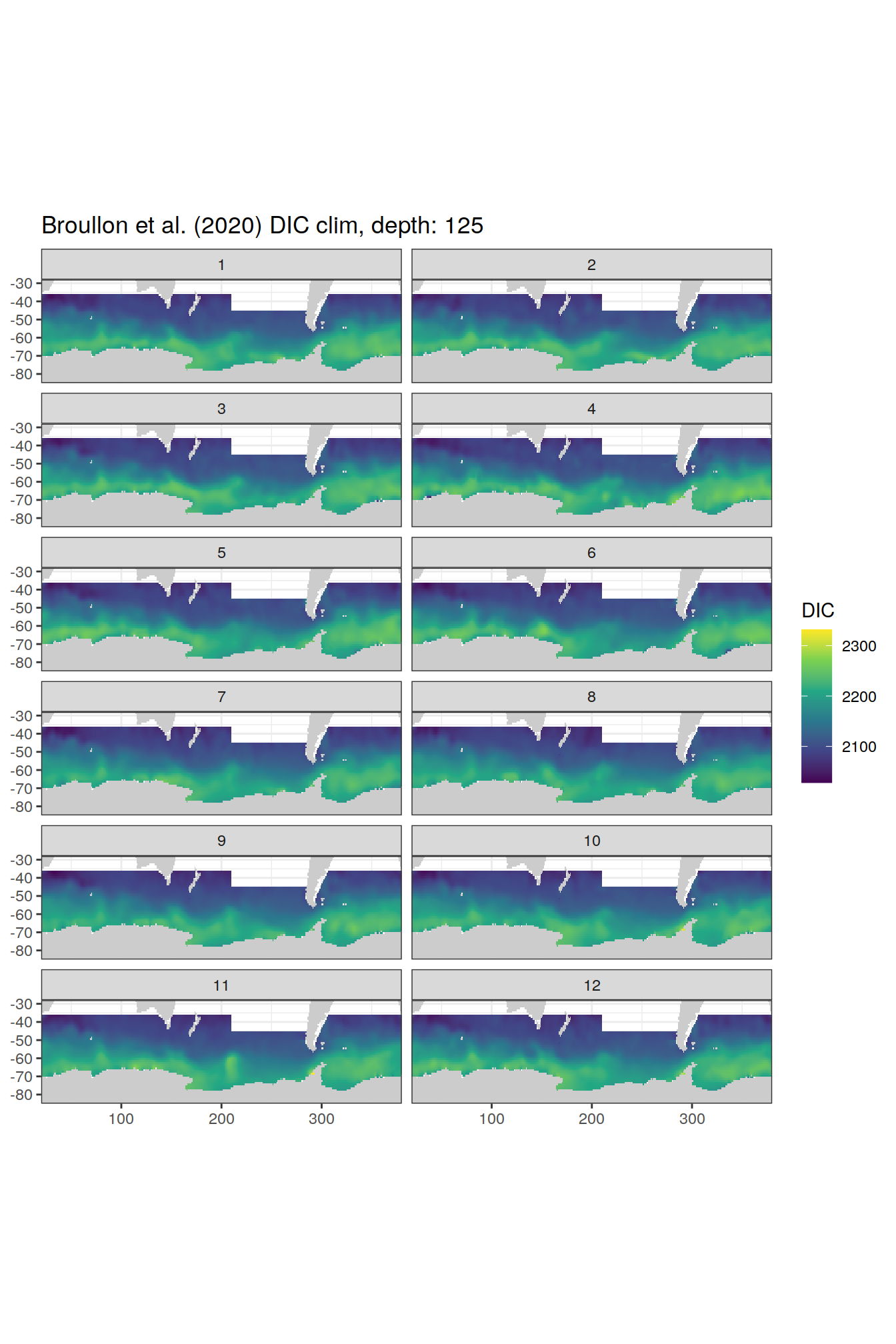
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[23]]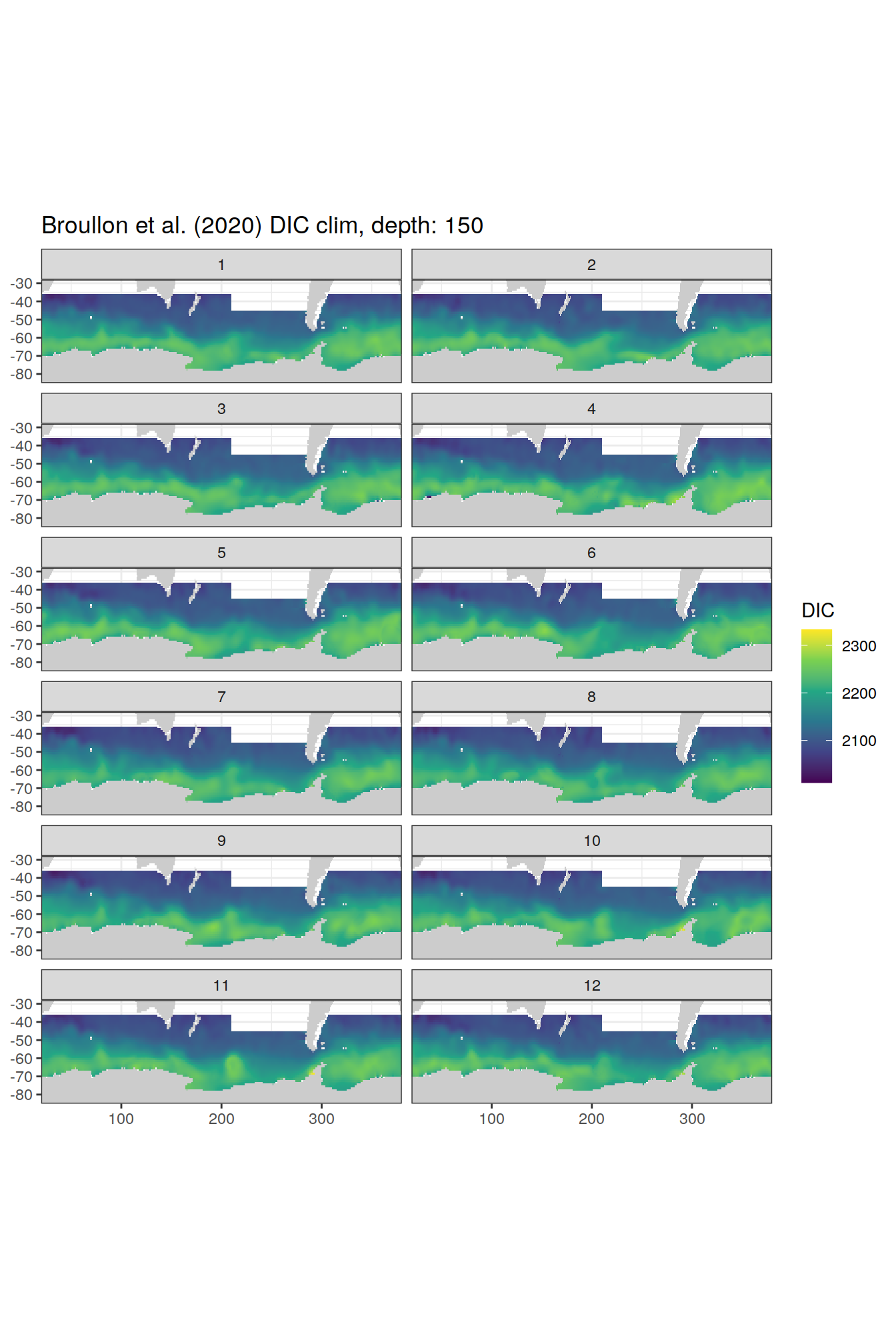
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[24]]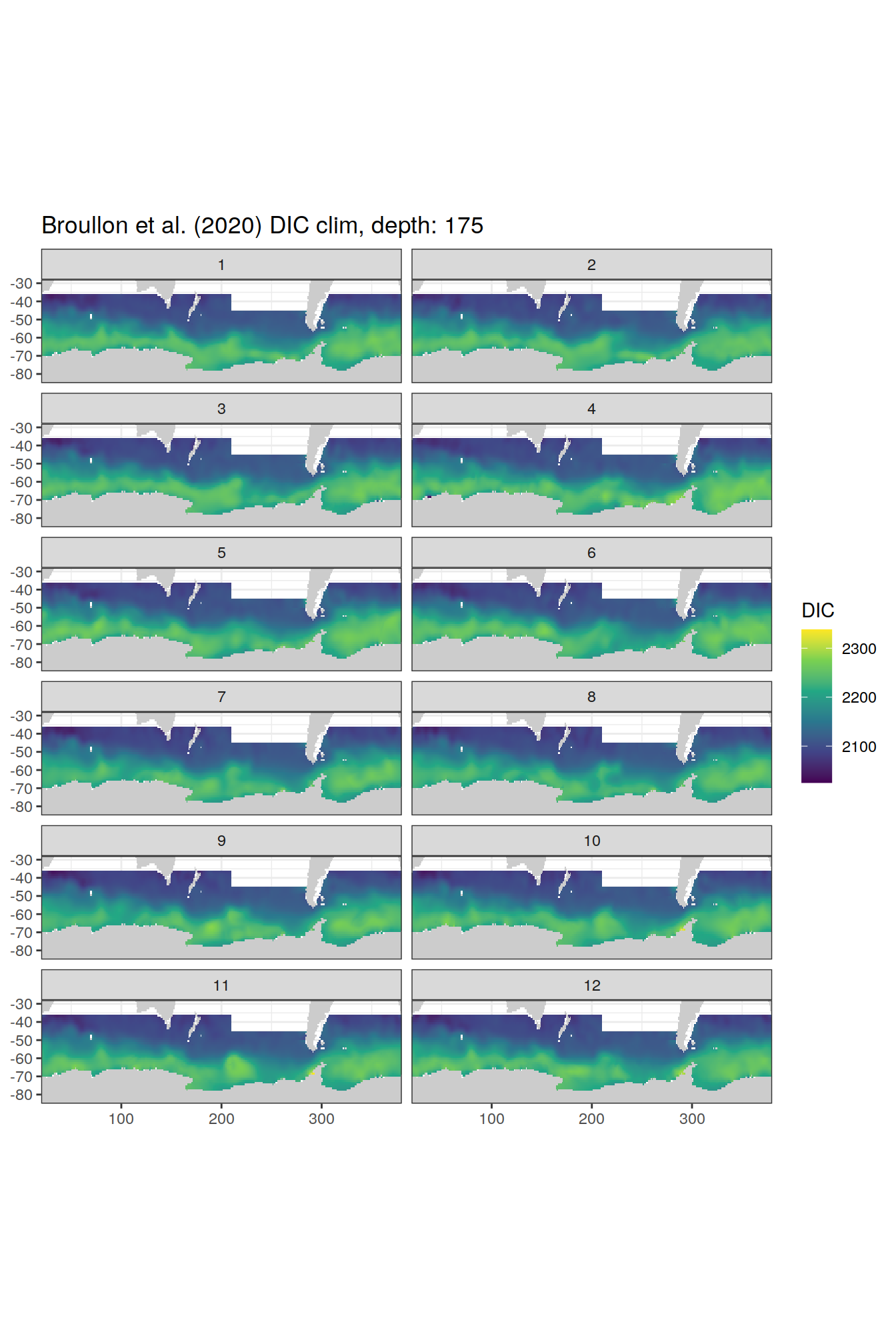
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[25]]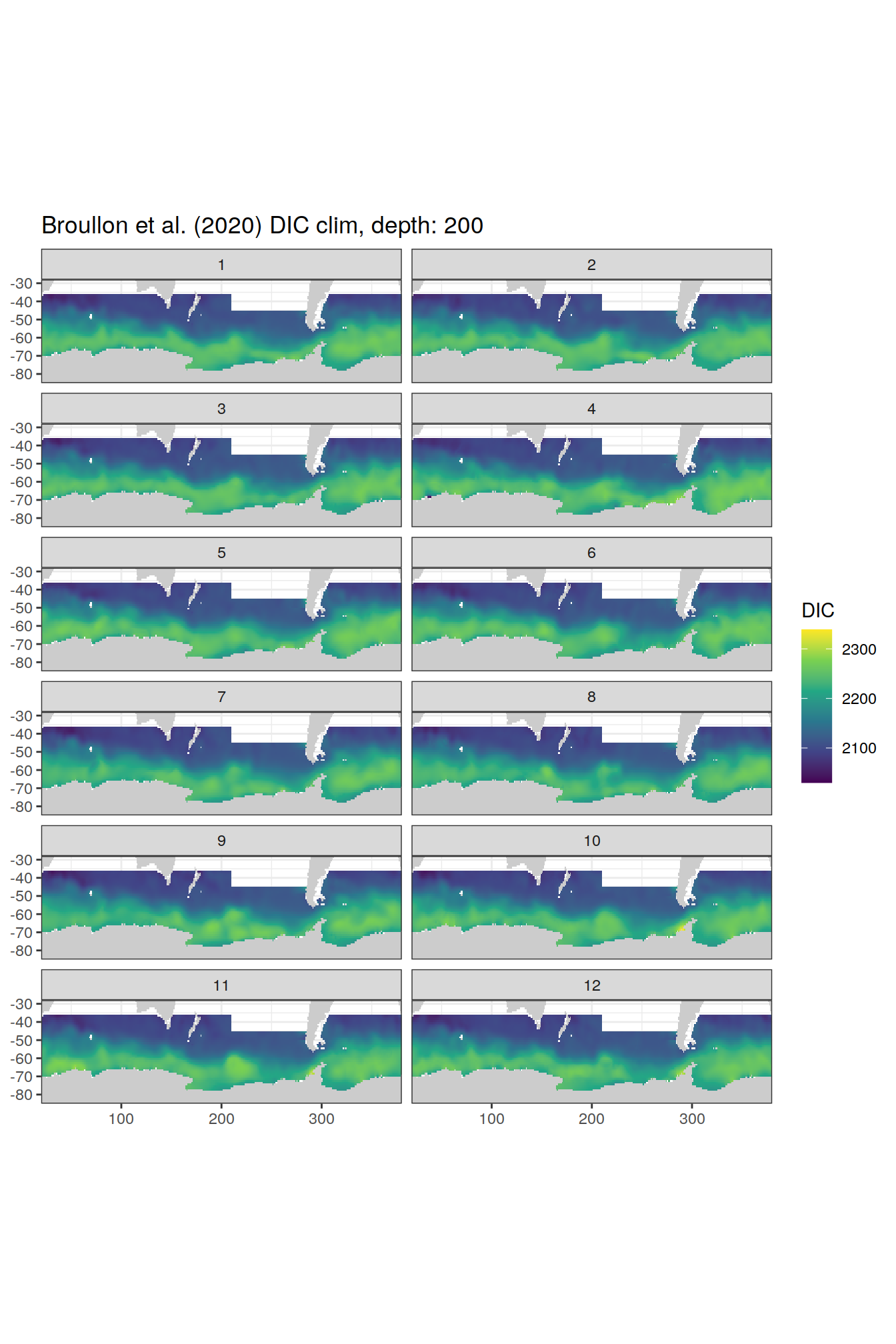
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[26]]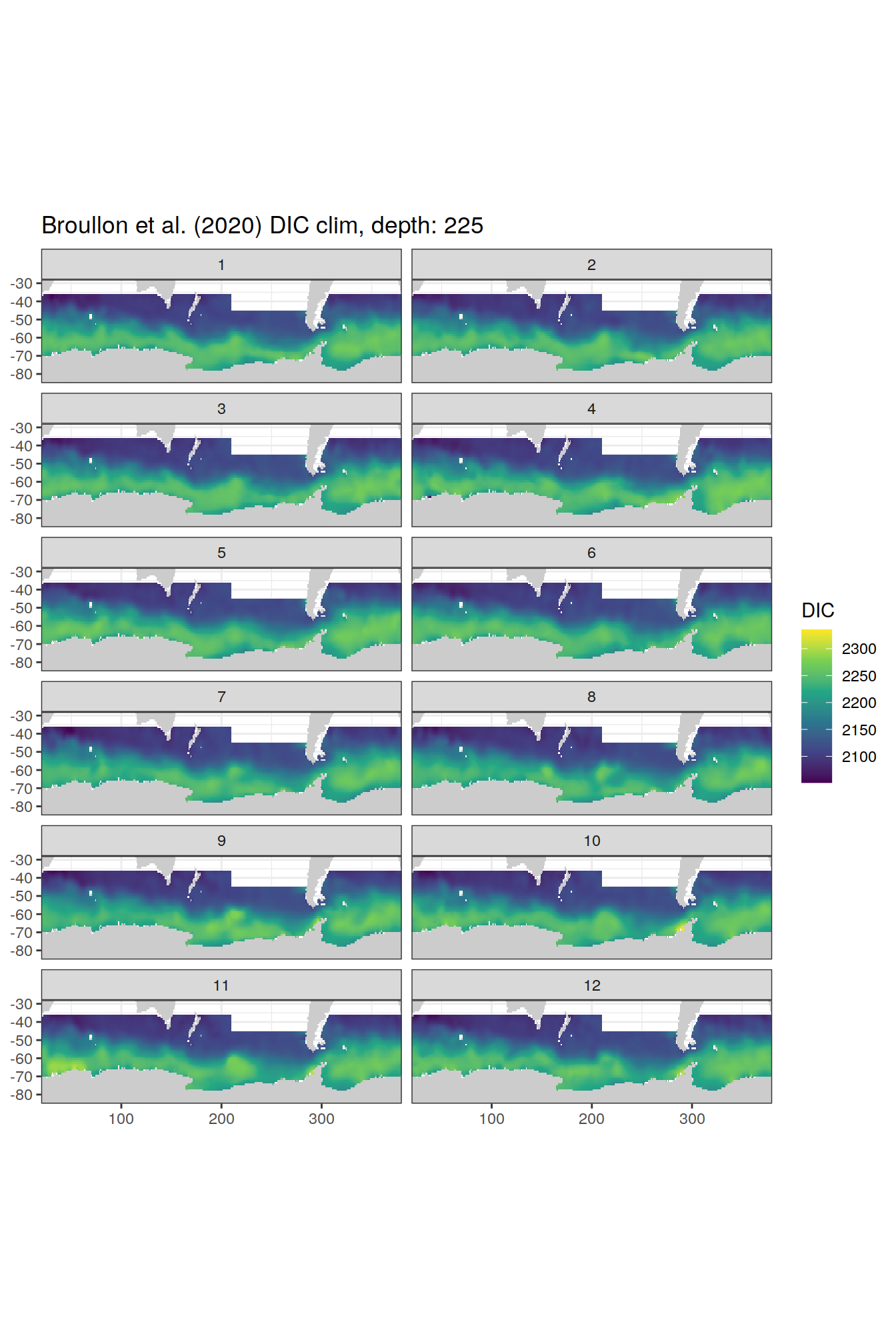
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[27]]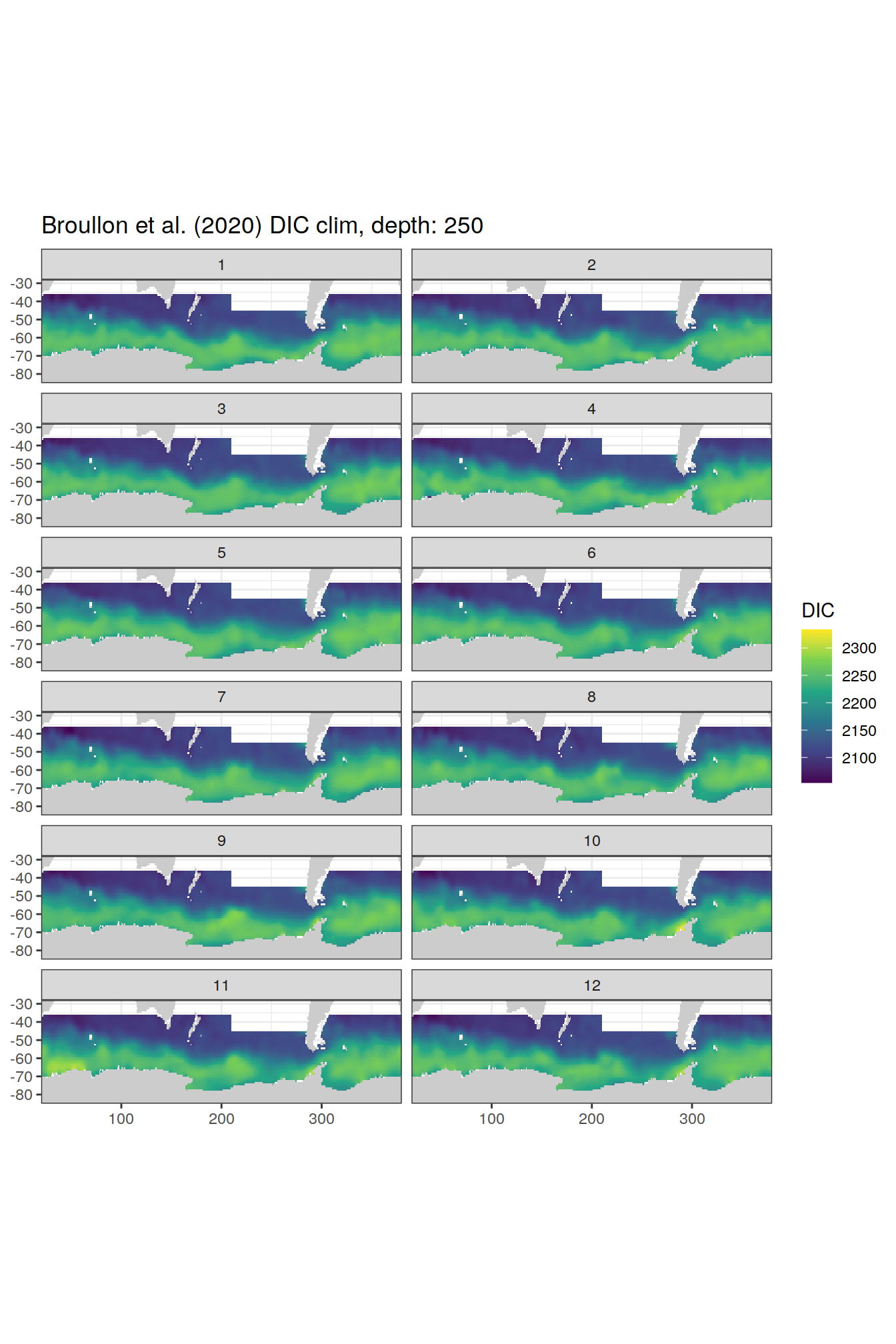
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[28]]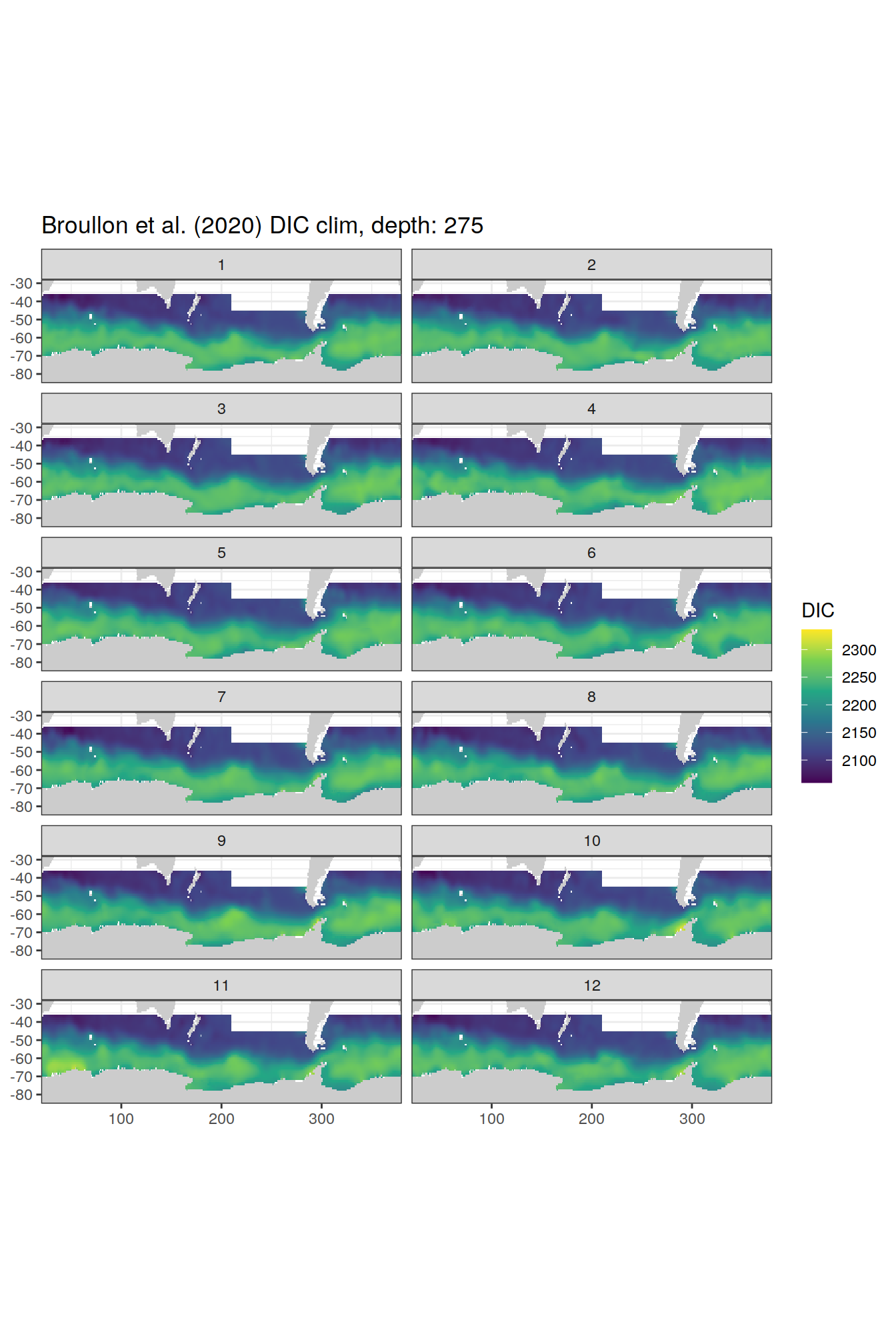
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[29]]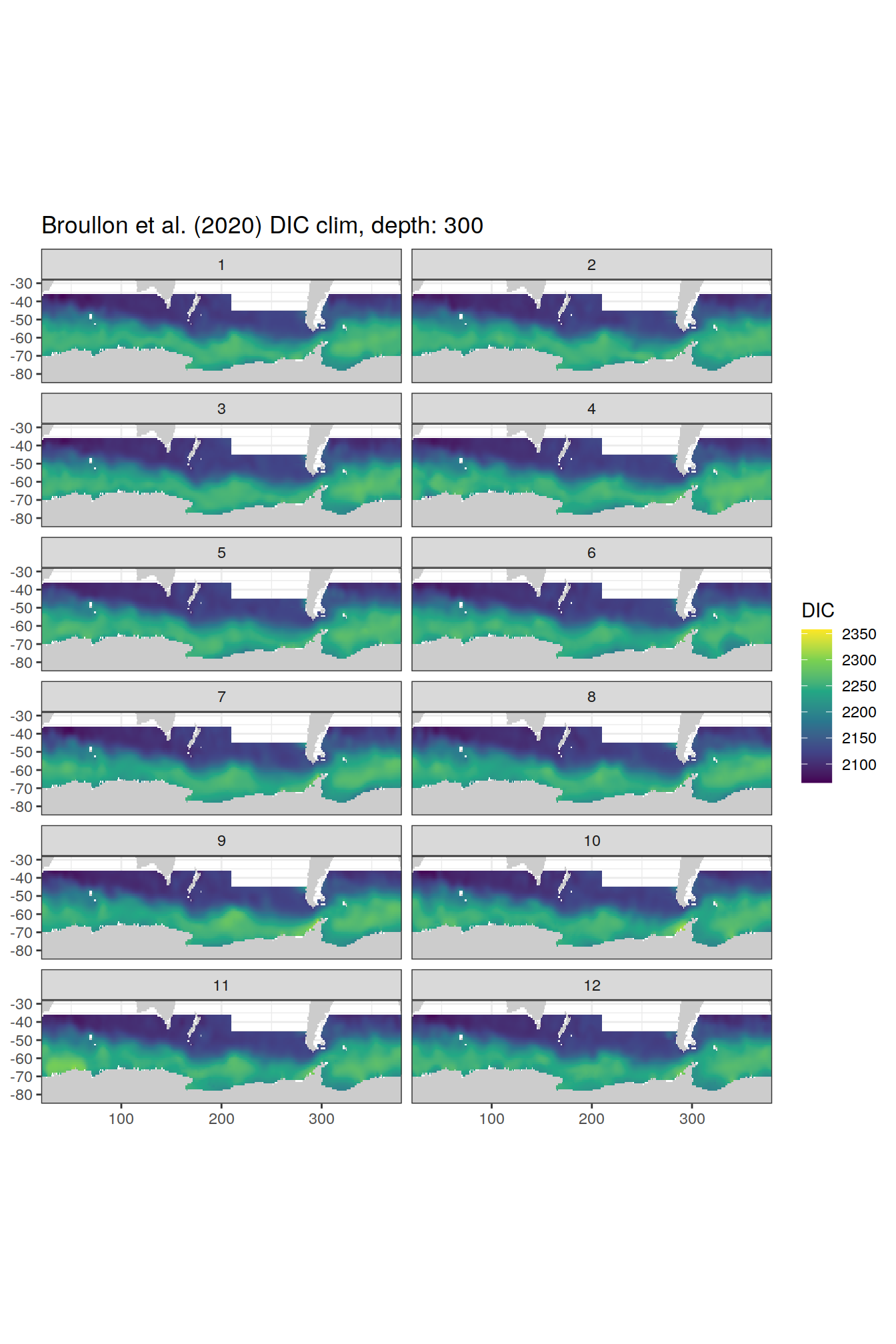
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[30]]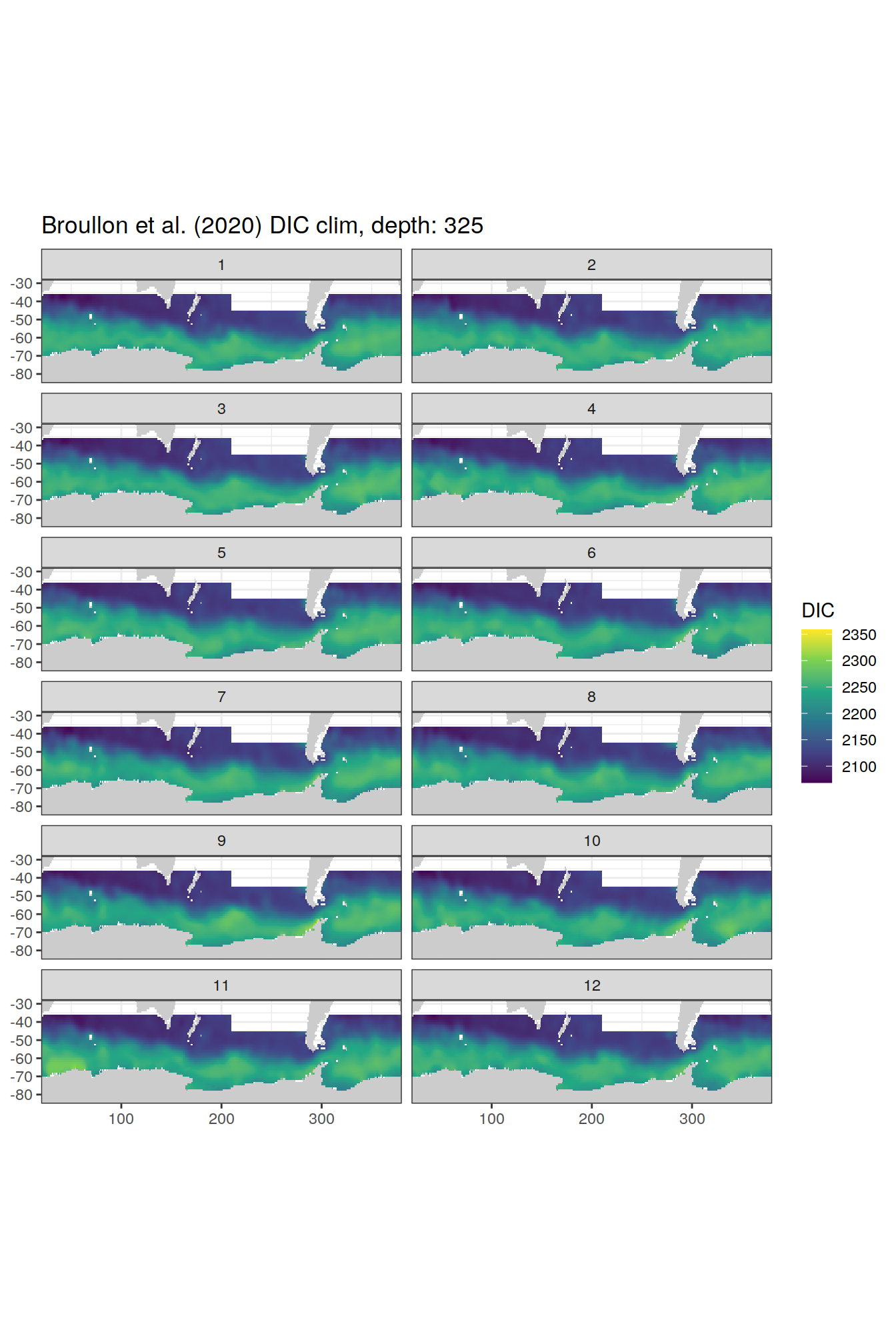
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[31]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[32]]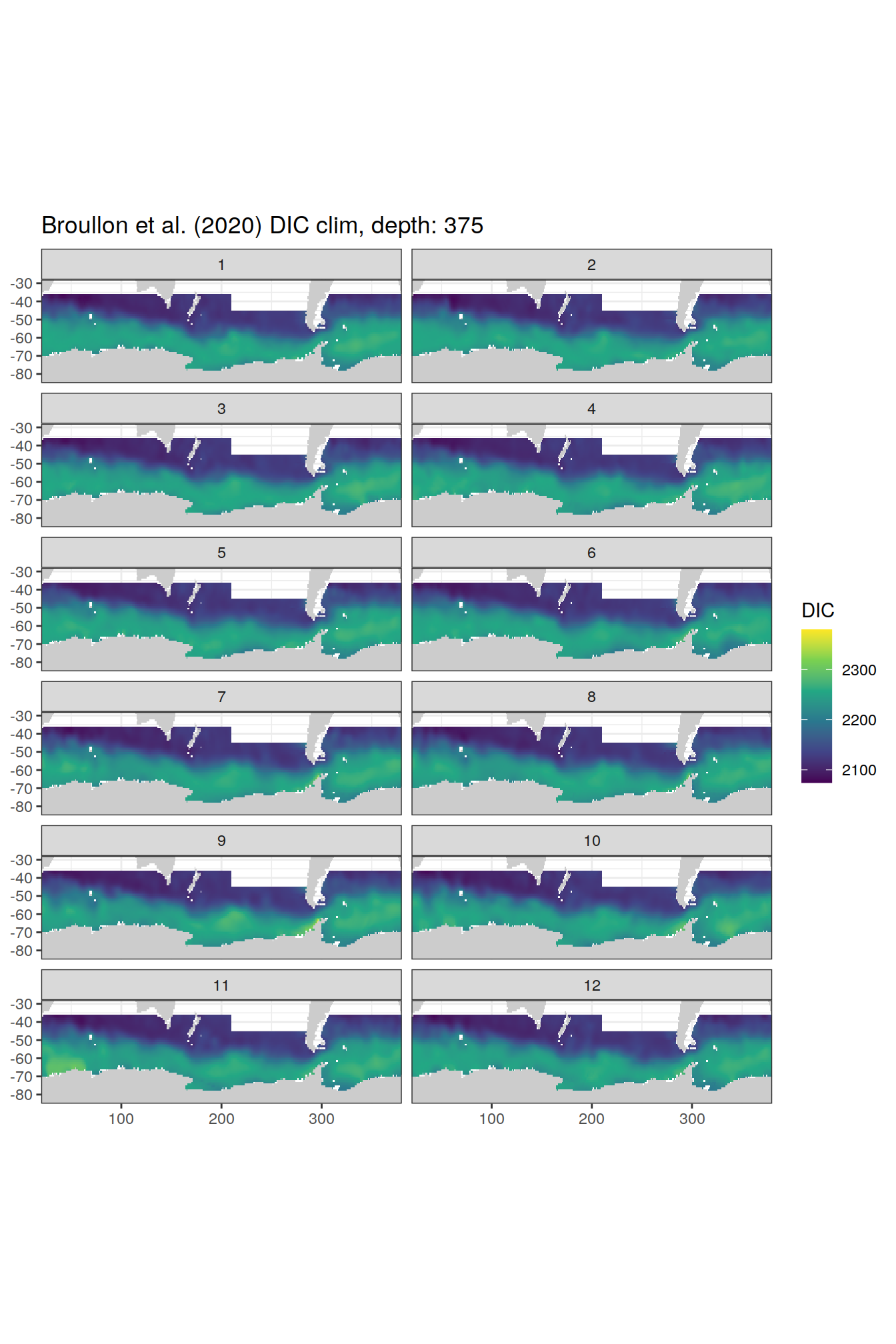
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[33]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[34]]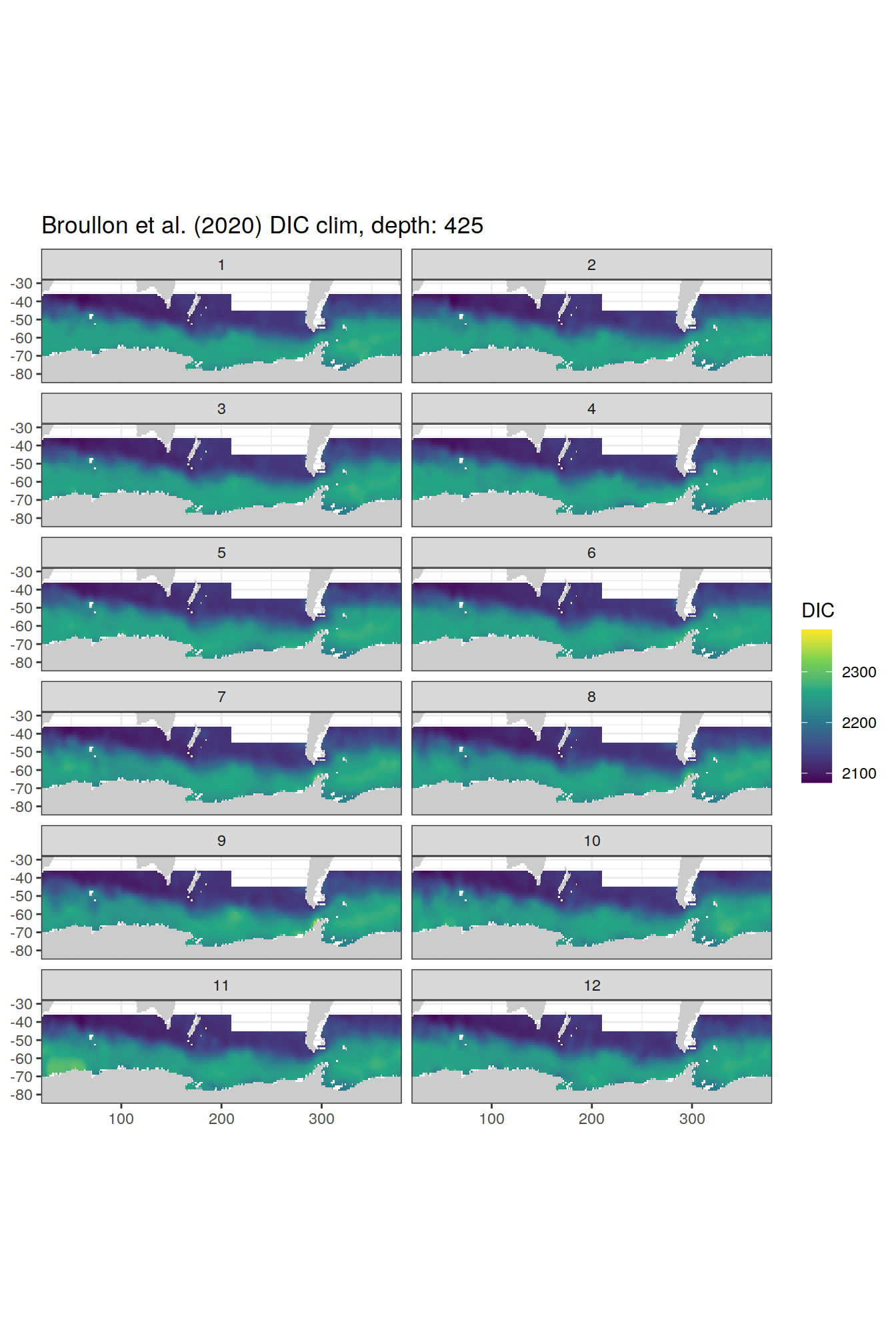
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[35]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[36]]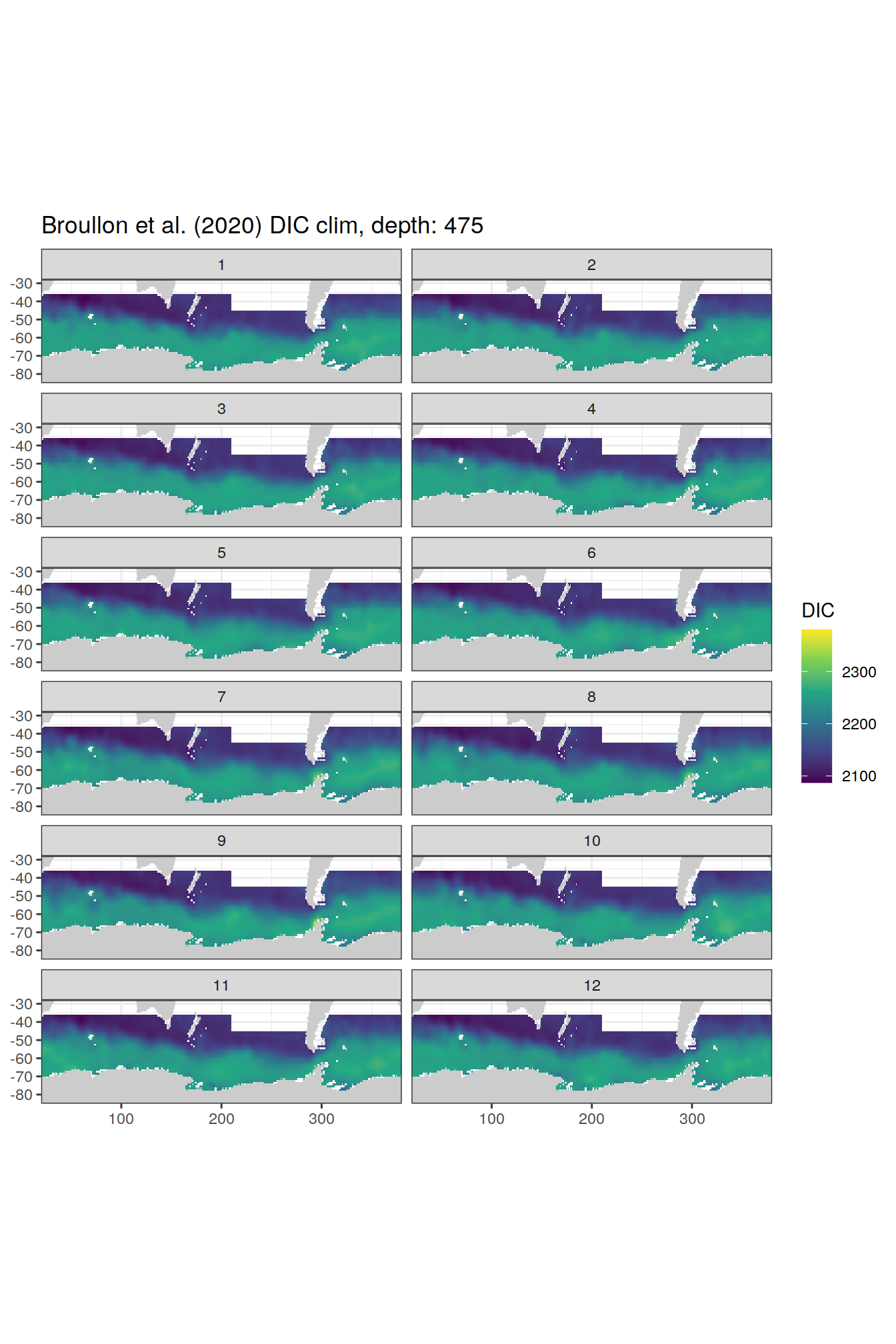
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[37]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[38]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[39]]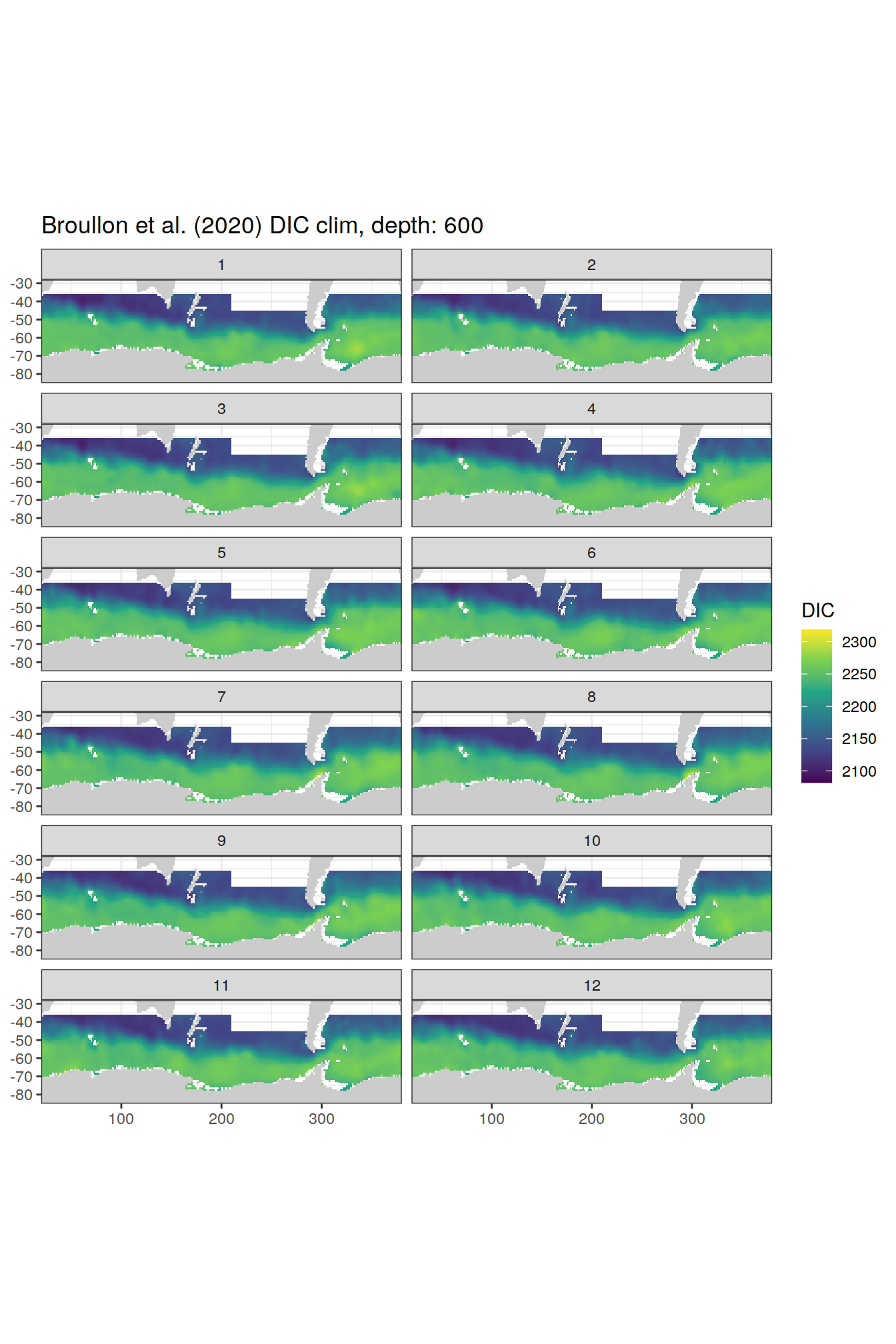
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[40]]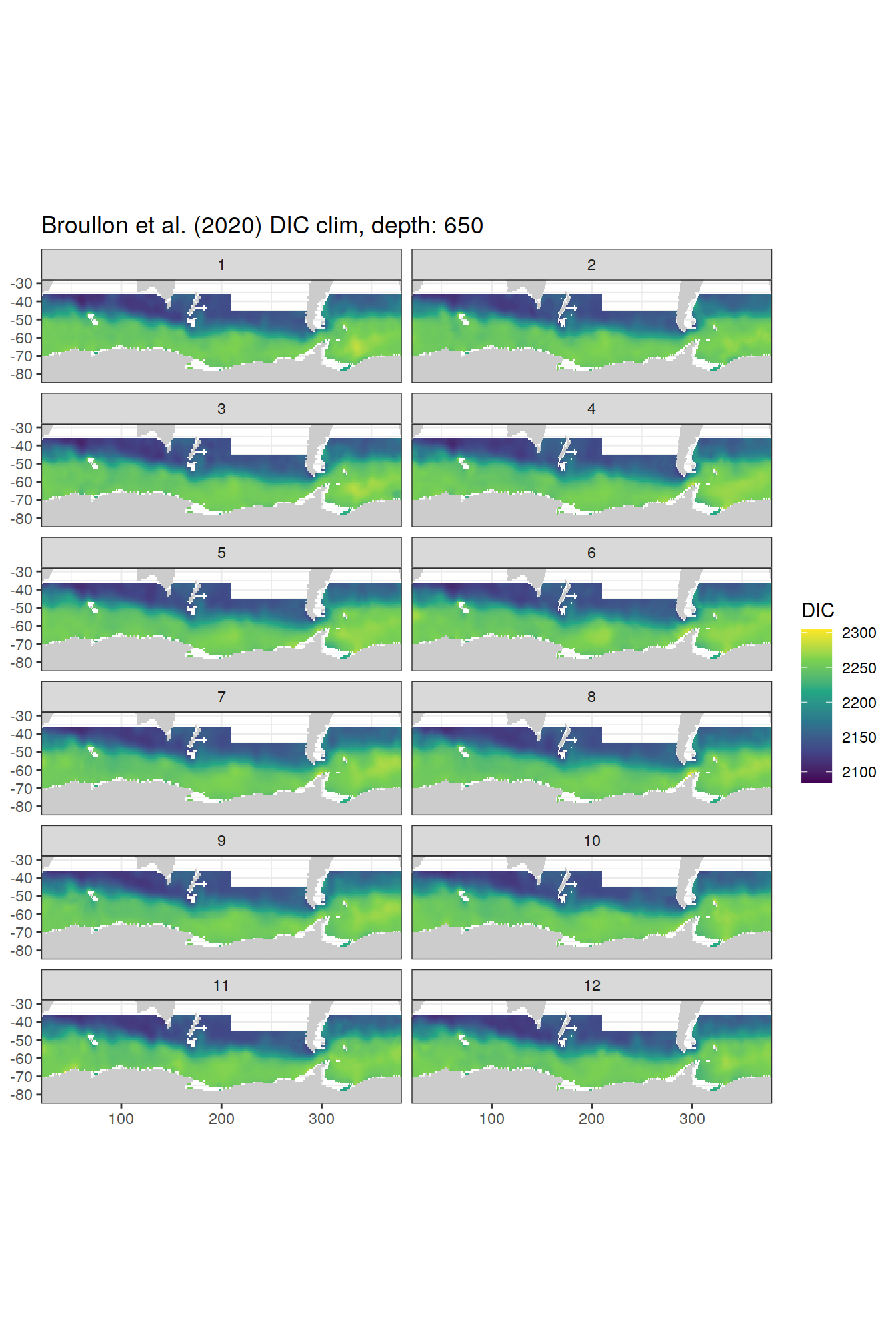
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[41]]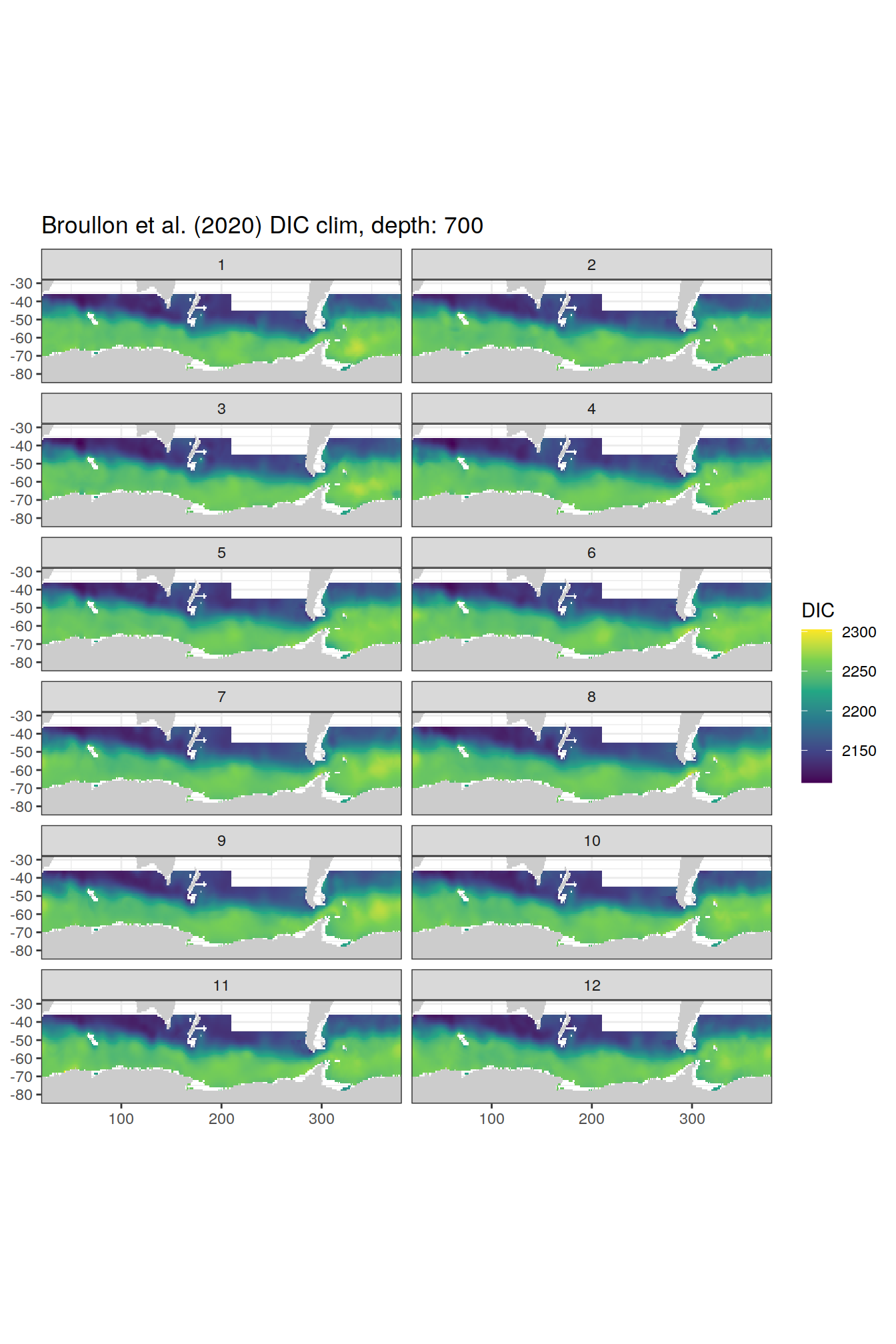
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[42]]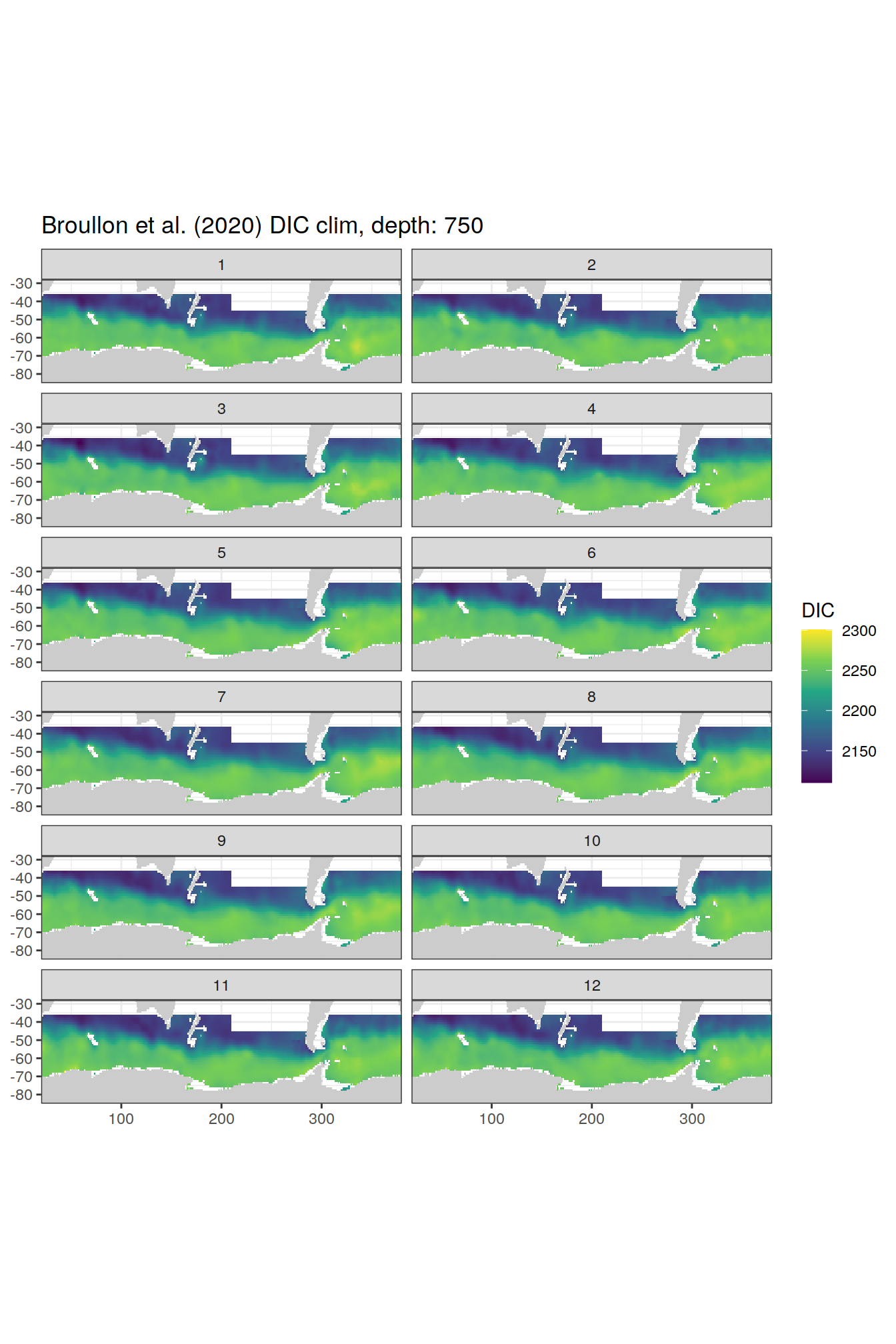
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[43]]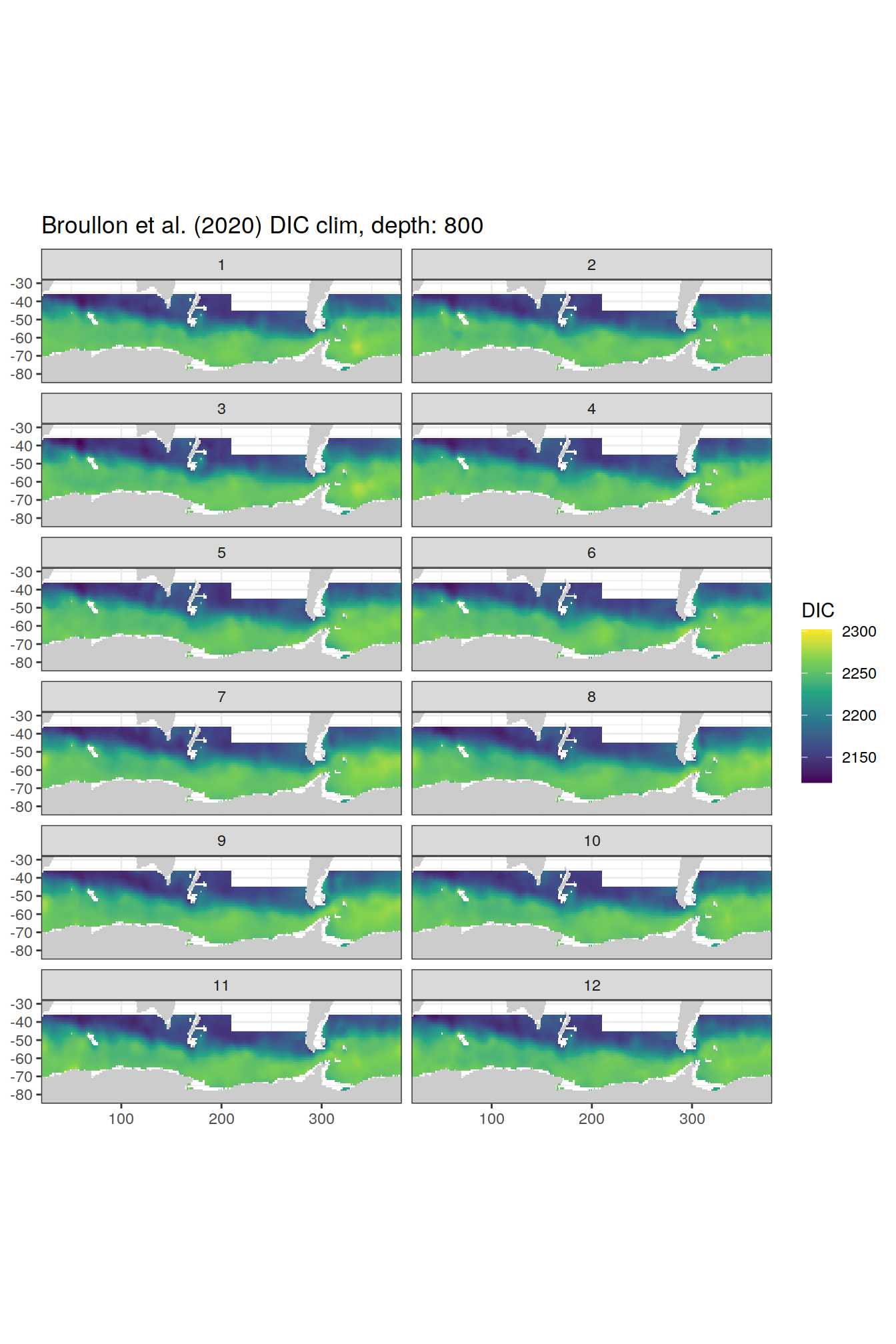
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[44]]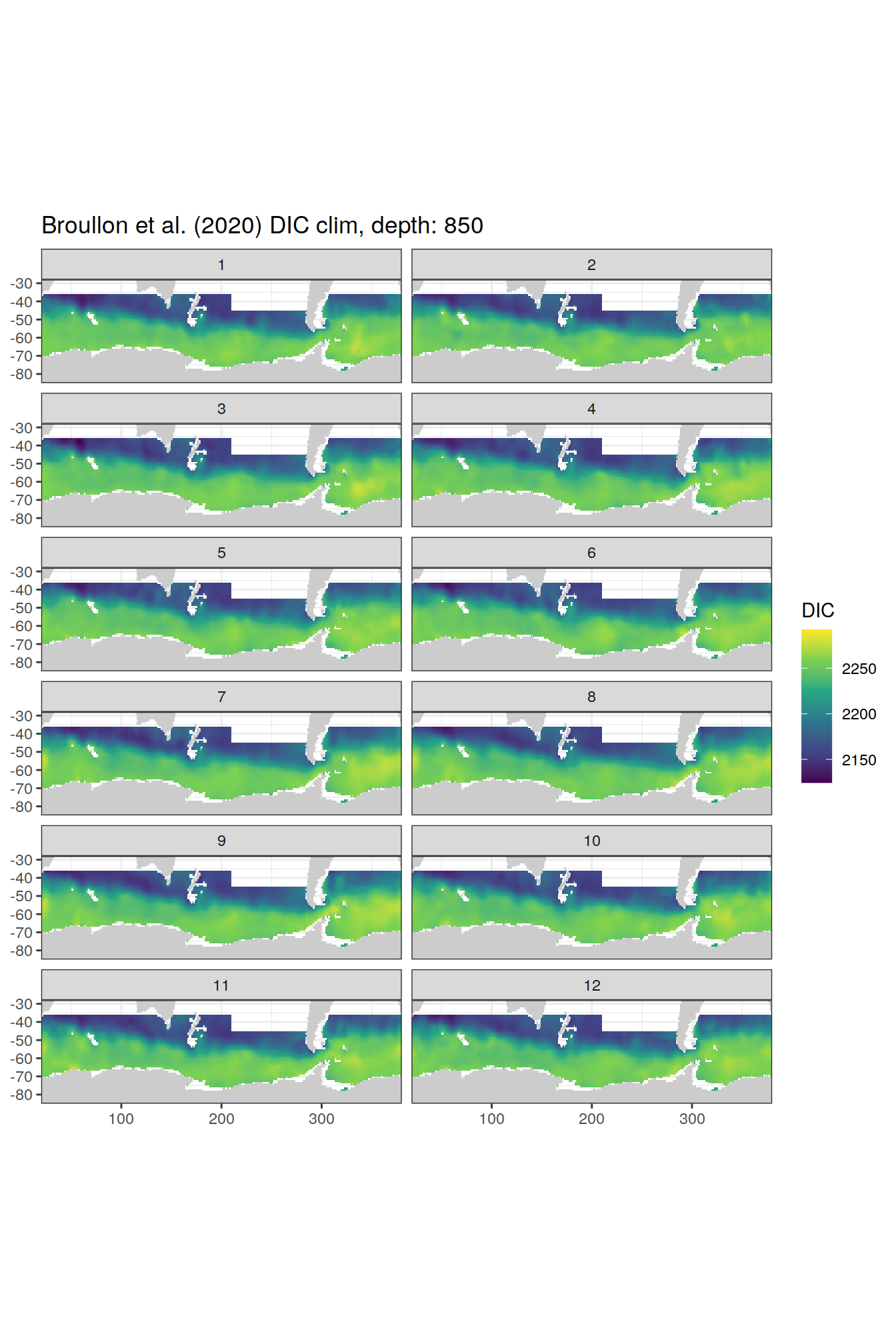
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[45]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[46]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[47]]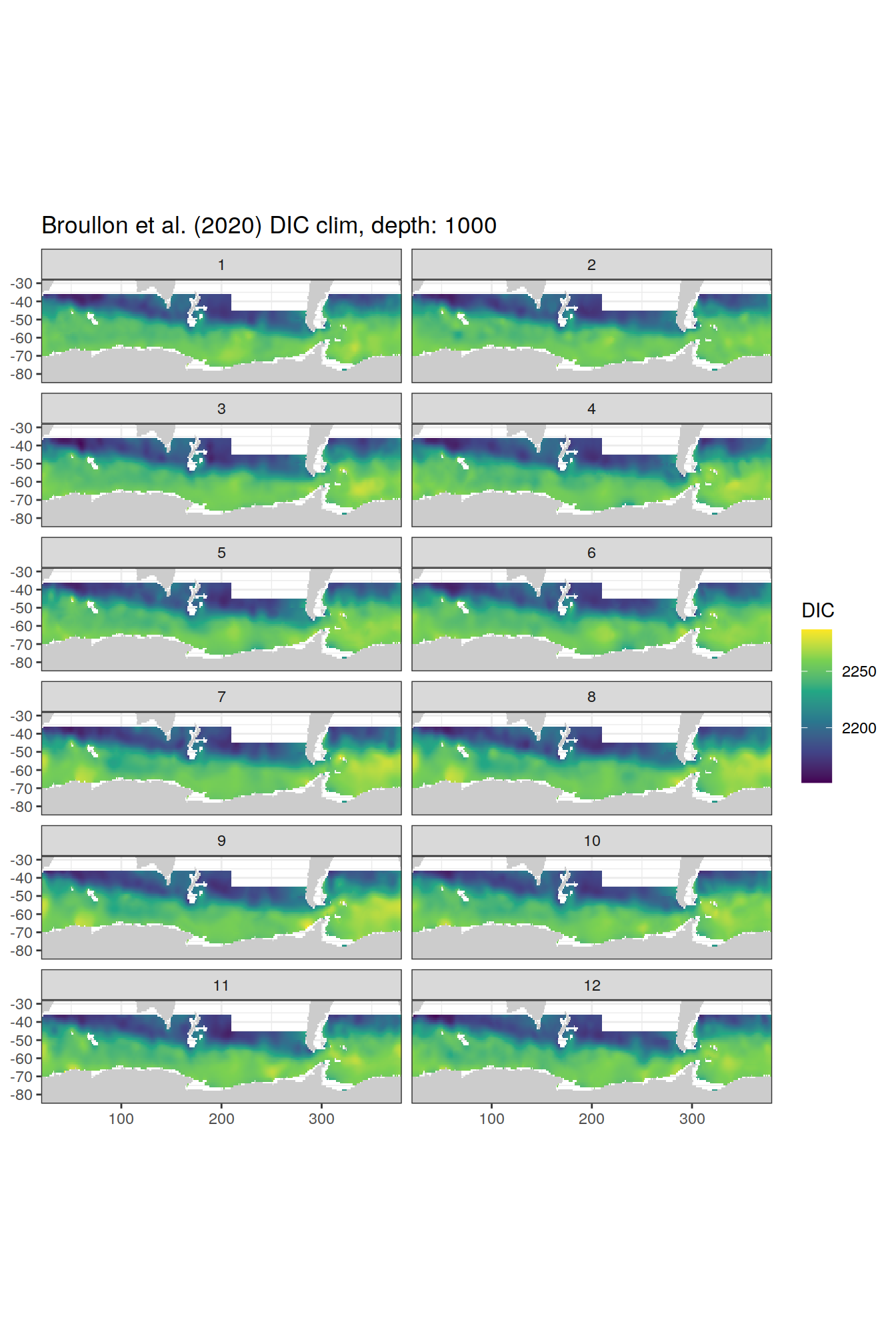
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[48]]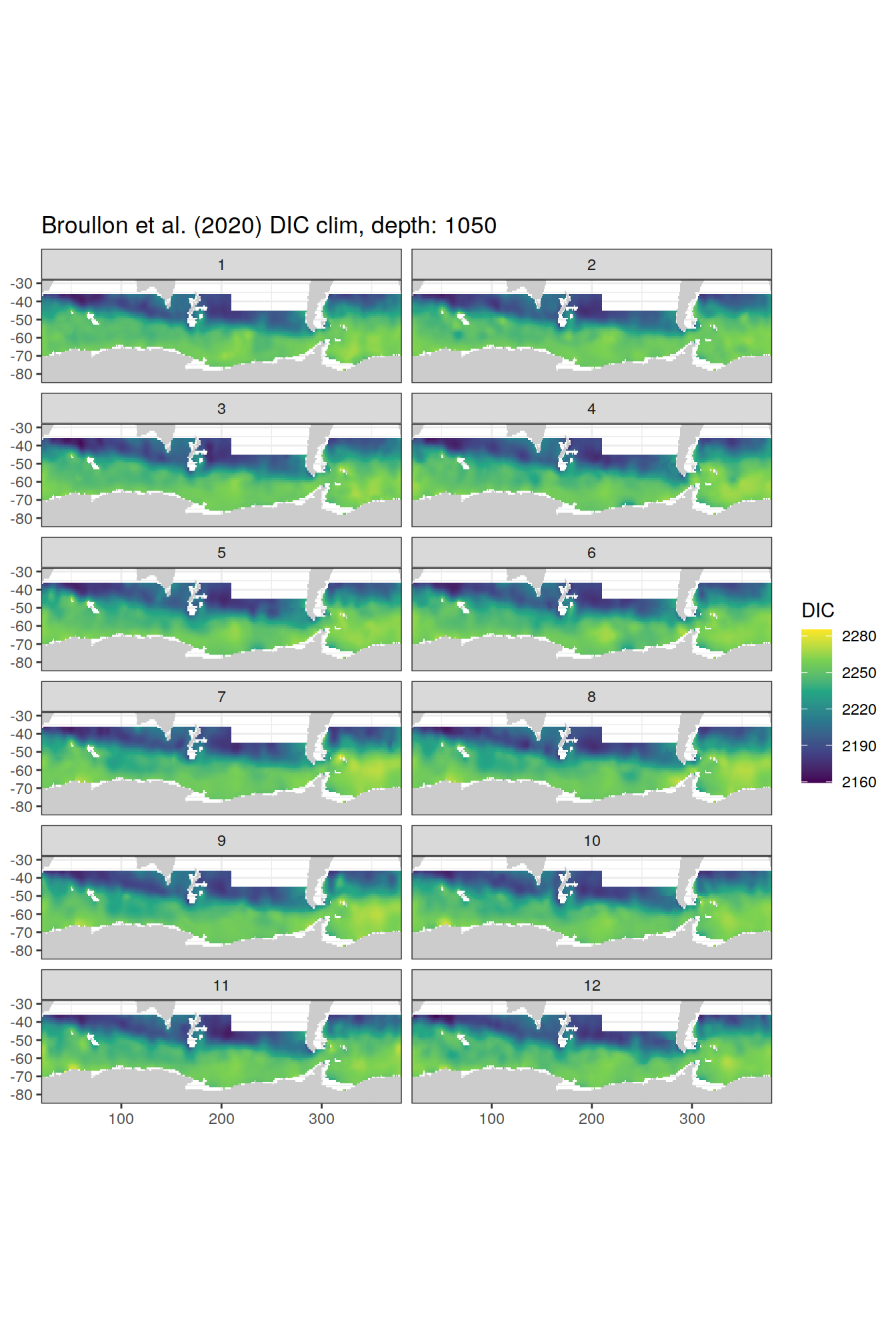
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[49]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[50]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[51]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[52]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[53]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[54]]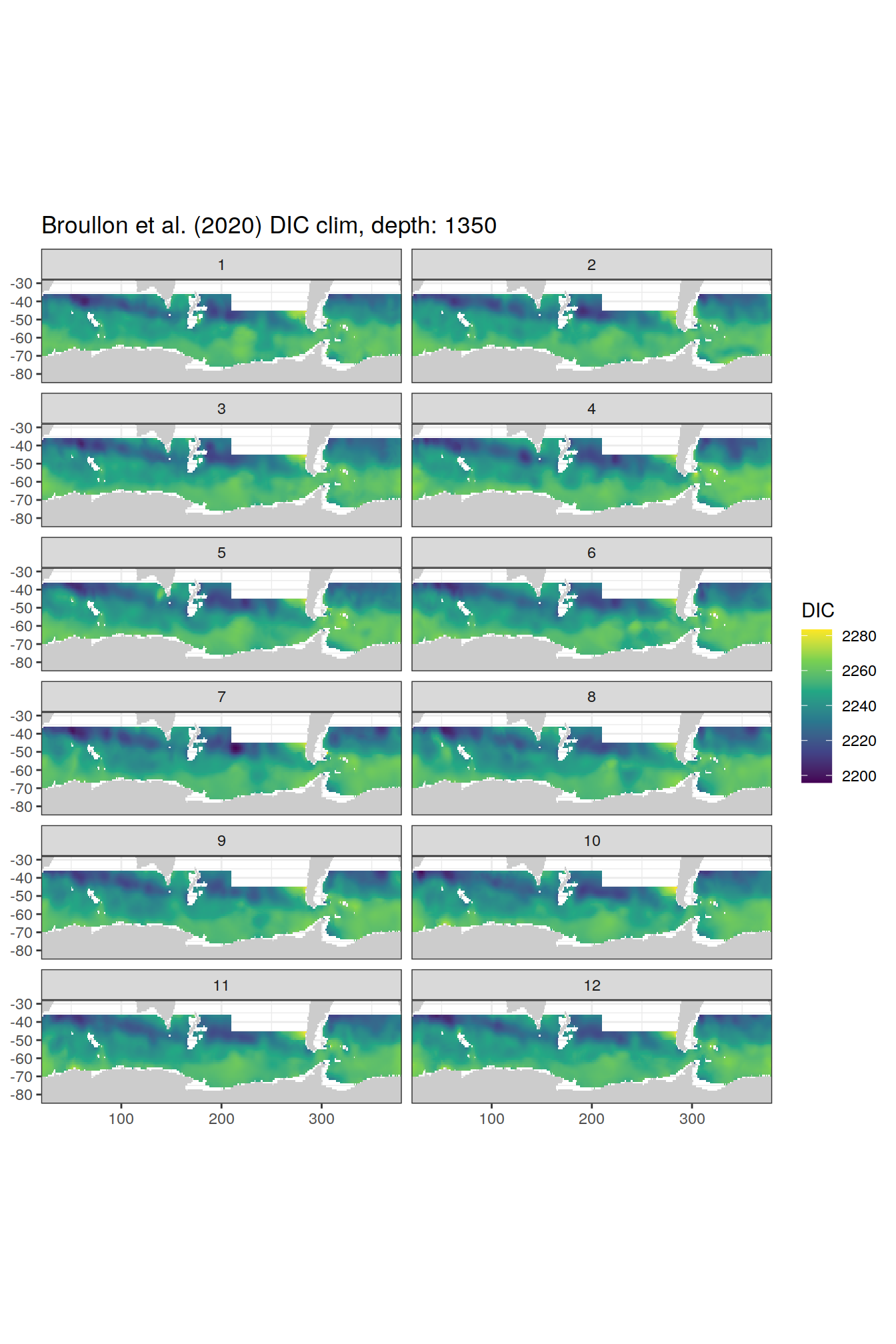
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[55]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[56]]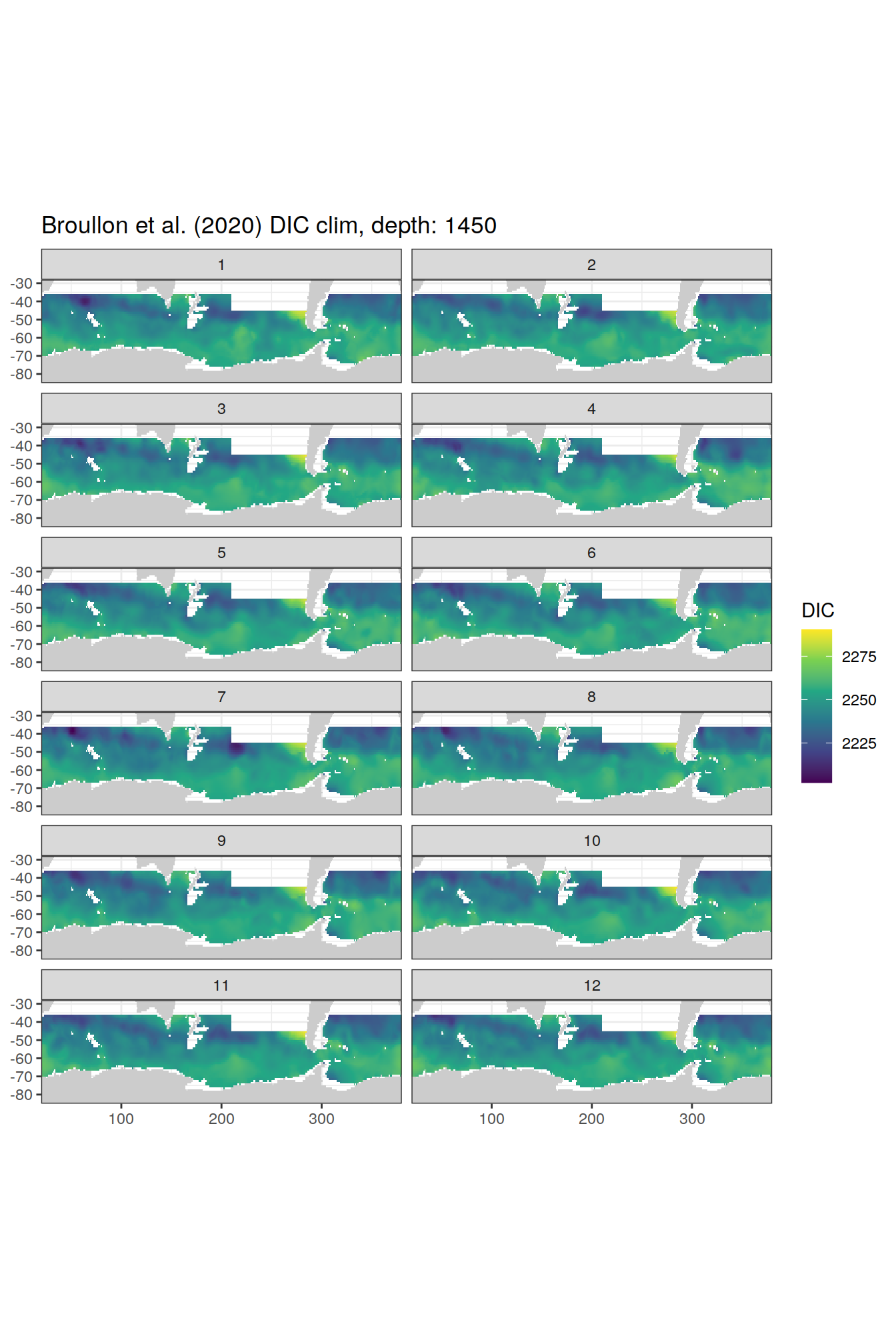
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[57]]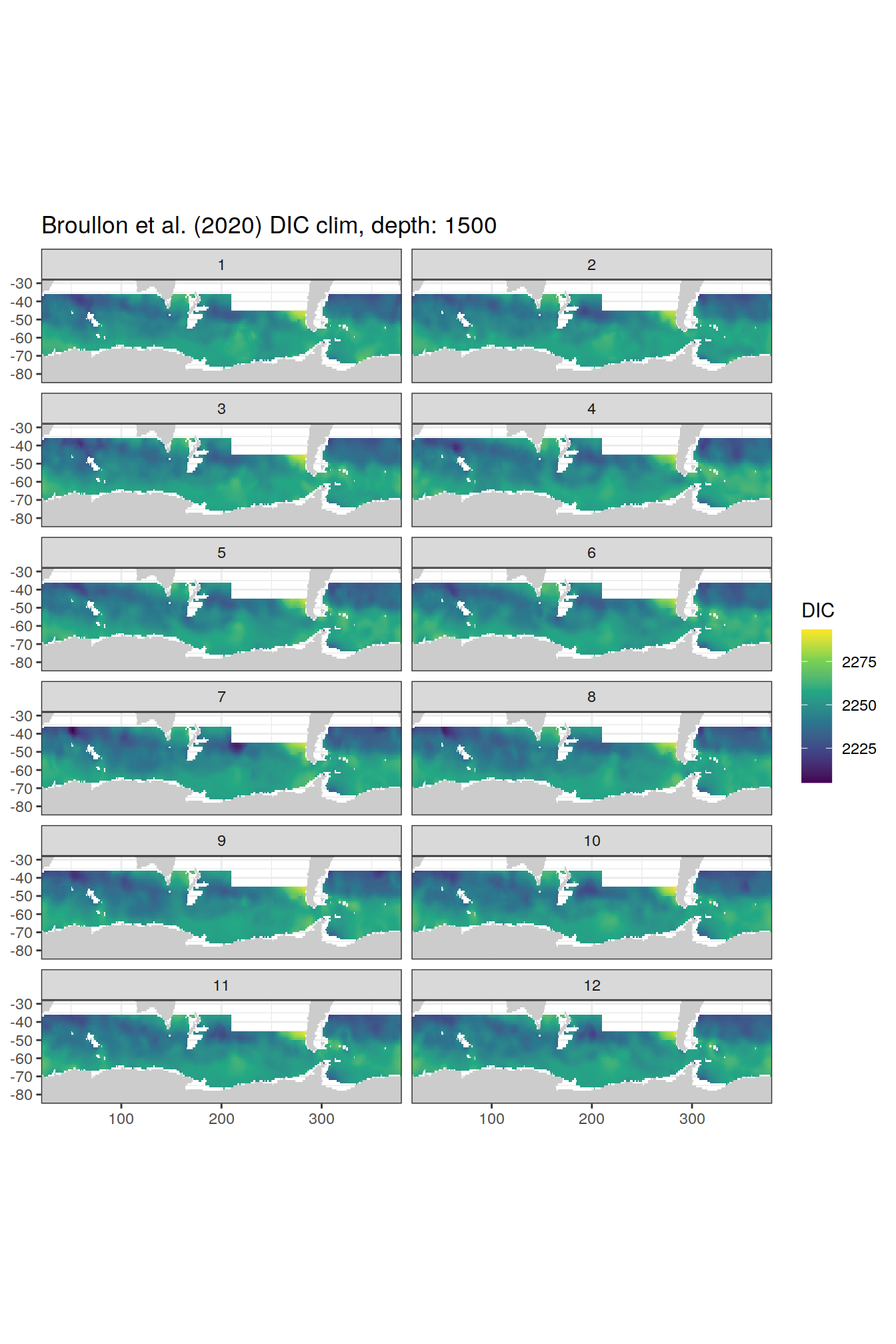
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[58]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[59]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[60]]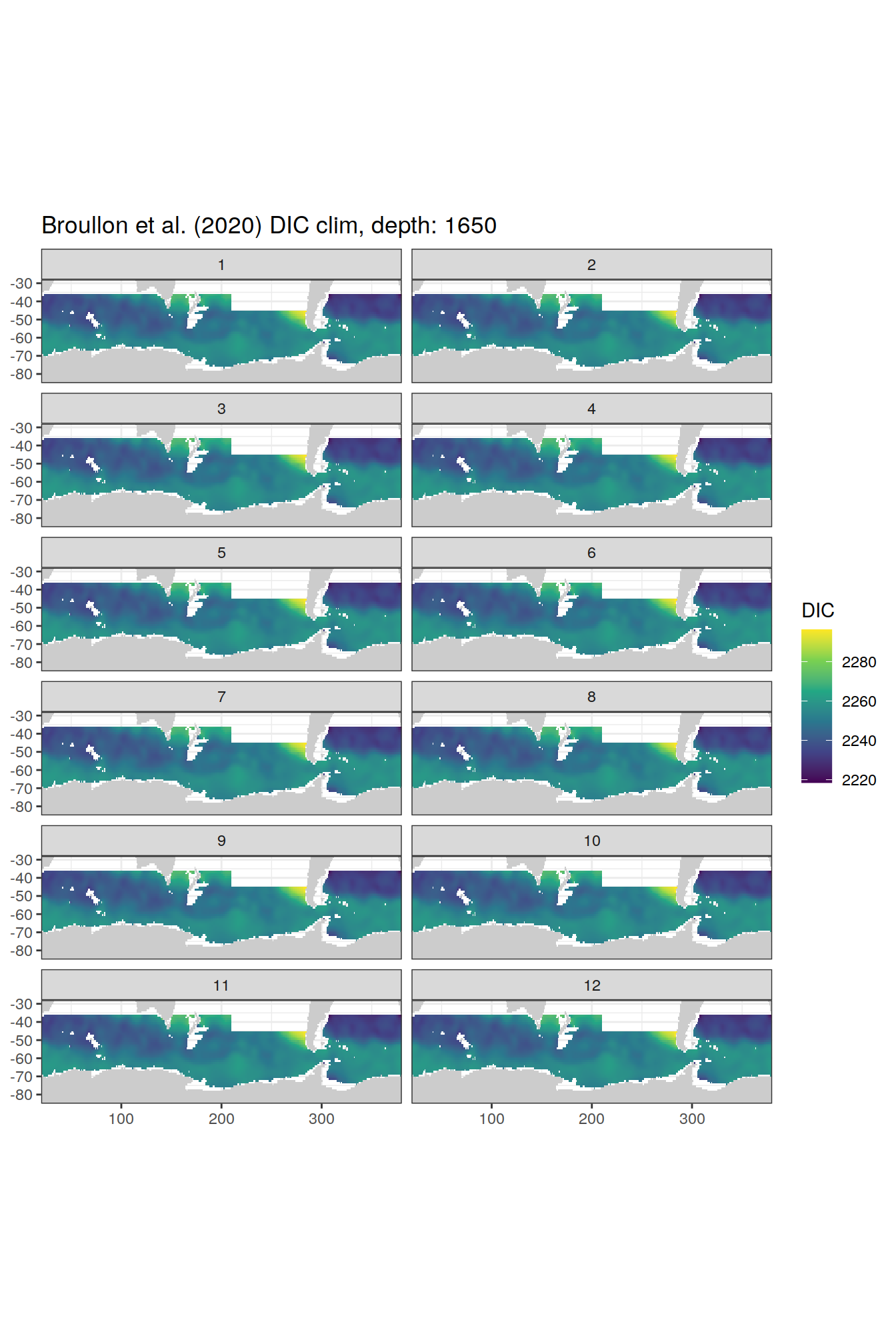
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[61]]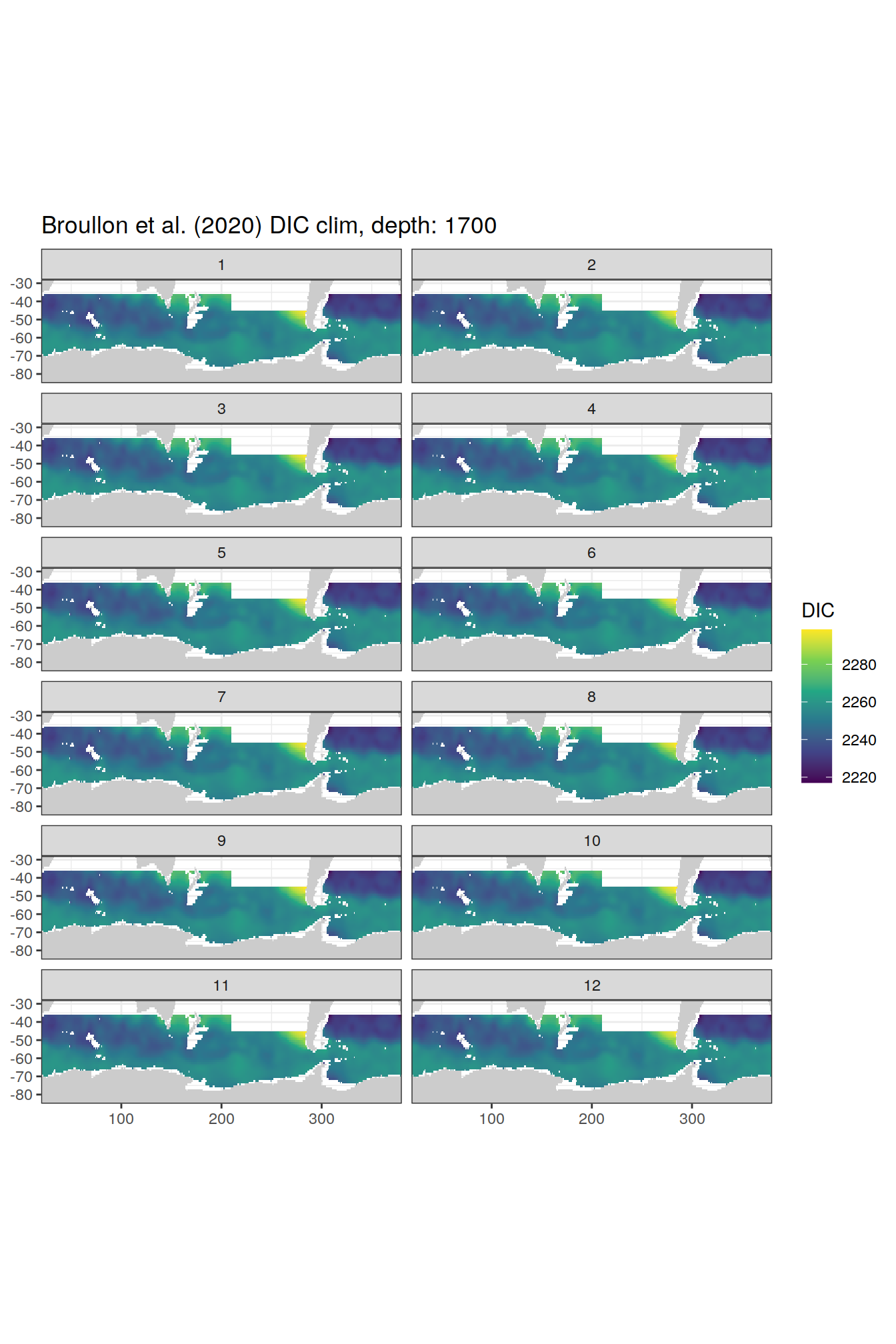
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[62]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[63]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[64]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[65]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[66]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[67]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[68]]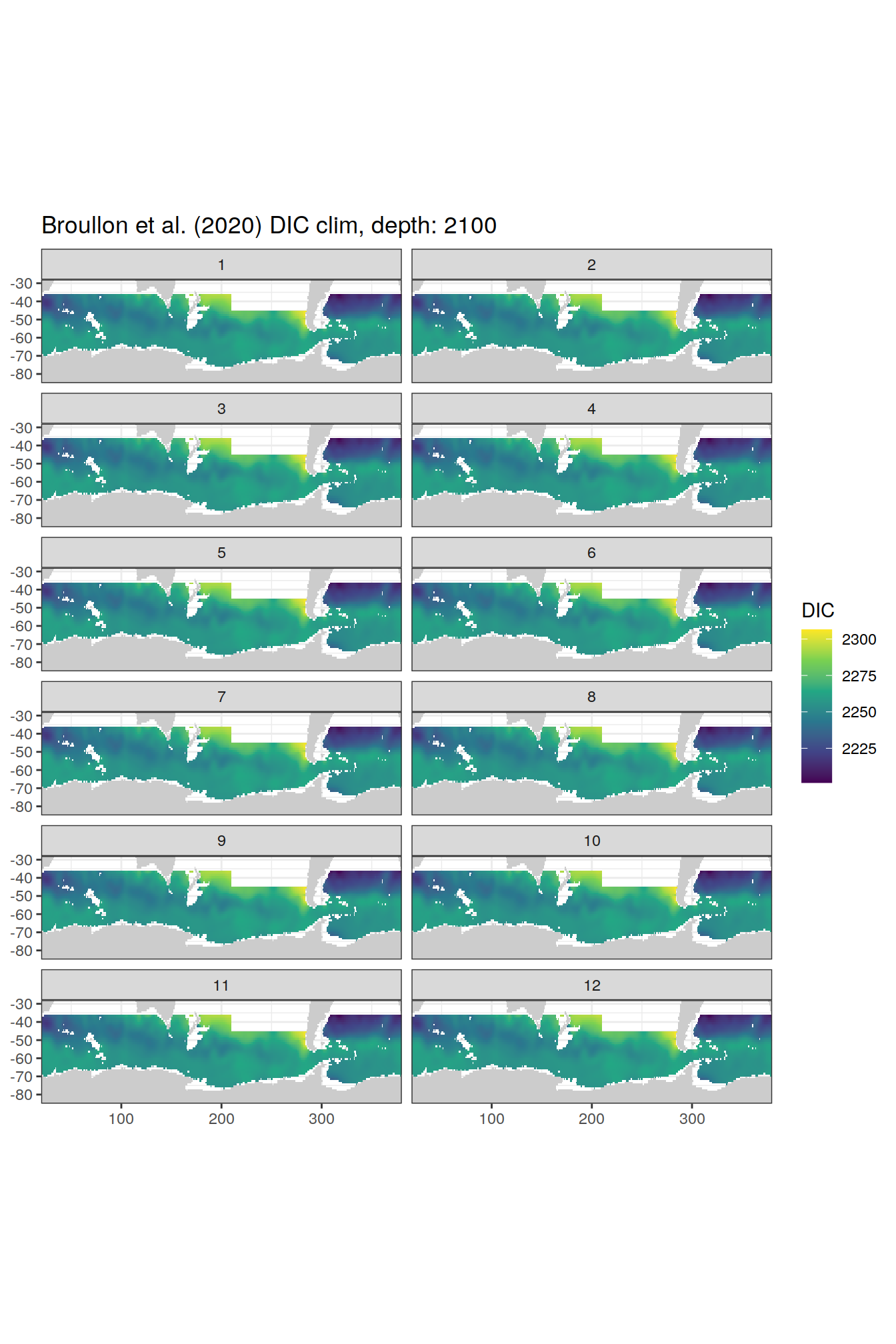
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[69]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[70]]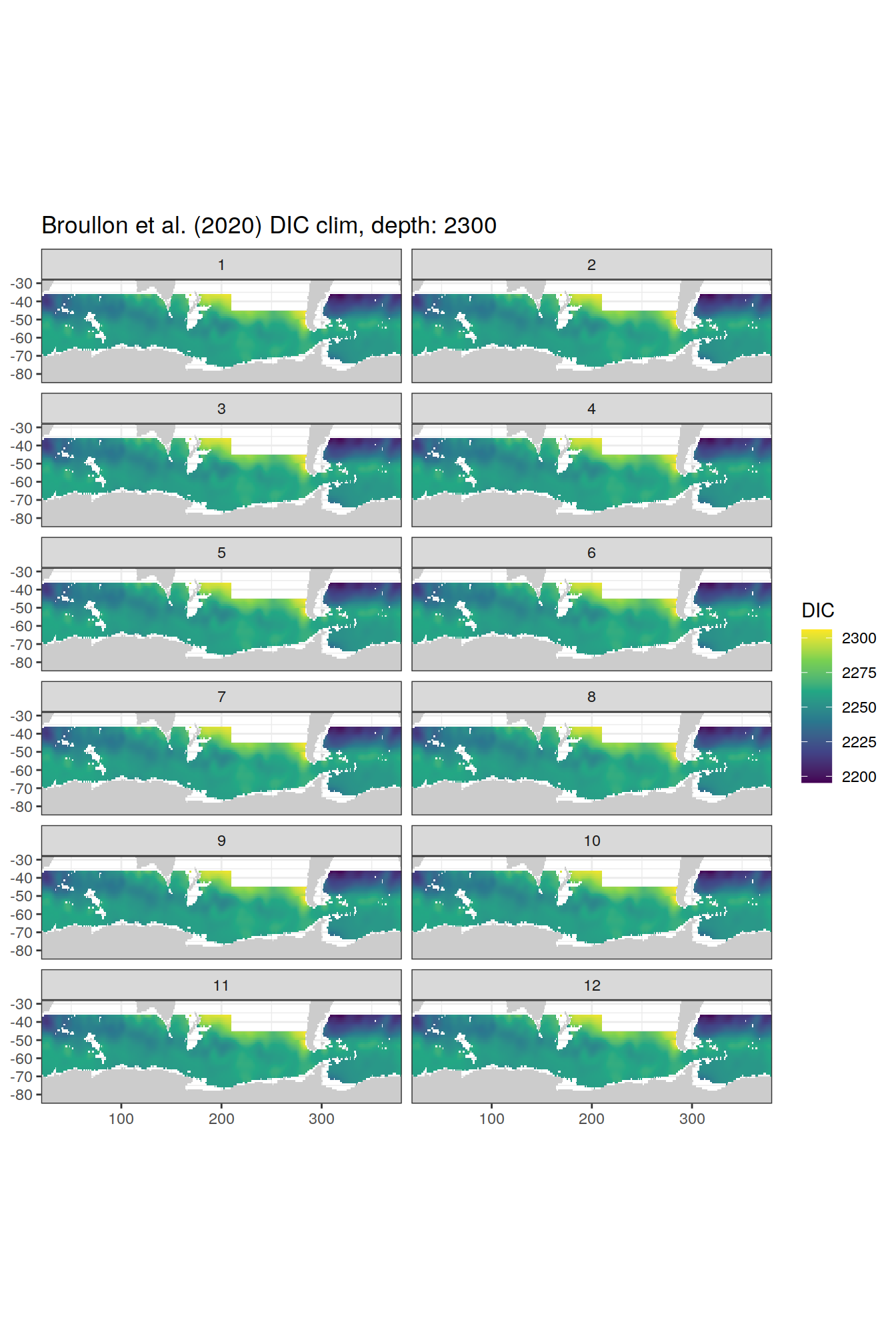
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[71]]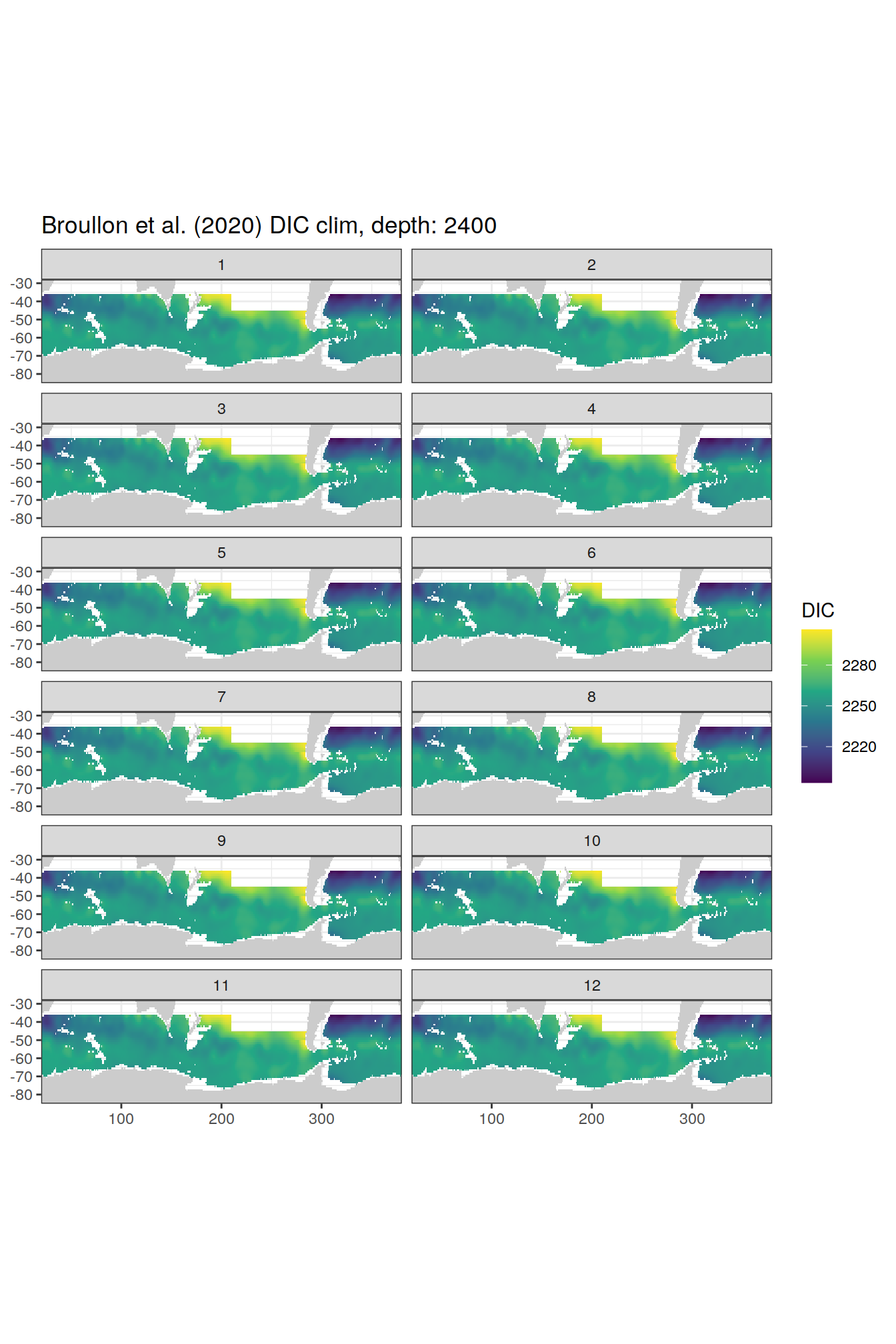
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[72]]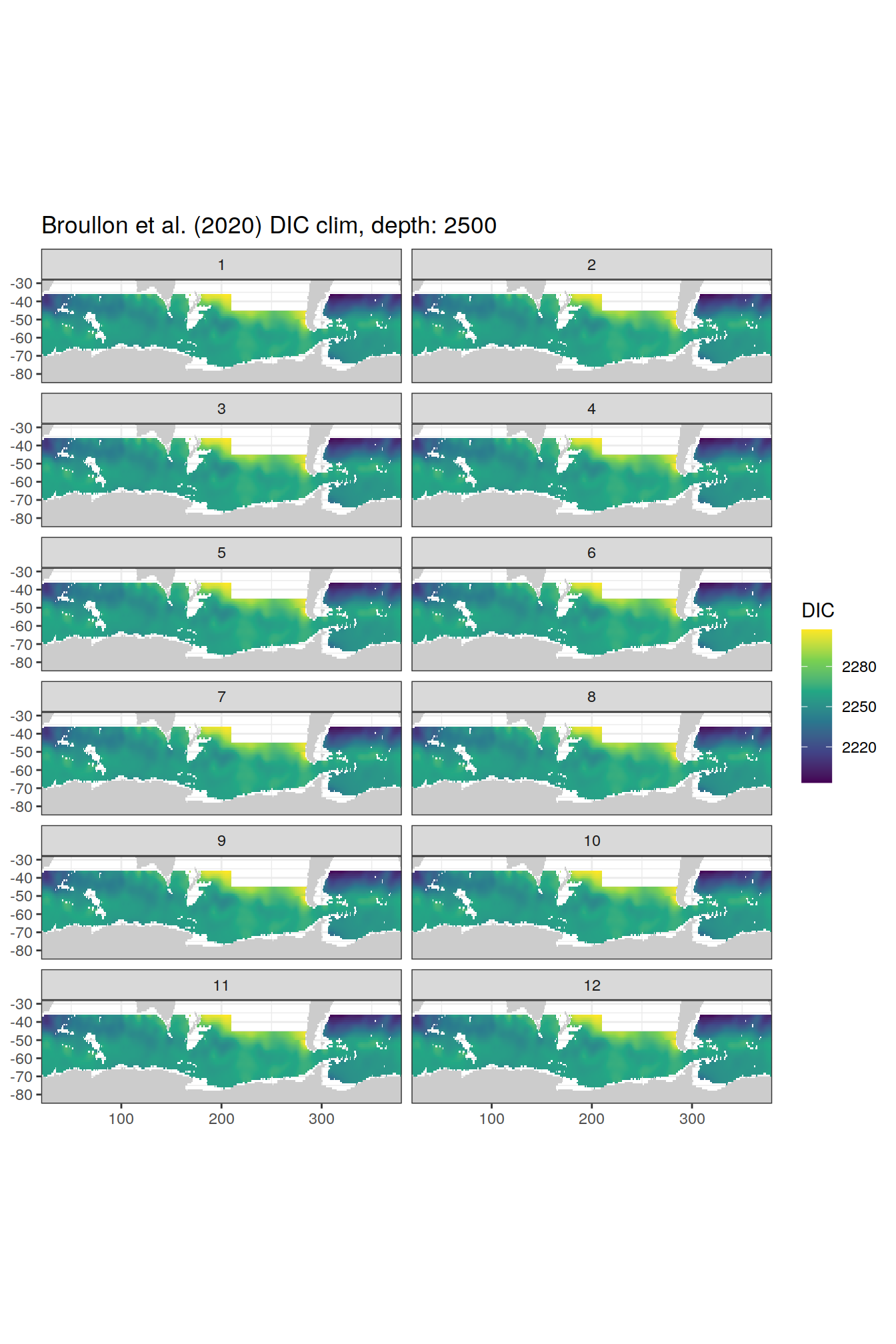
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[73]]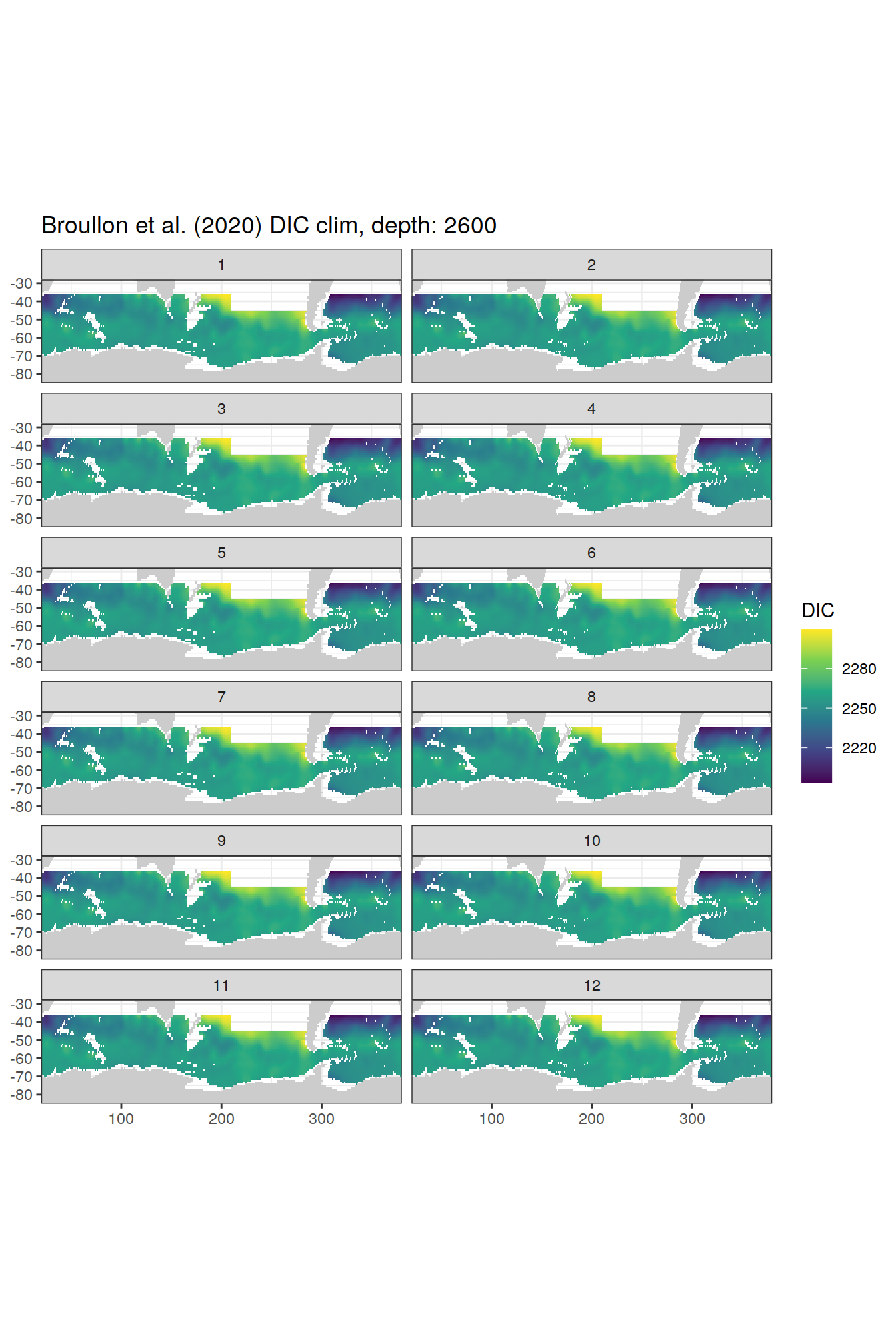
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[74]]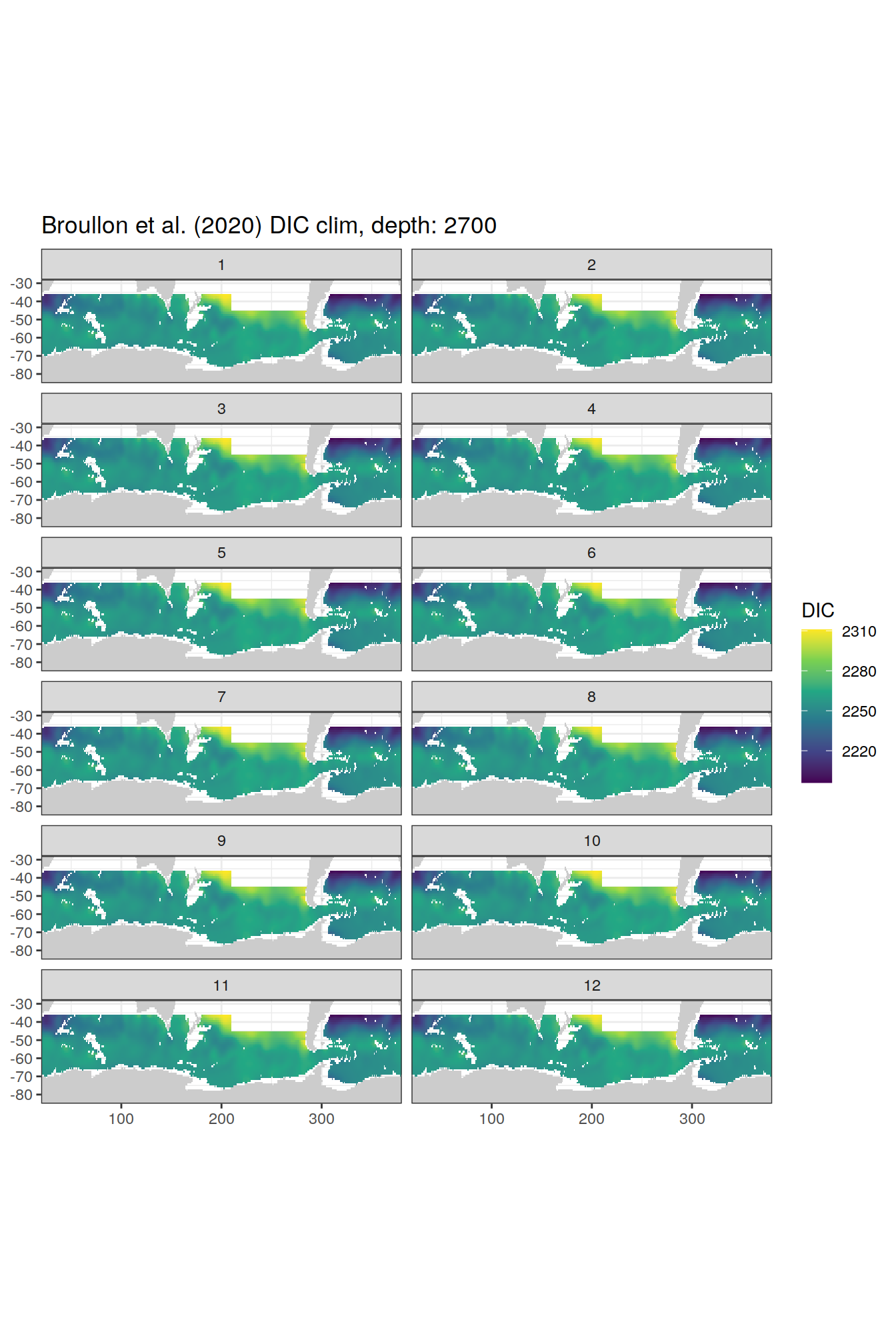
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[75]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[76]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[77]]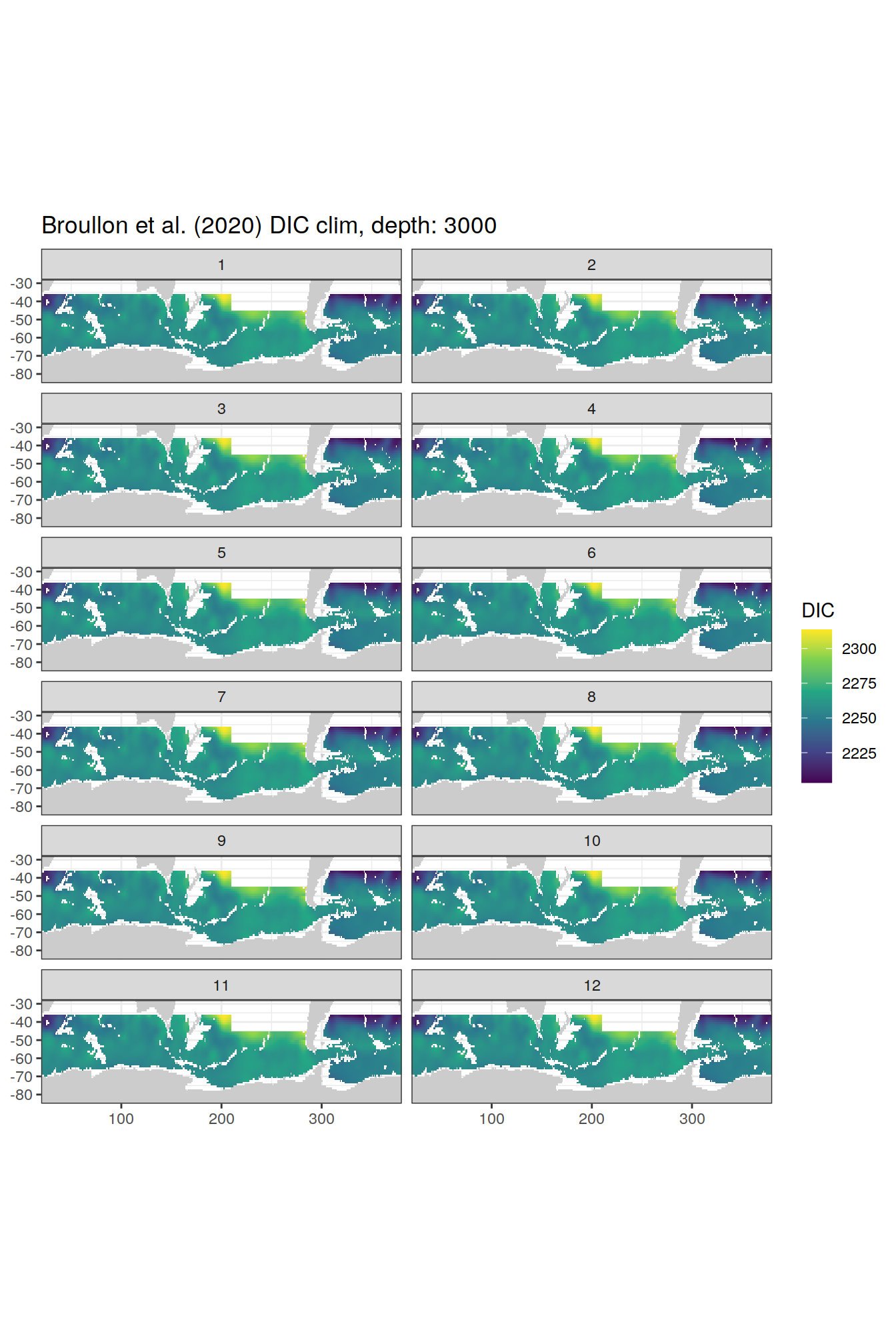
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[78]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[79]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[80]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[81]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[82]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[83]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[84]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[85]]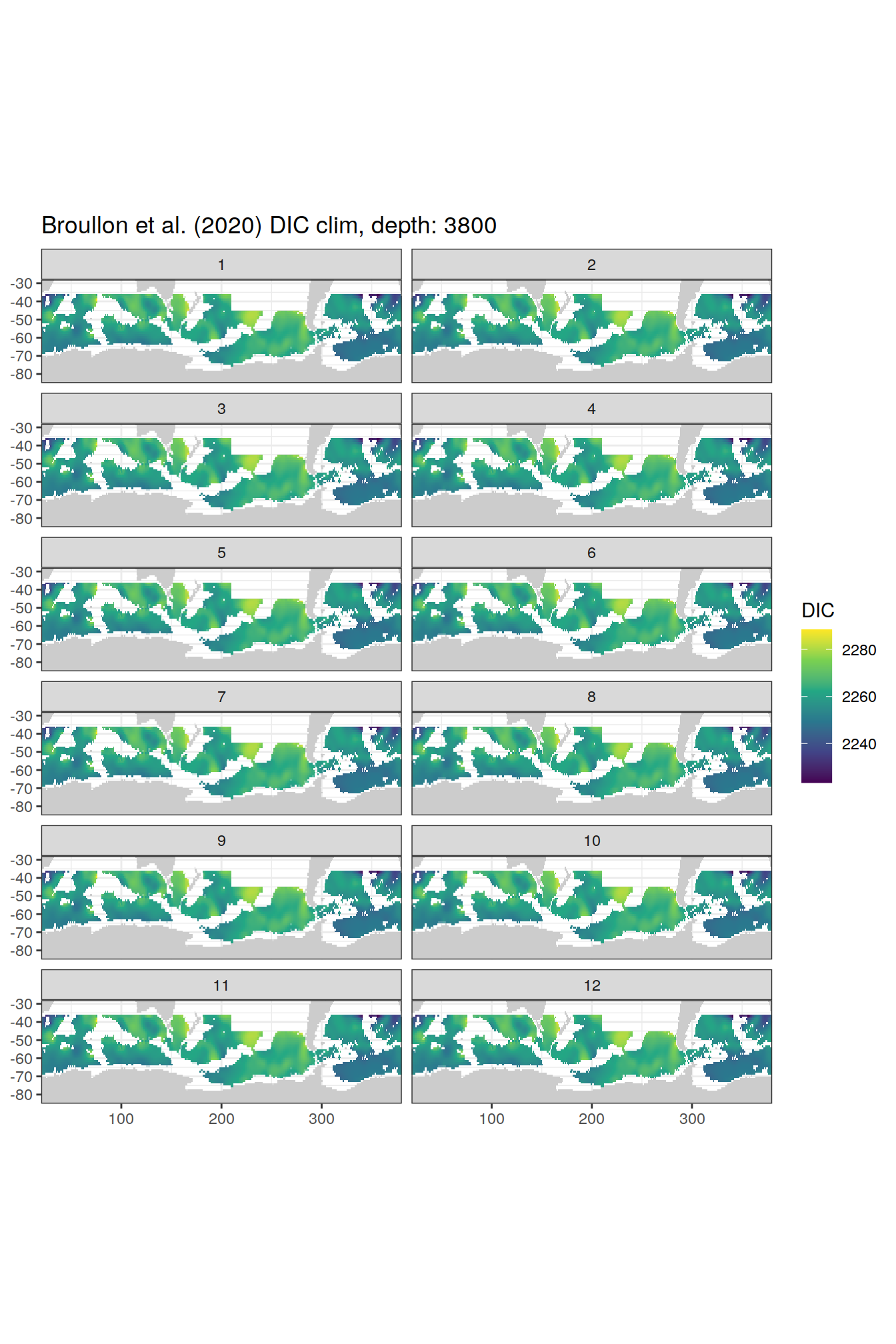
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[86]]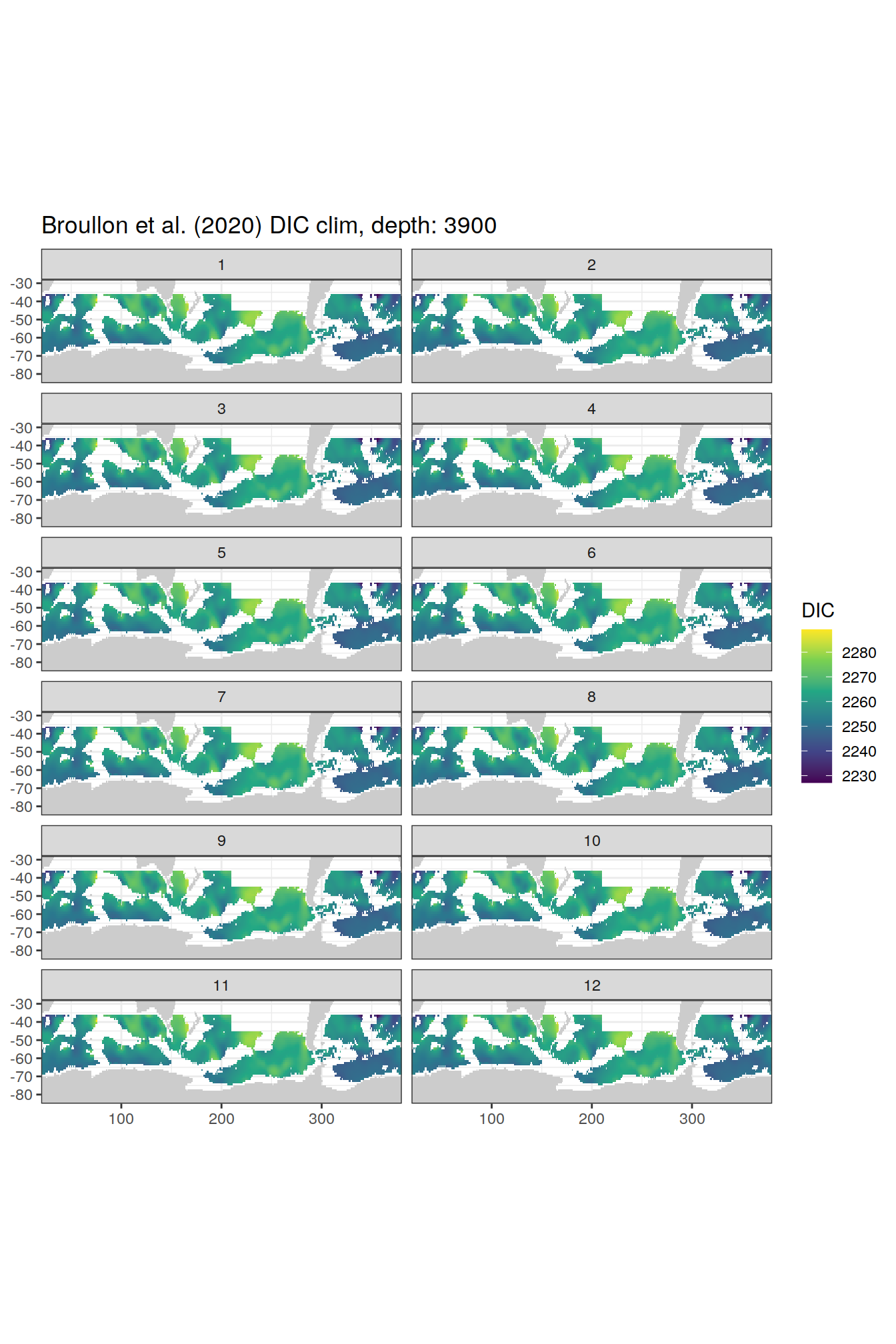
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[87]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[88]]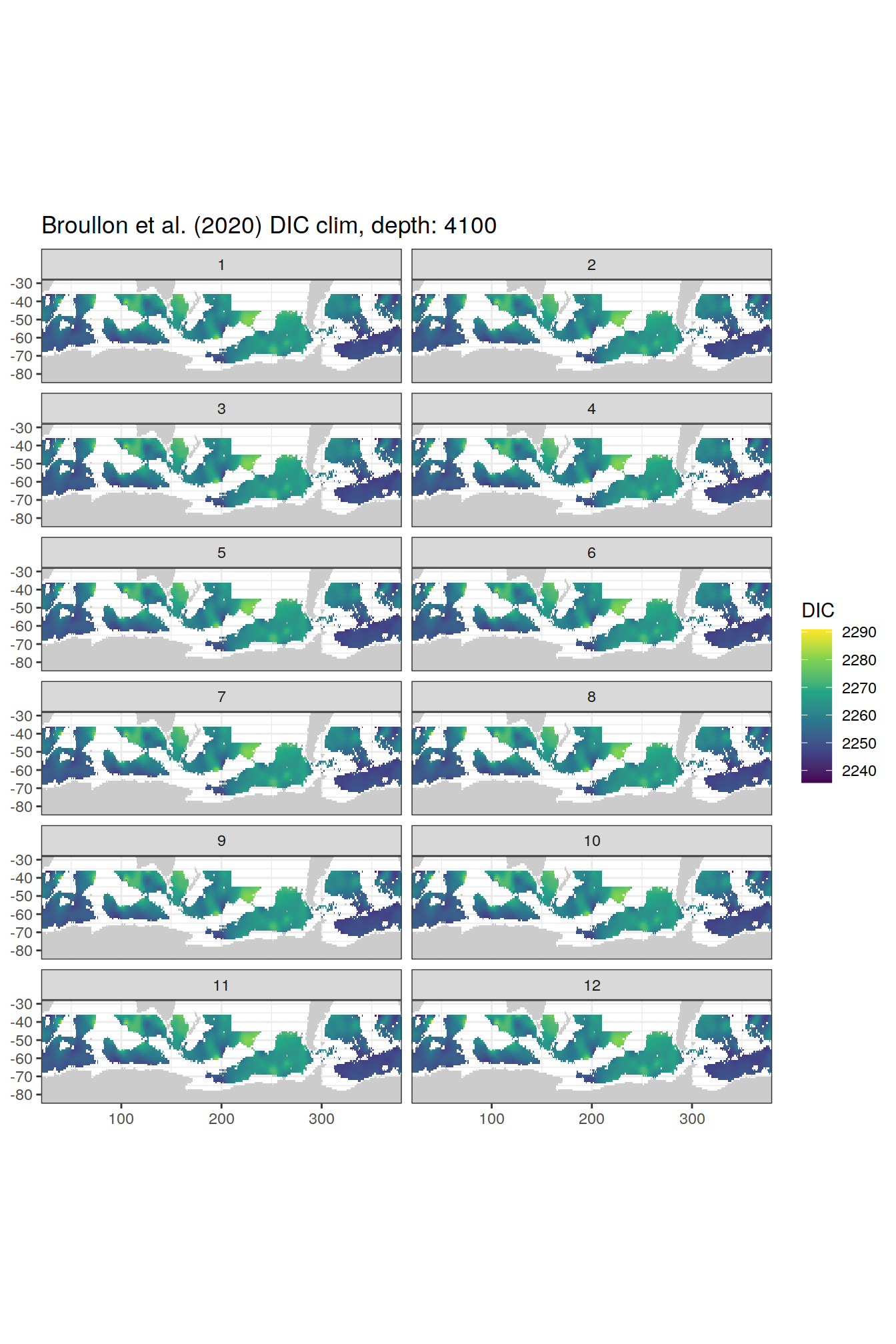
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[89]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[90]]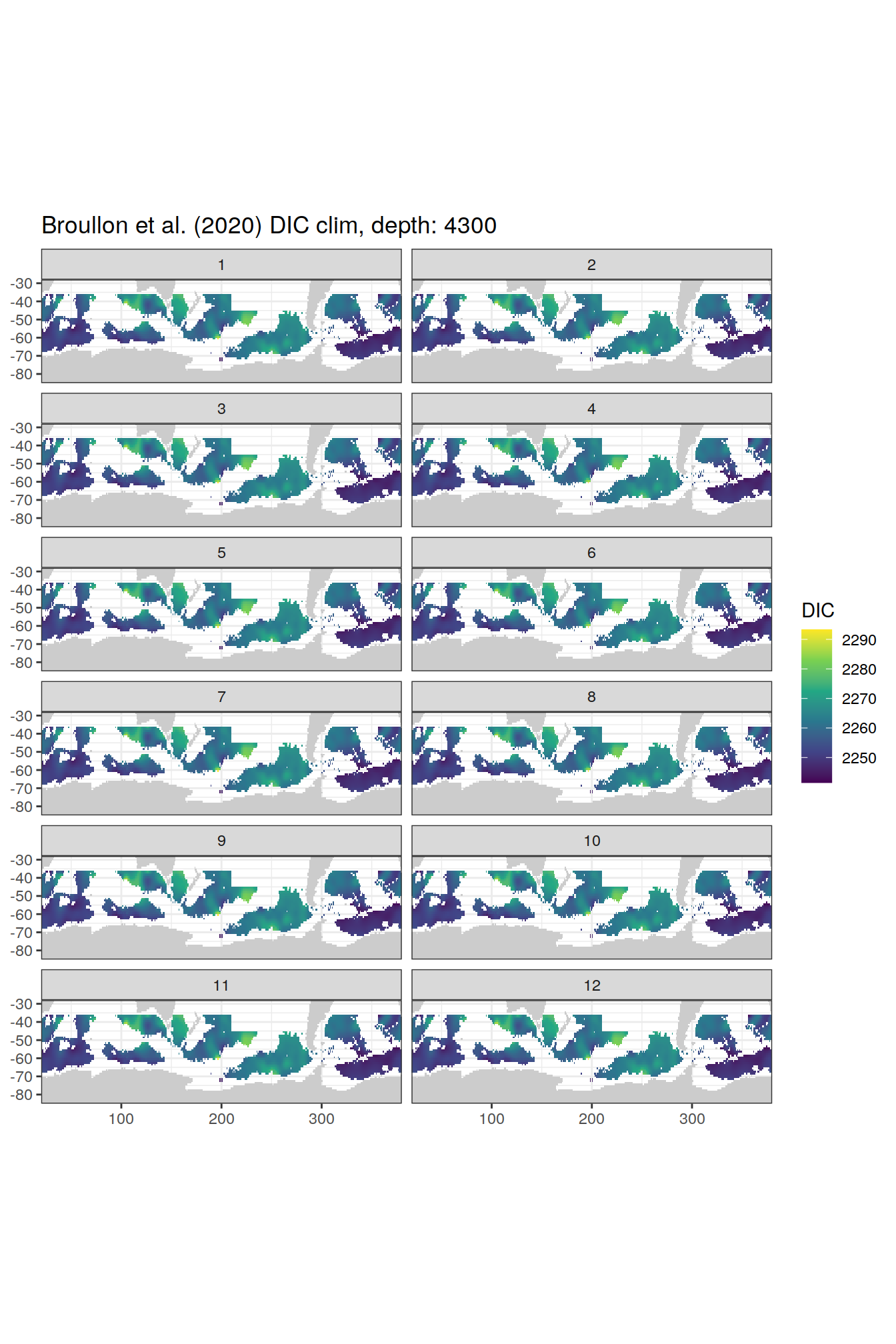
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[91]]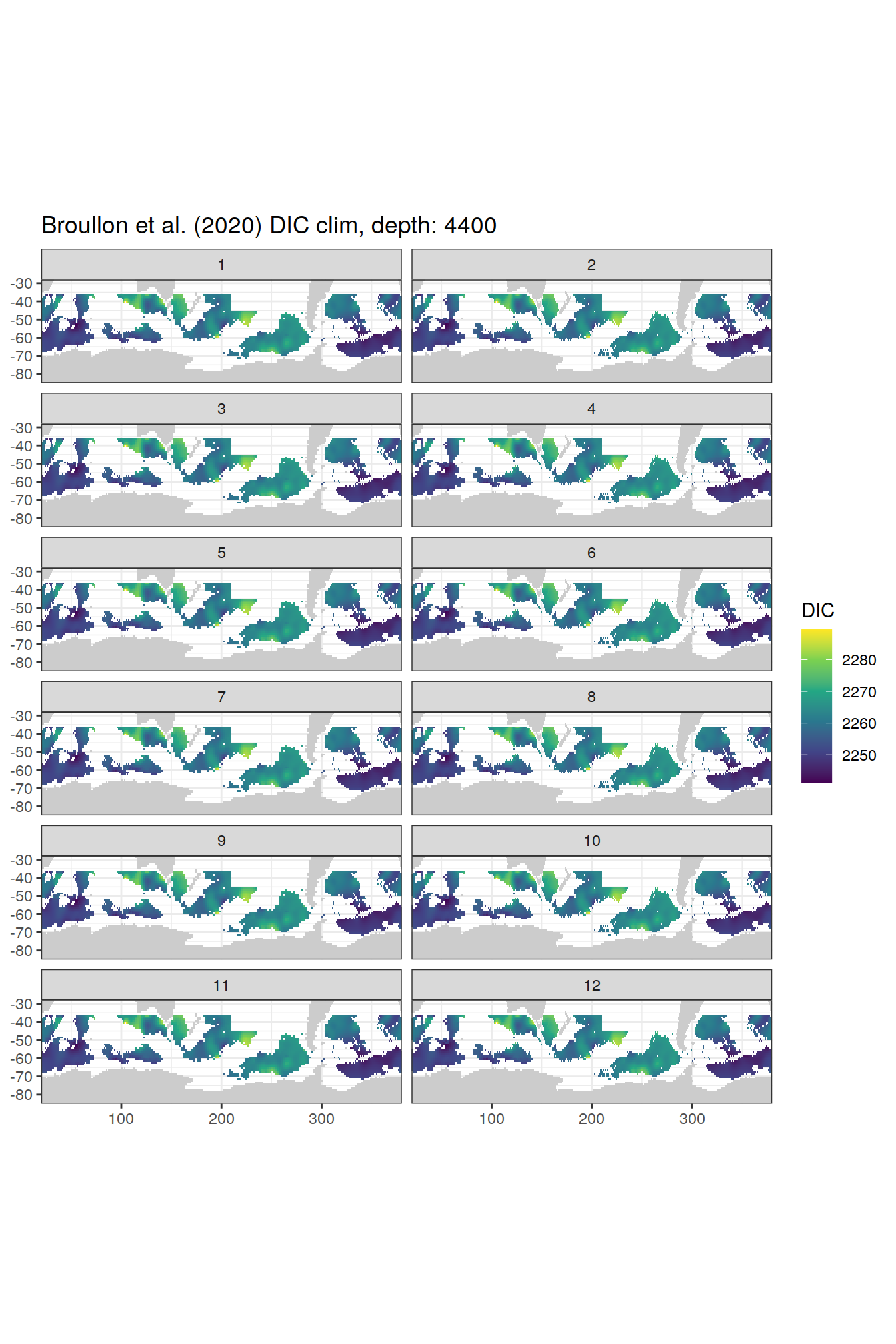
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[92]]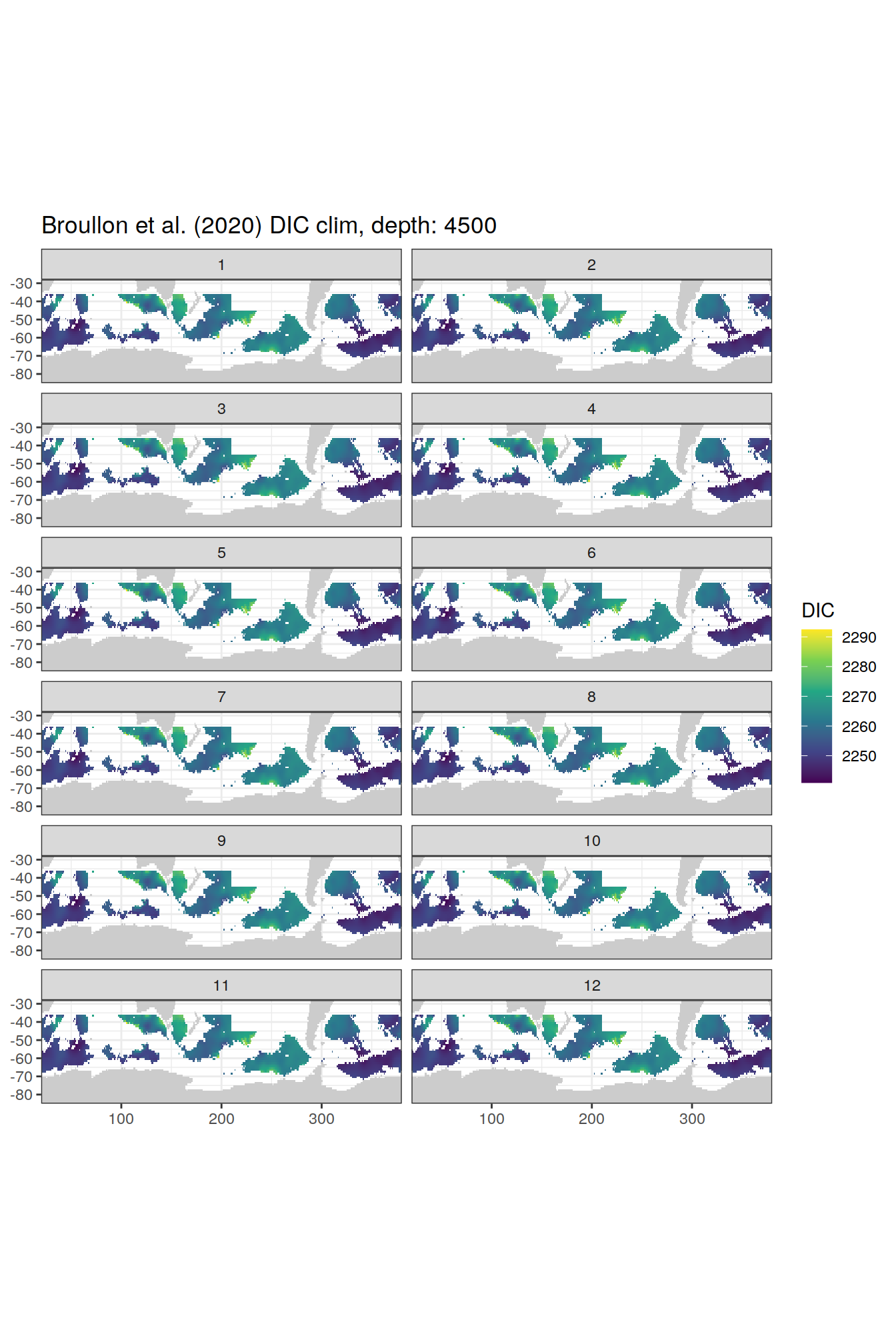
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[93]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[94]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[95]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[96]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[97]]
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[98]]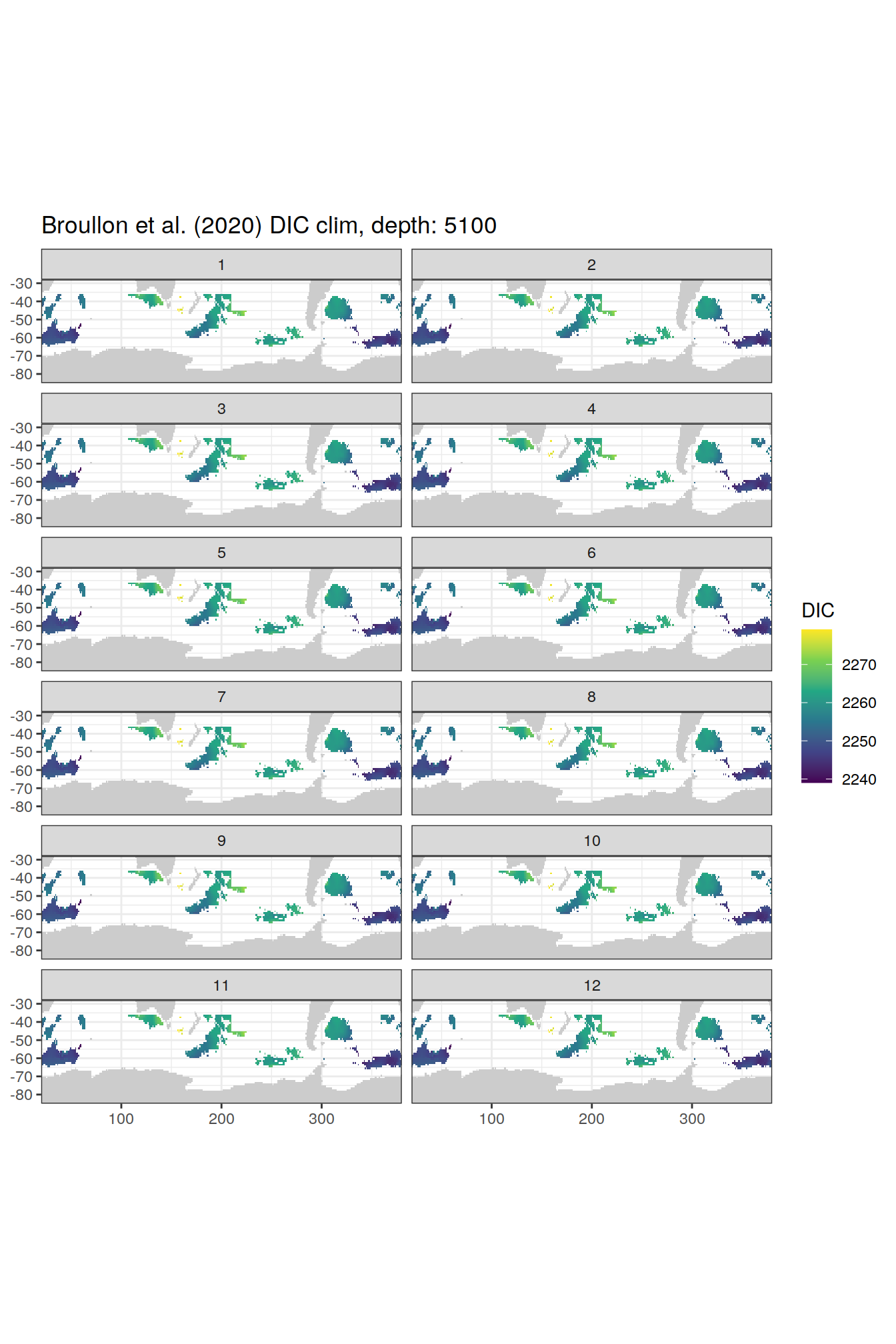
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[99]]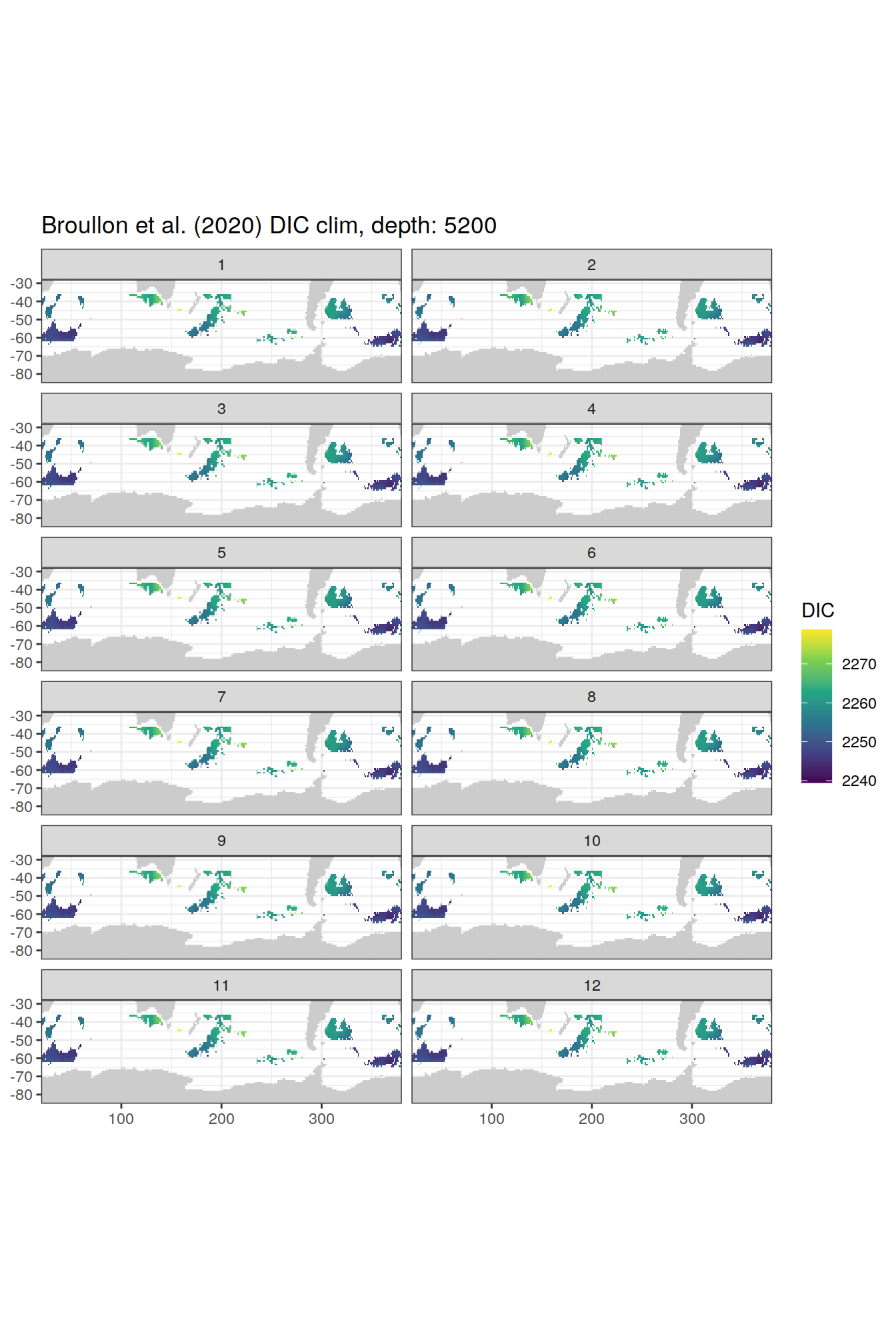
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[100]]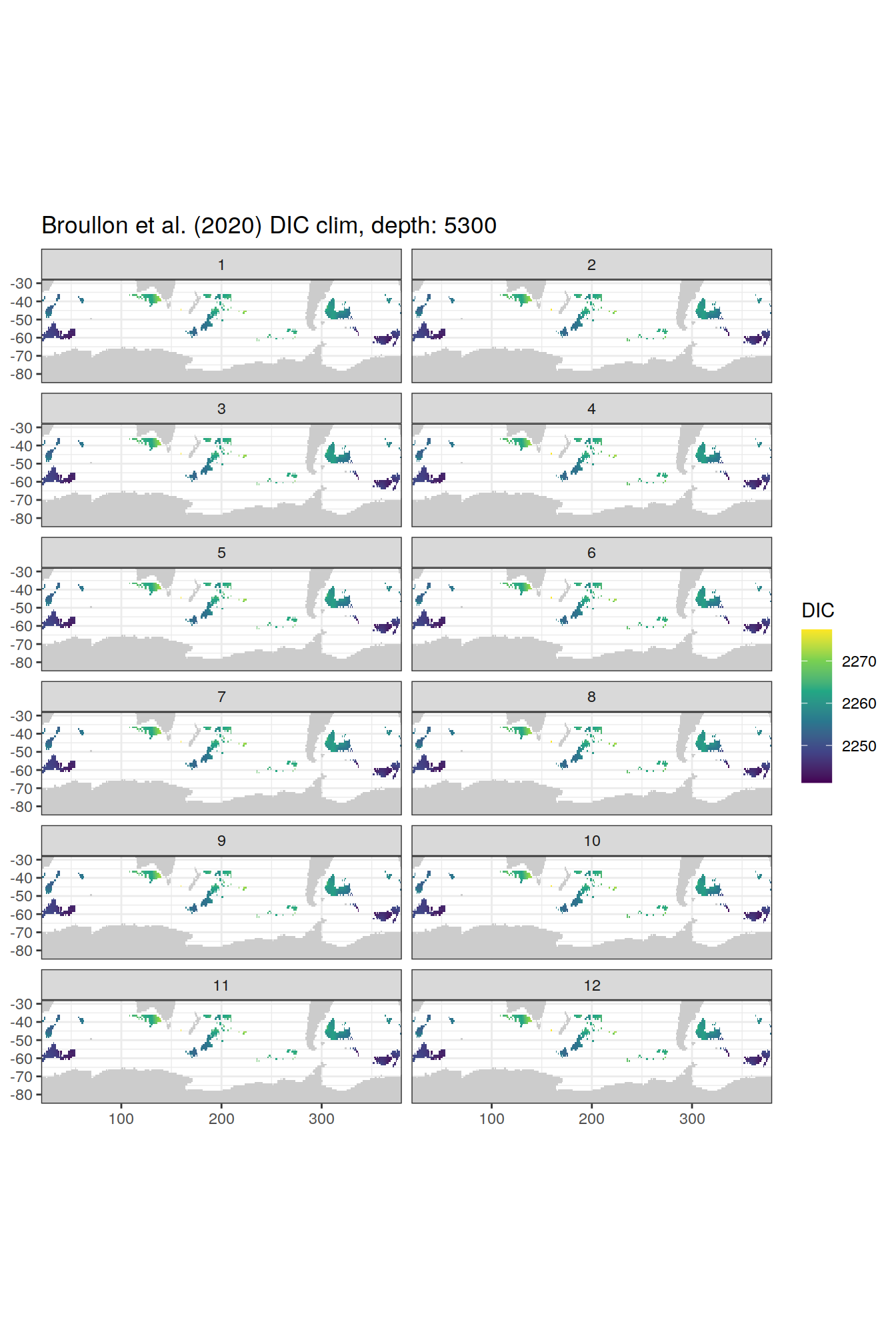
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[101]]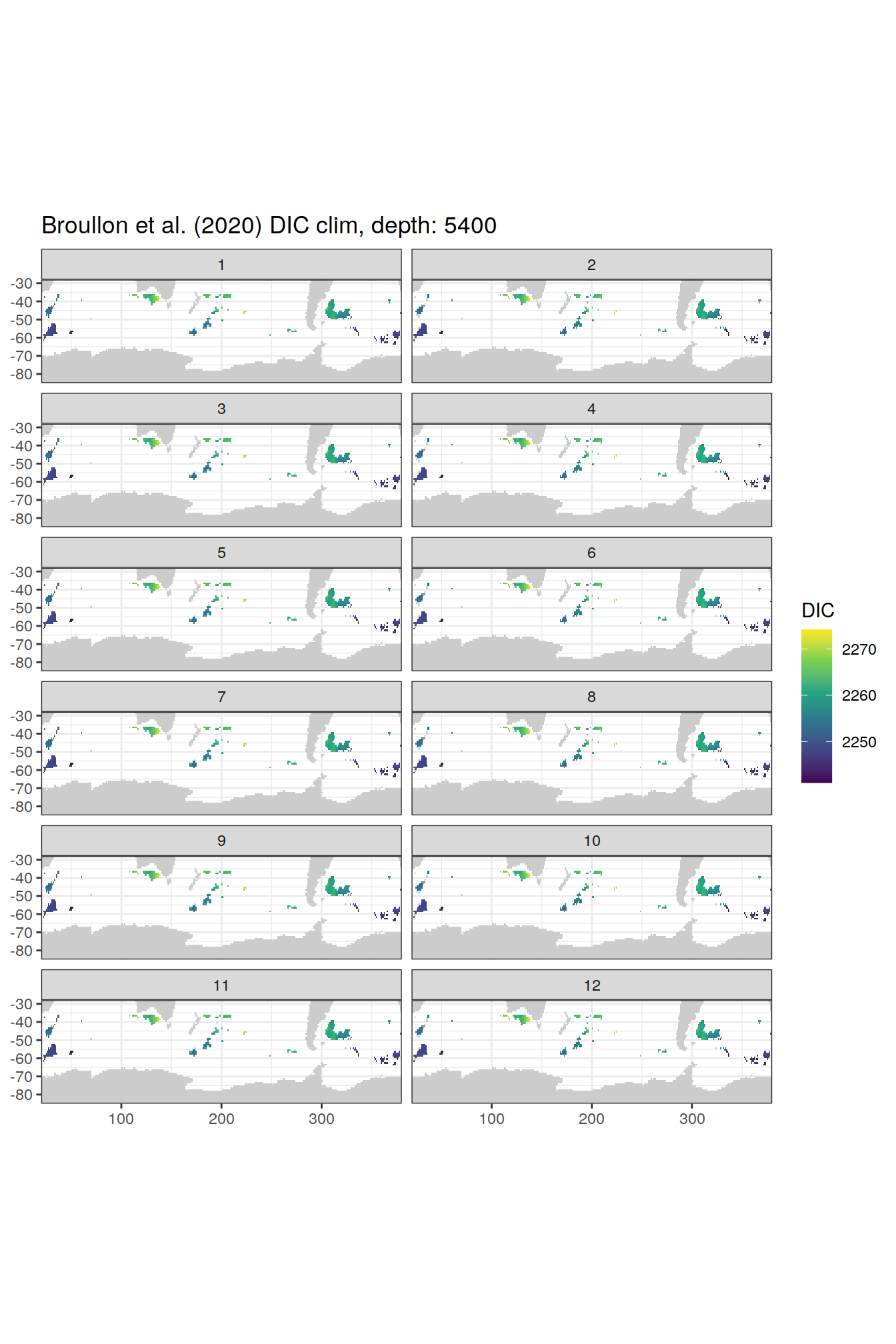
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[102]]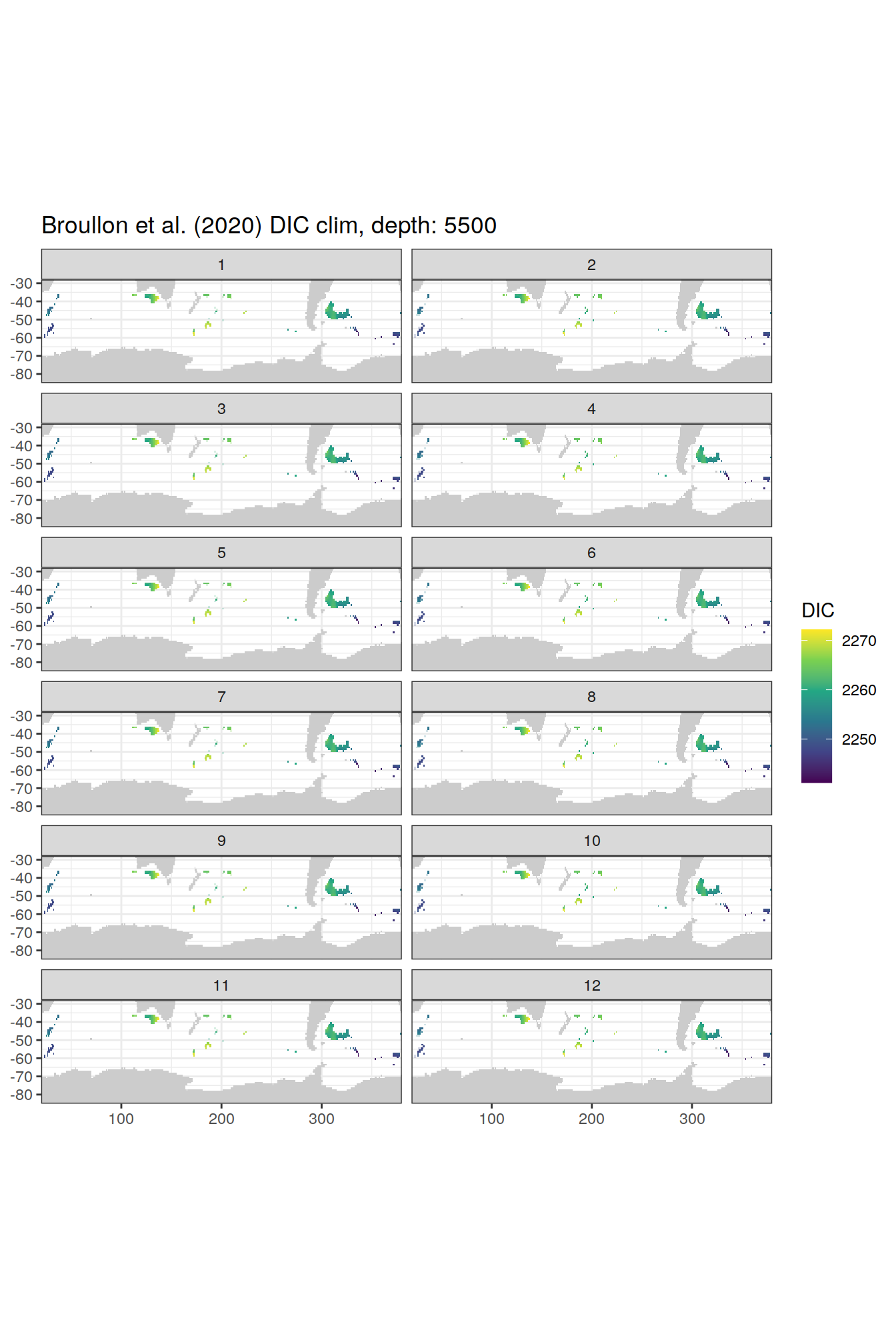
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
broullon_clim_SO %>%
# filter(depth <= 1500) %>%
group_split(month) %>%
map(
~ ggplot(data = .x,
aes(x = DIC,
y = depth))+
geom_point(data = .x,
aes(x = DIC,
y = depth),
size = 0.2,
pch = 1)+
scale_y_reverse()+
facet_grid(biome_name~basin_AIP)+
labs(title = paste0('Broullon et al. clim DIC, month: ', unique(.x$month)),
x = 'DIC')
)[[1]]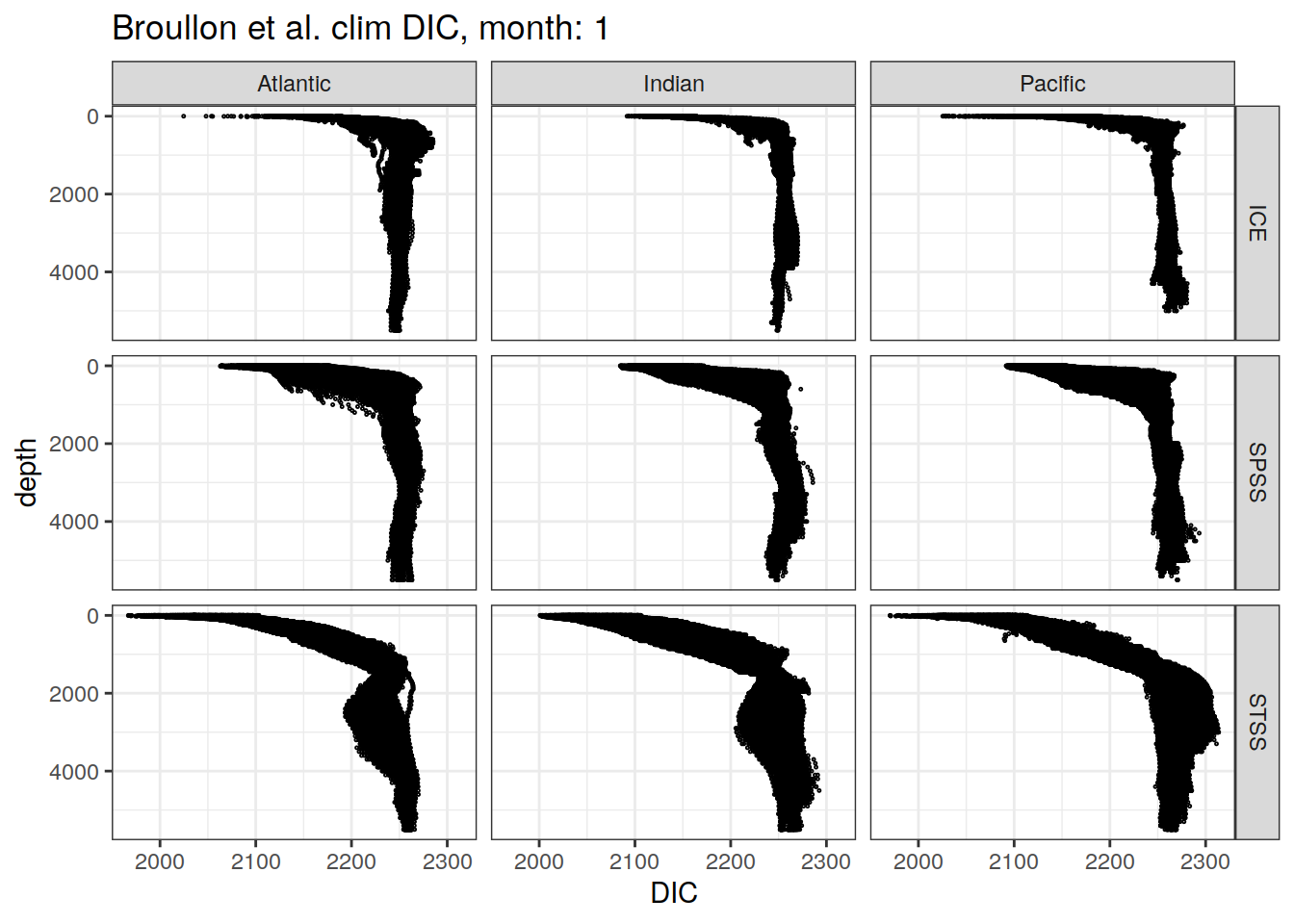
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[2]]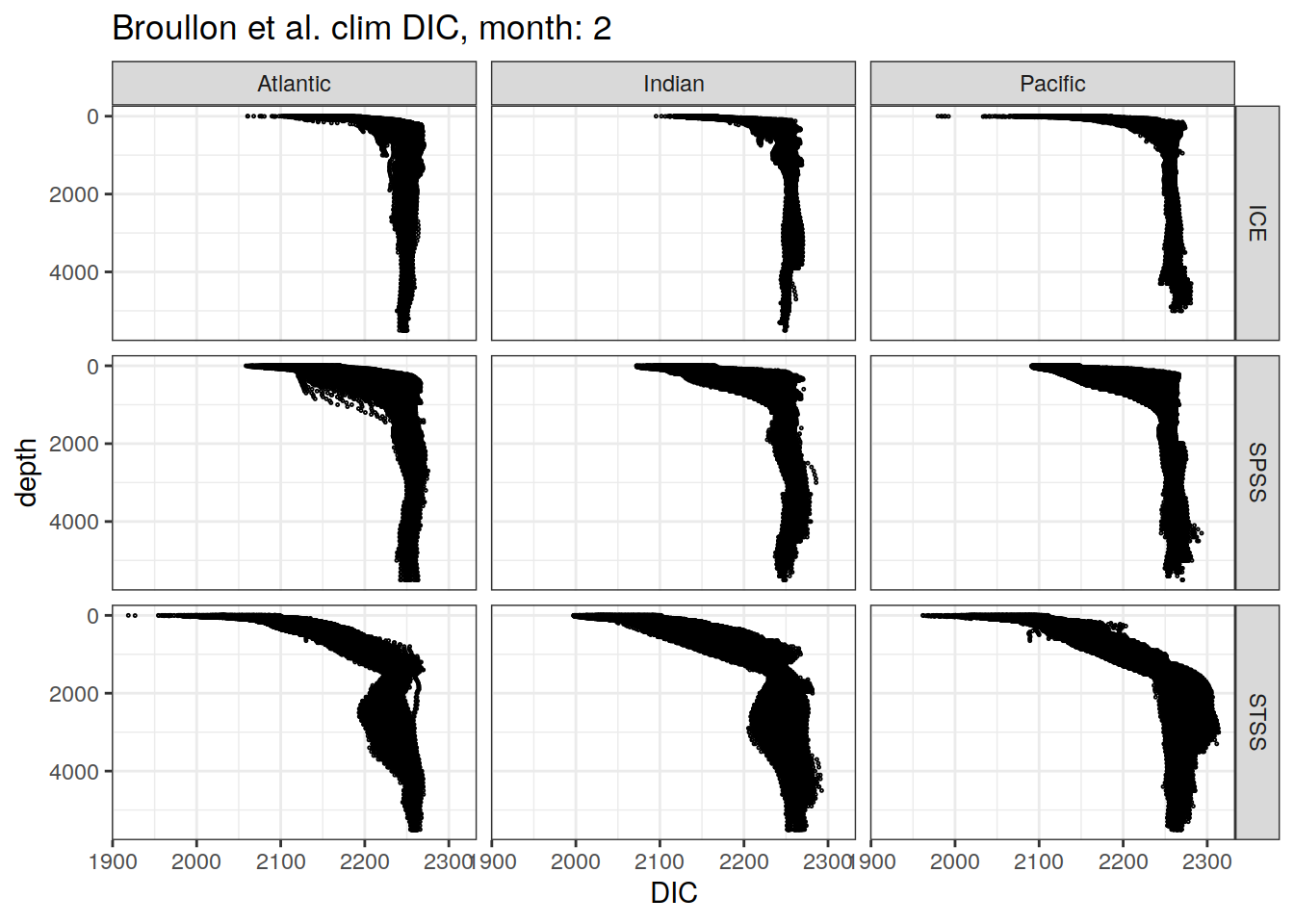
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[3]]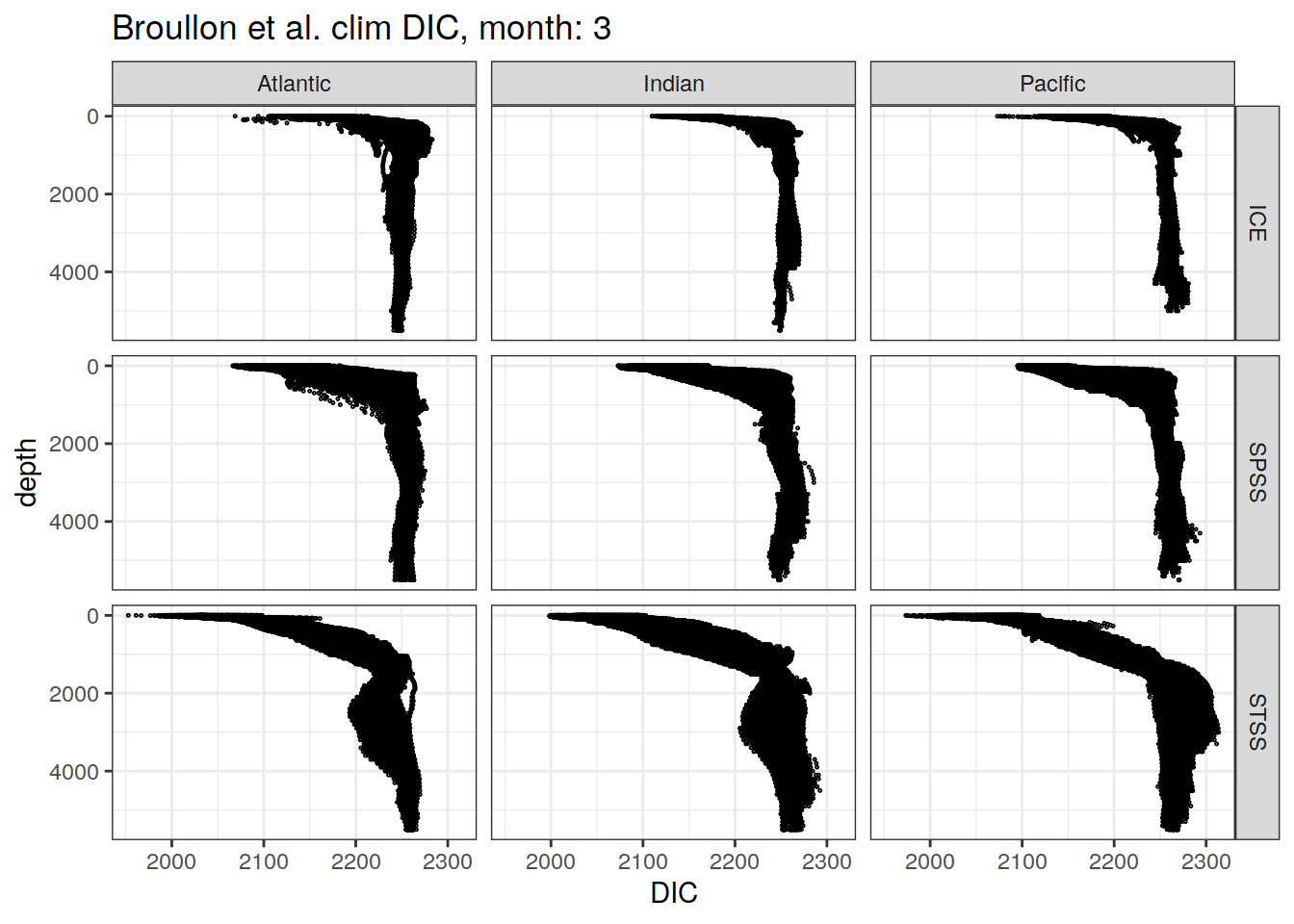
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[4]]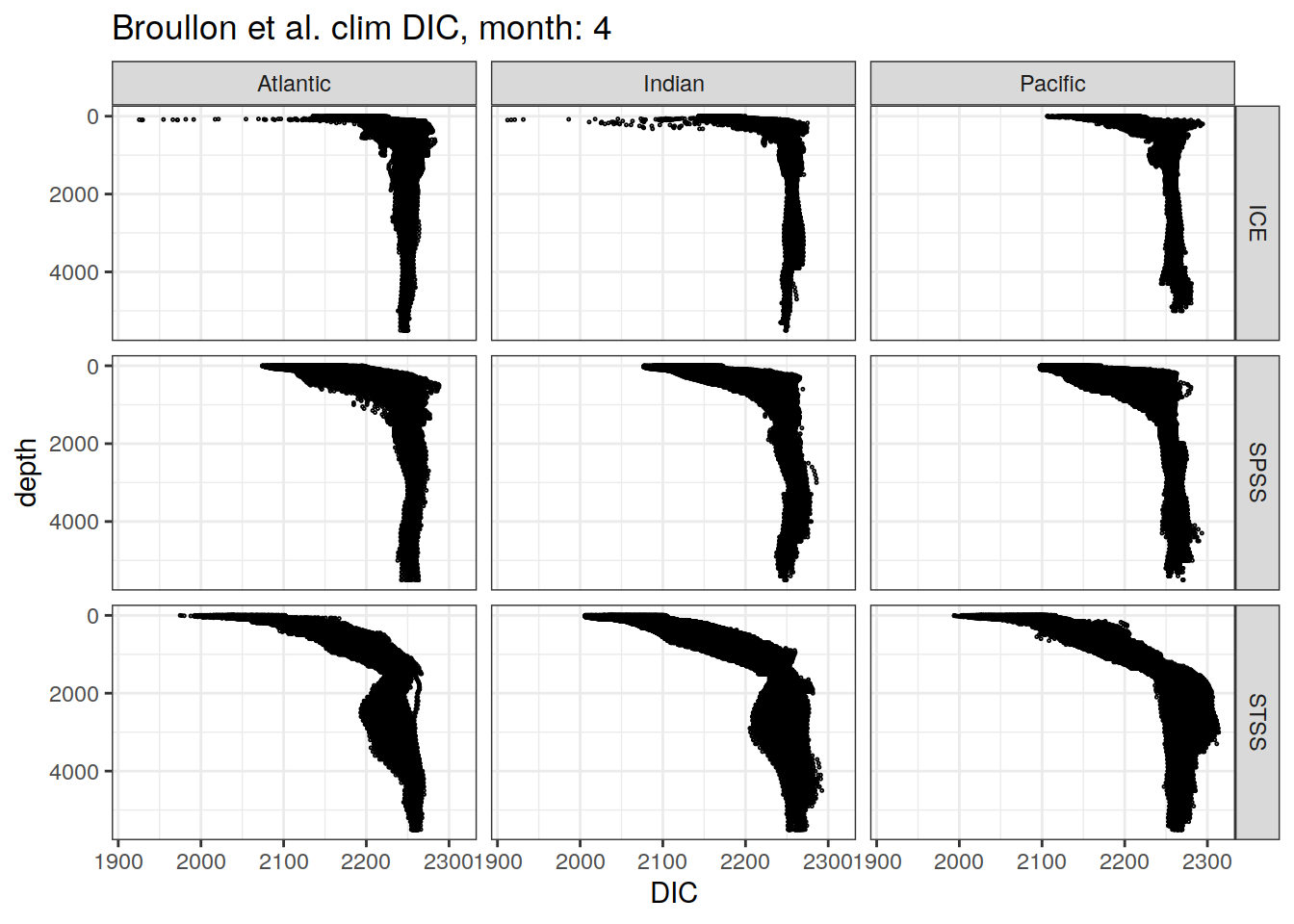
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[5]]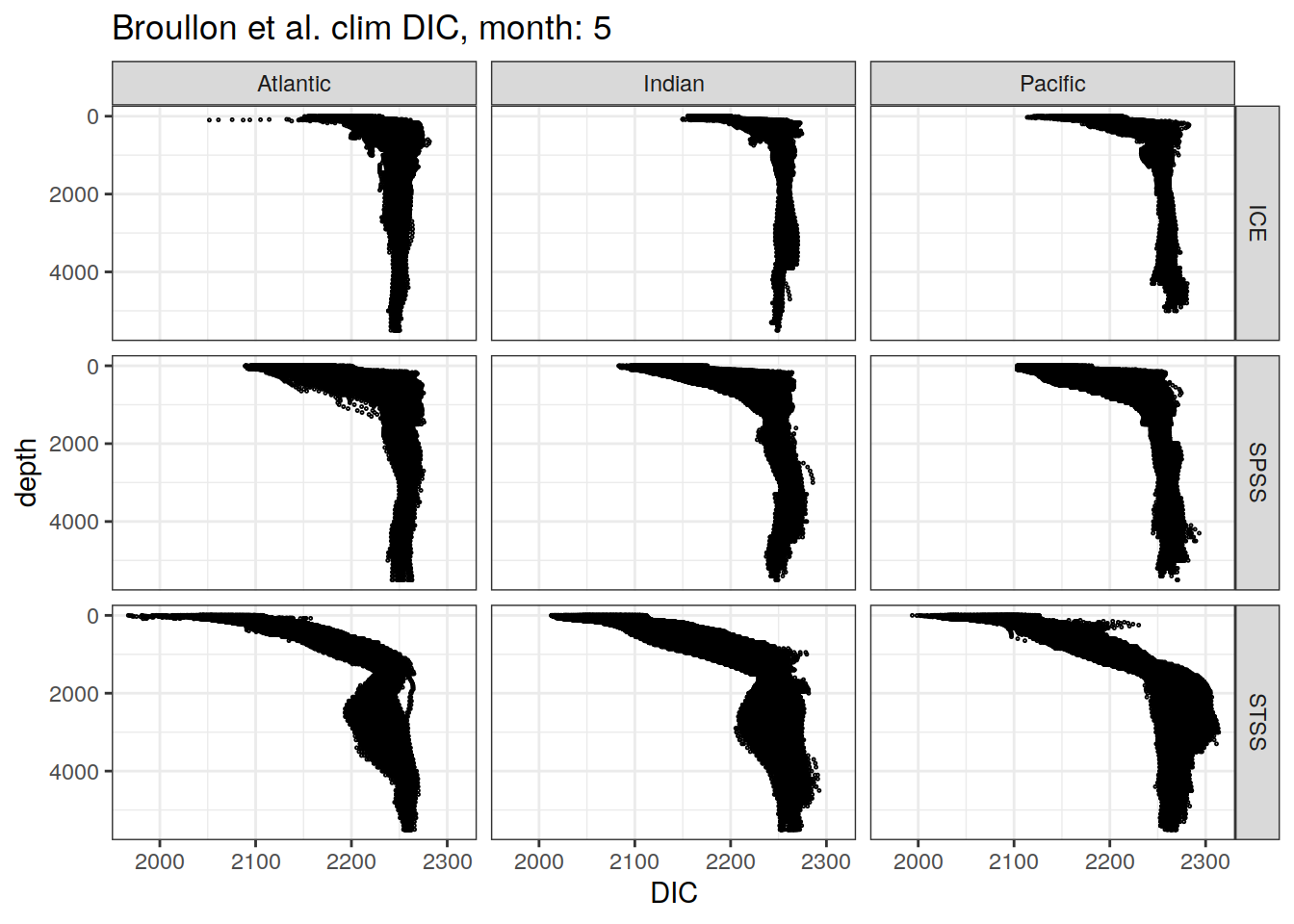
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[6]]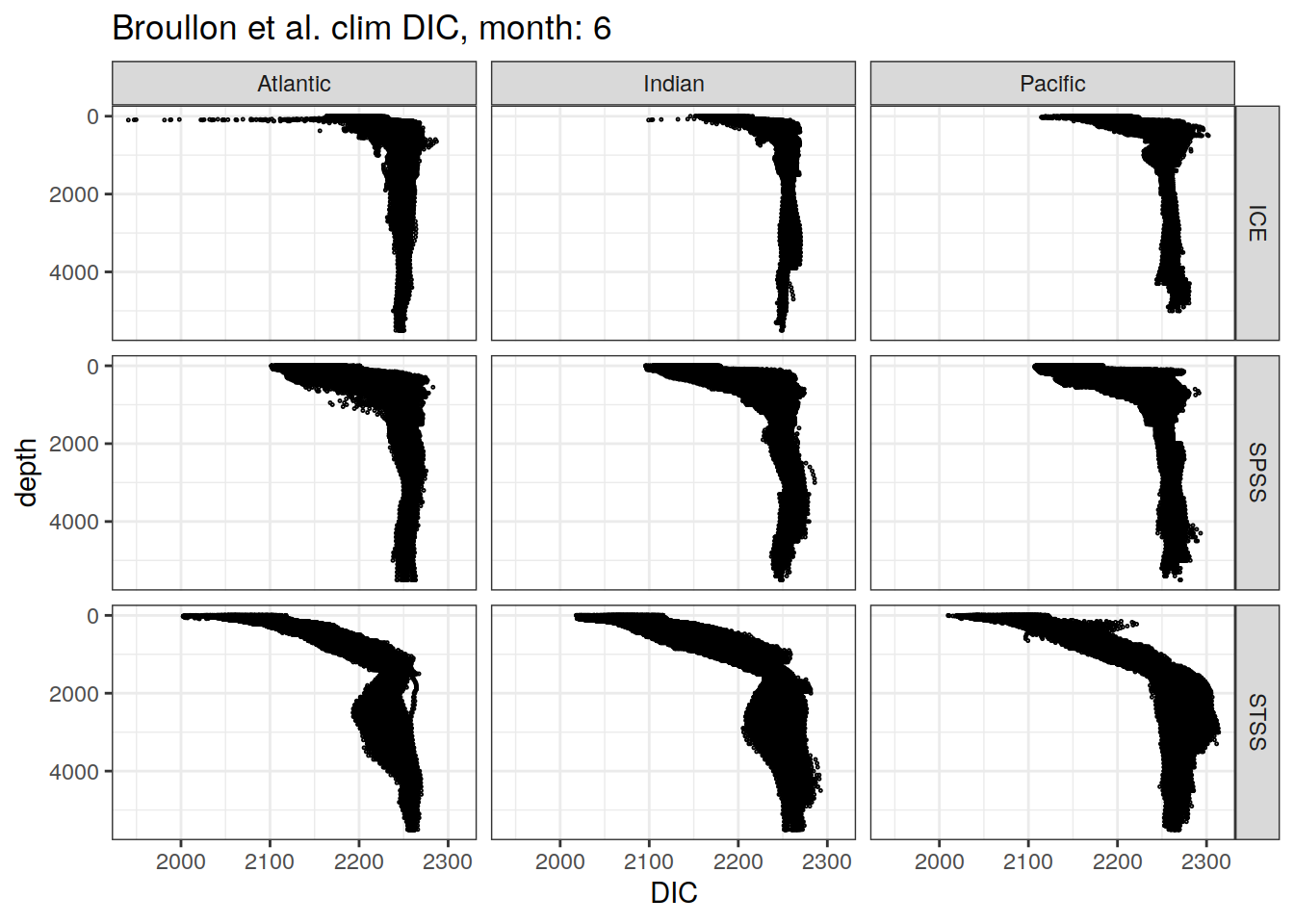
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[7]]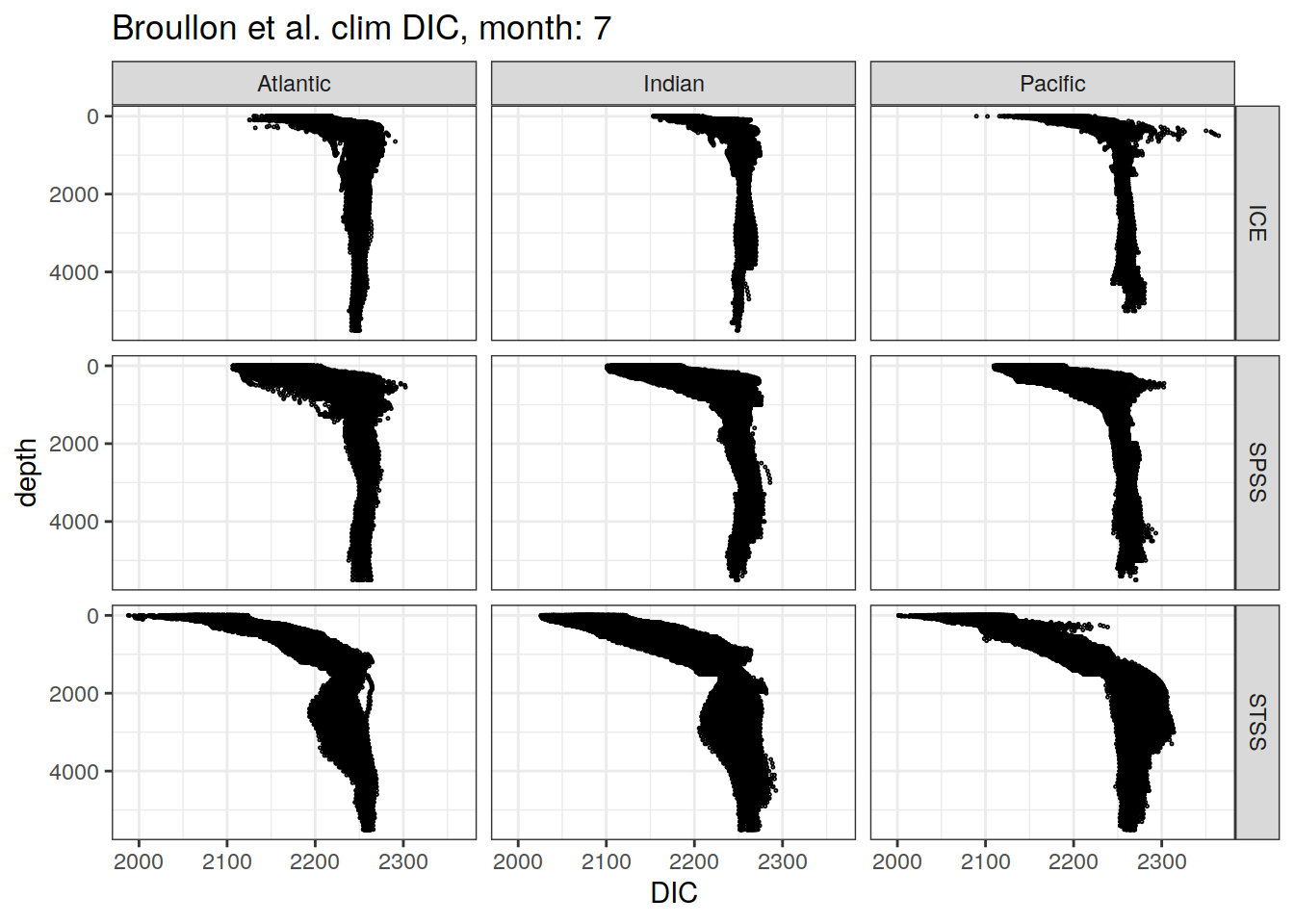
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[8]]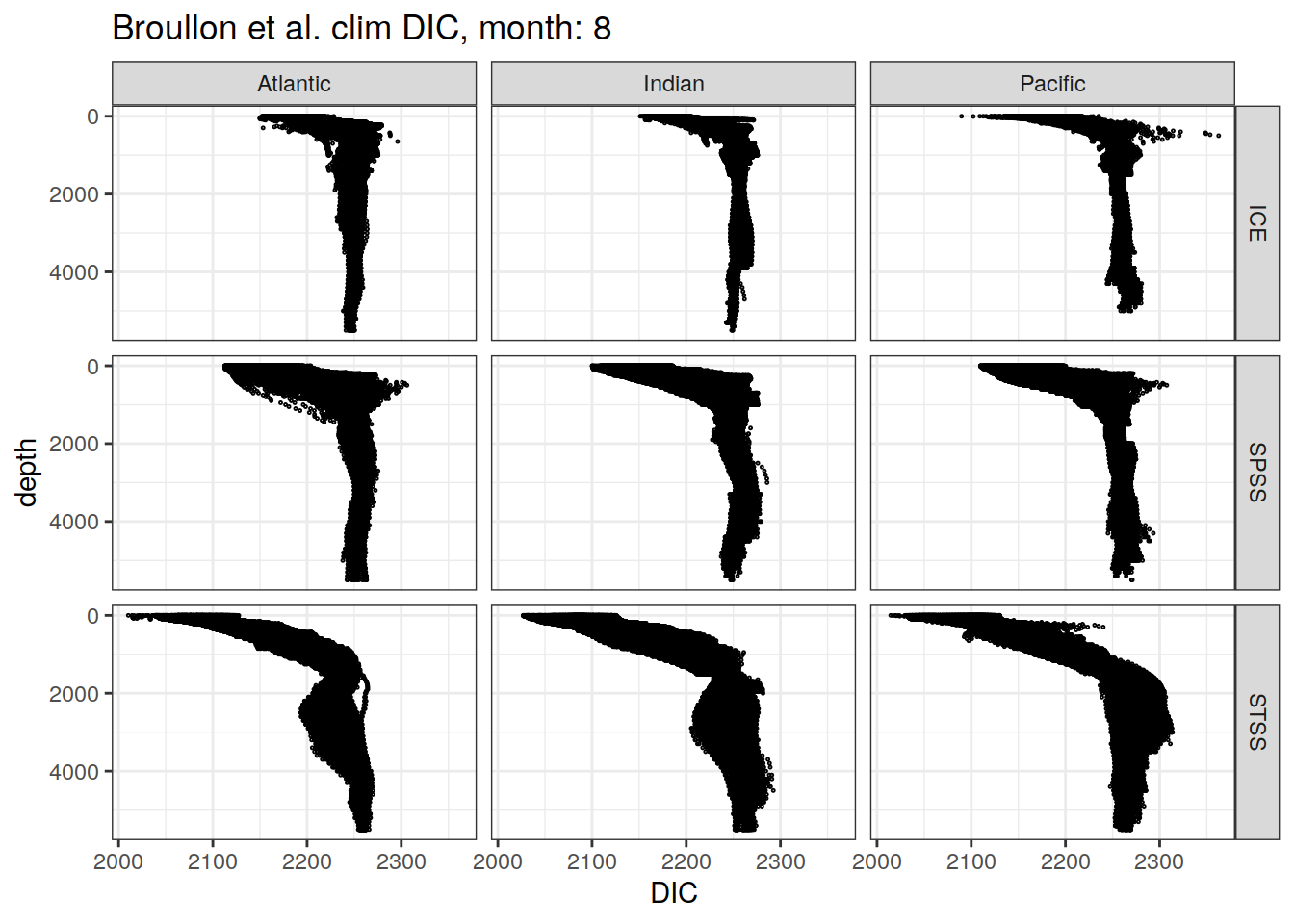
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[9]]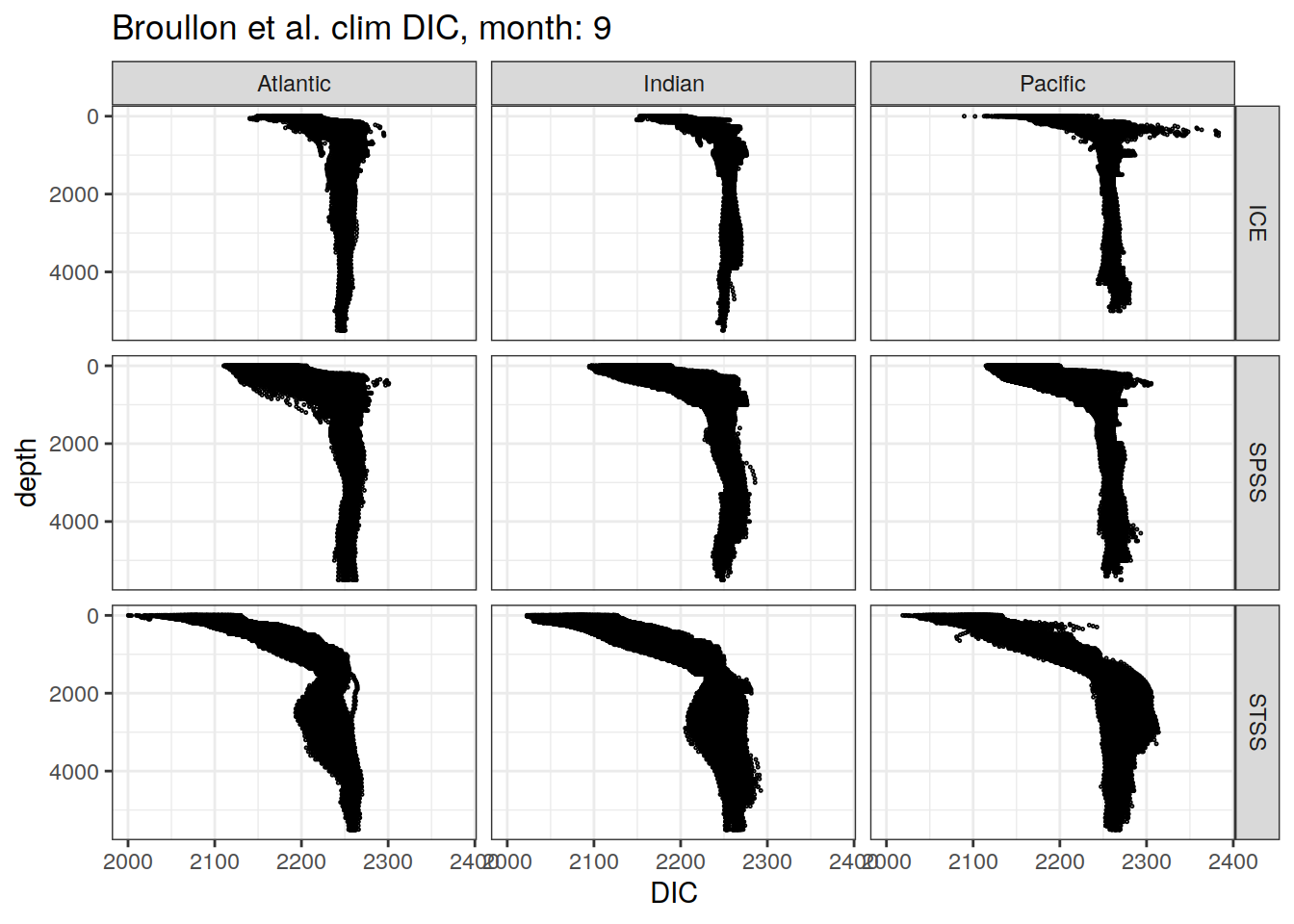
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[10]]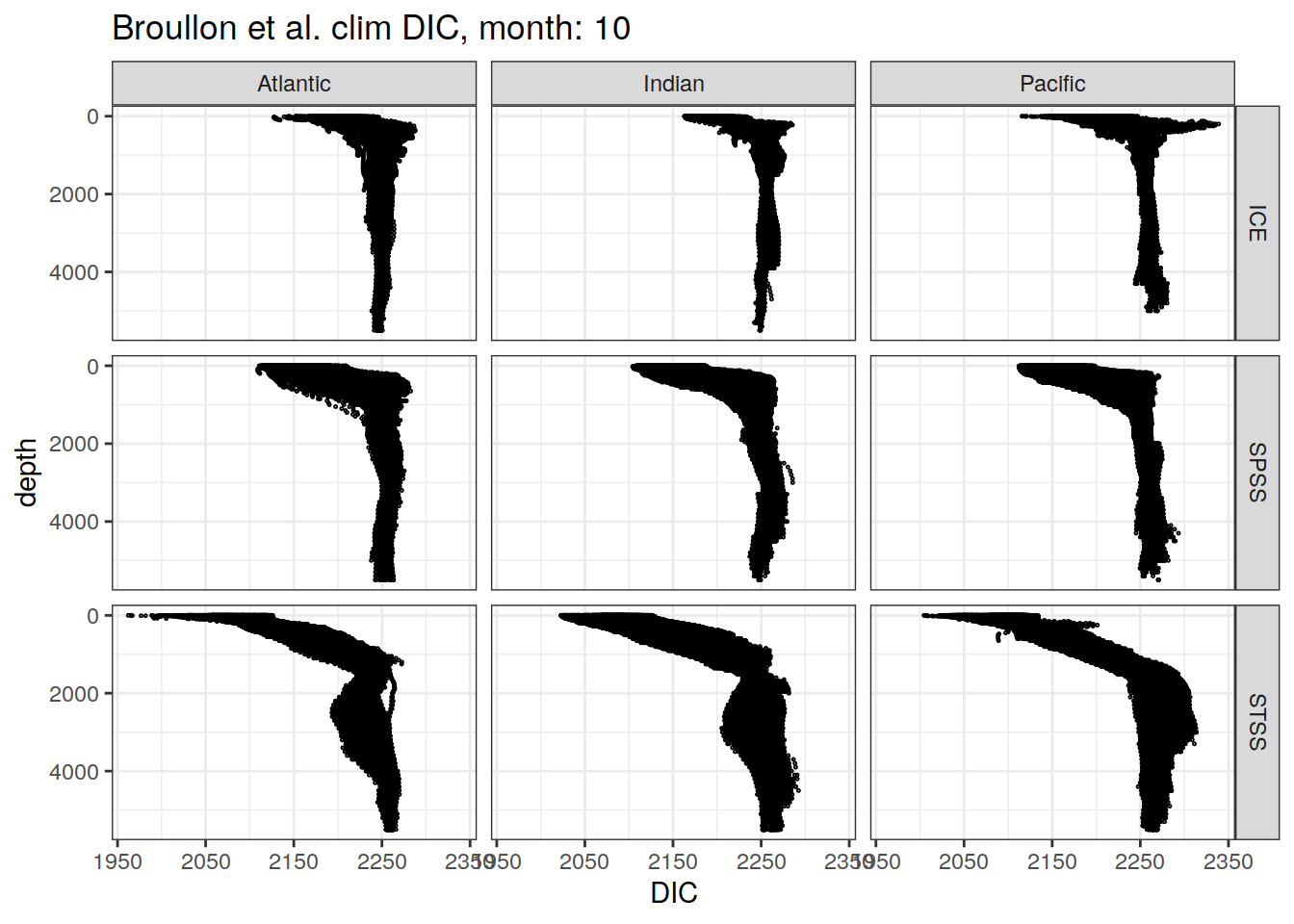
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[11]]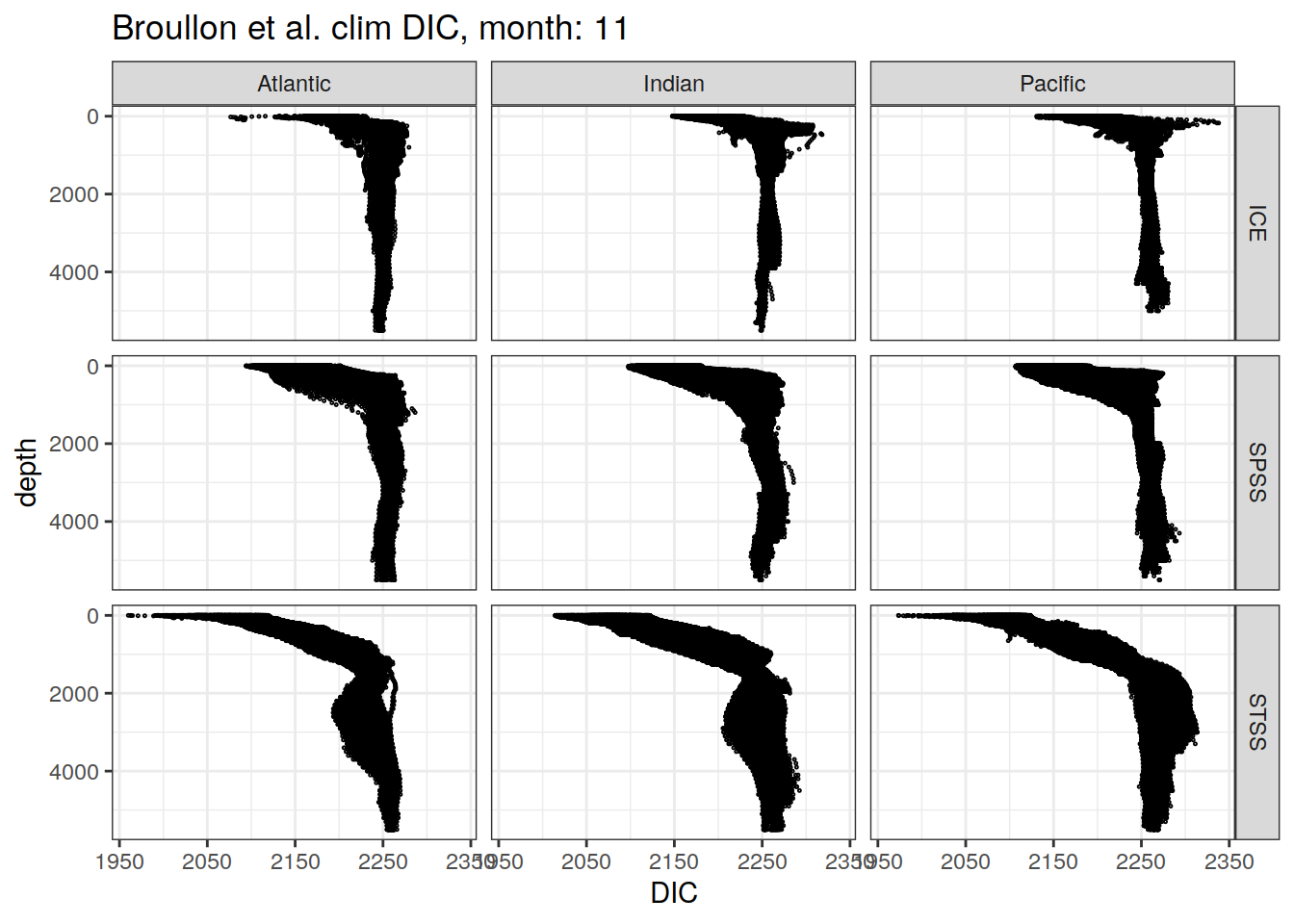
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[12]]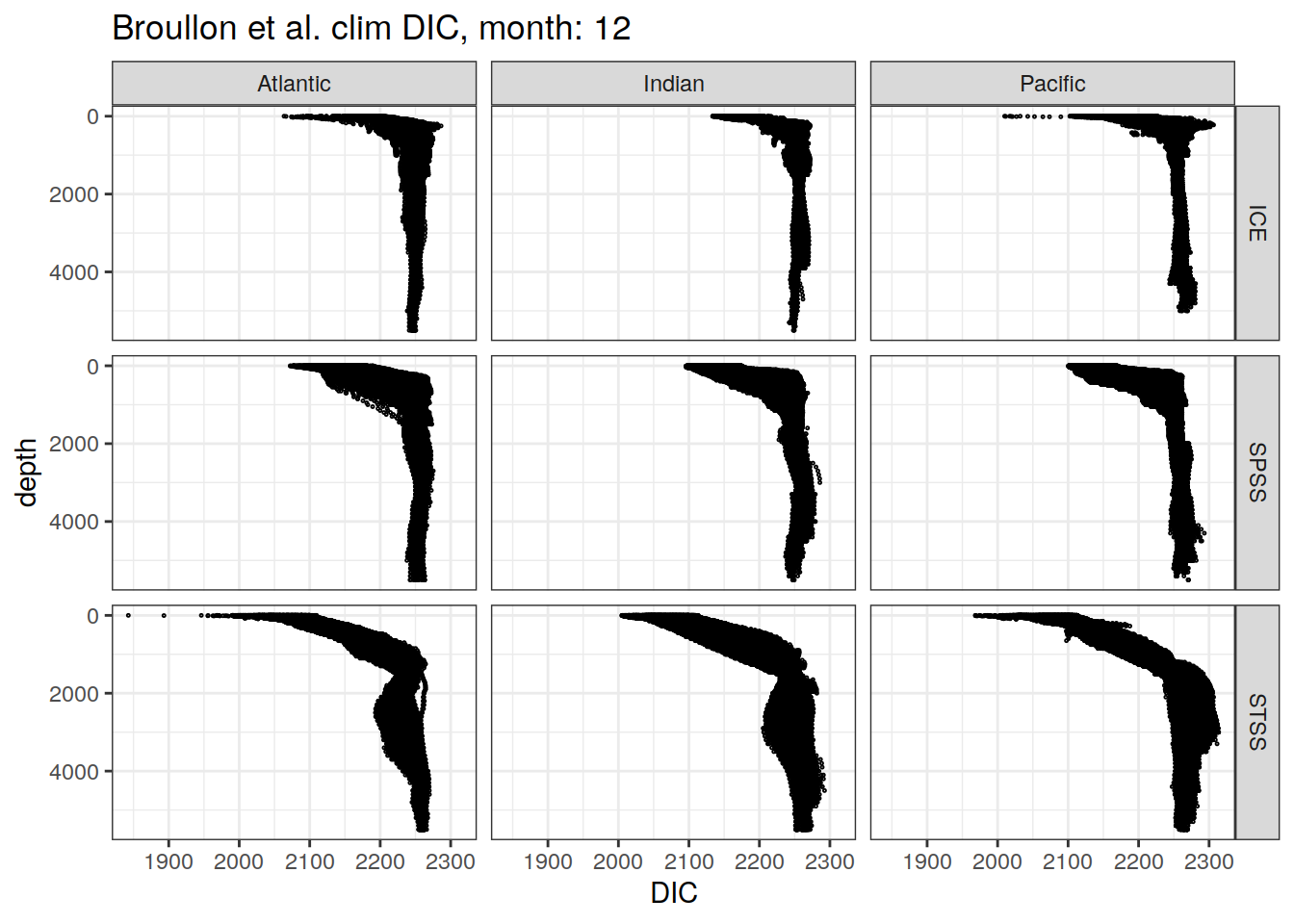
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
TA
broullon_clim_SO %>%
group_split(depth) %>%
map(
~map +
geom_tile(data = .x,
aes(x = lon,
y = lat,
fill = TA))+
scale_fill_viridis_c()+
lims(y = c(-85, -28))+
facet_wrap(~month, ncol = 2)+
labs(title = paste0('Broullon et al. (2020) TA clim, depth: ', unique(.x$depth)))
)[[1]]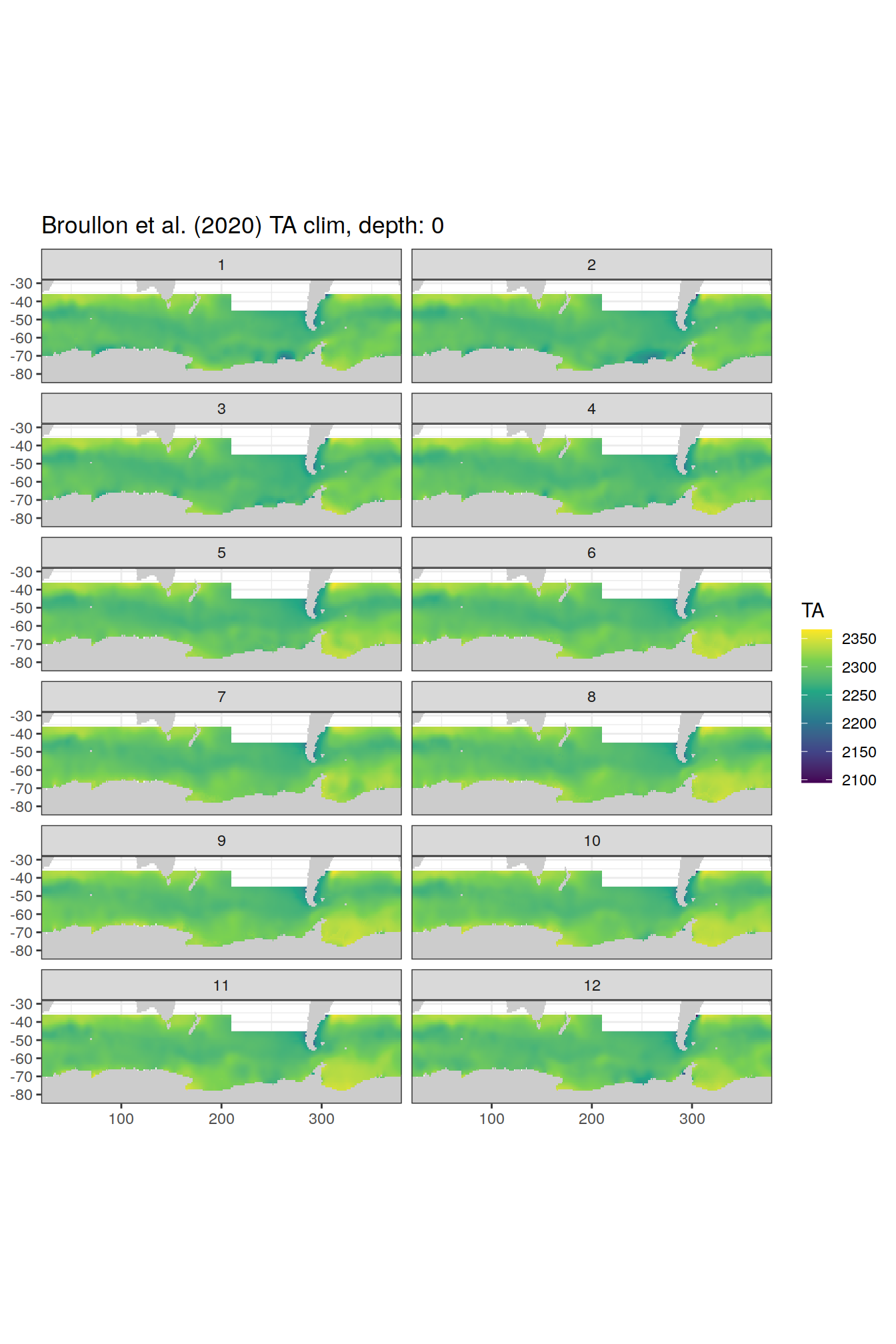
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[2]]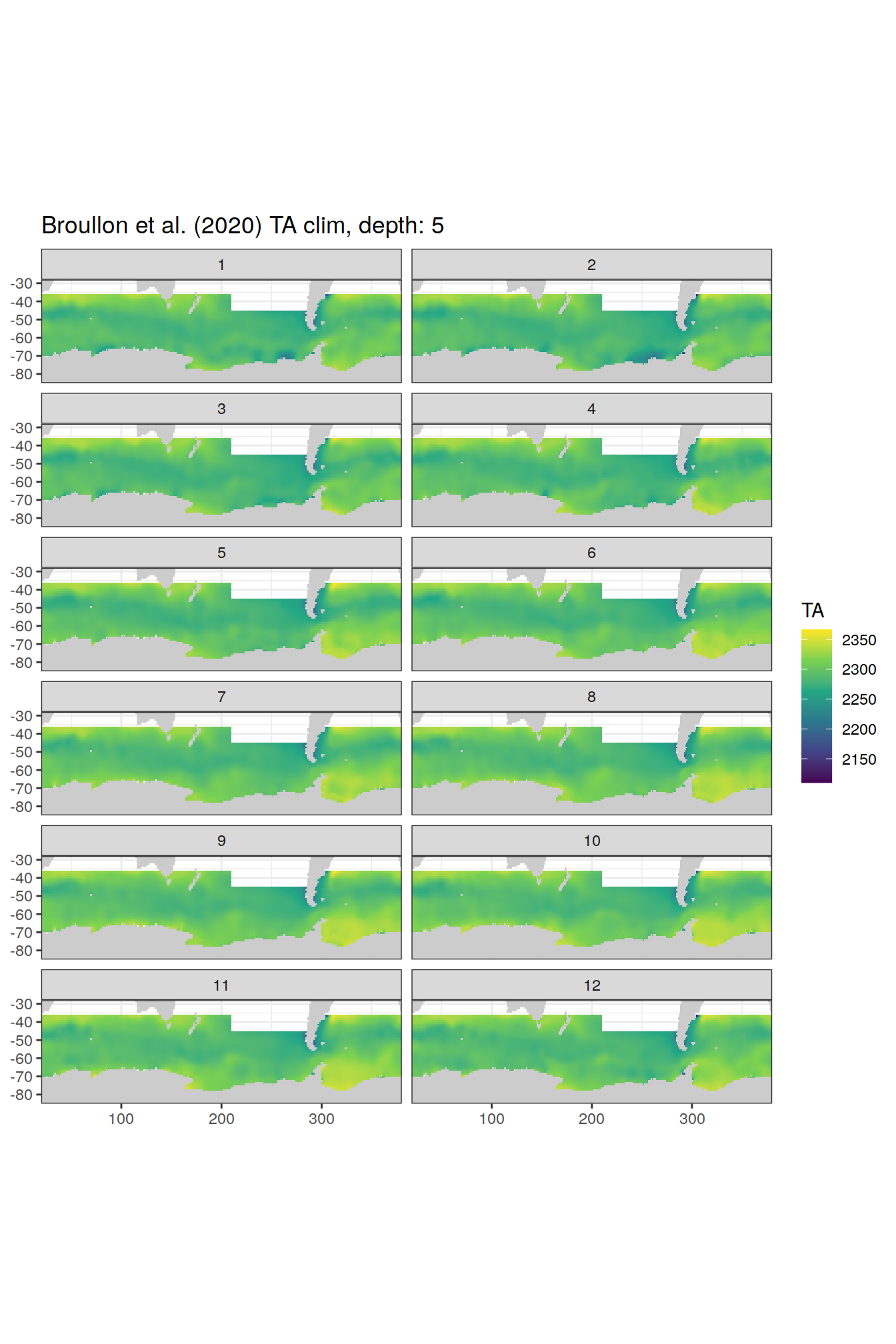
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[3]]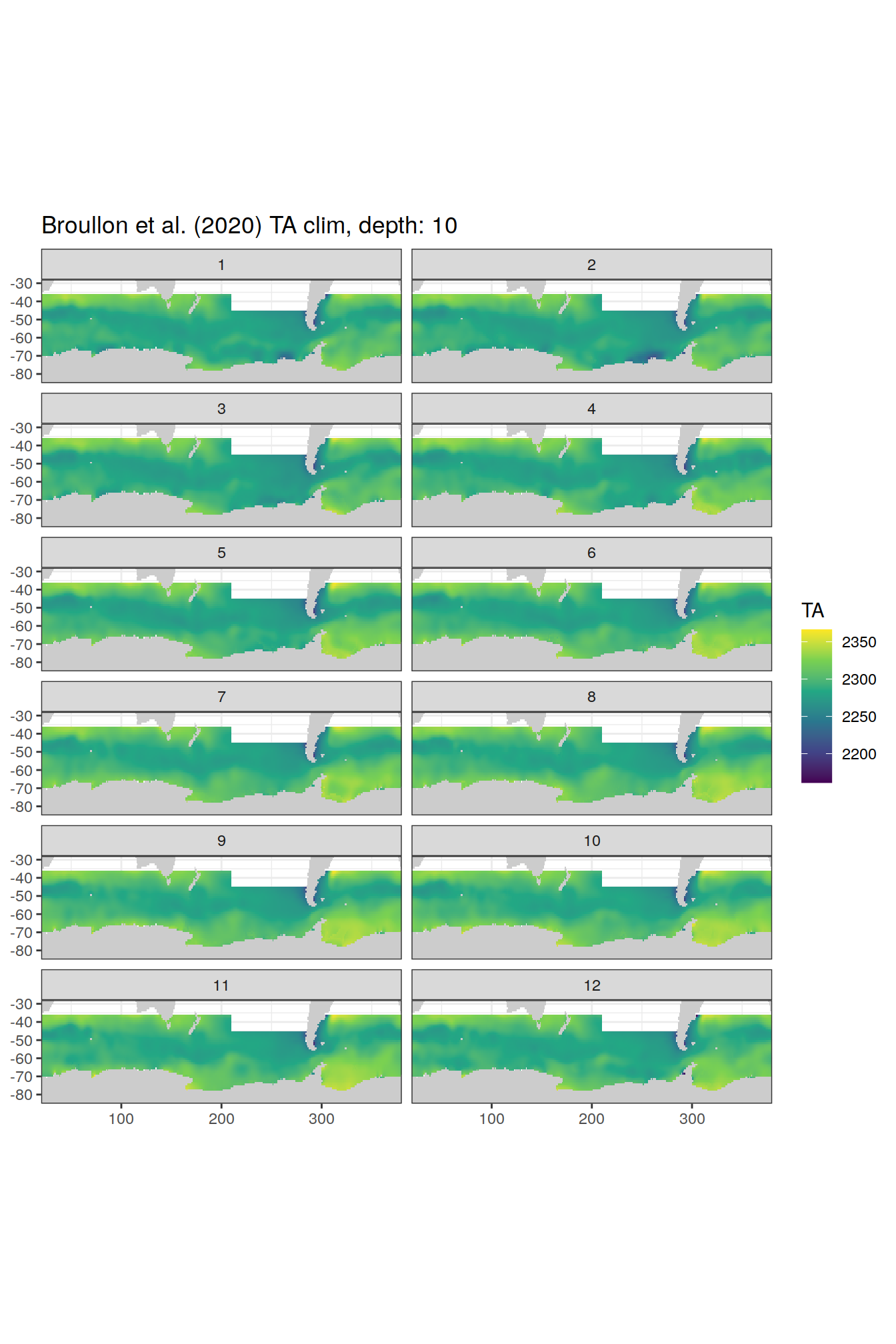
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[4]]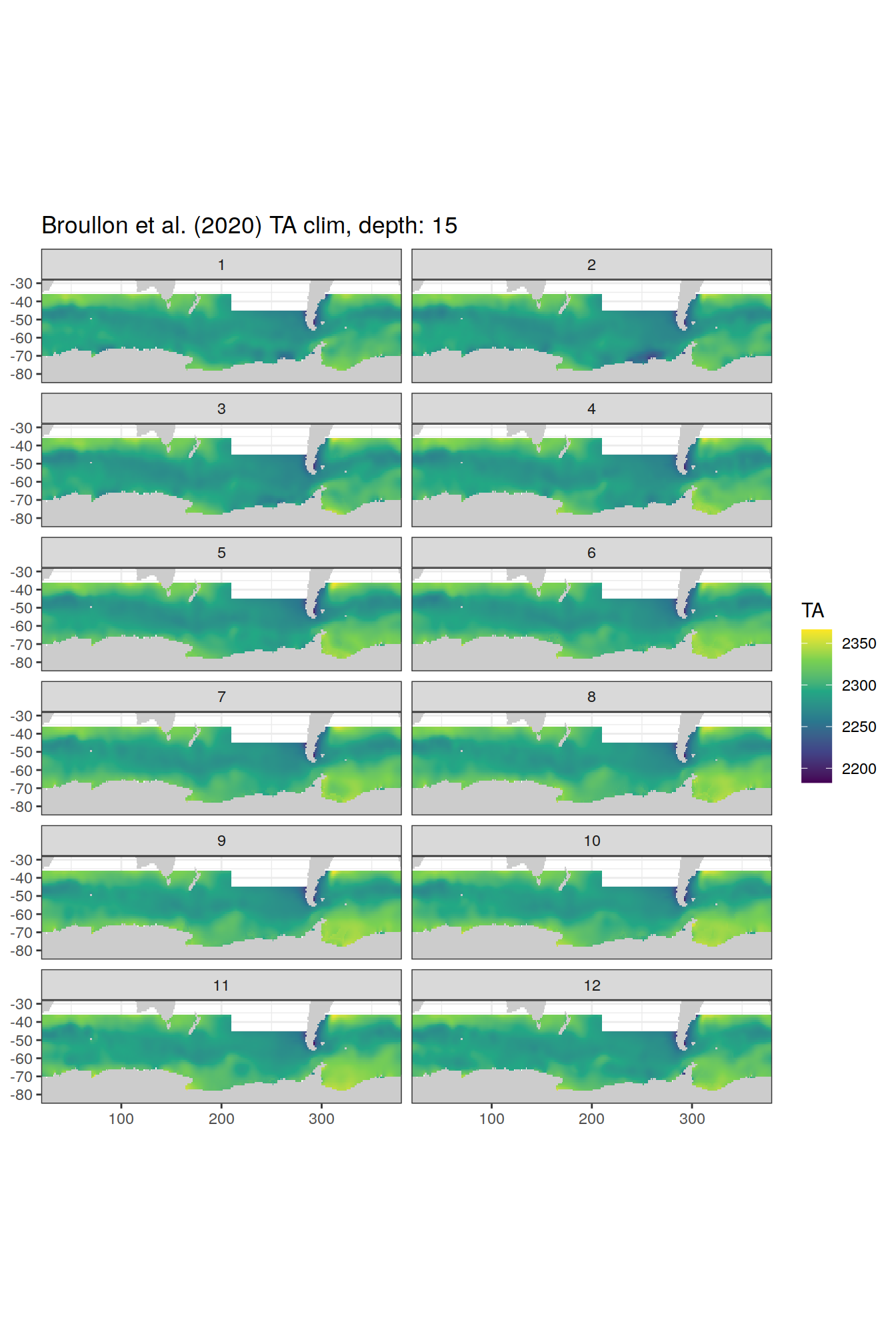
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[5]]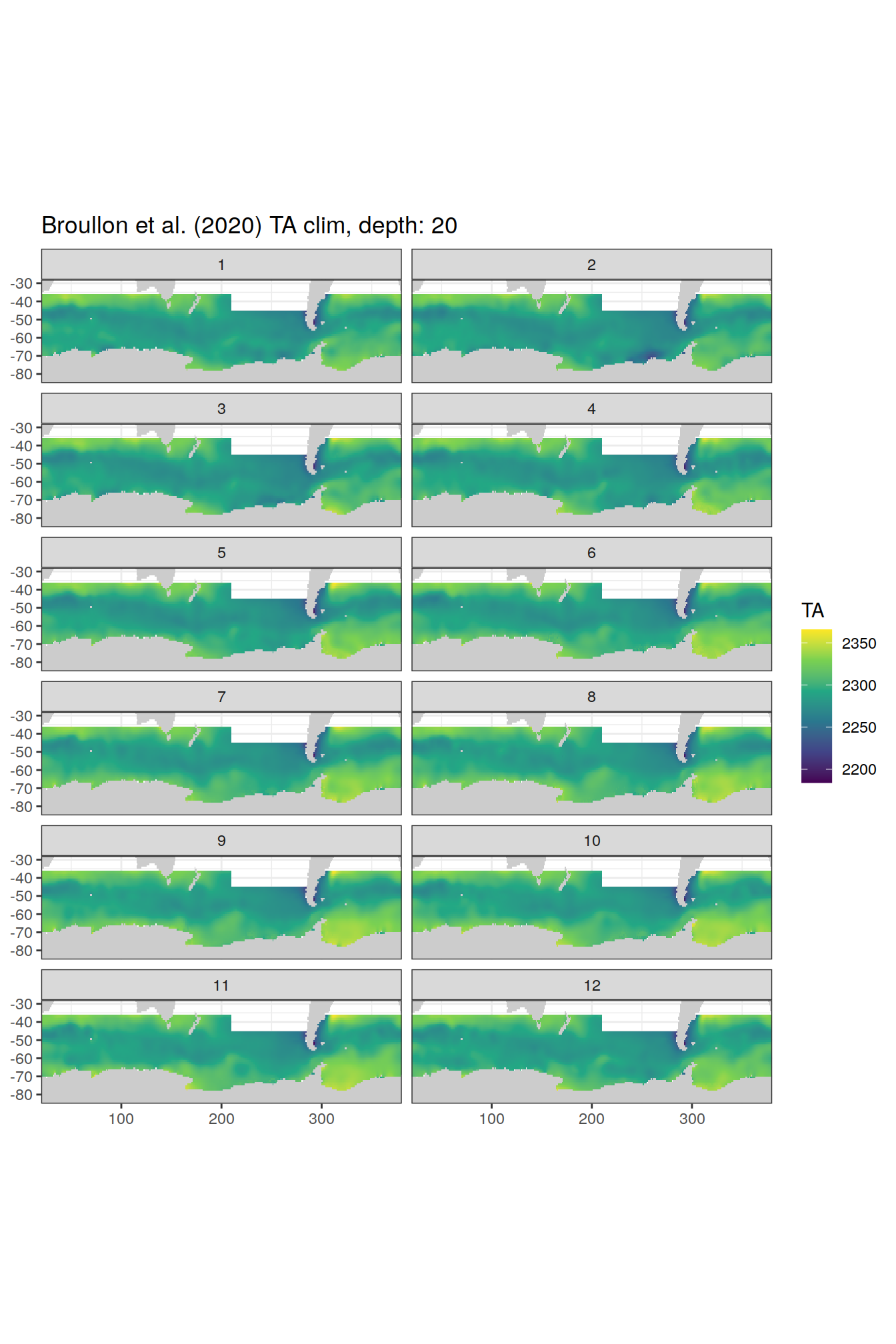
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[6]]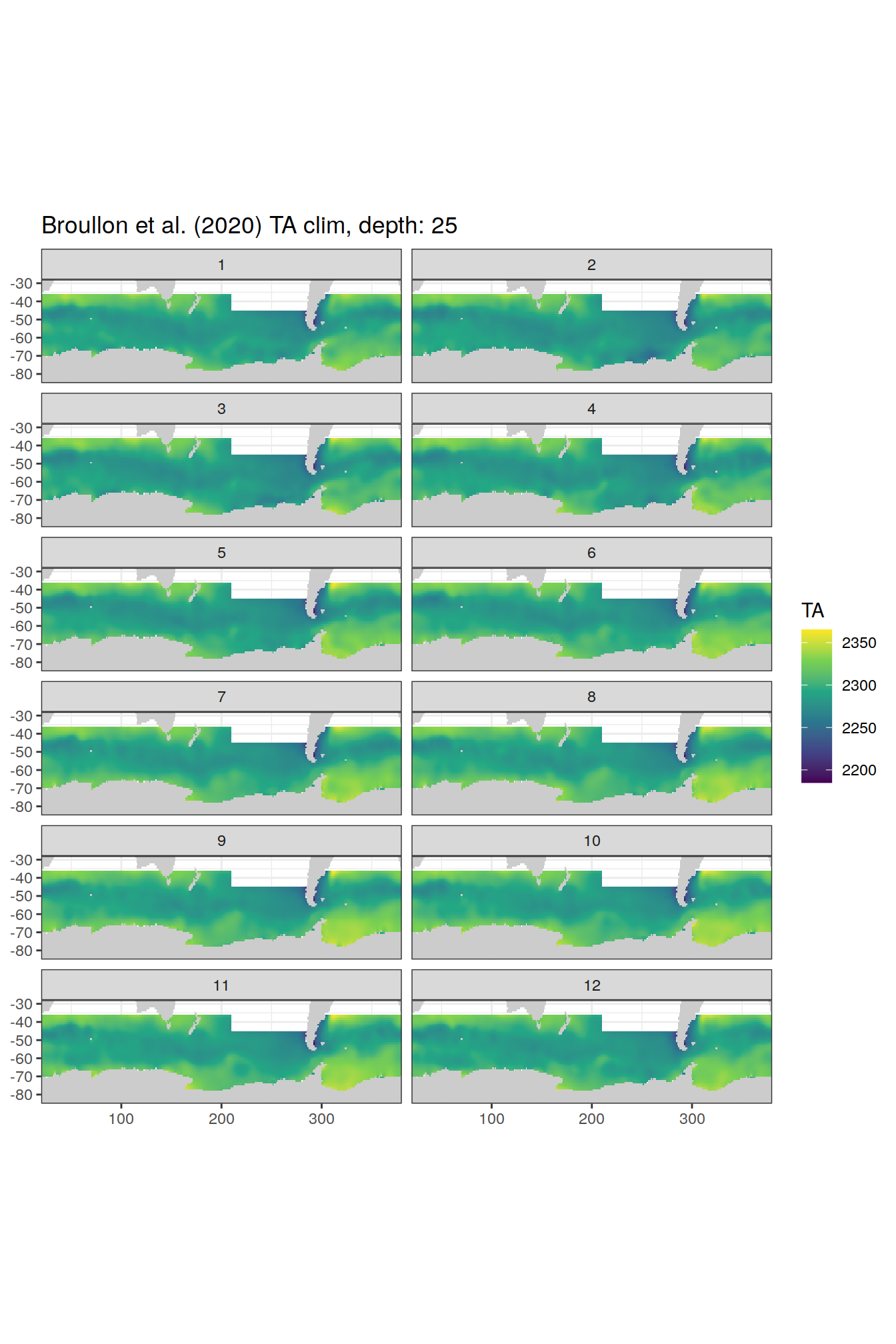
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[7]]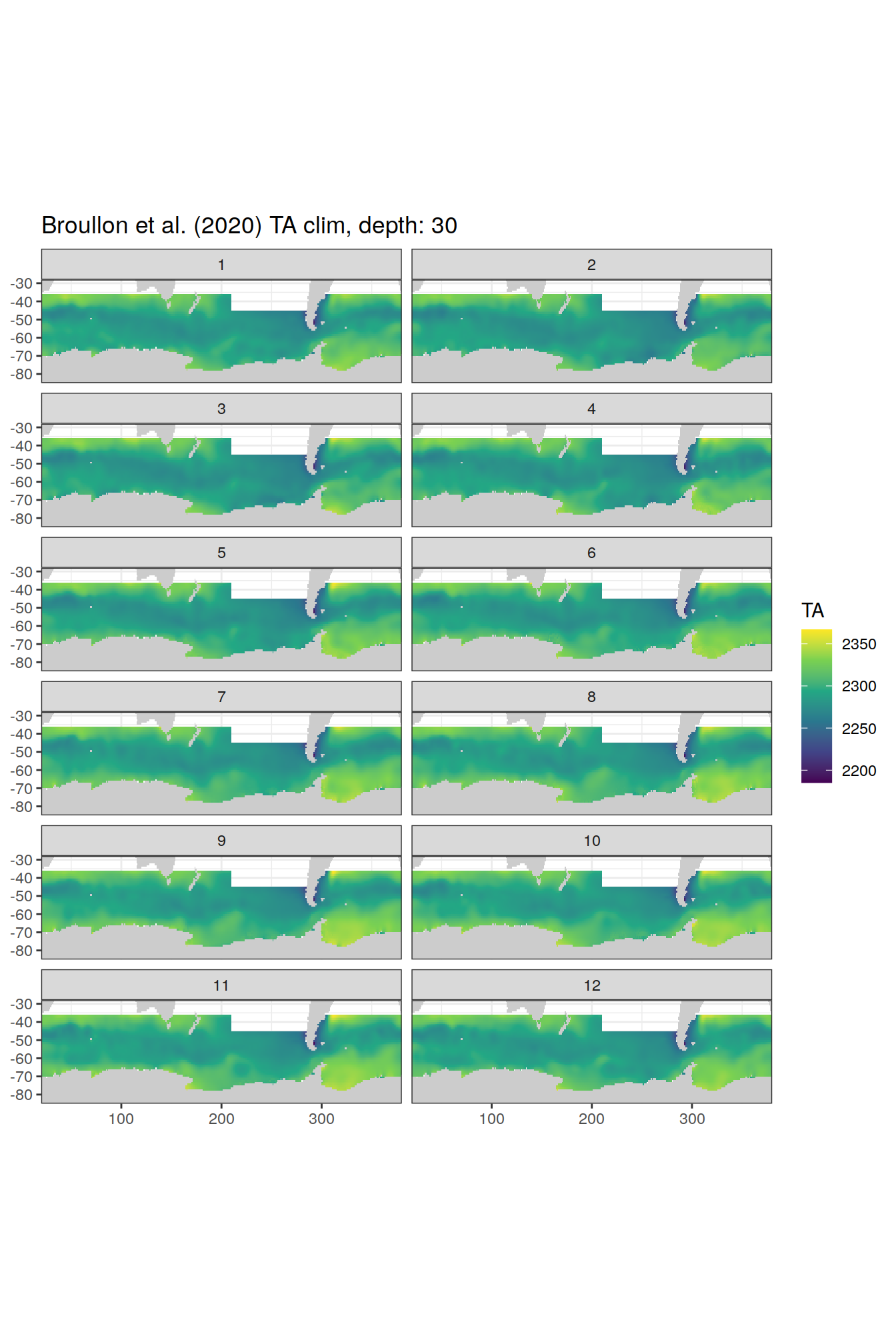
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[8]]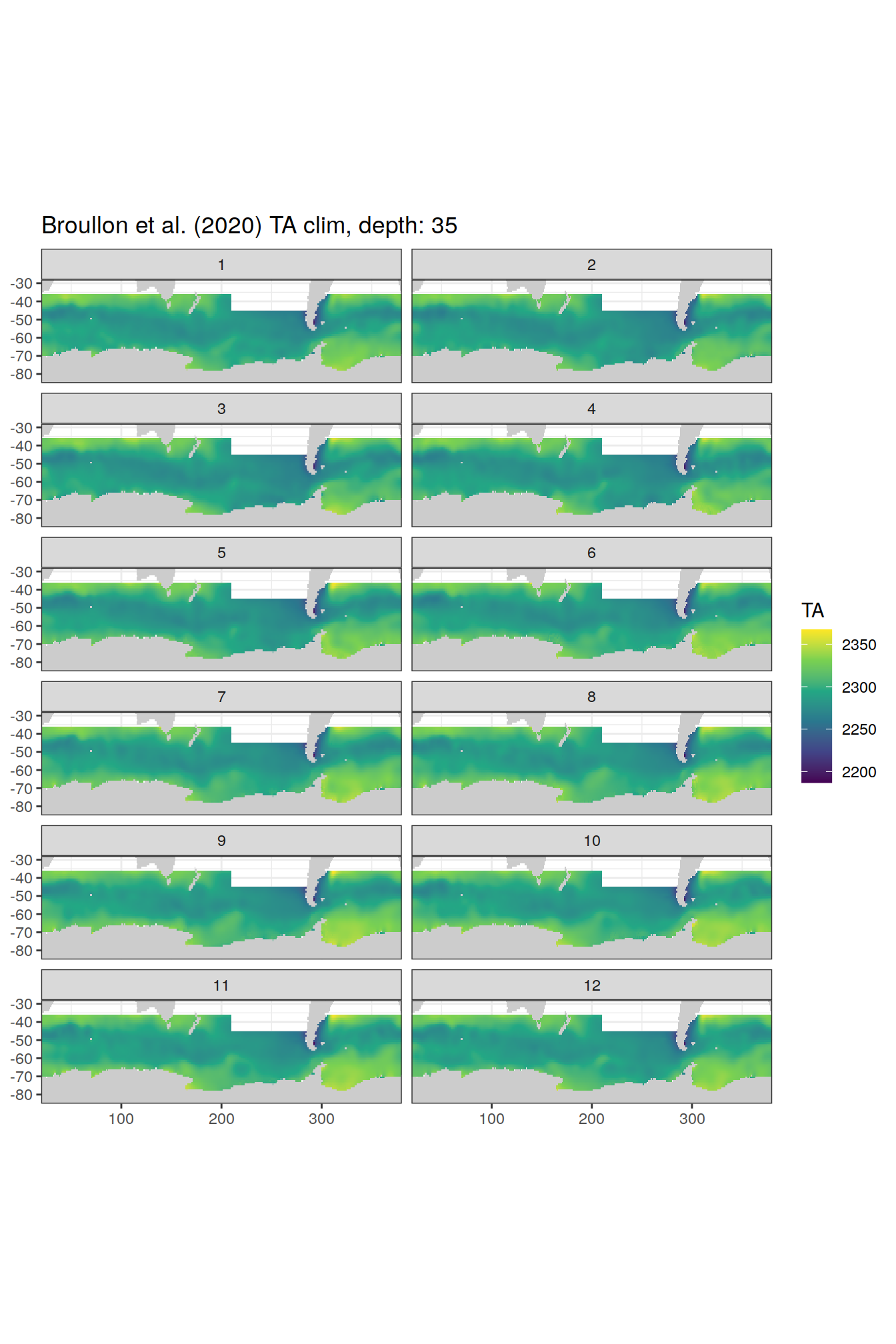
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[9]]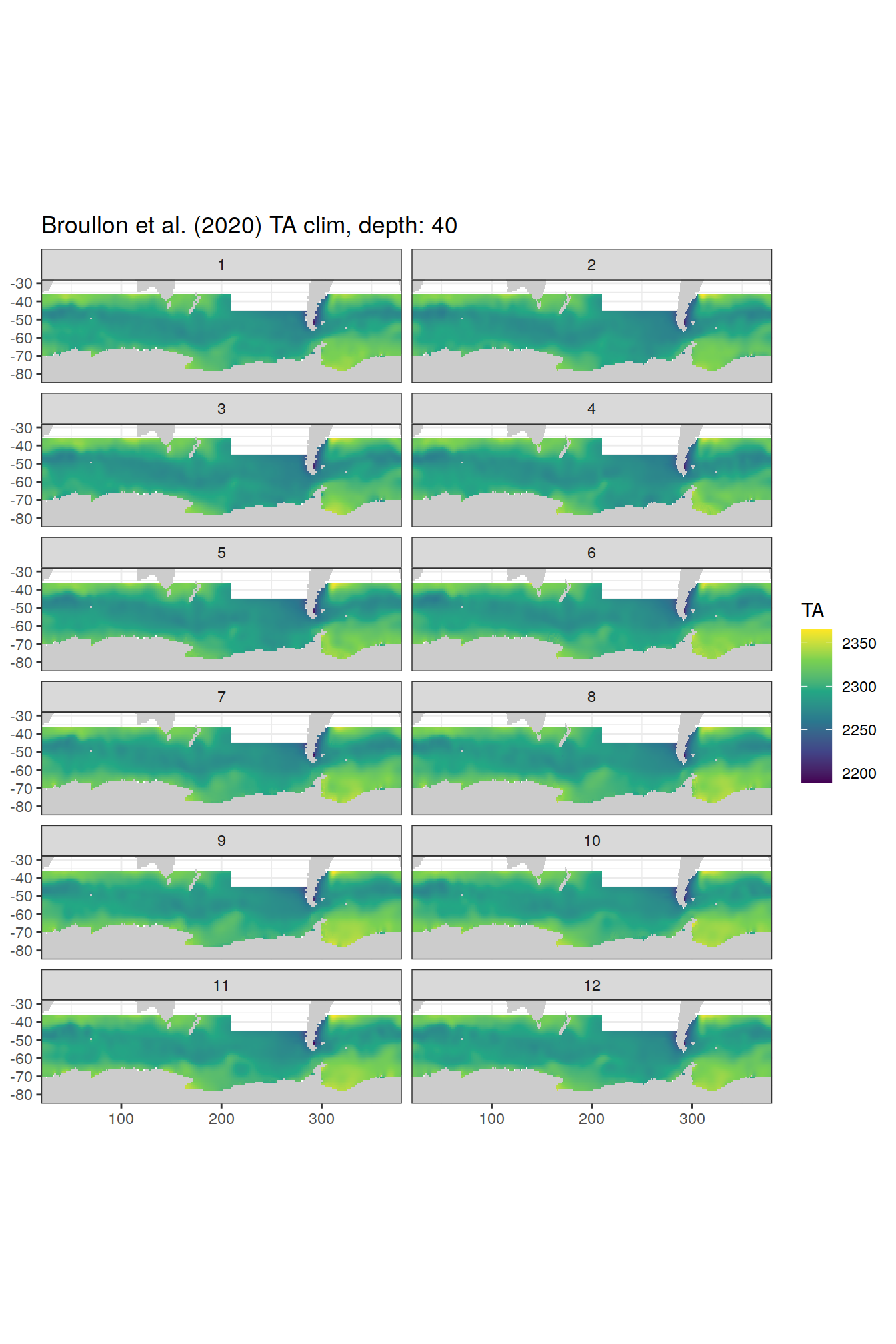
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[10]]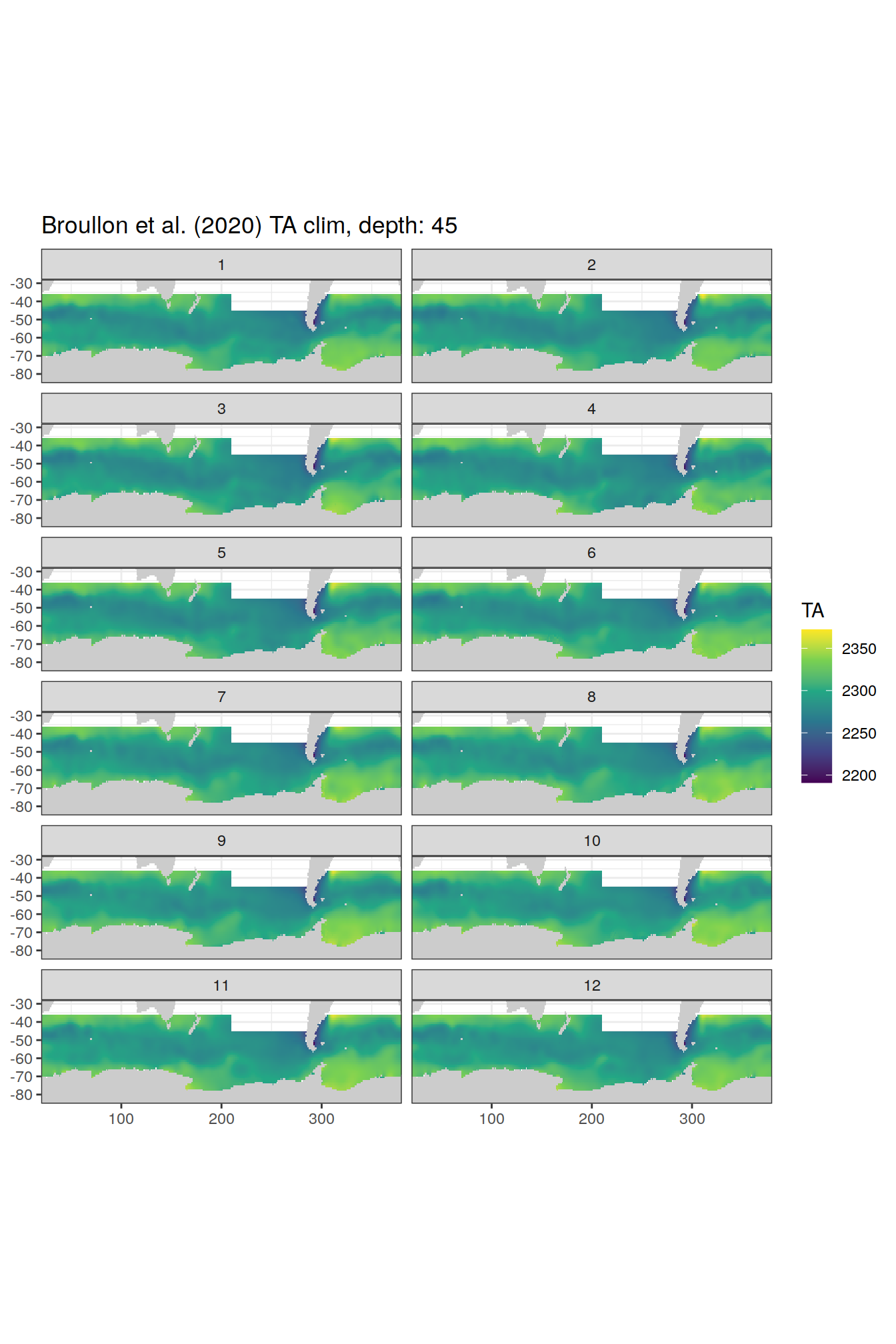
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[11]]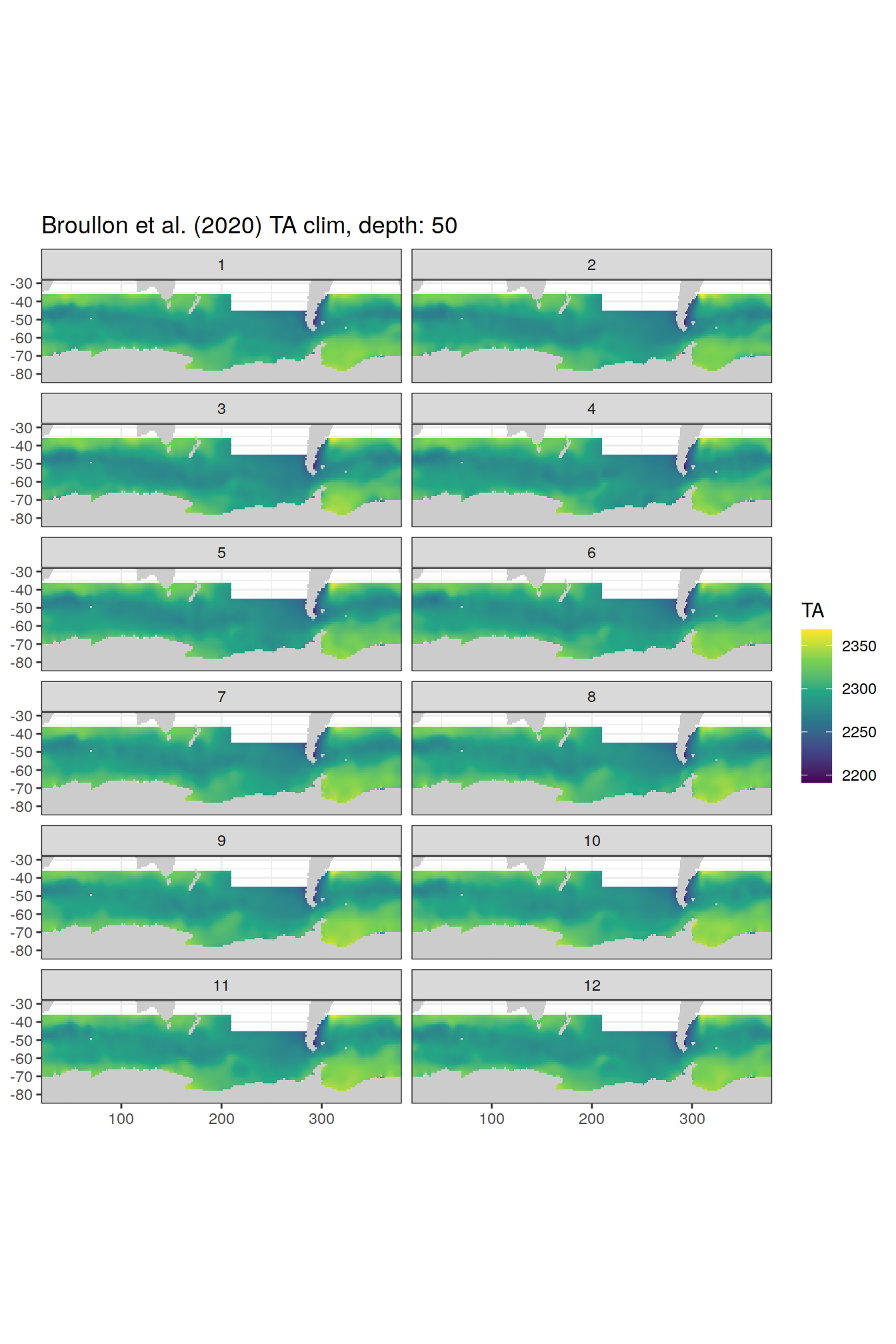
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[12]]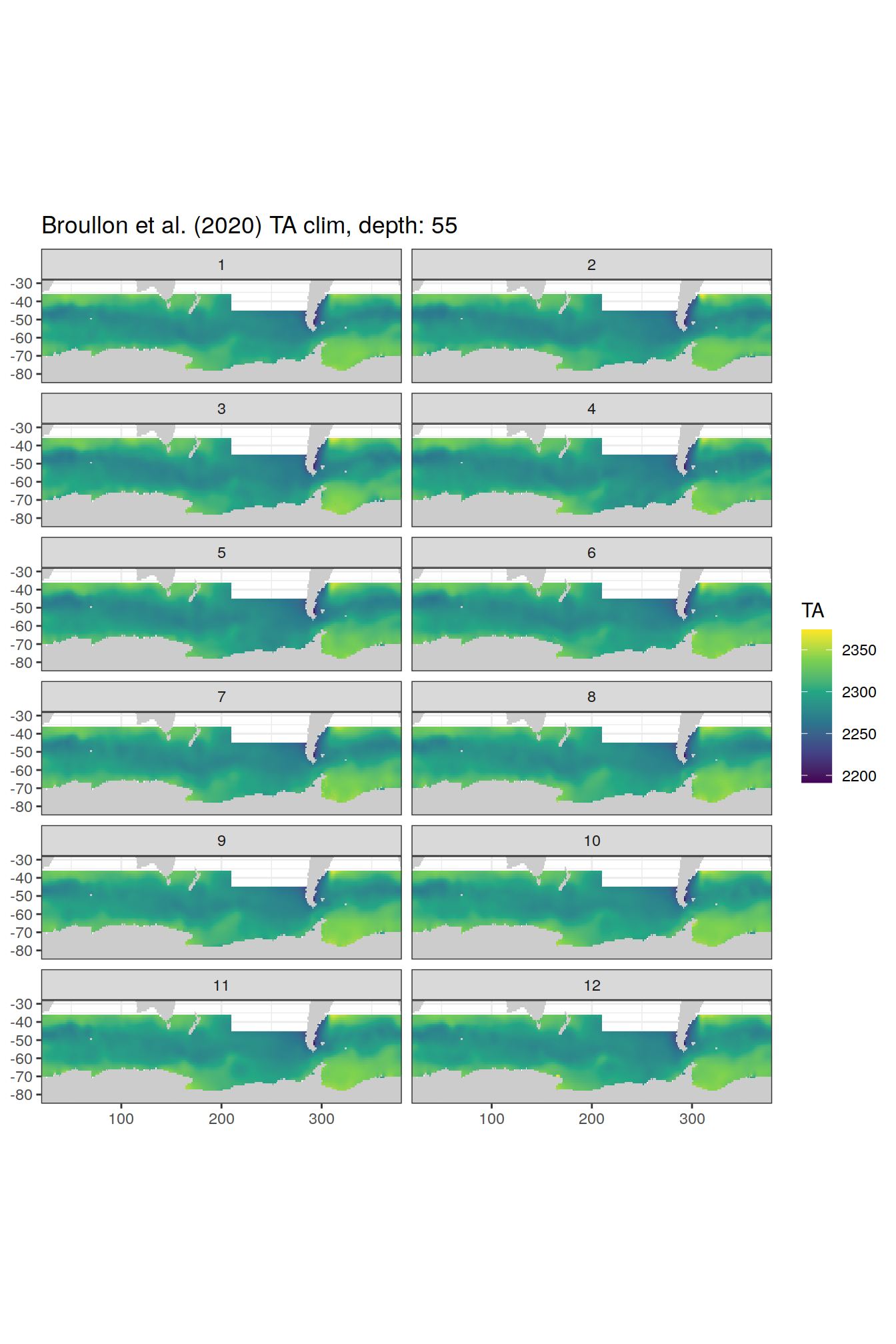
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[13]]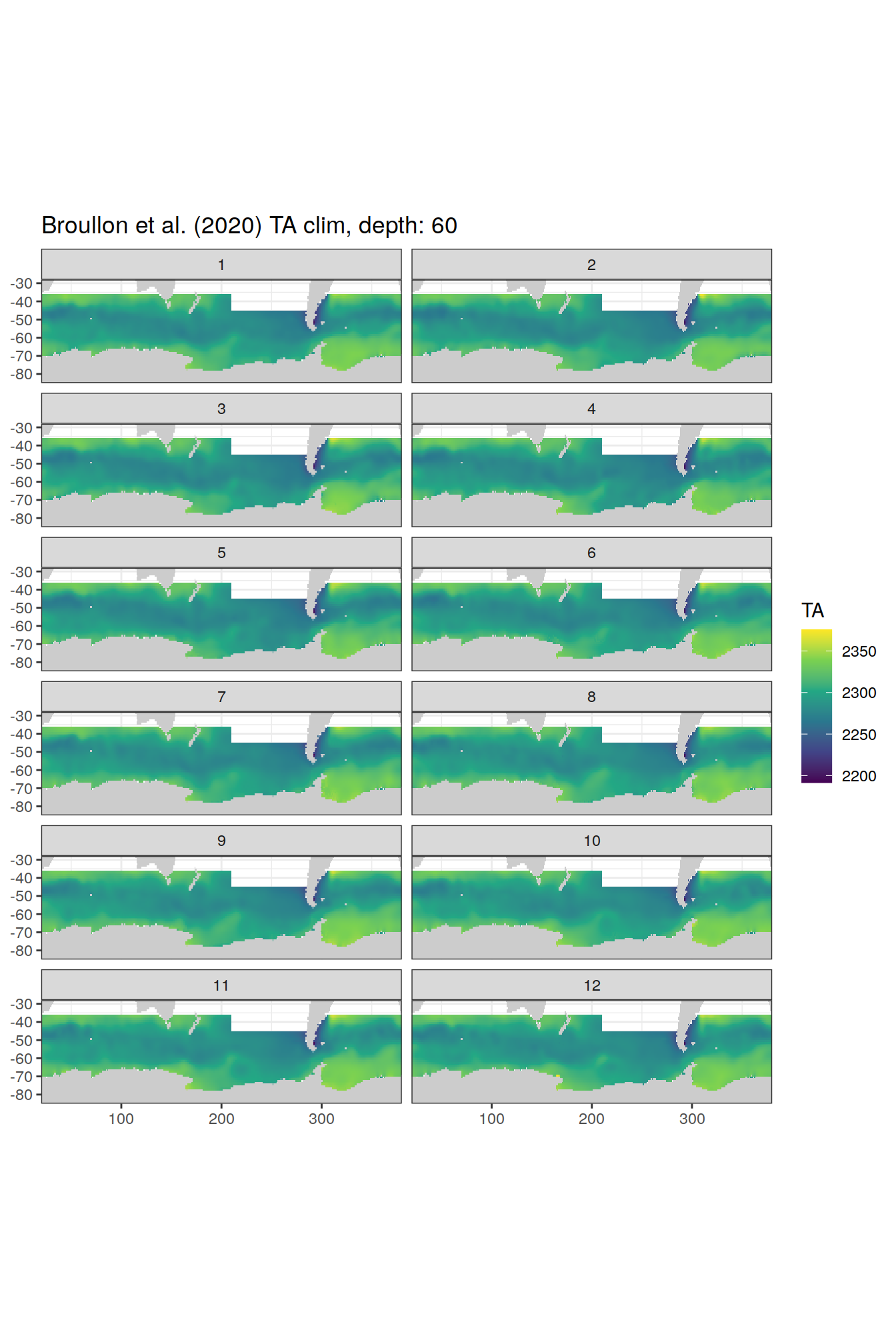
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[14]]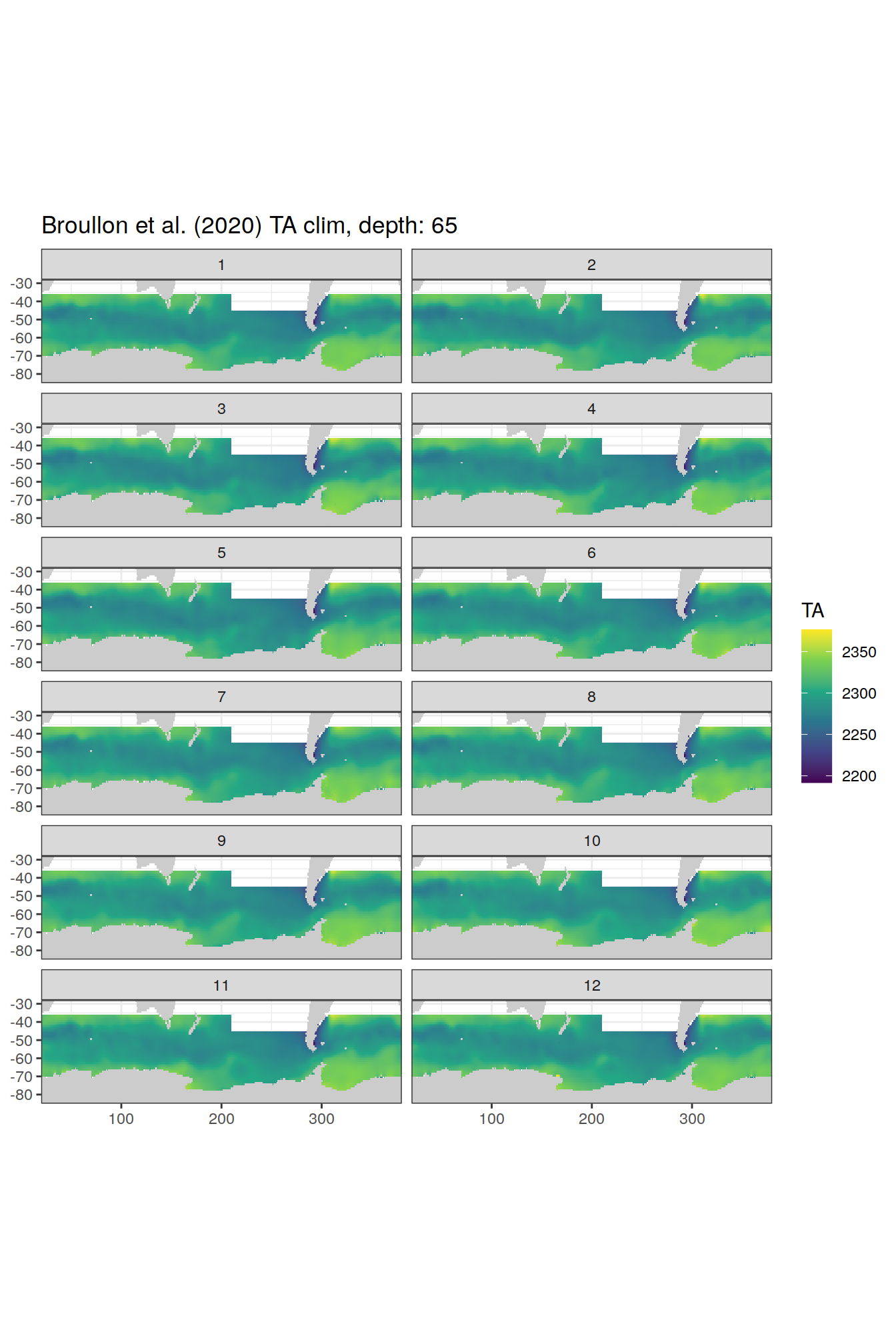
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[15]]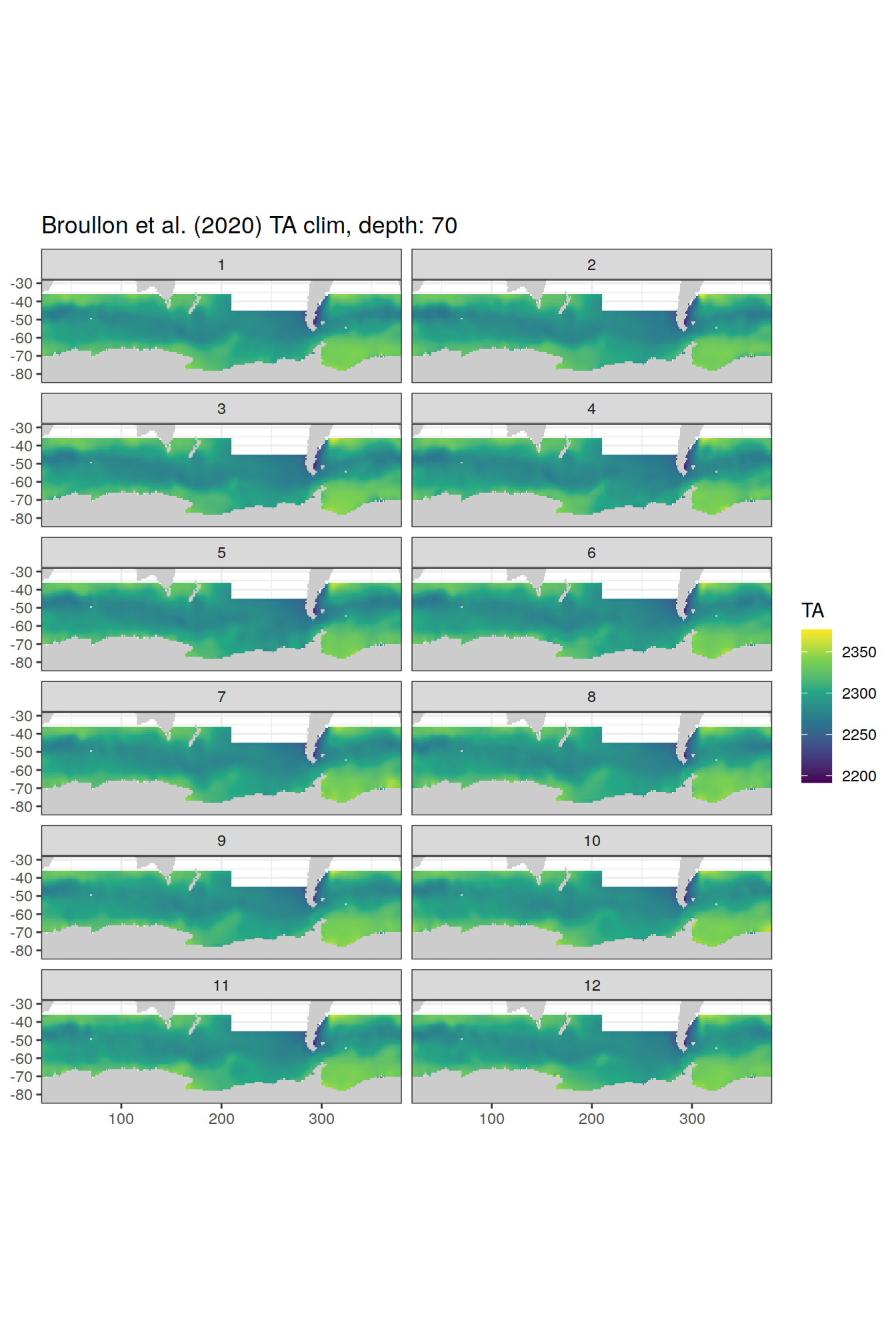
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[16]]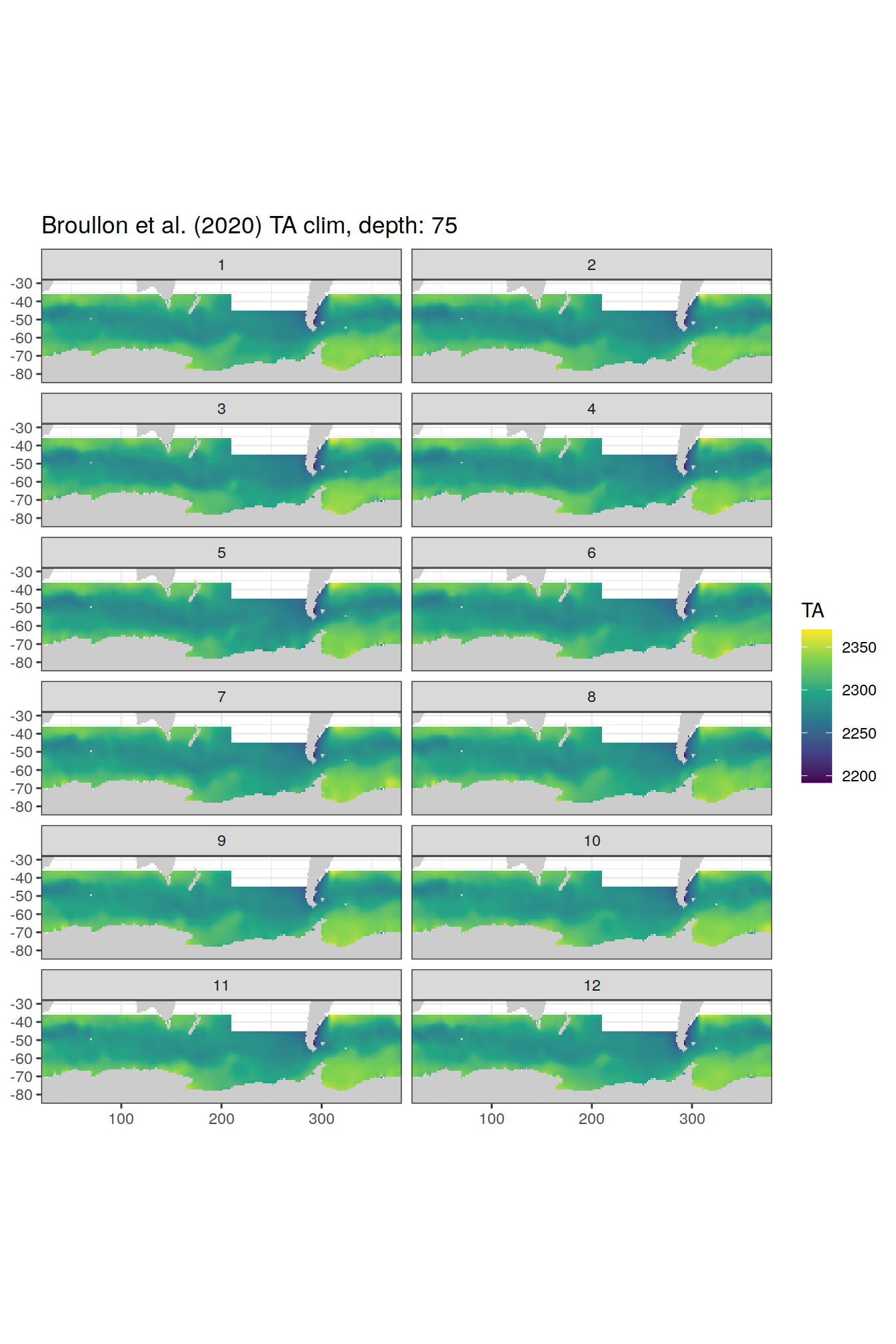
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[17]]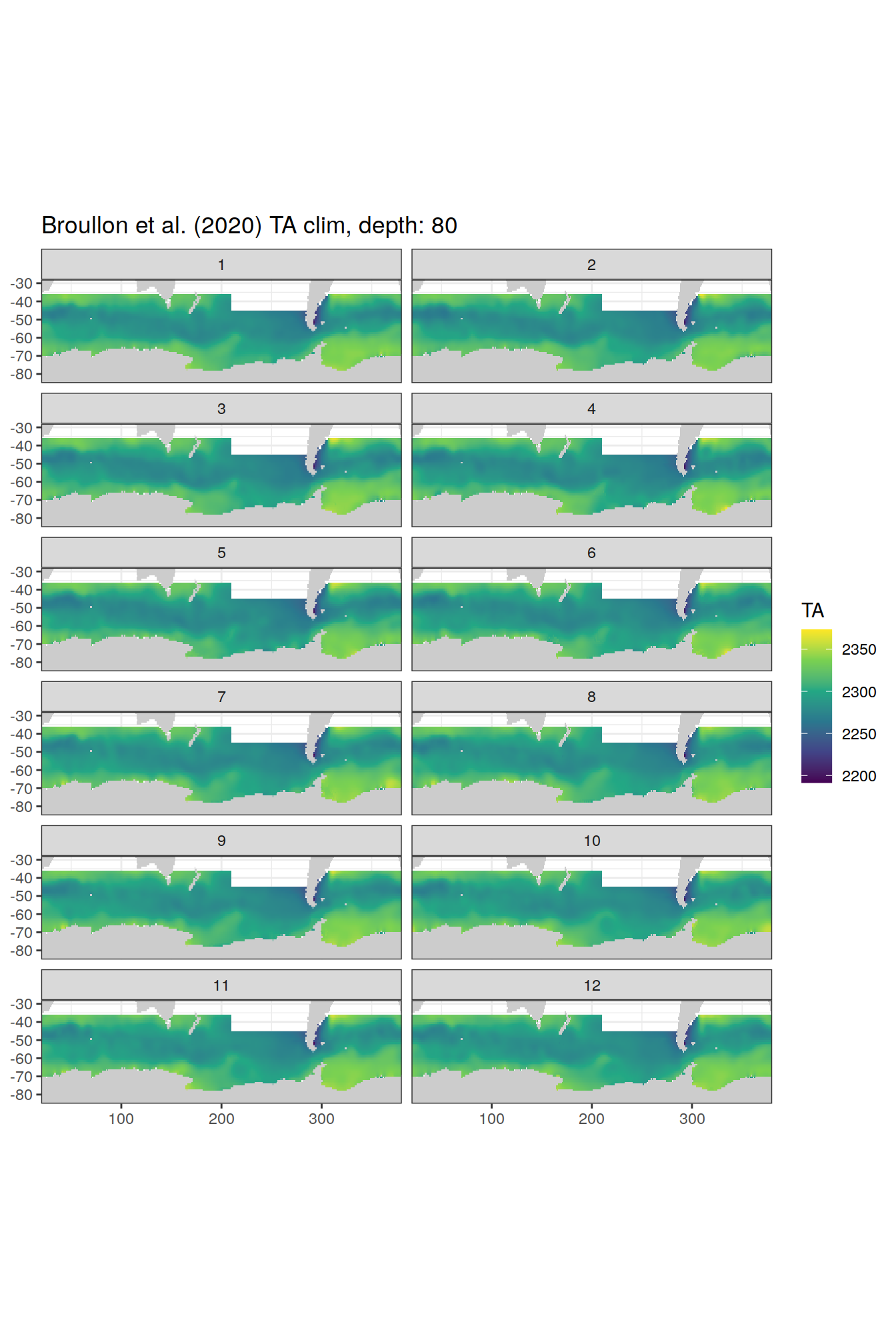
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[18]]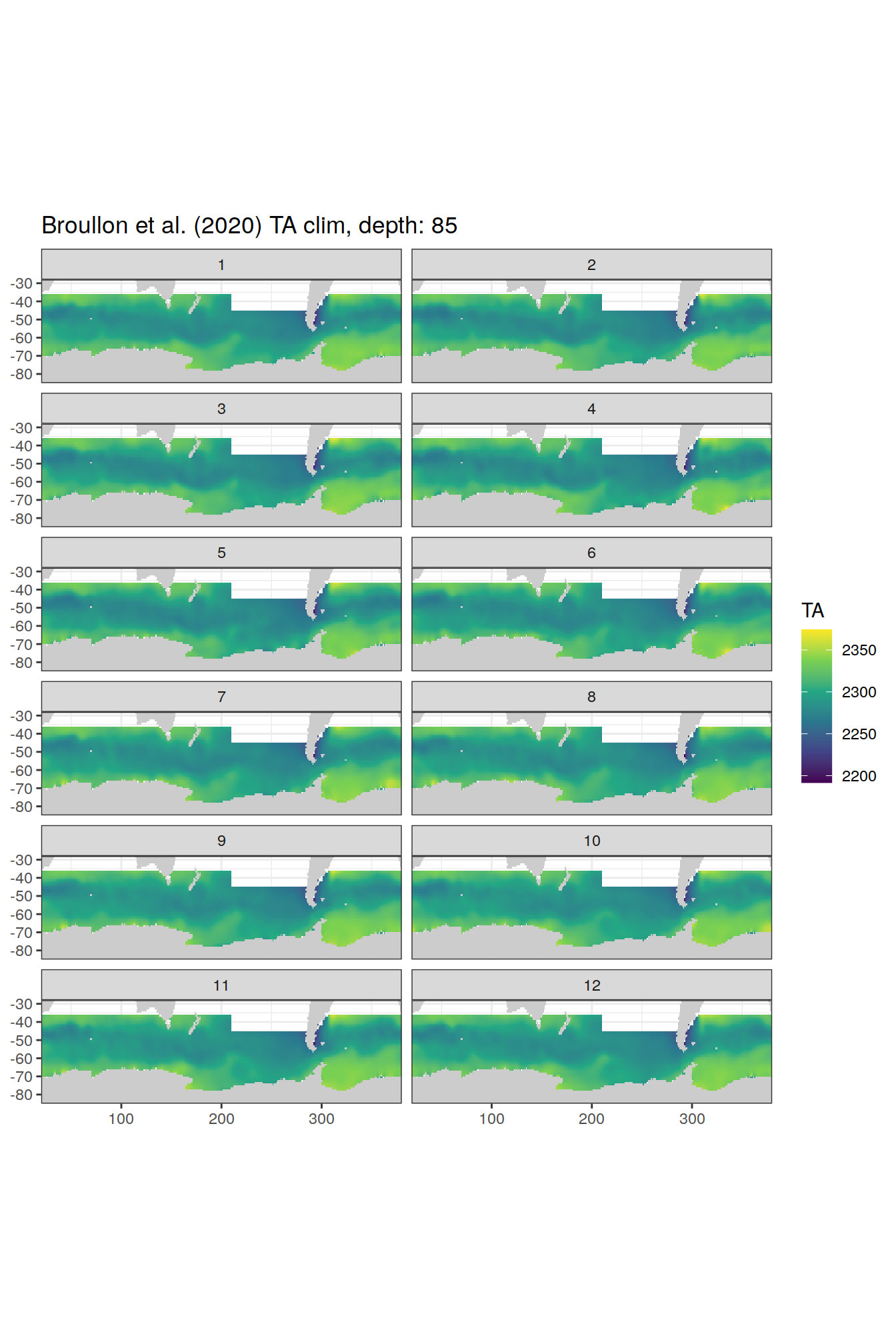
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[19]]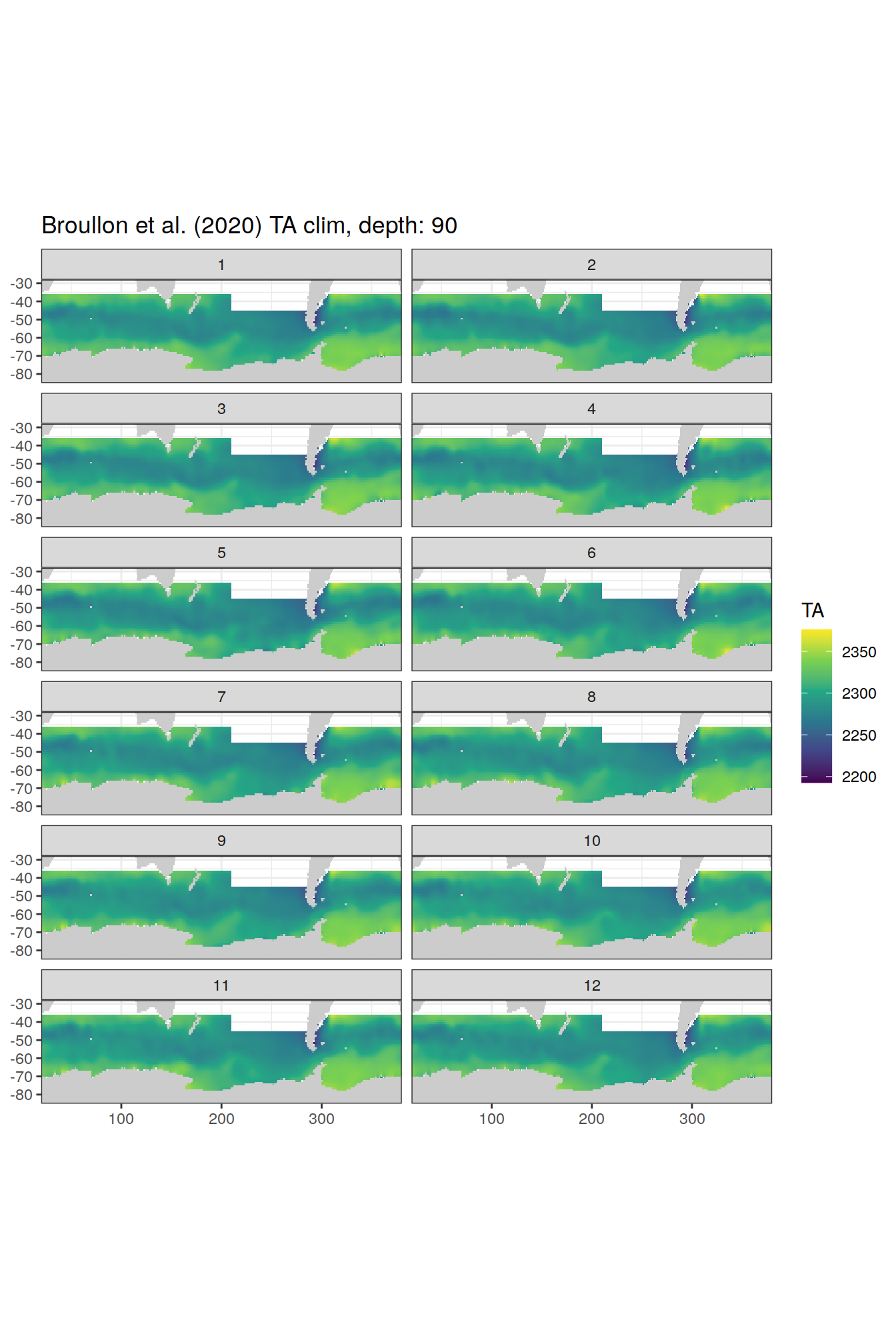
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[20]]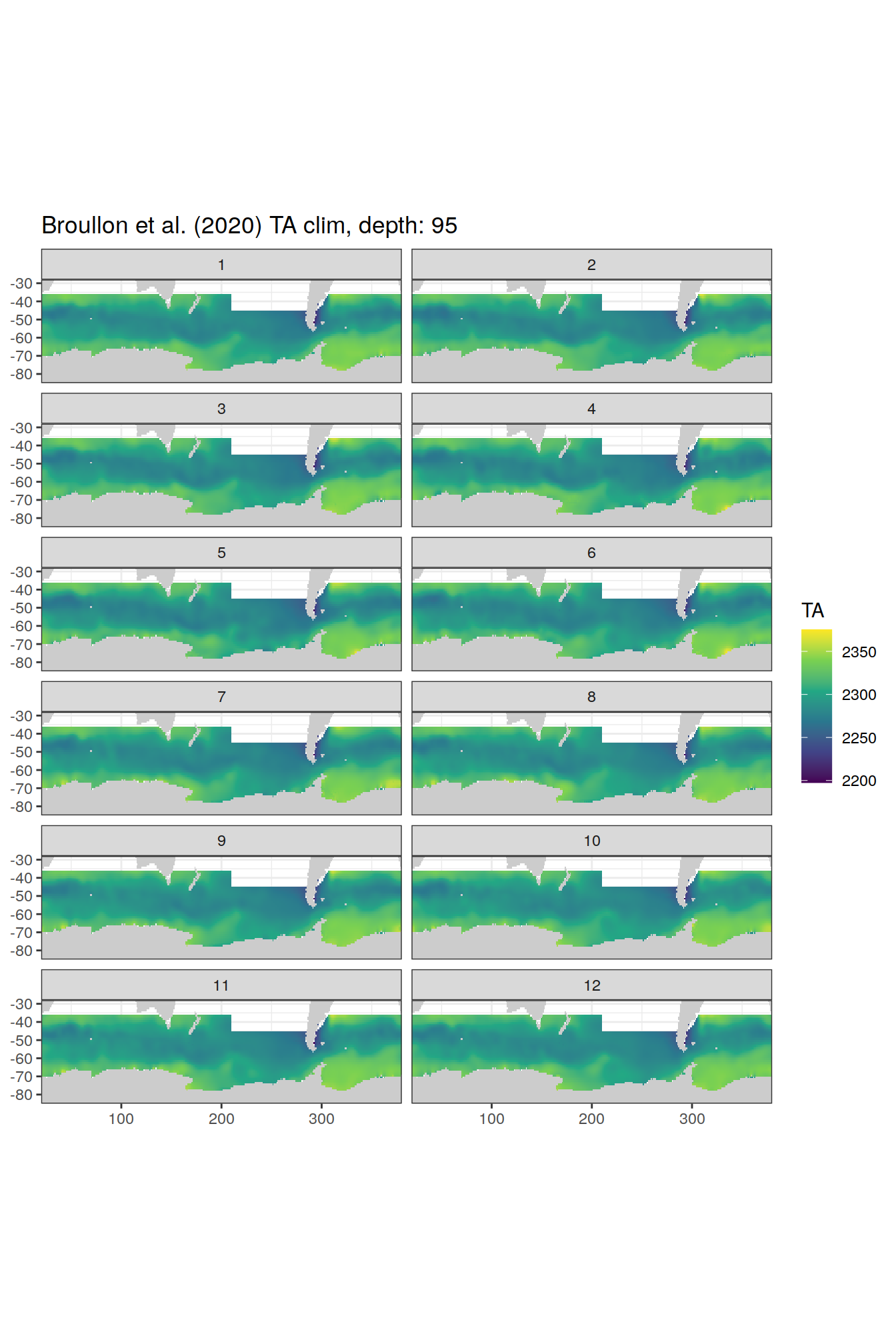
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[21]]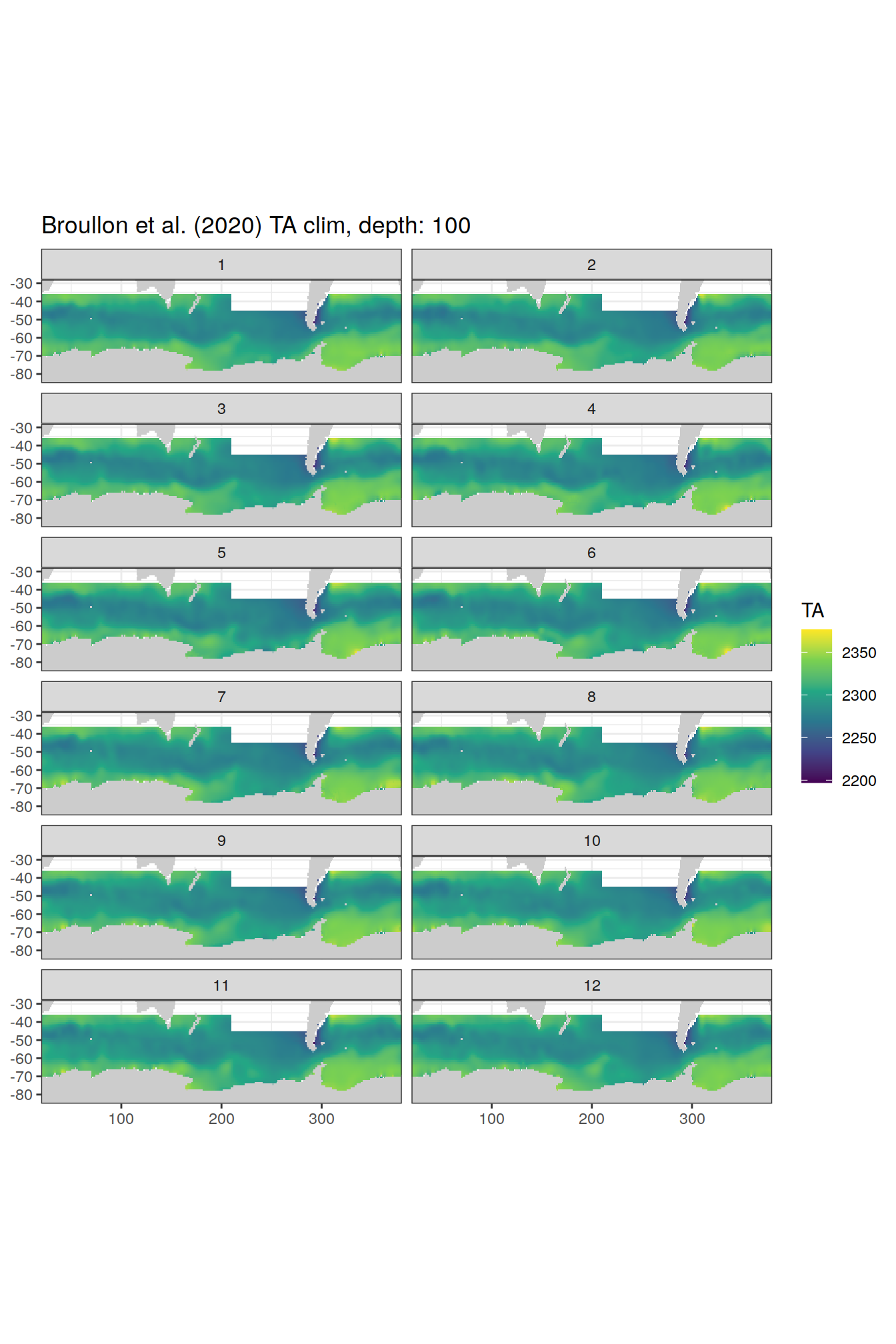
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[22]]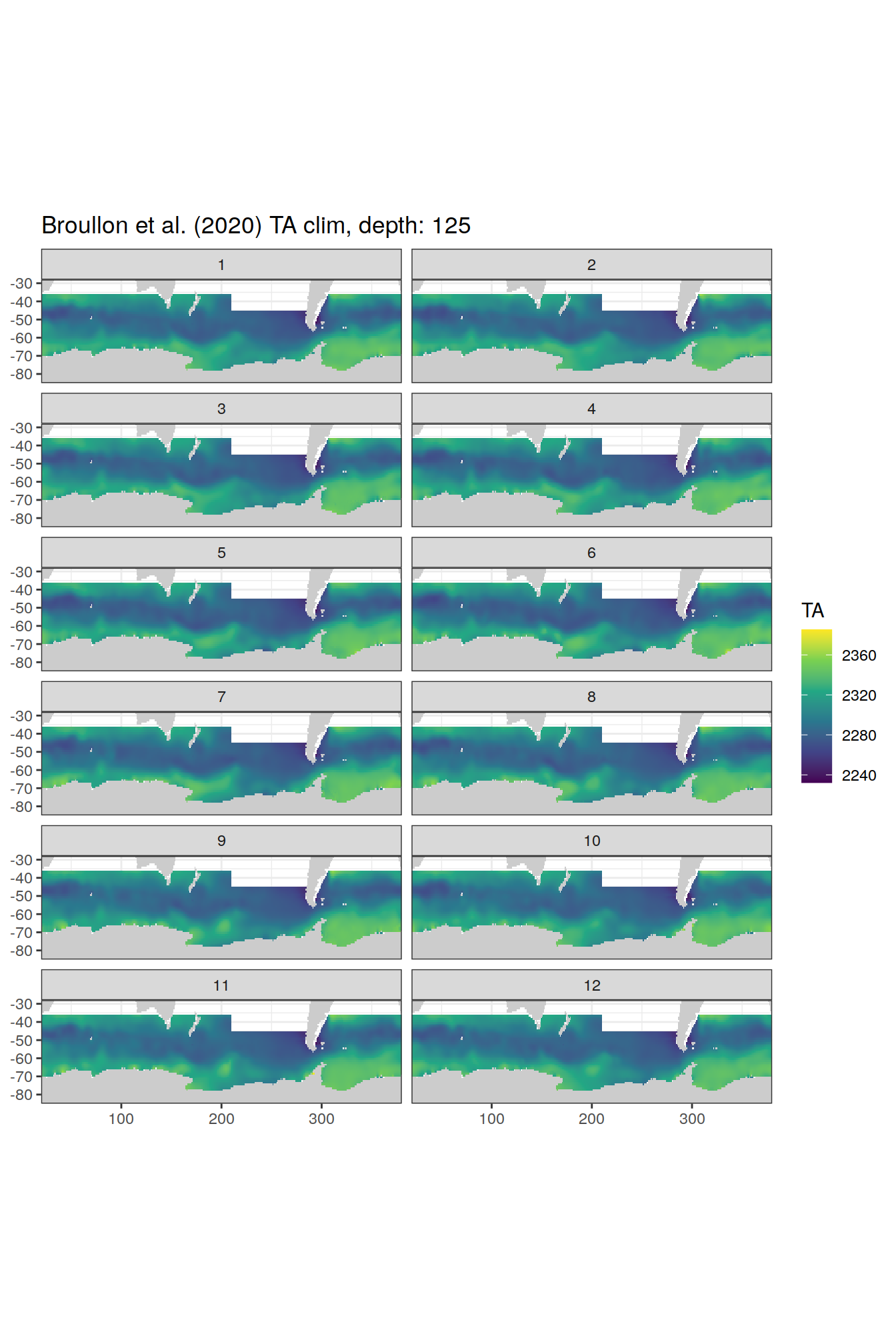
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[23]]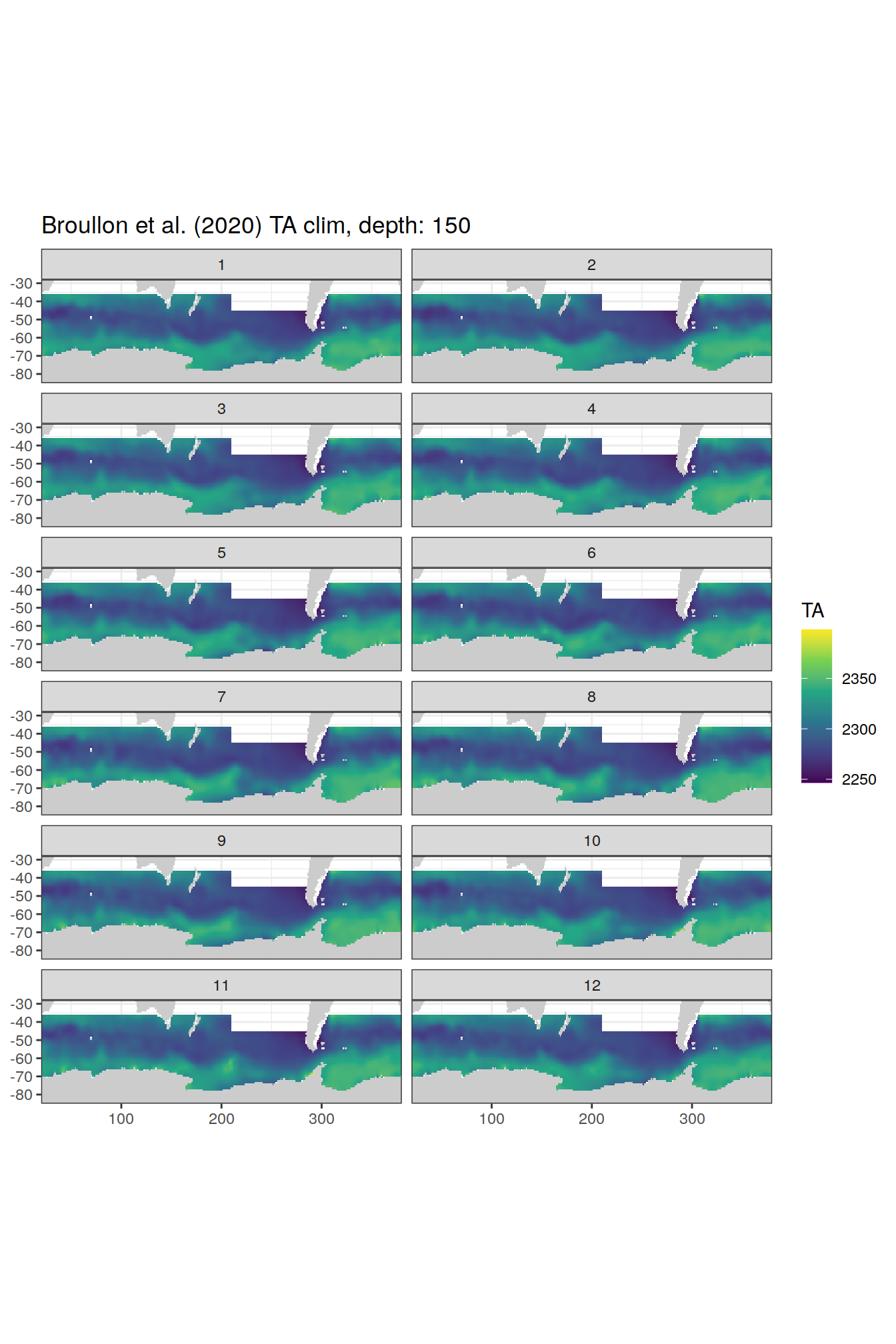
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[24]]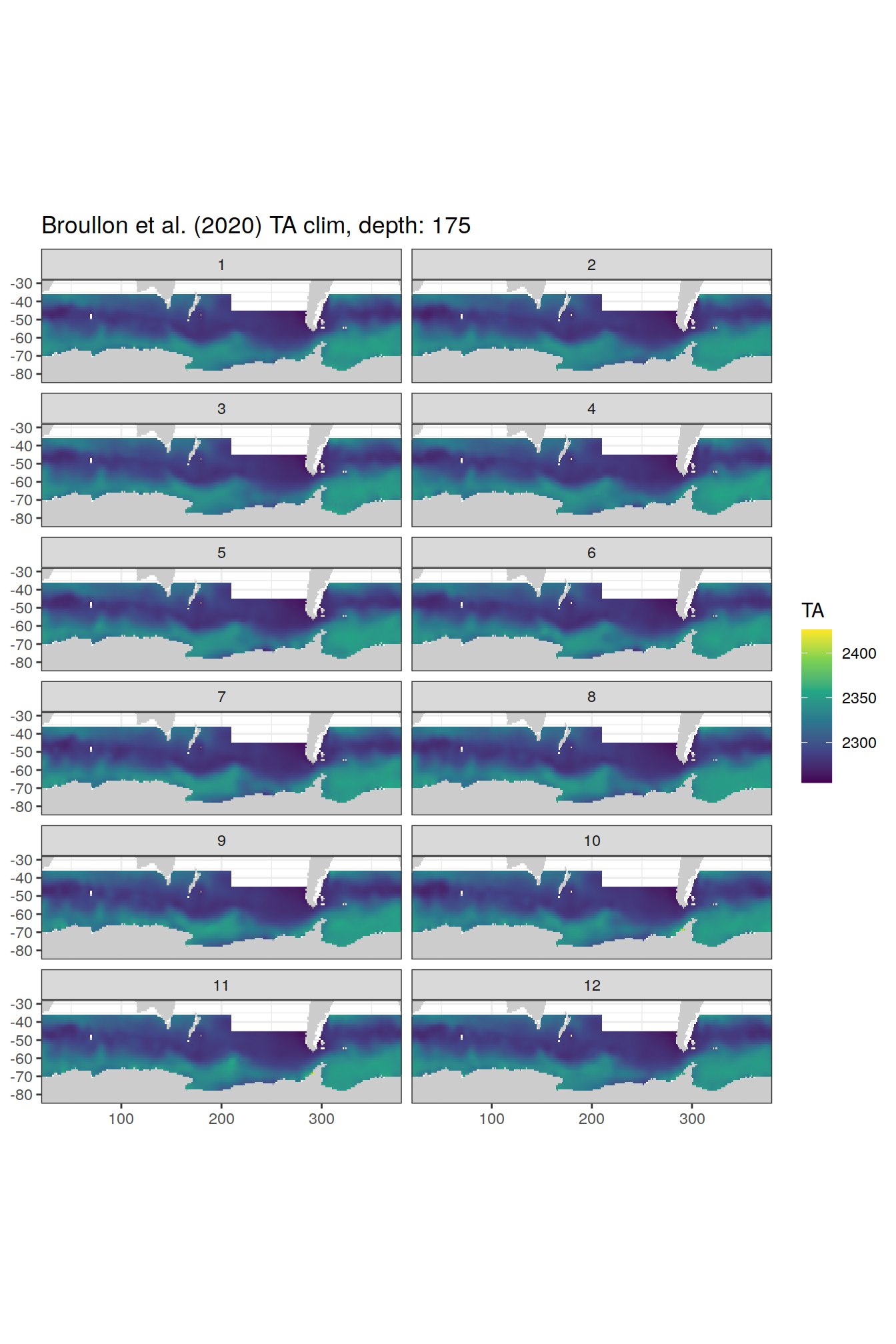
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[25]]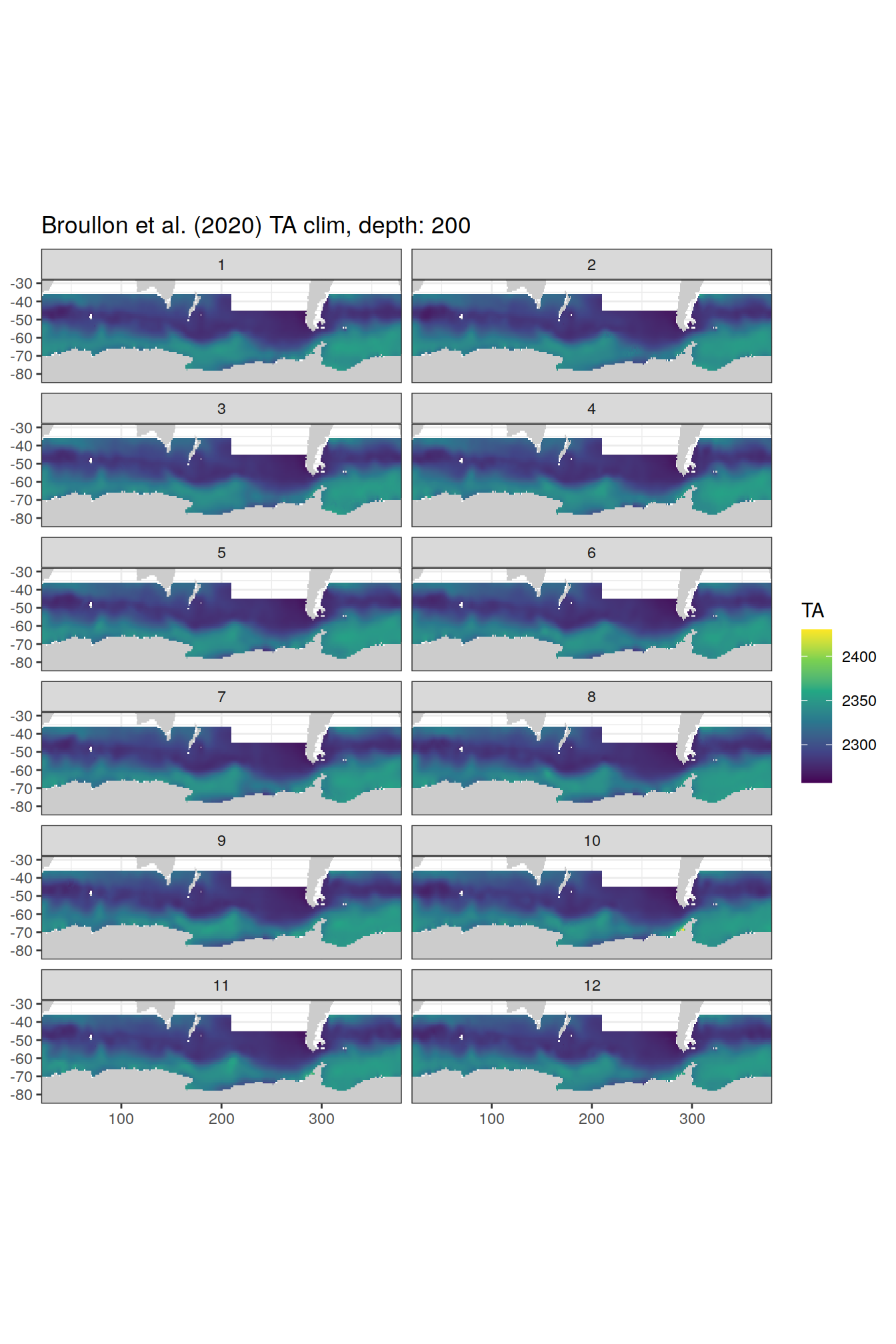
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[26]]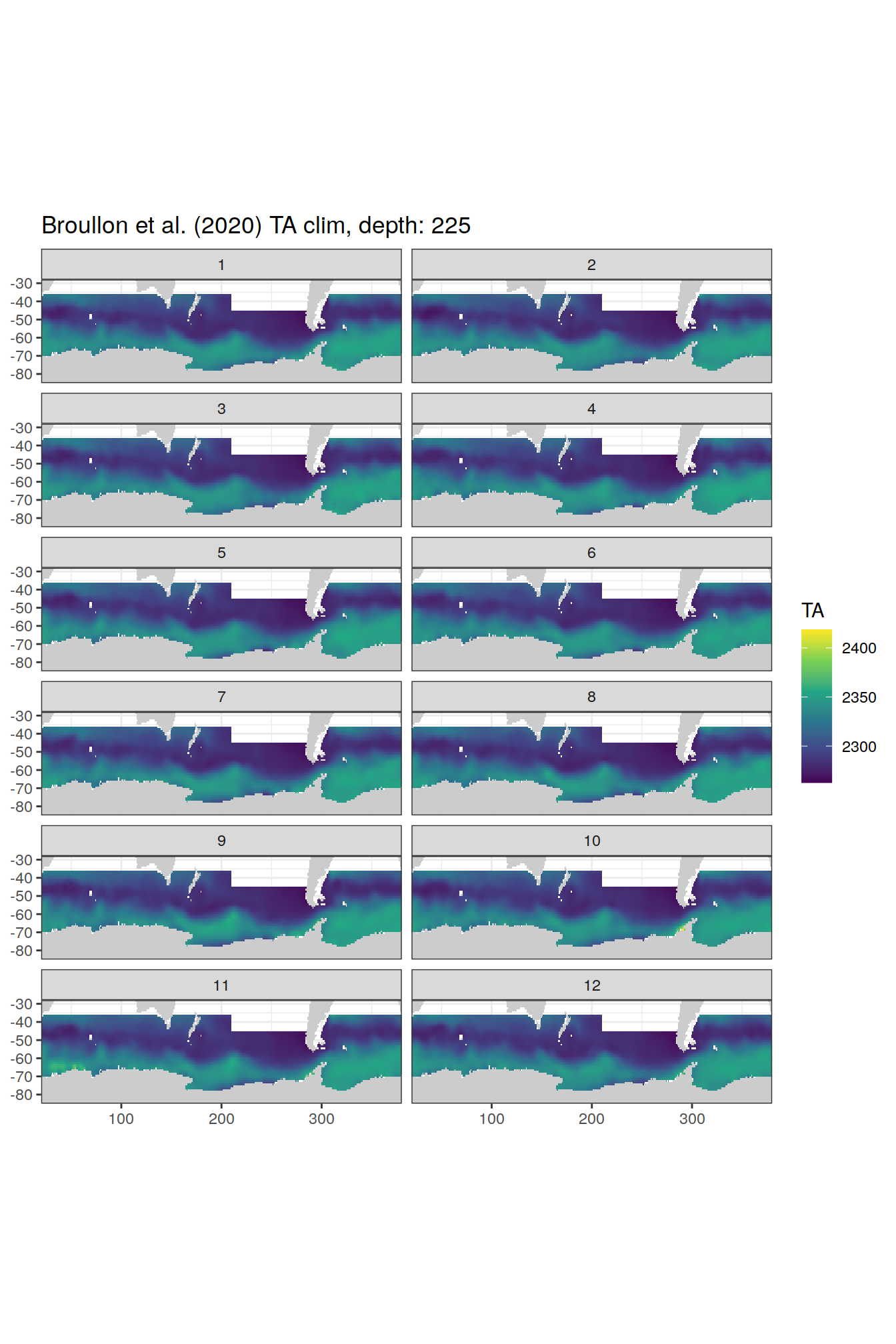
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[27]]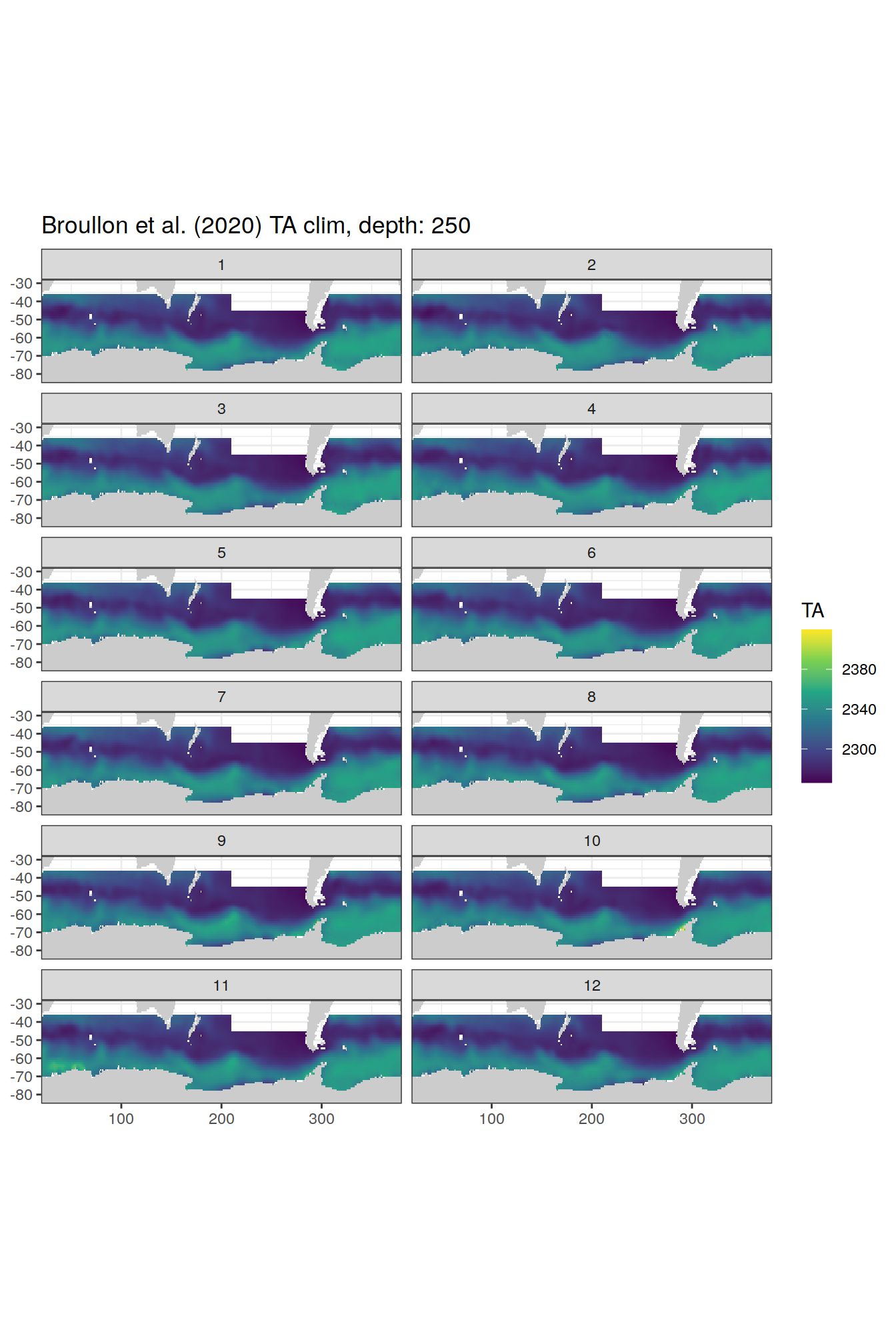
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[28]]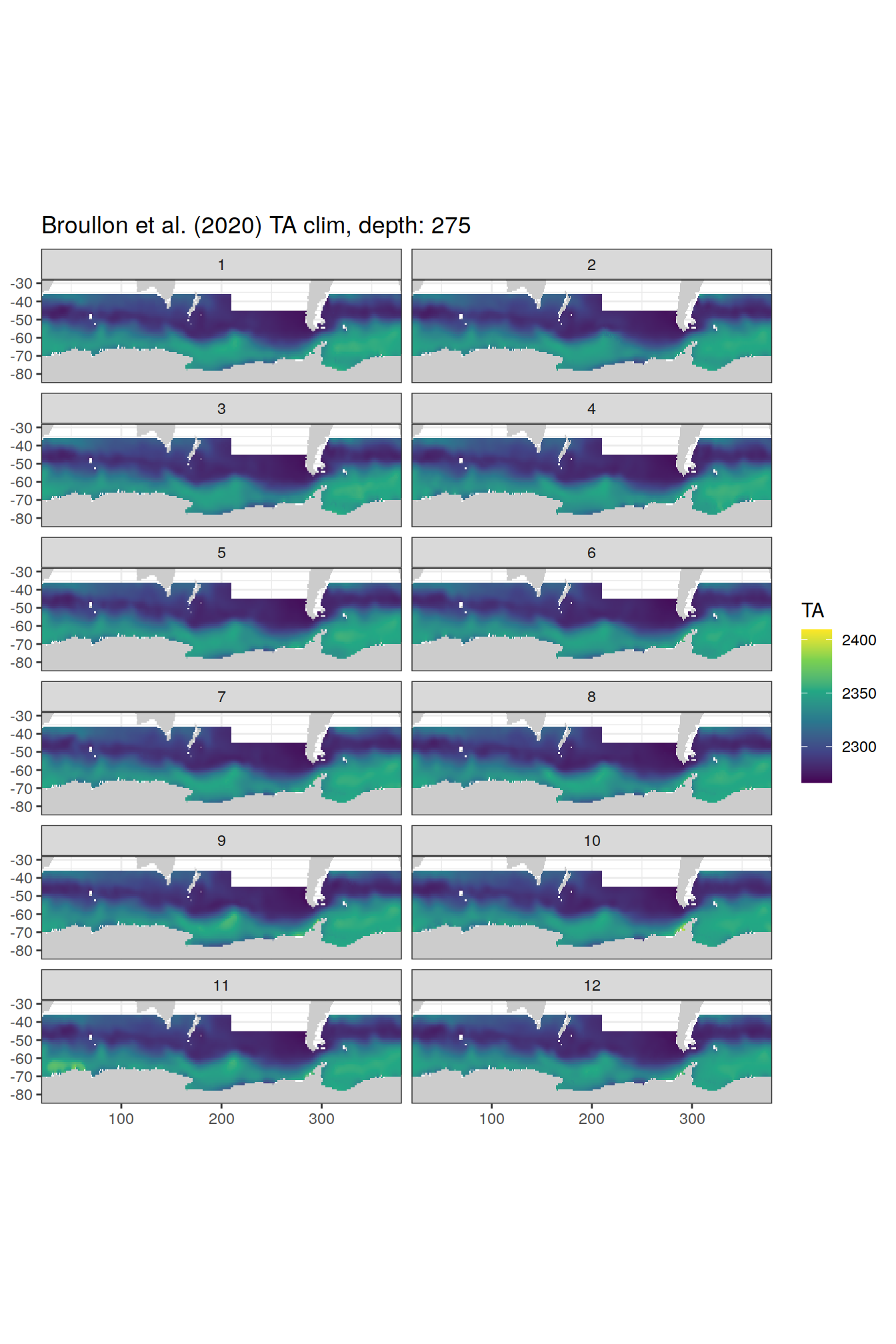
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[29]]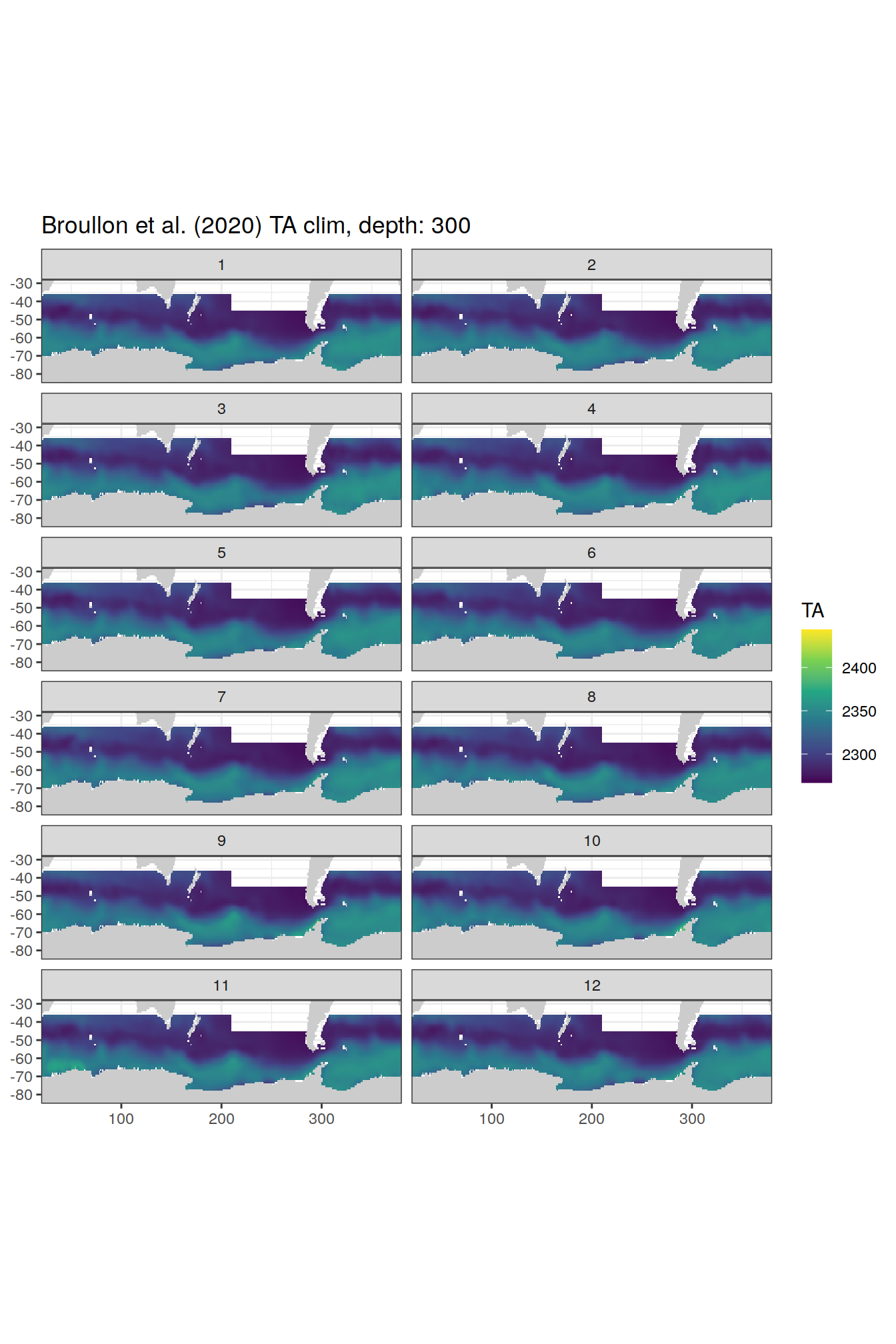
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[30]]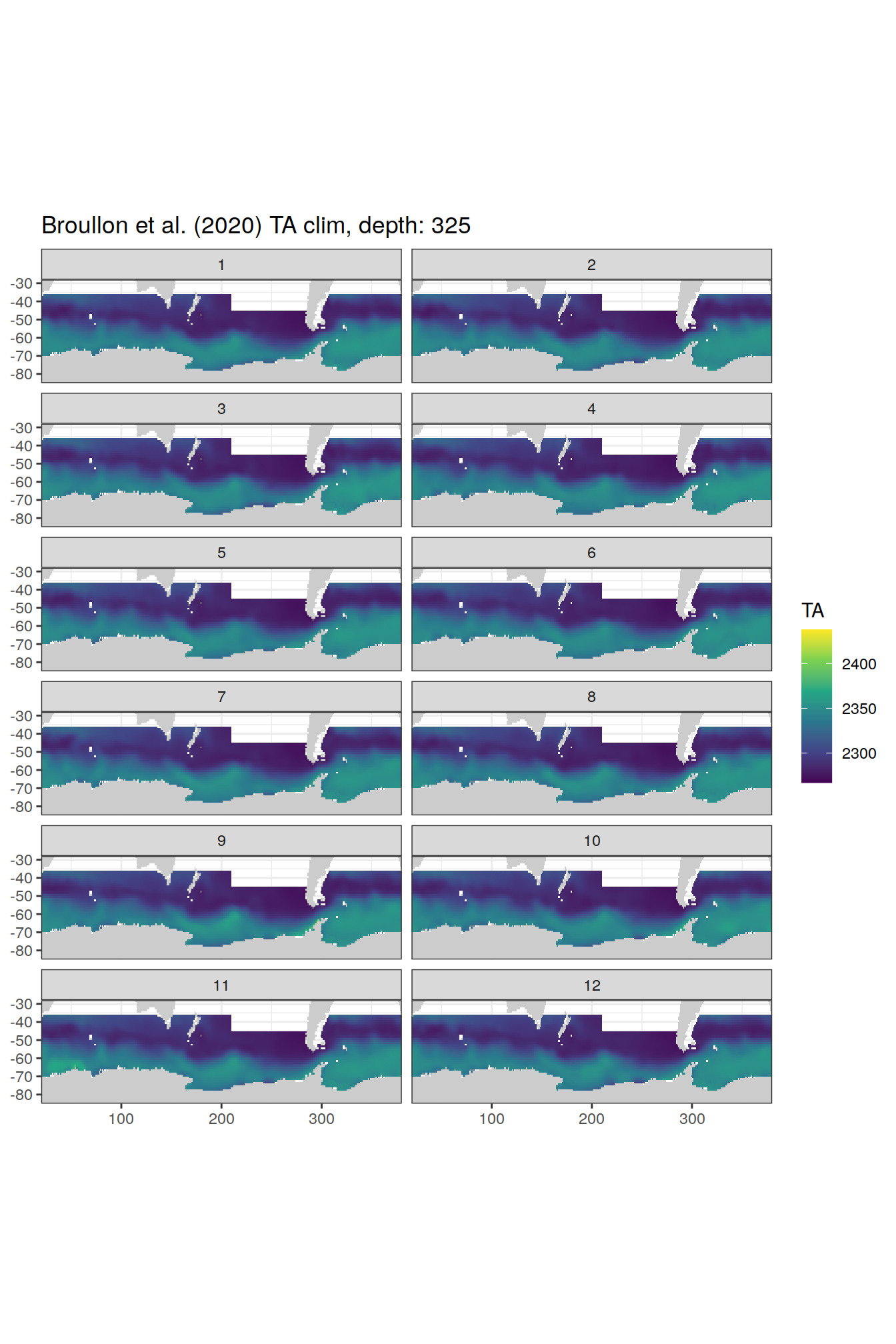
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[31]]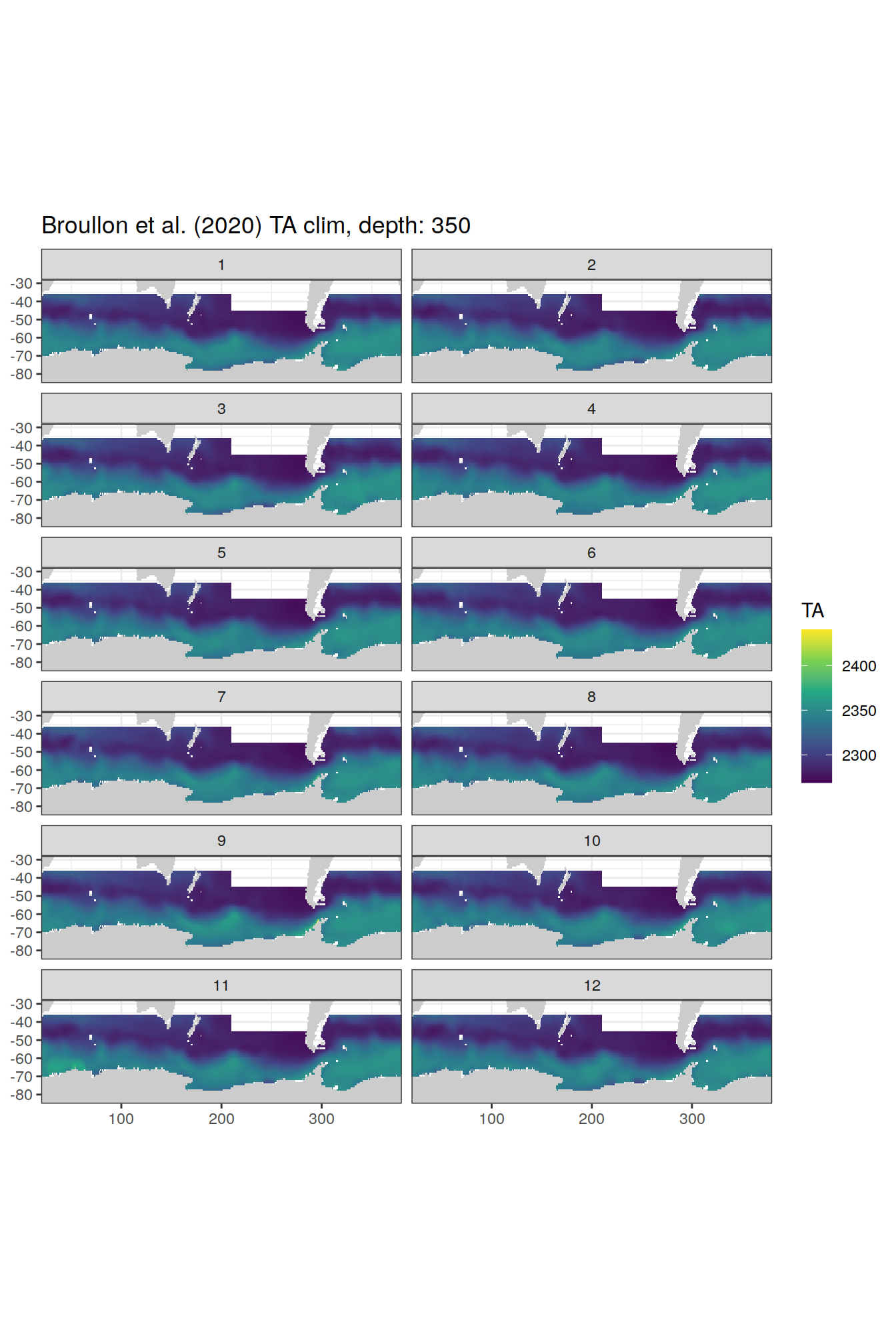
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[32]]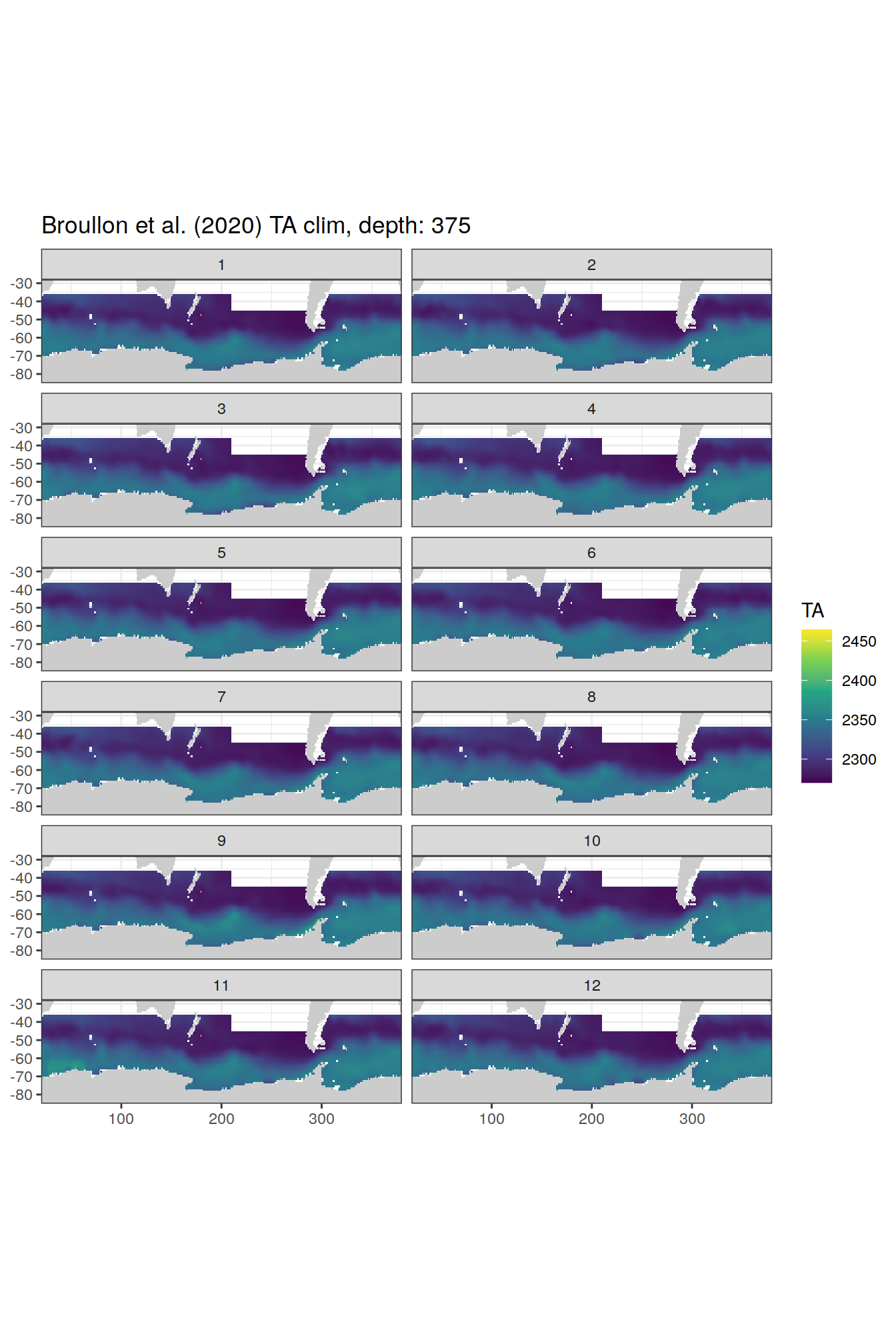
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[33]]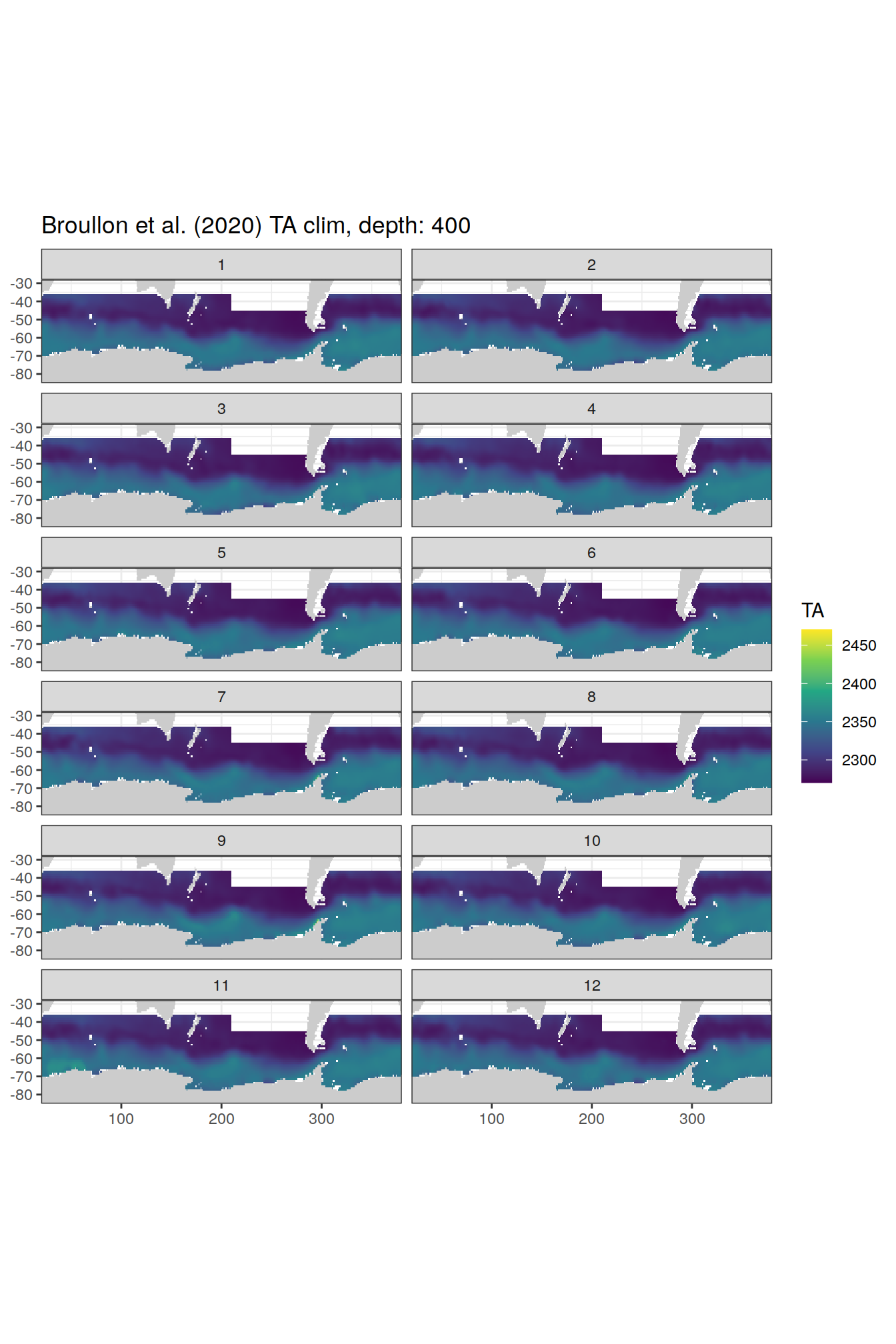
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[34]]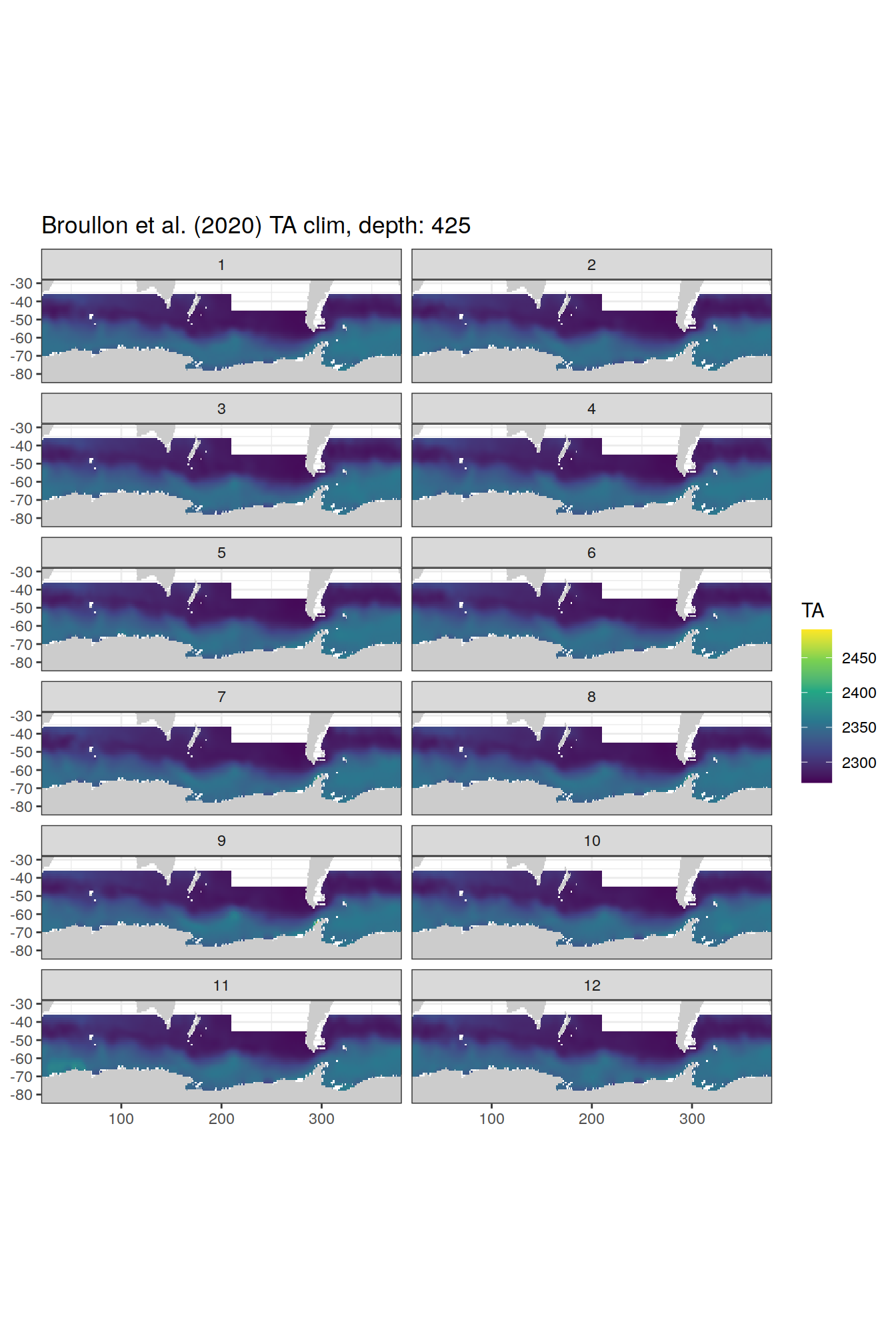
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[35]]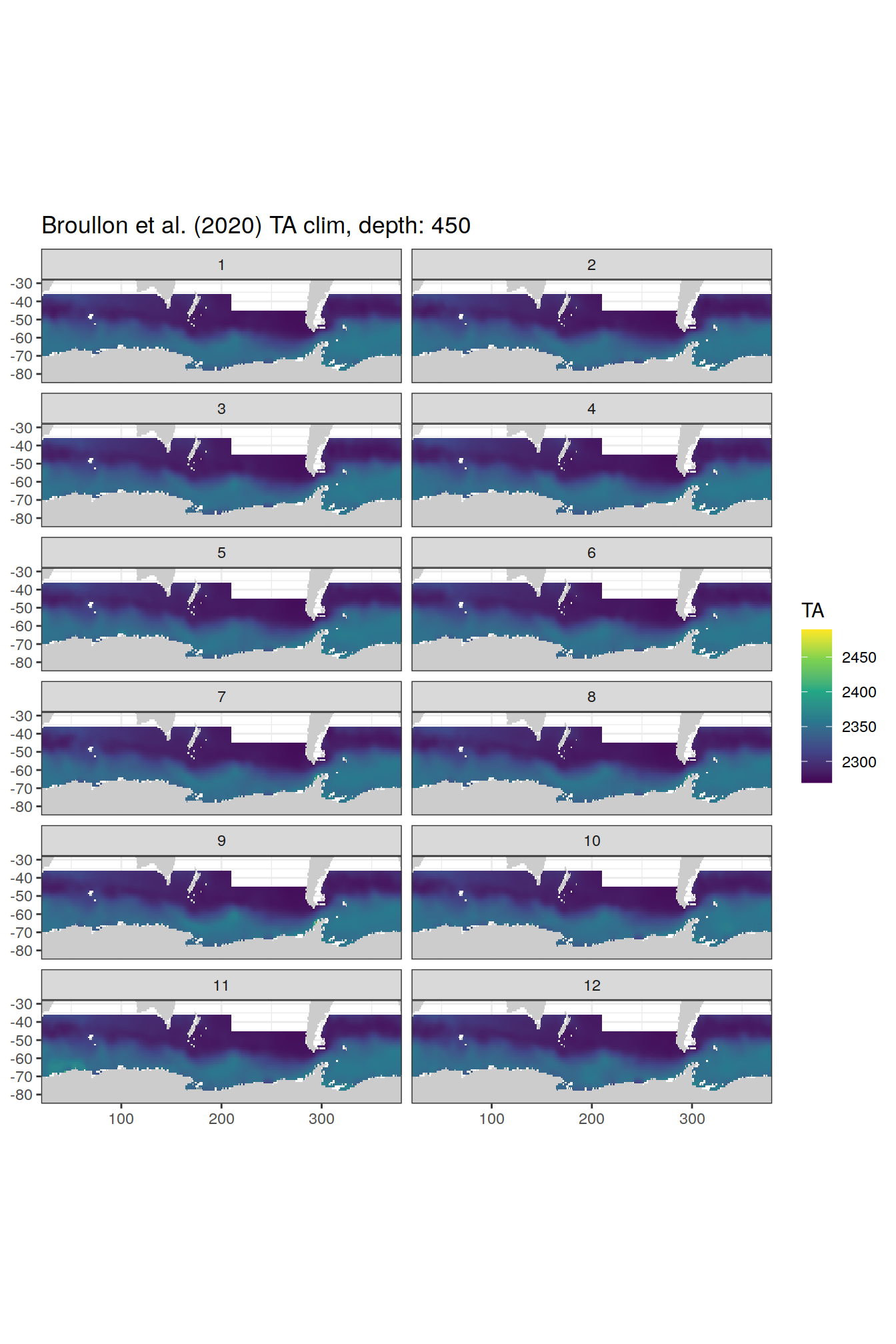
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[36]]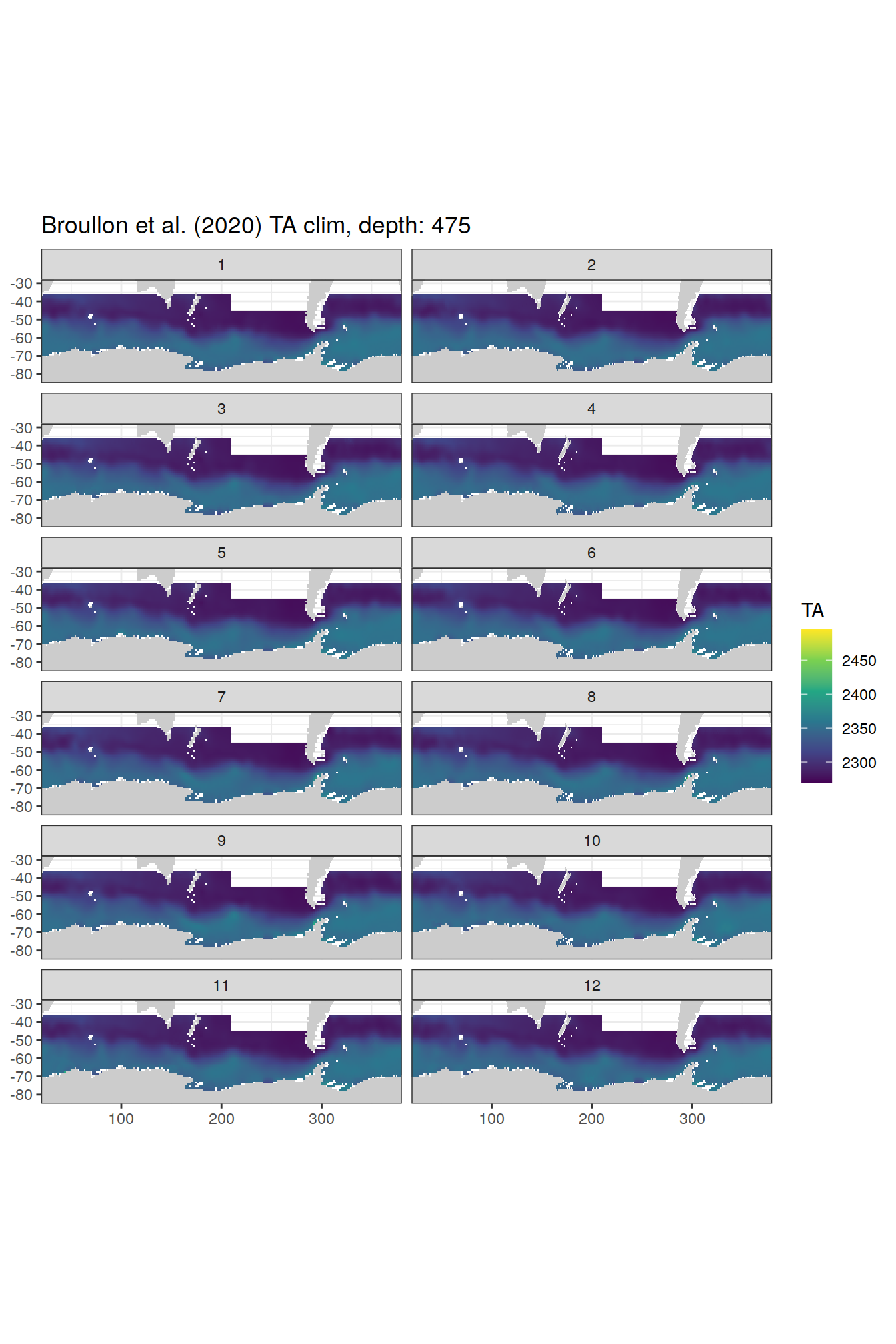
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[37]]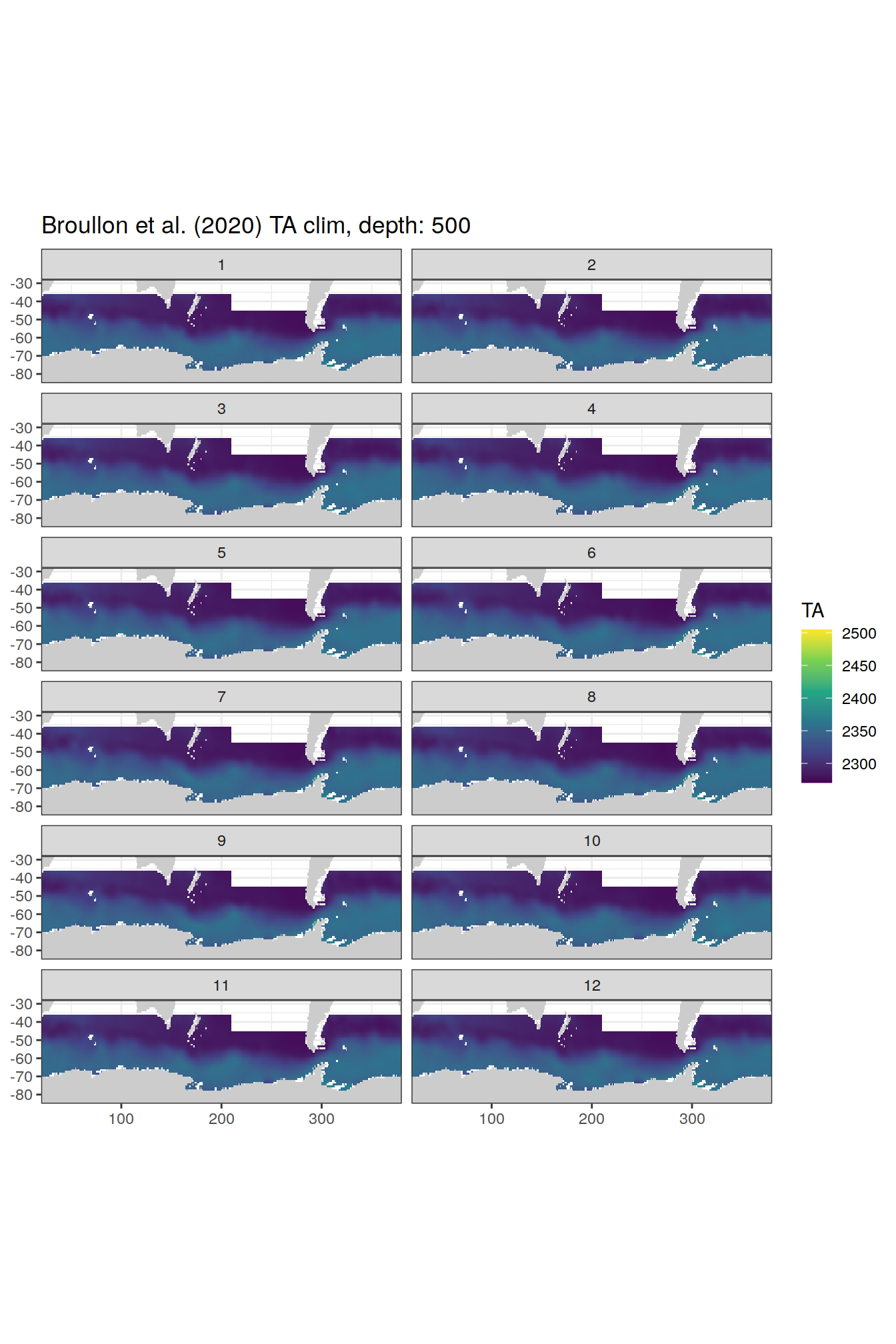
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[38]]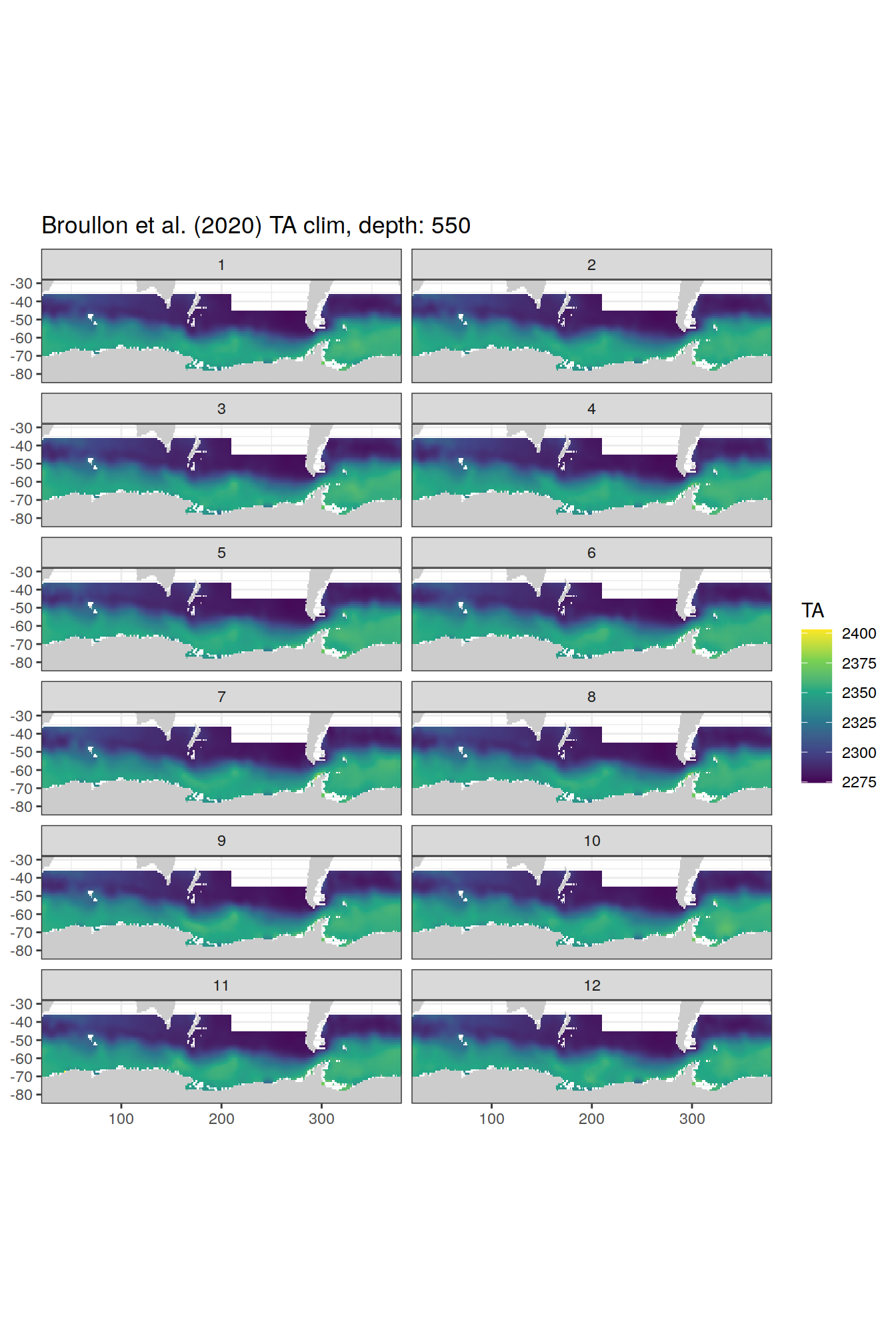
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[39]]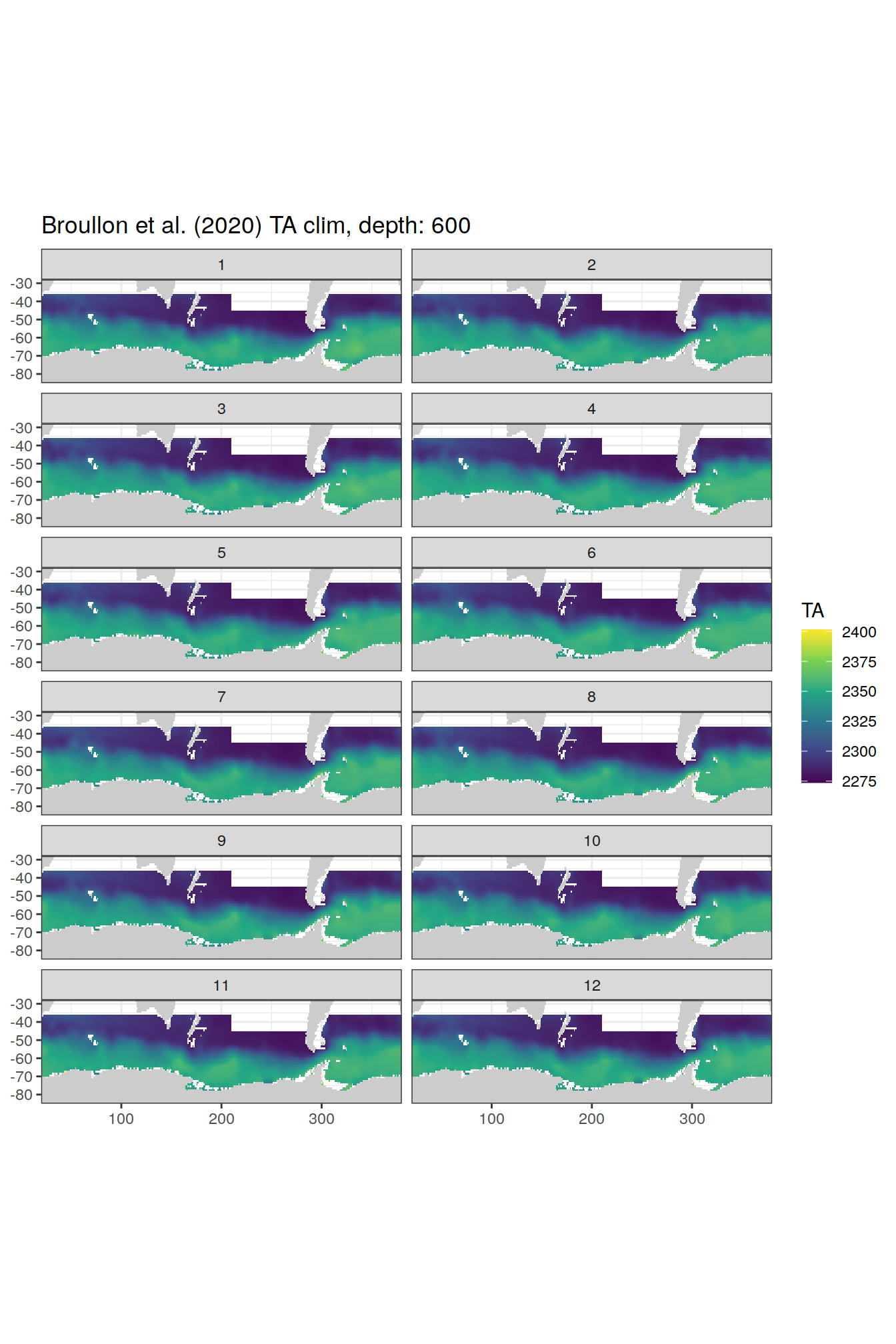
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[40]]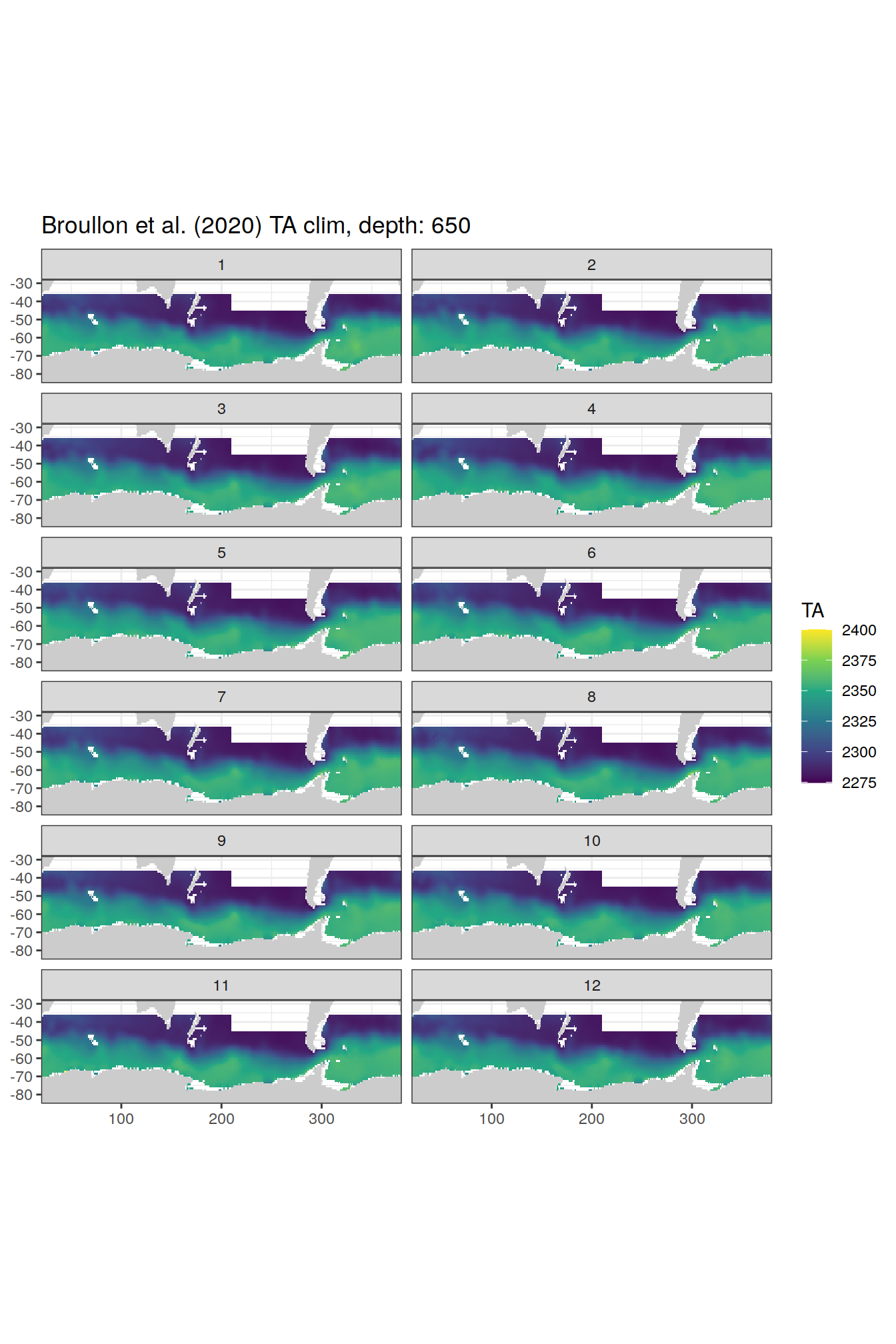
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[41]]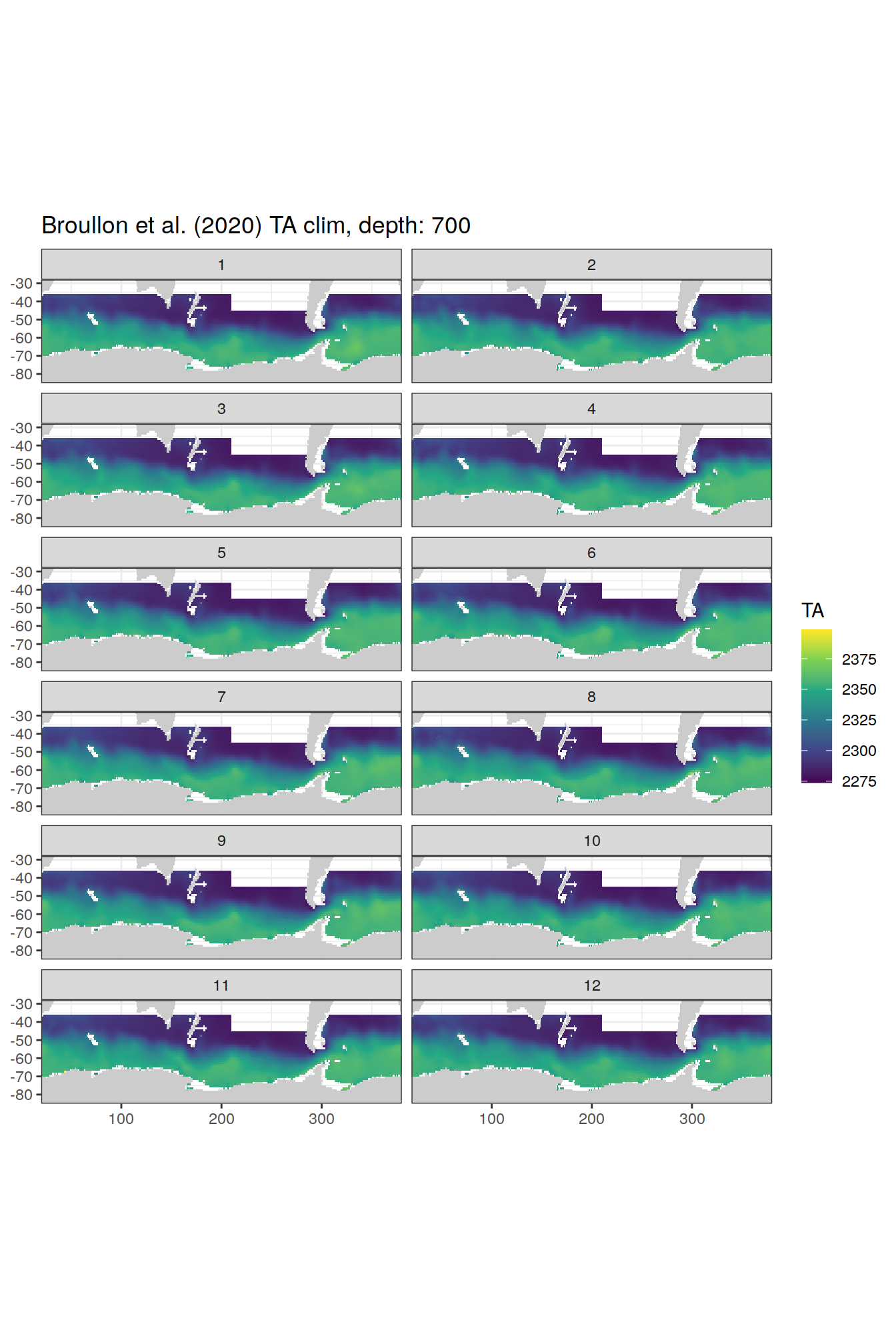
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[42]]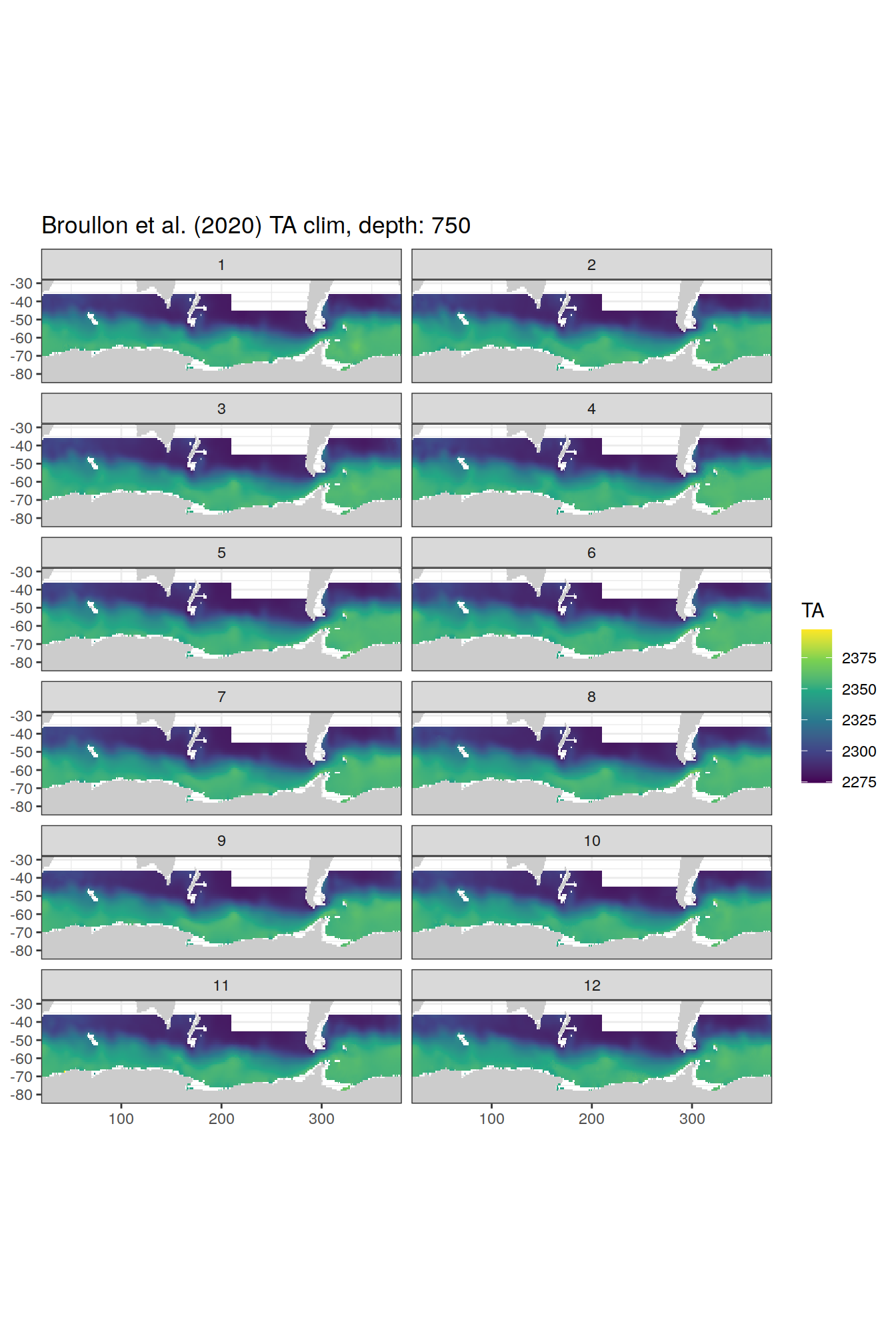
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[43]]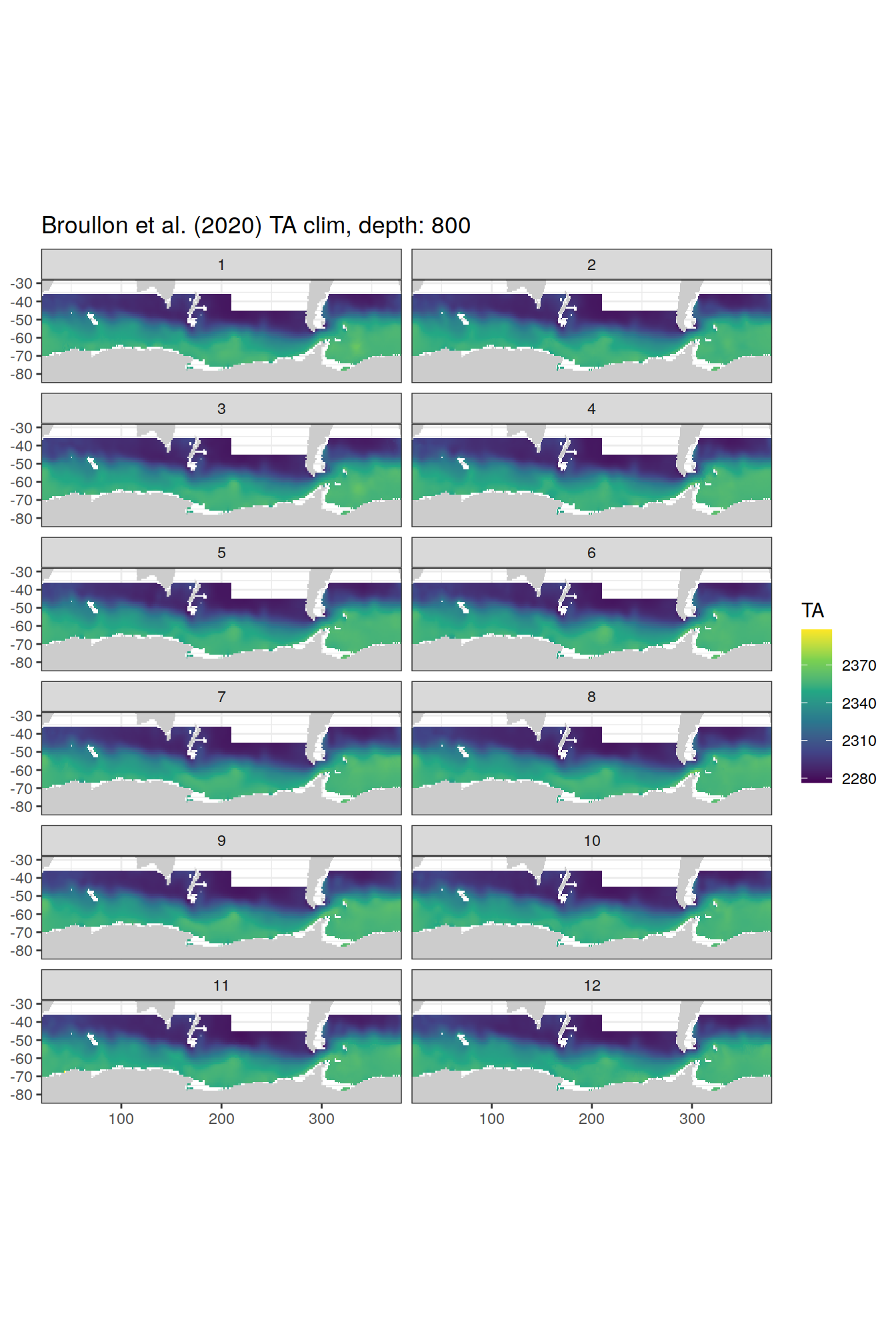
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[44]]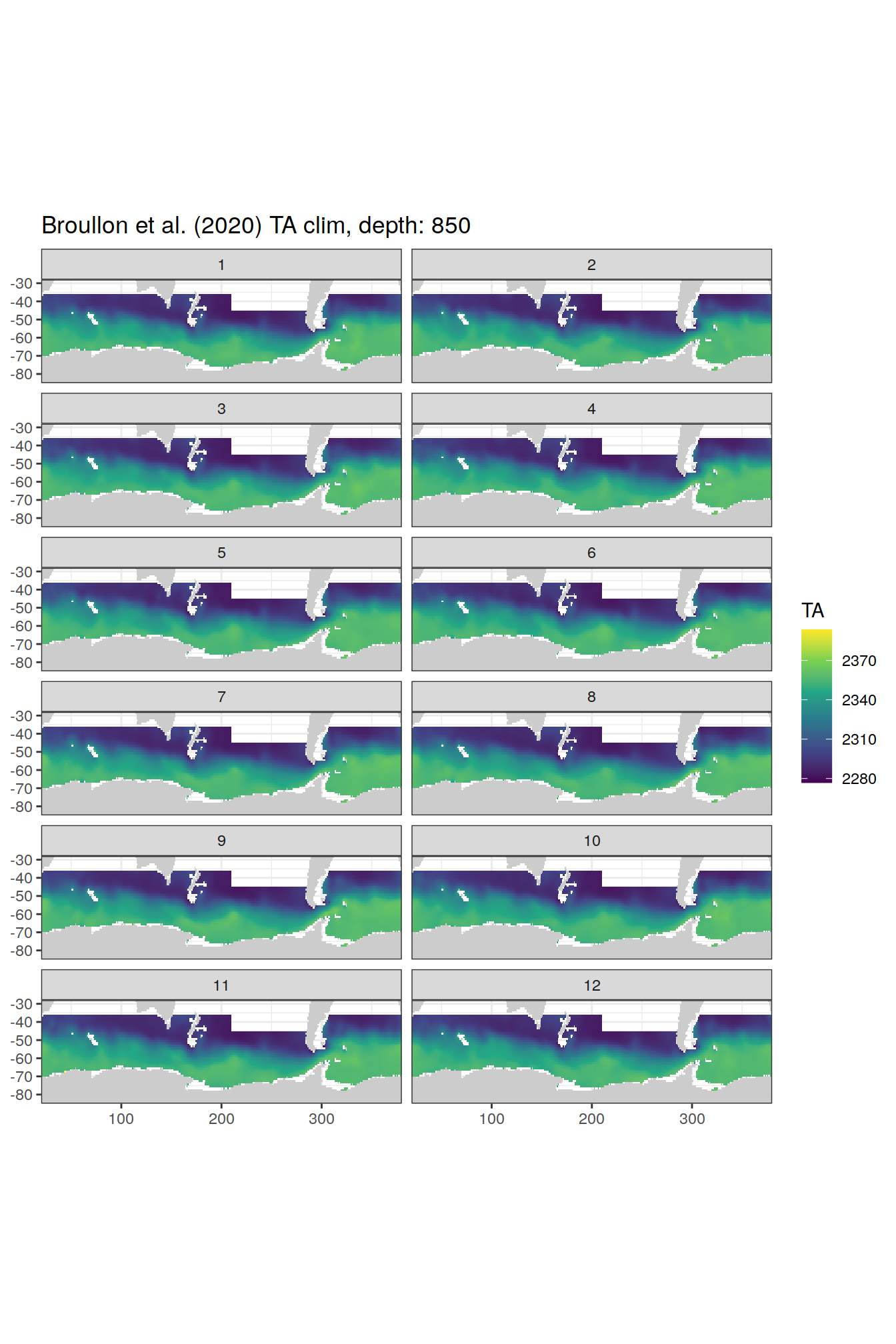
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[45]]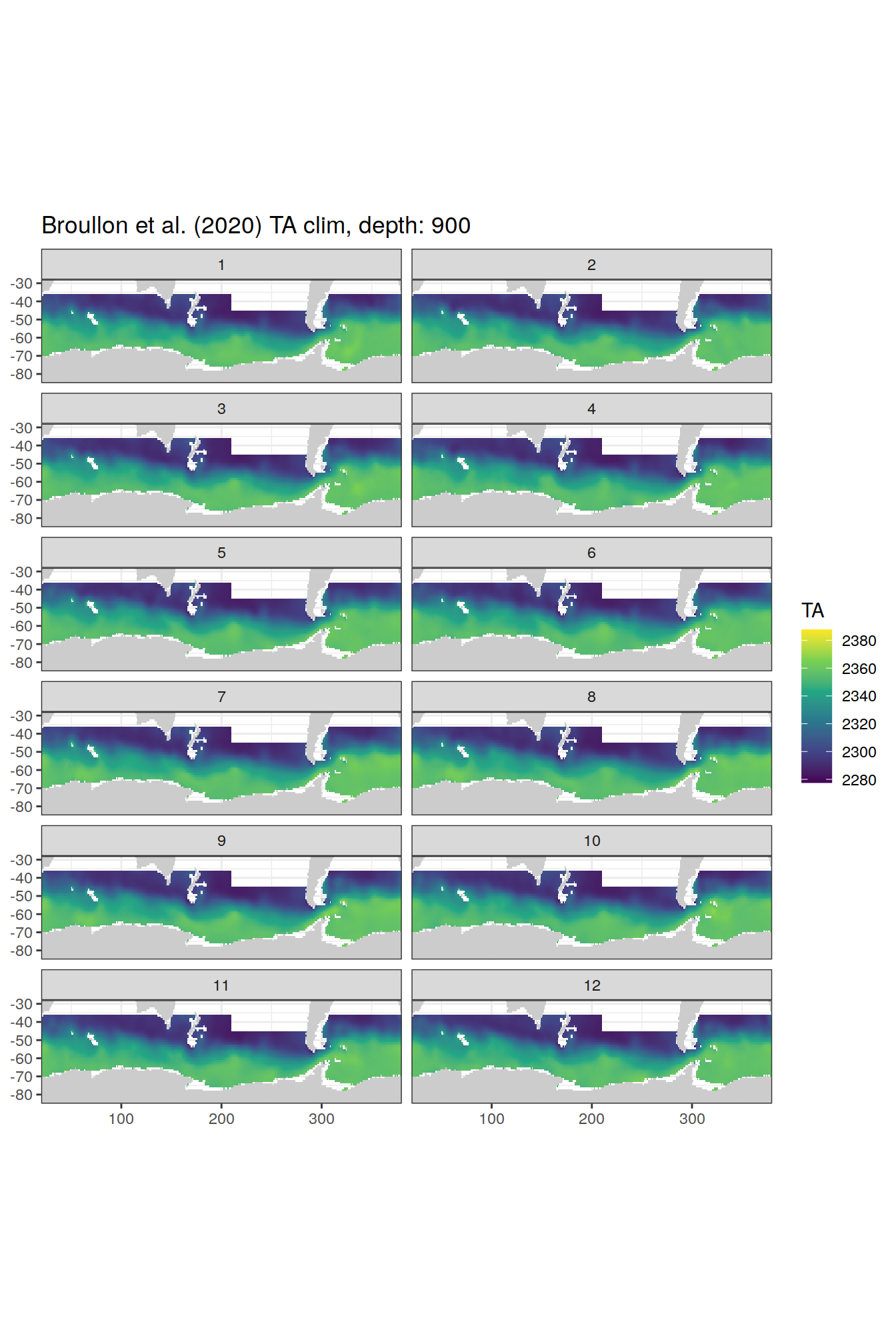
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[46]]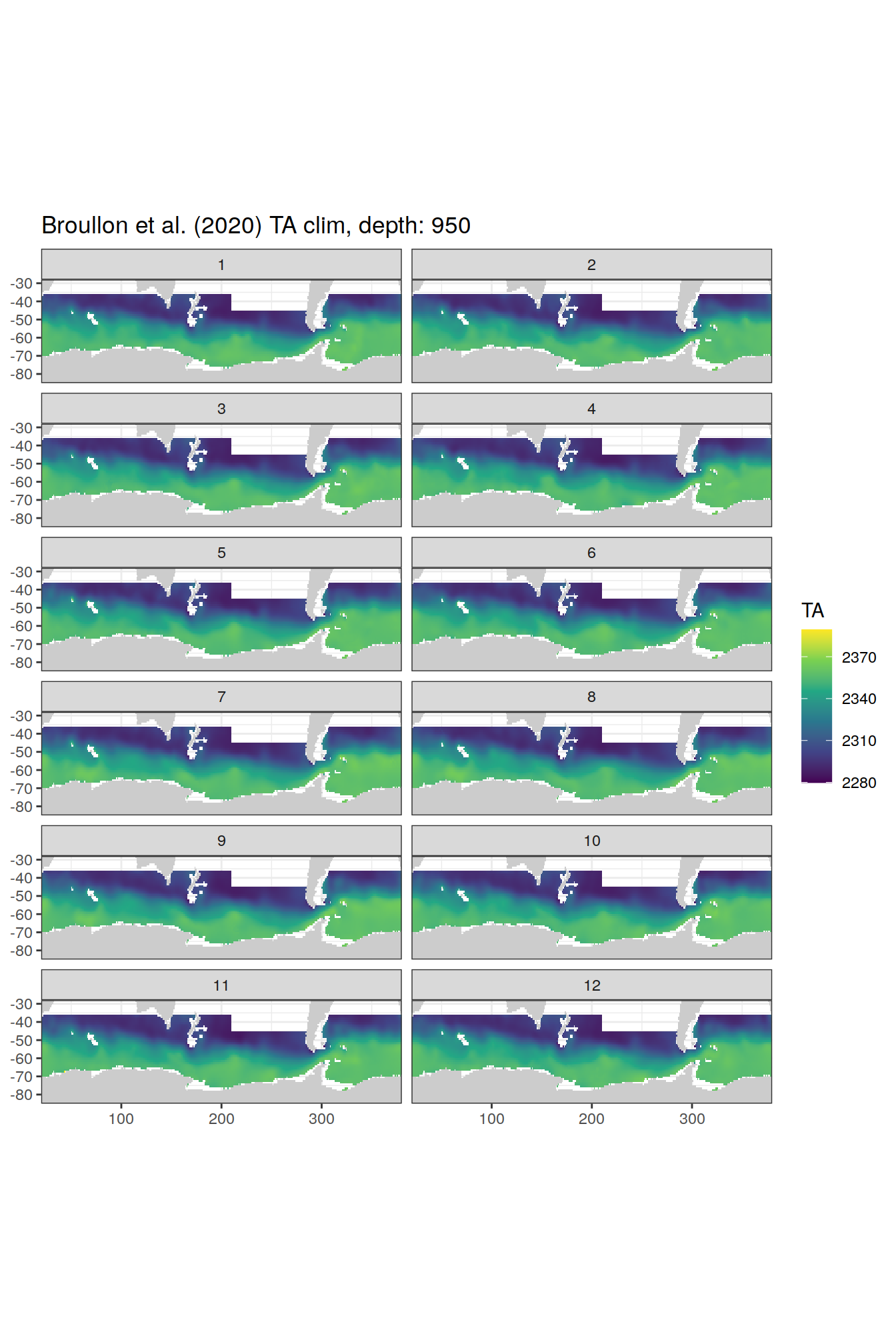
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[47]]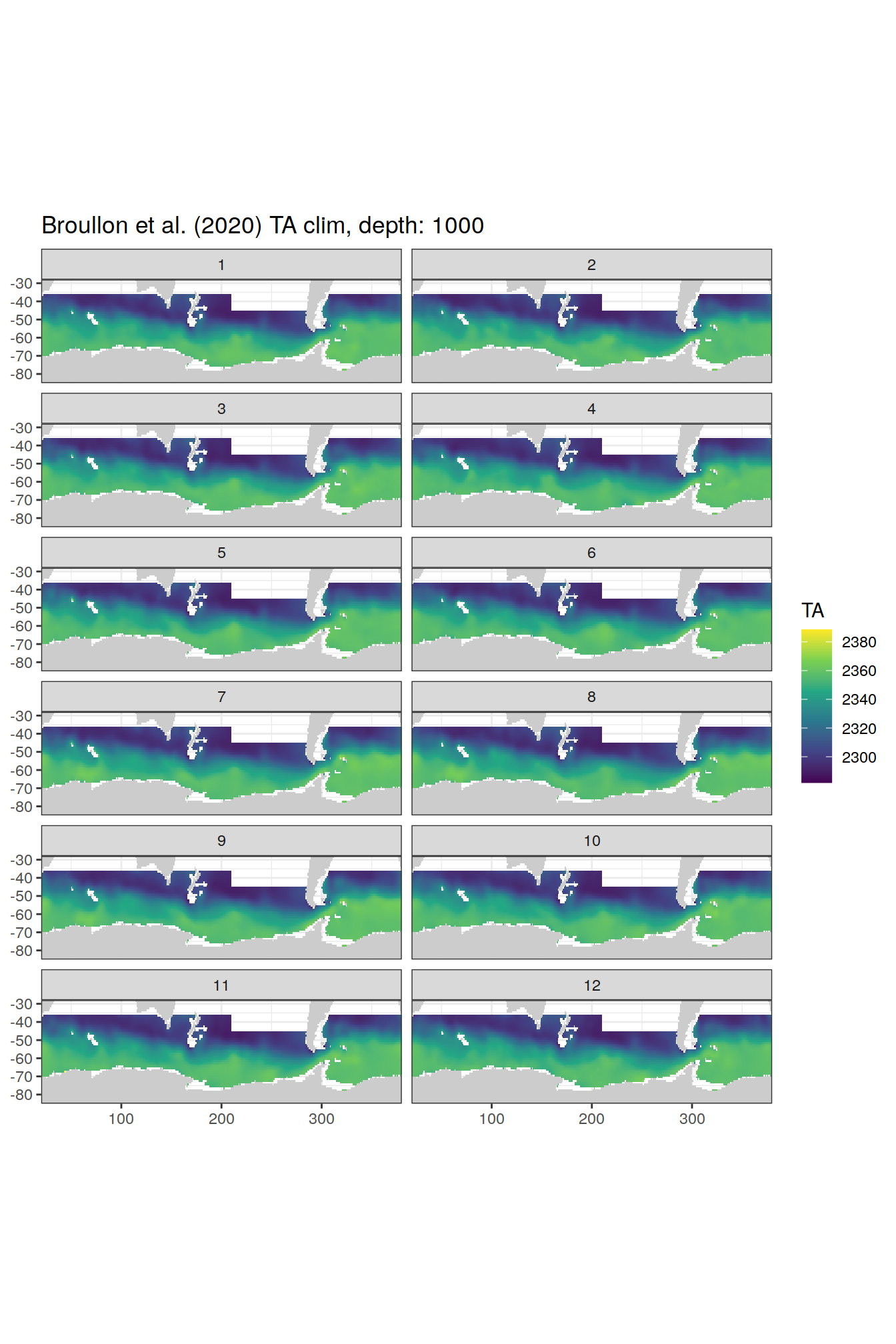
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[48]]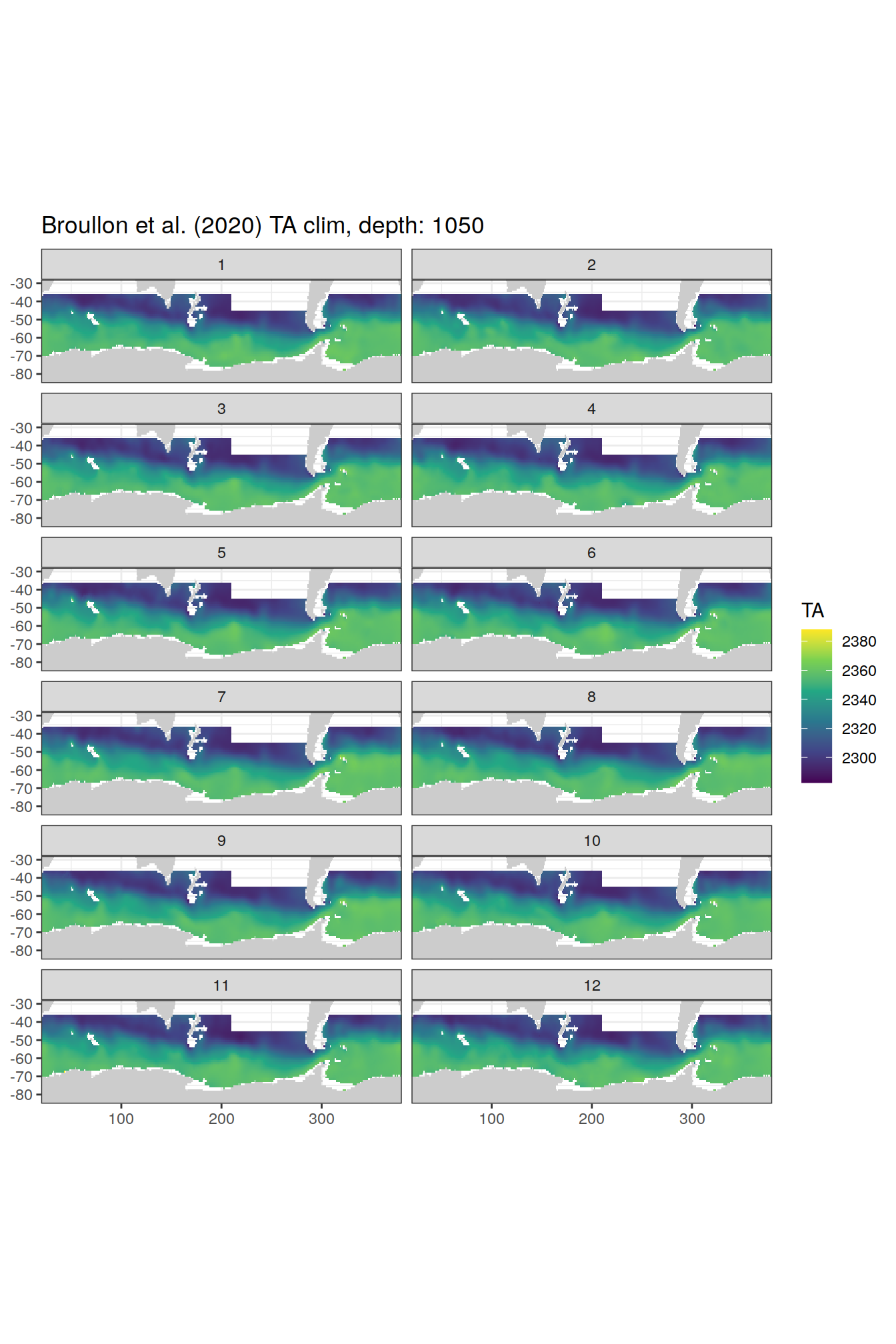
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[49]]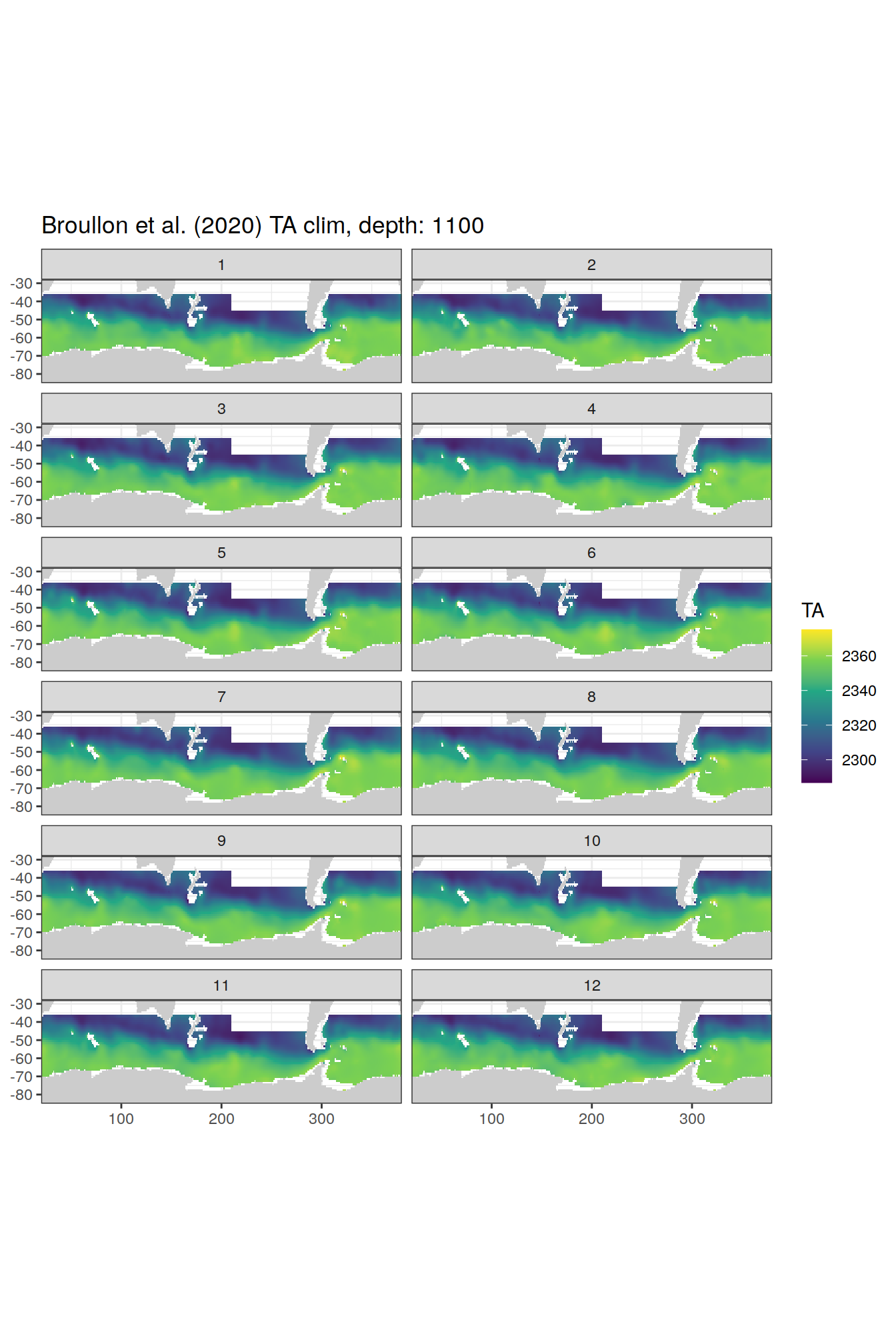
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[50]]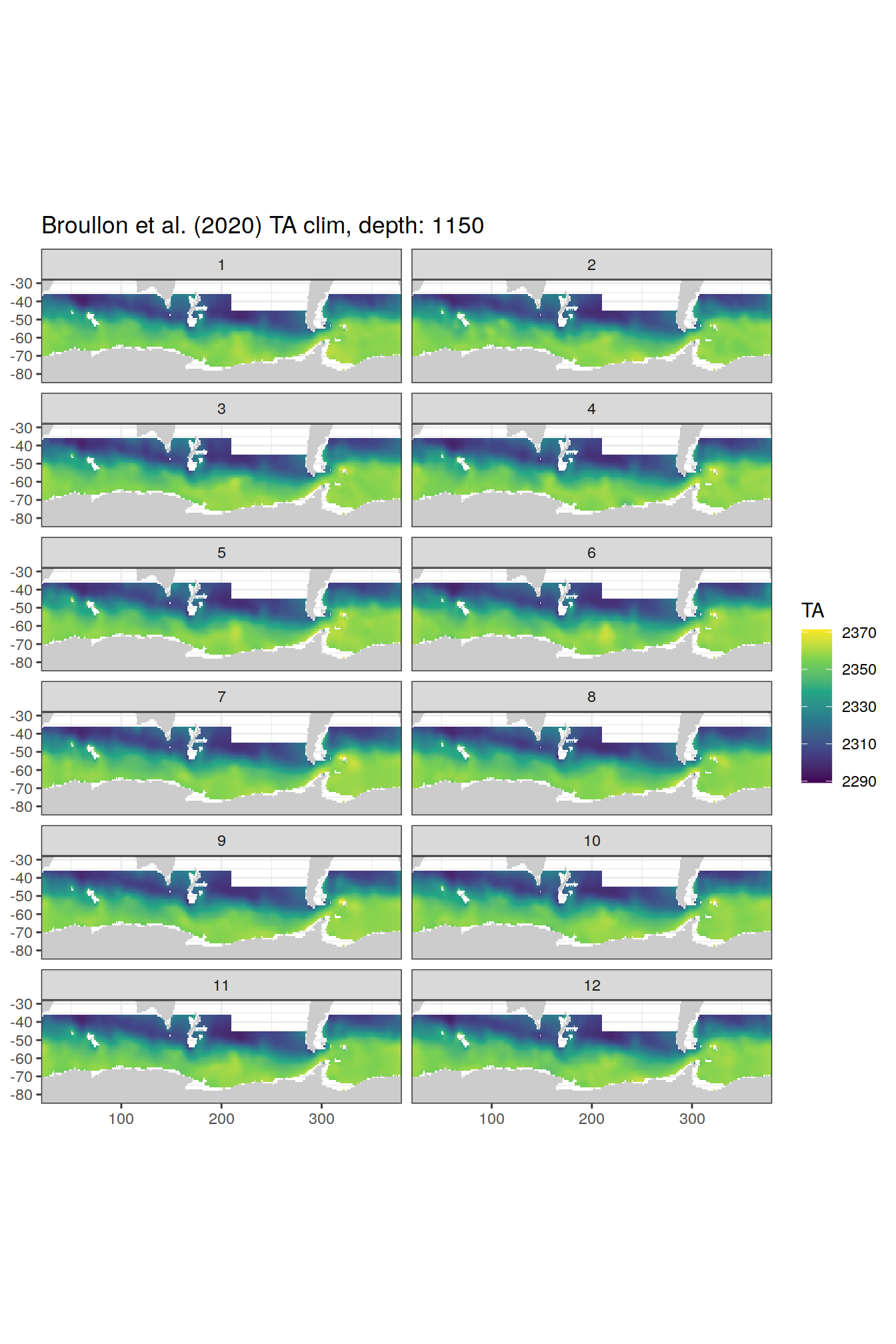
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[51]]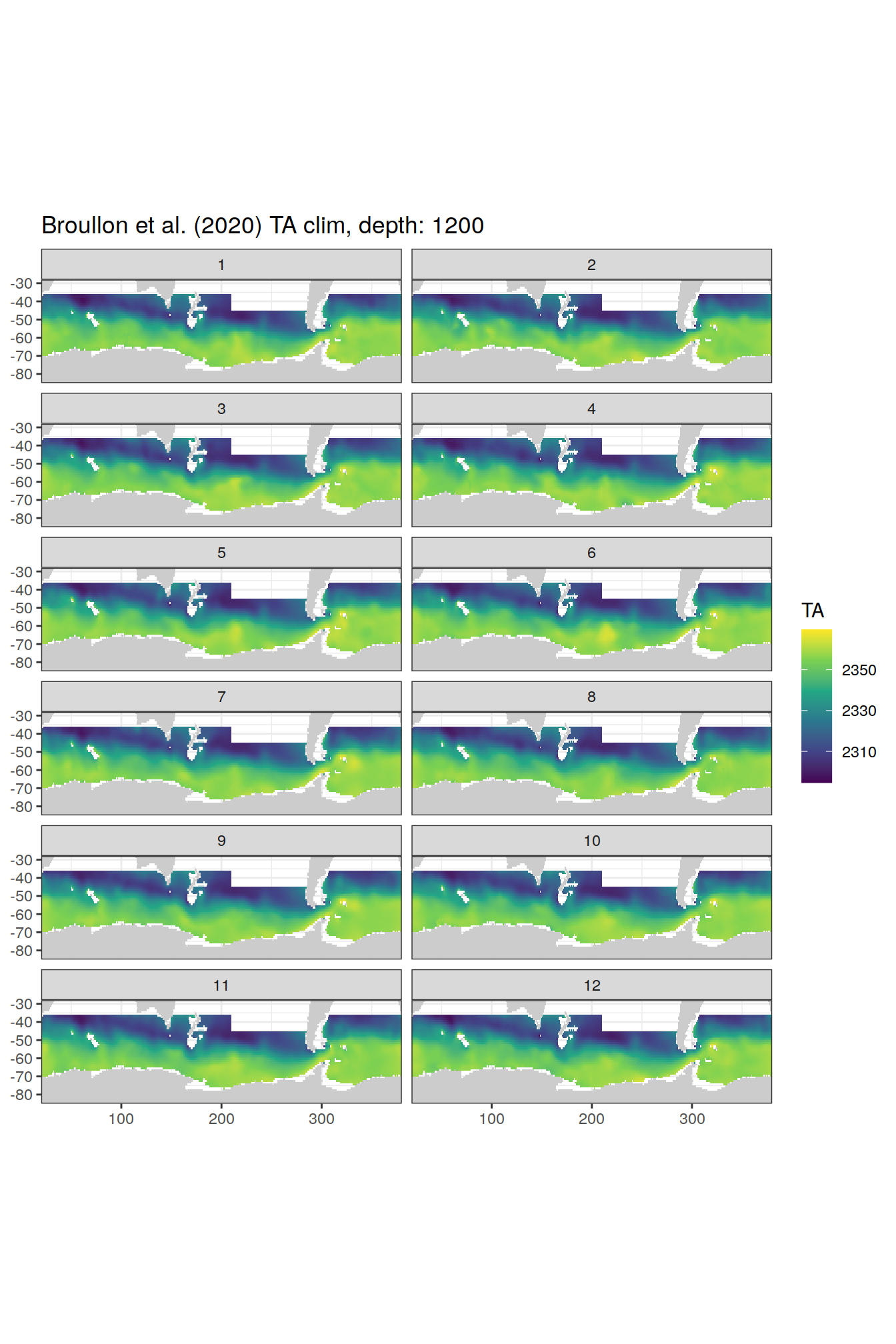
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[52]]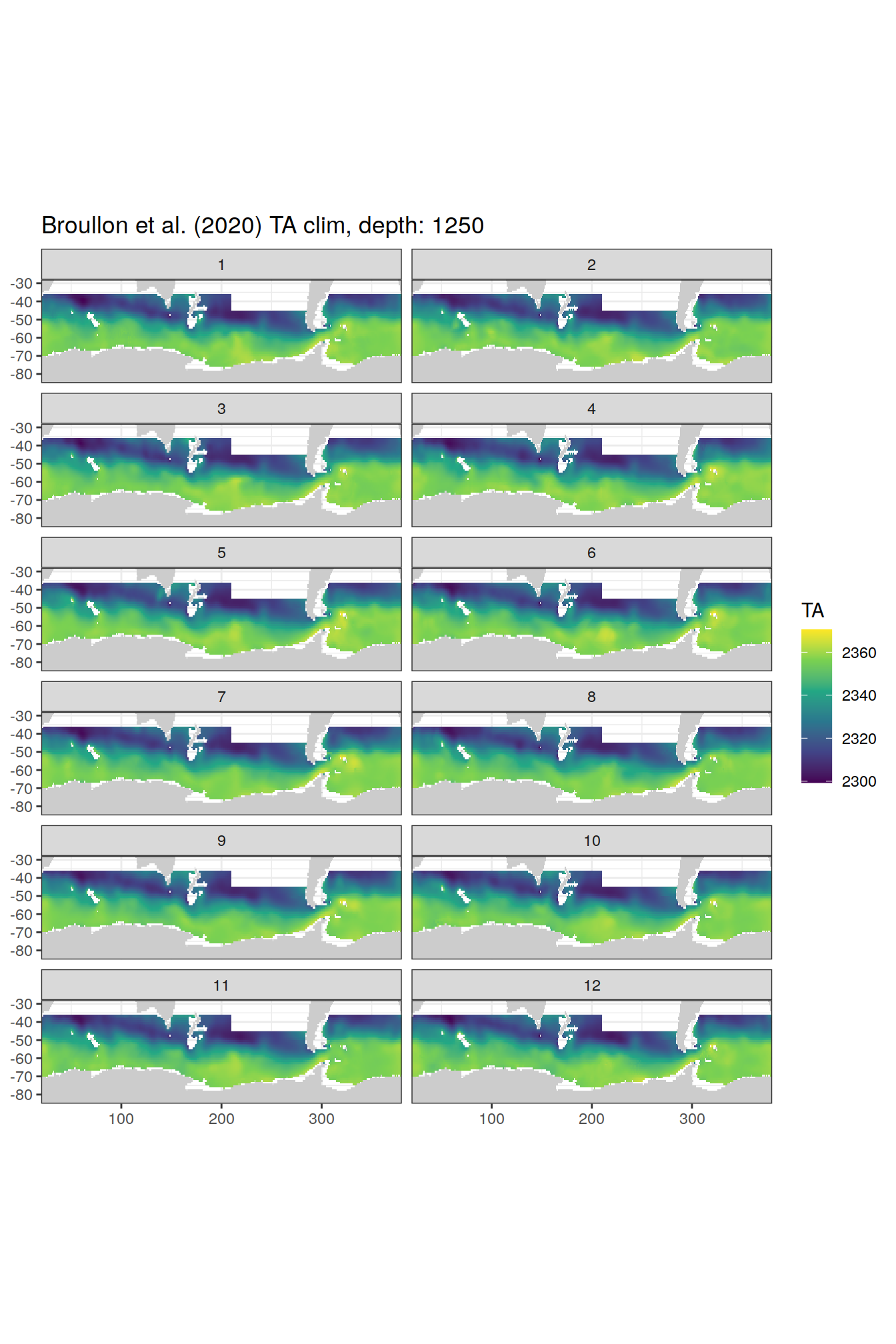
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[53]]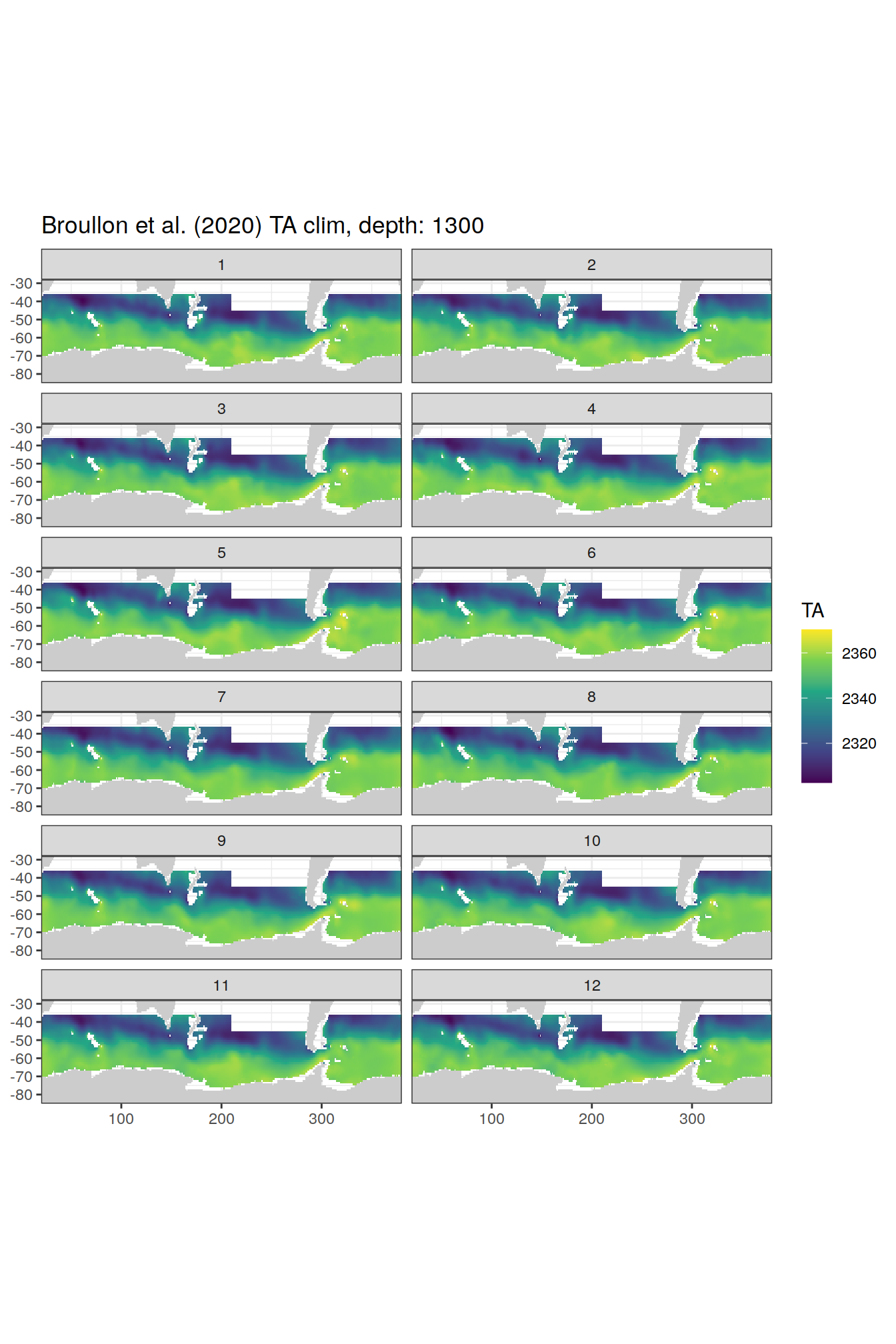
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[54]]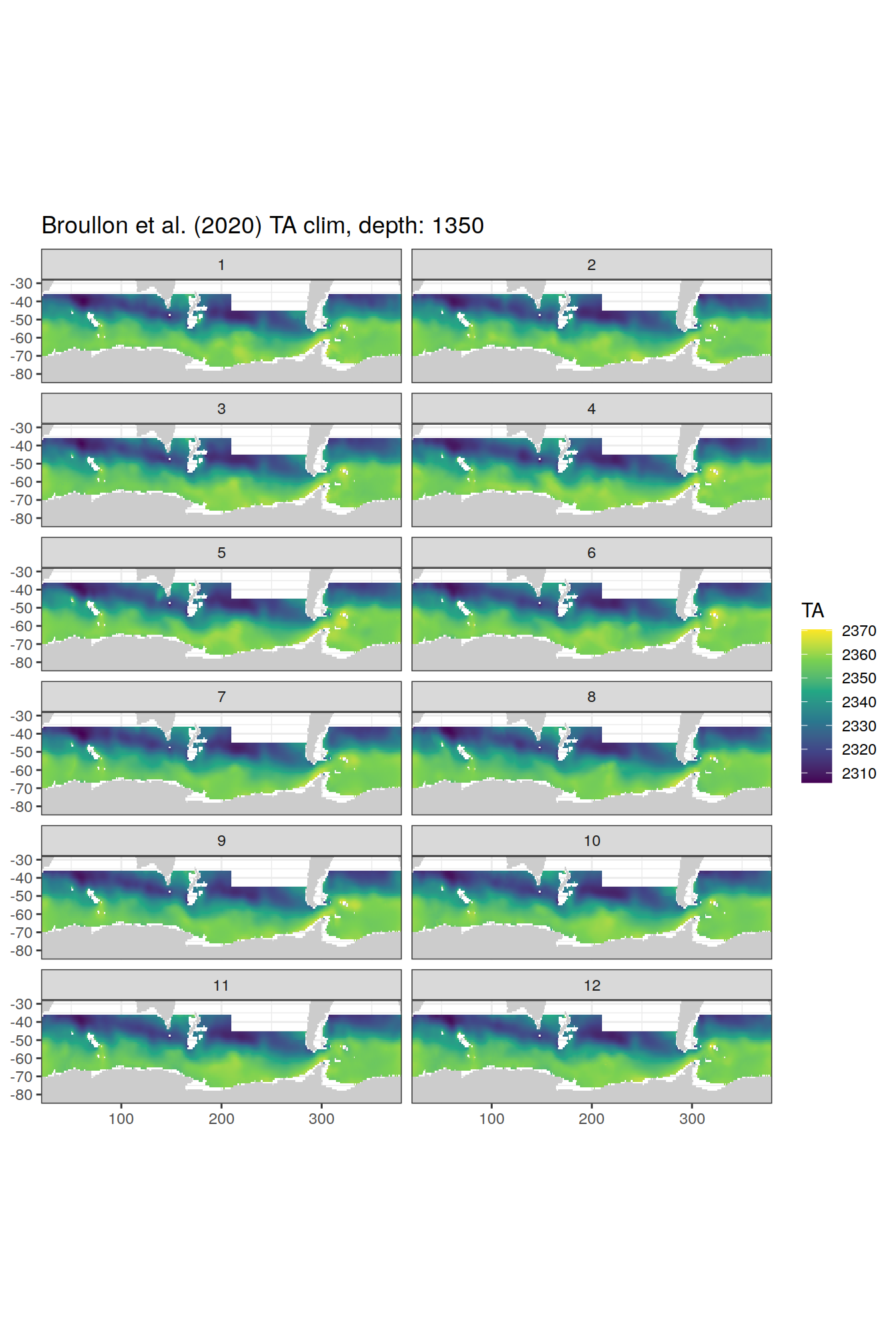
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[55]]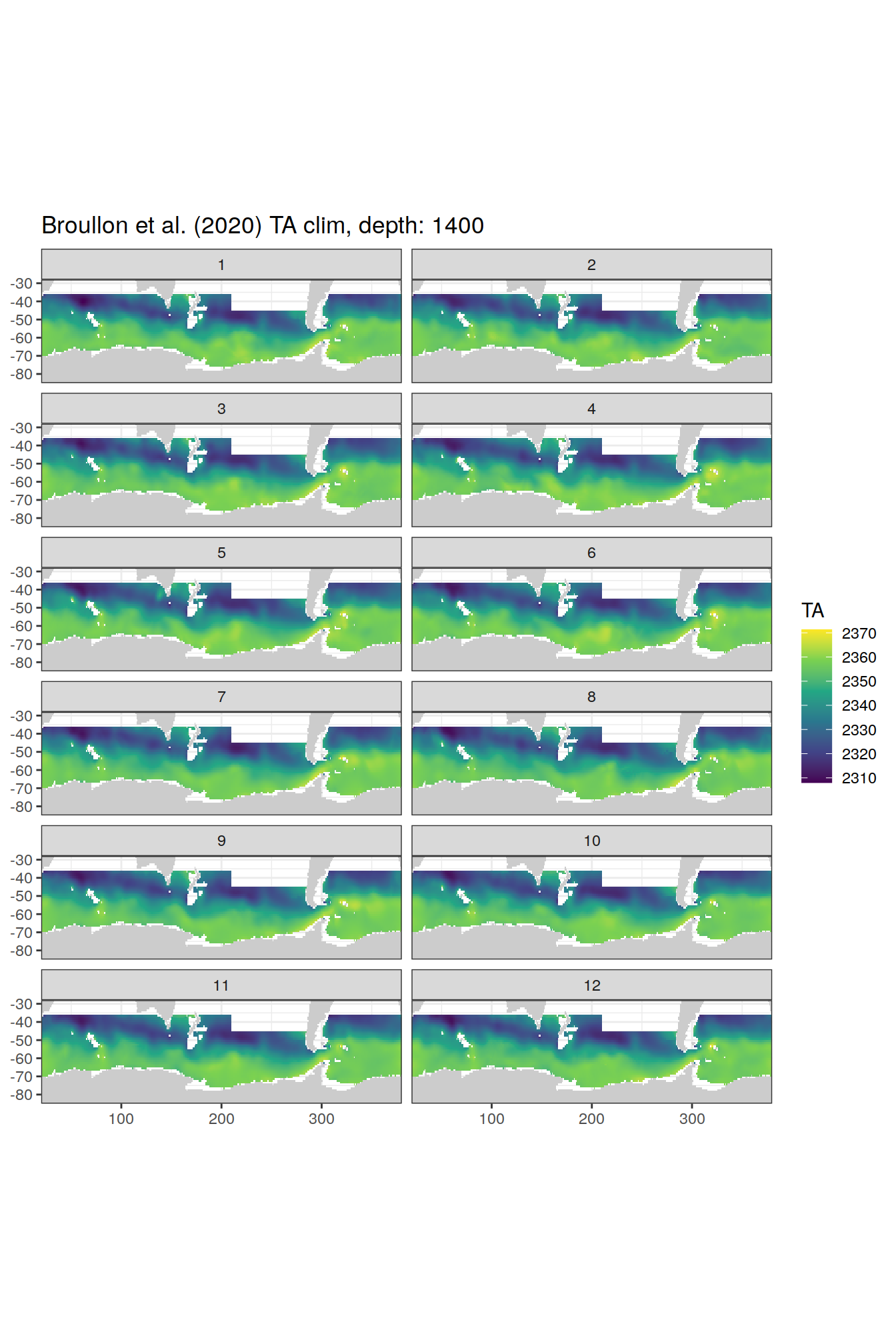
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[56]]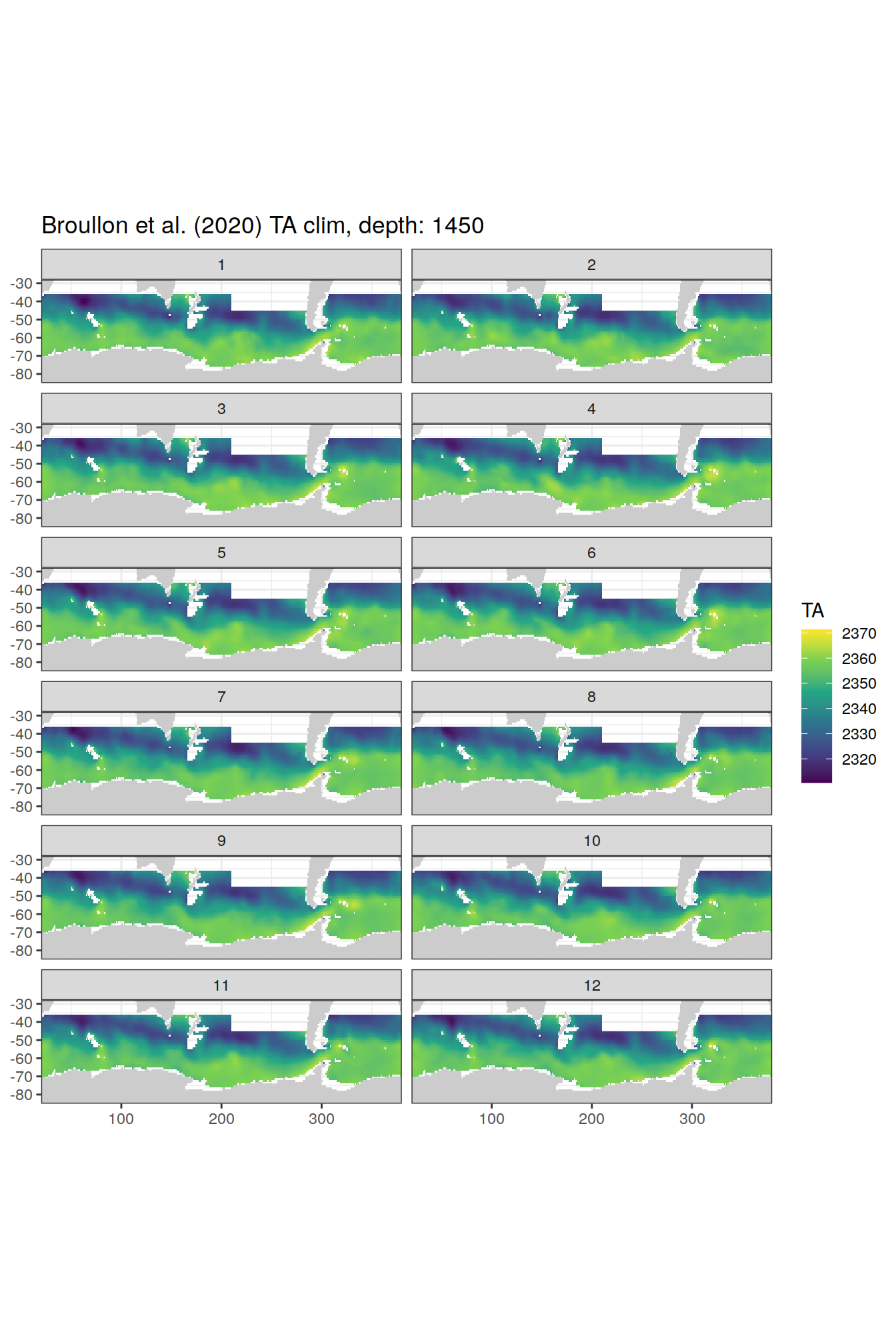
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[57]]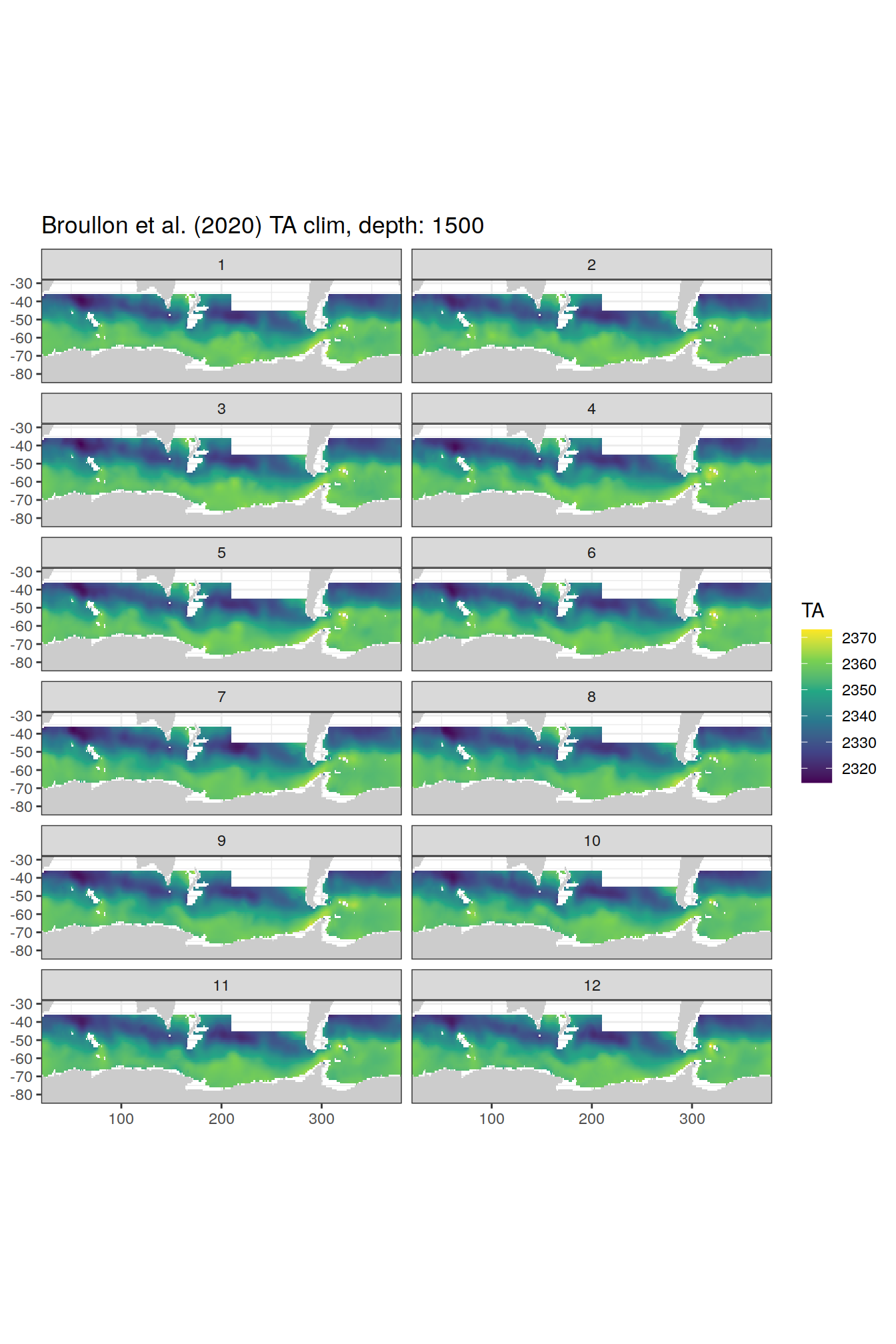
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[58]]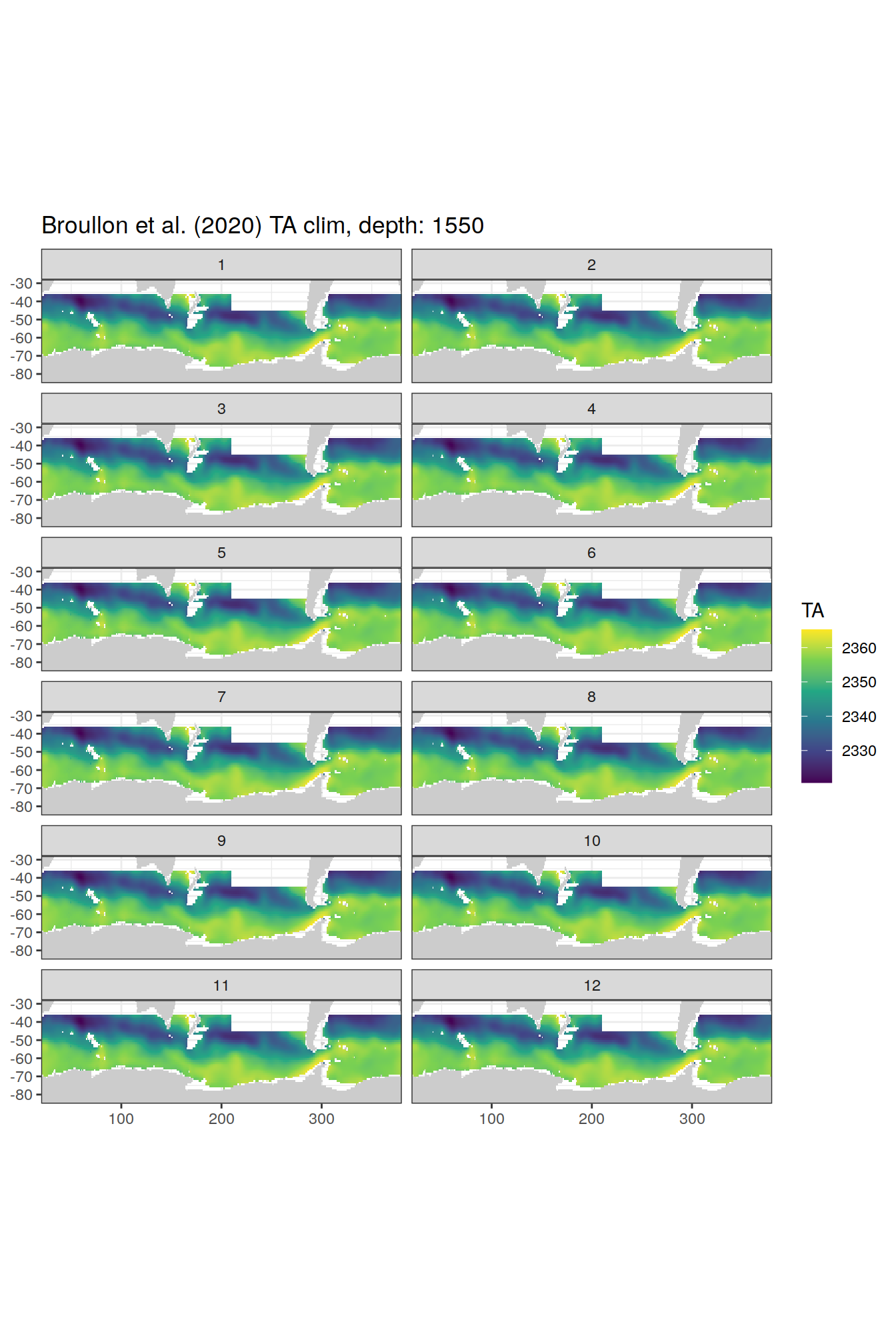
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[59]]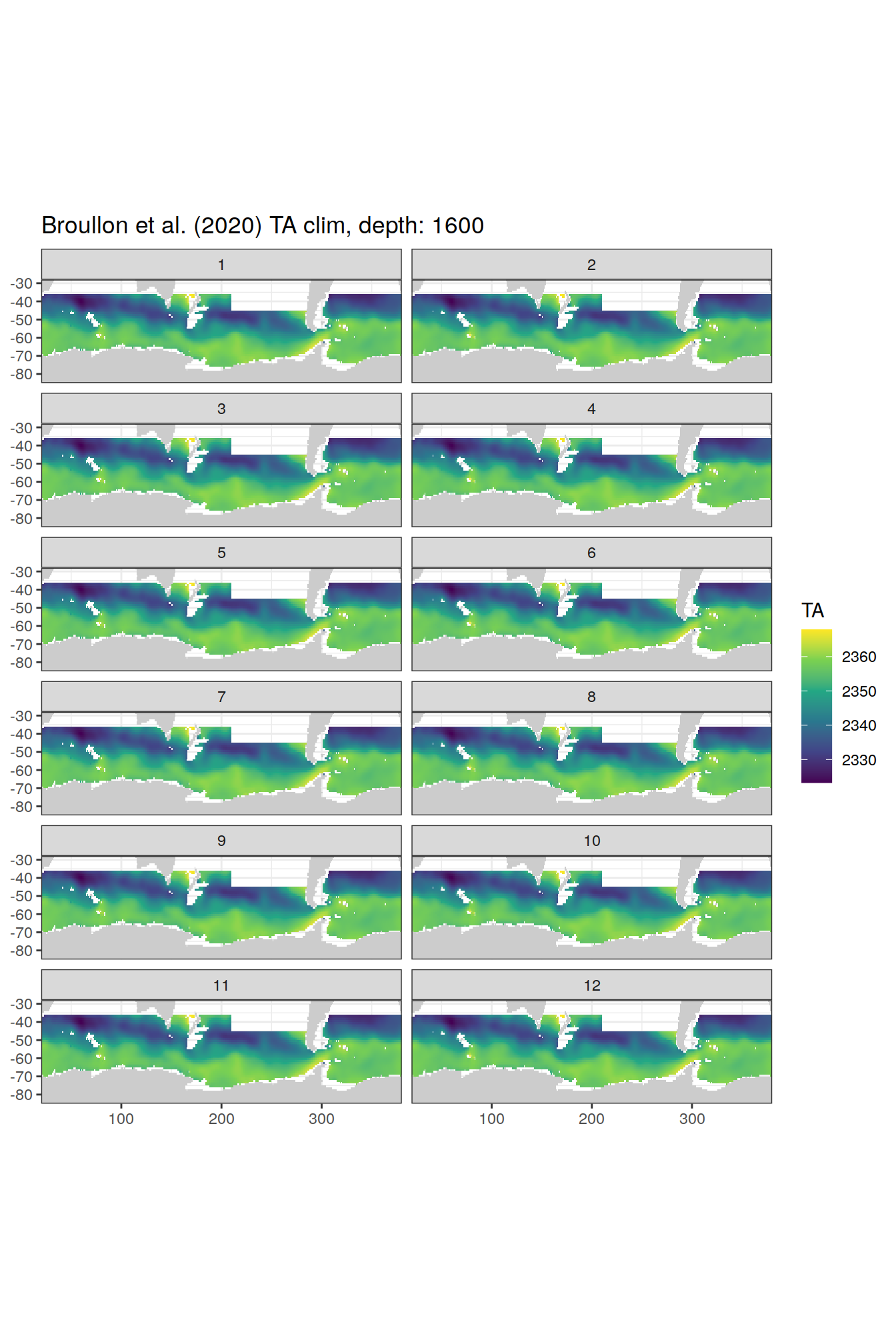
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[60]]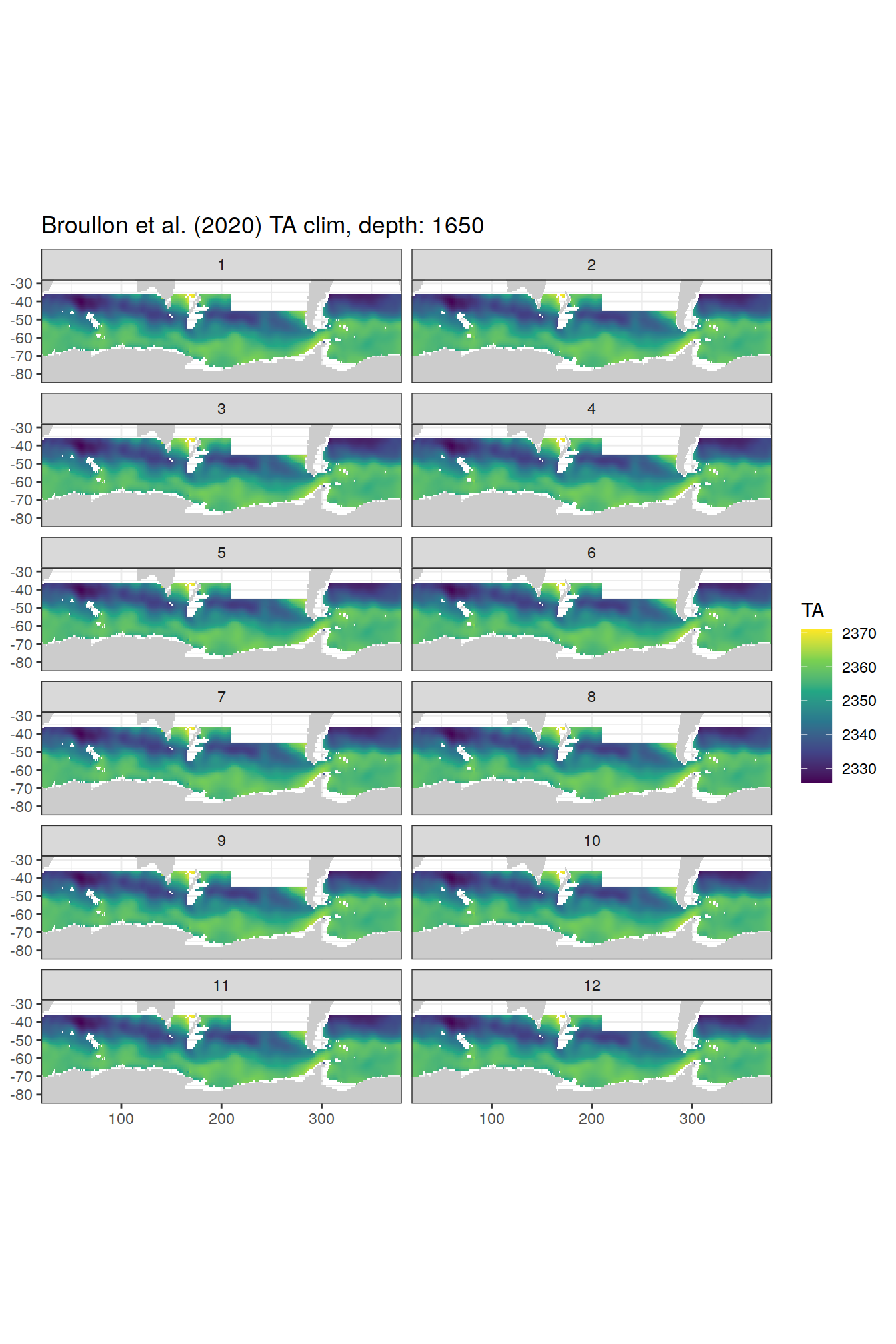
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[61]]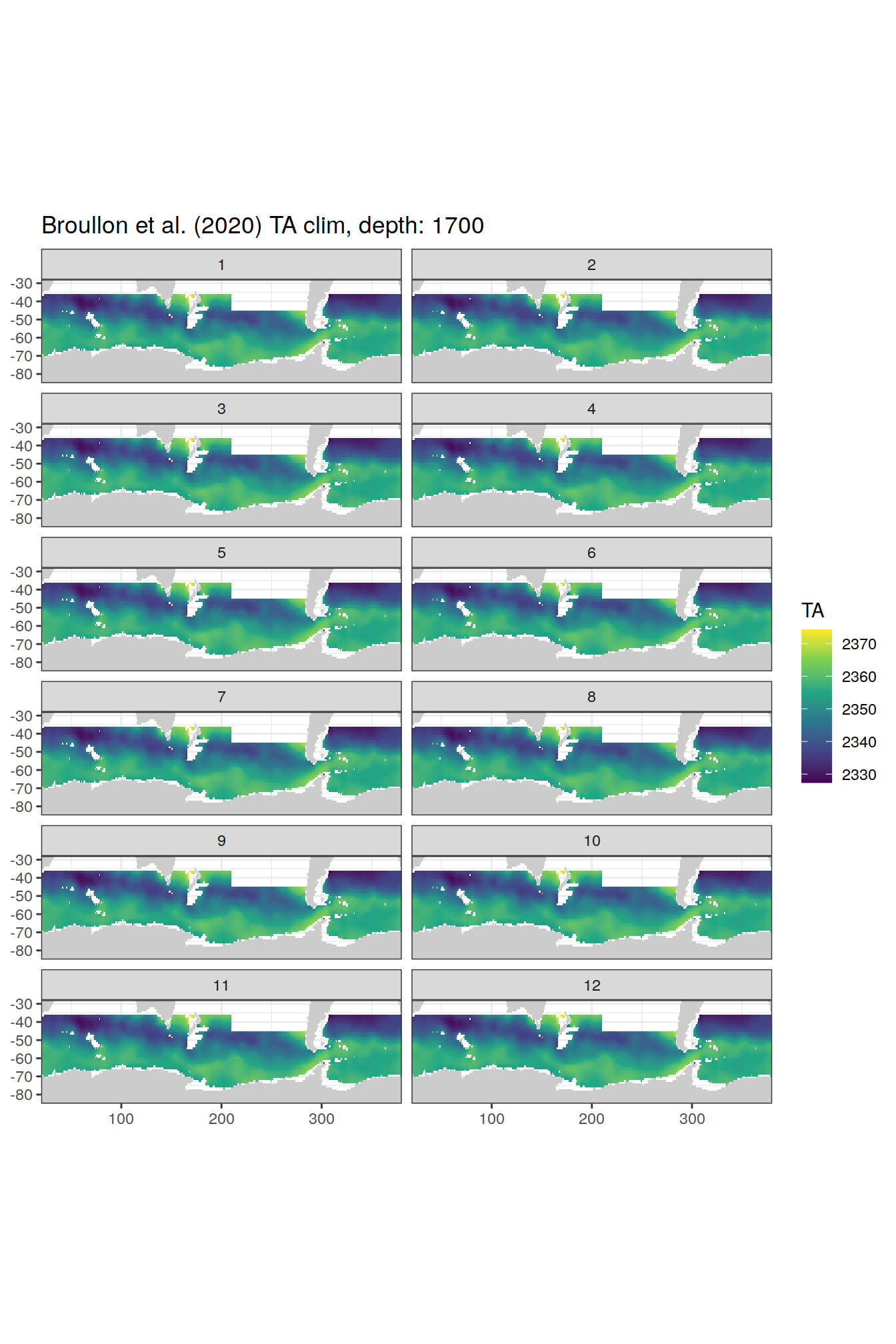
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[62]]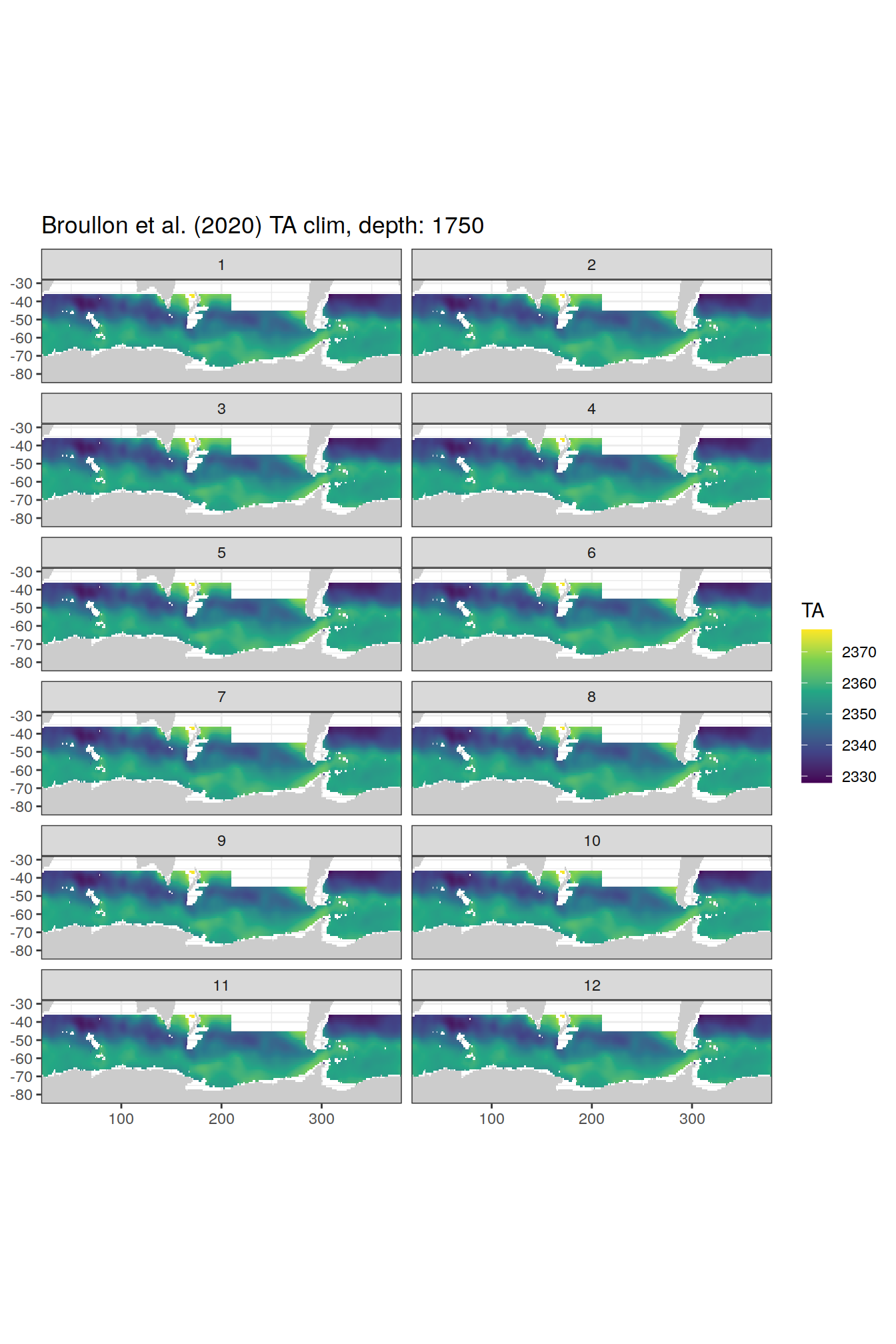
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[63]]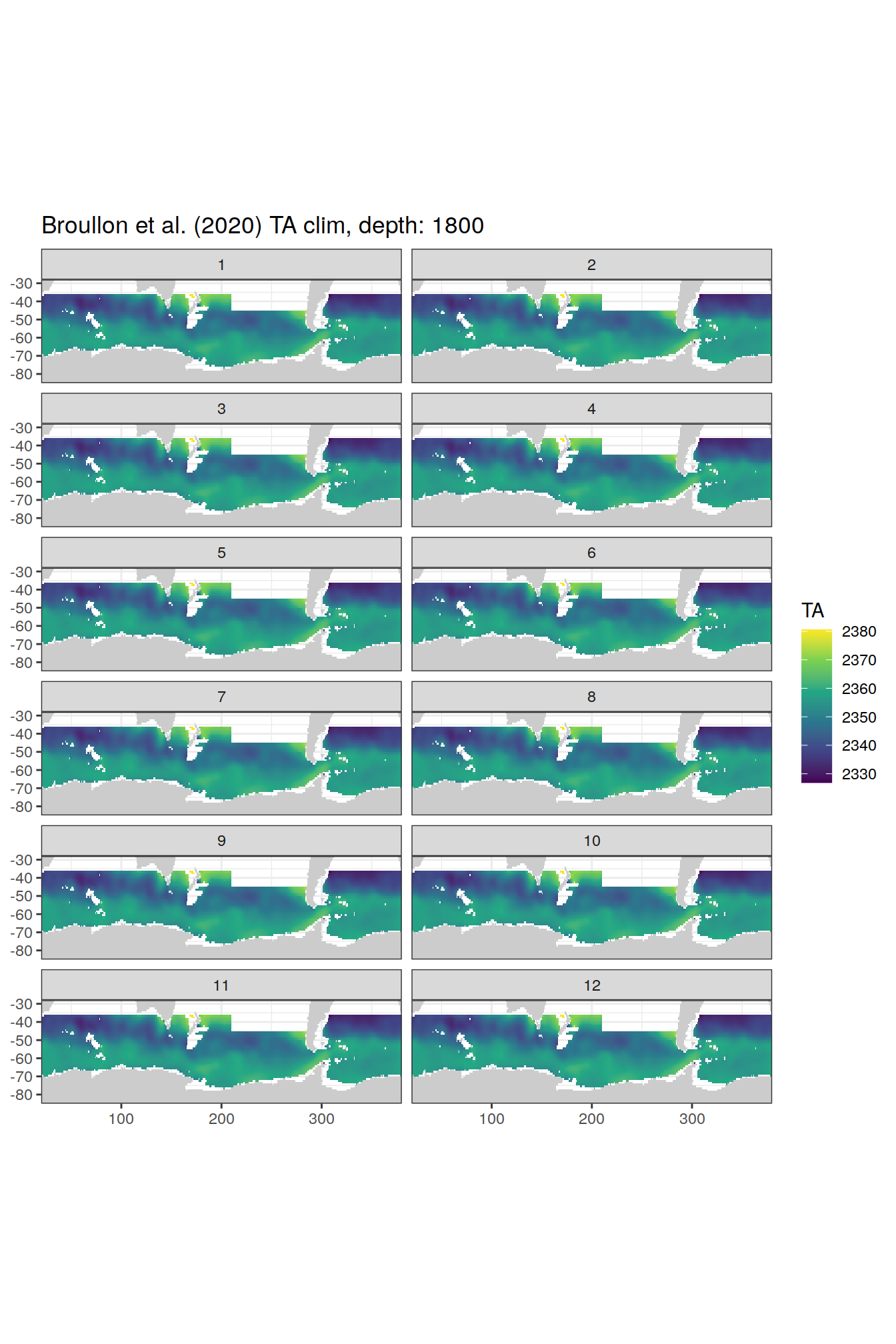
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[64]]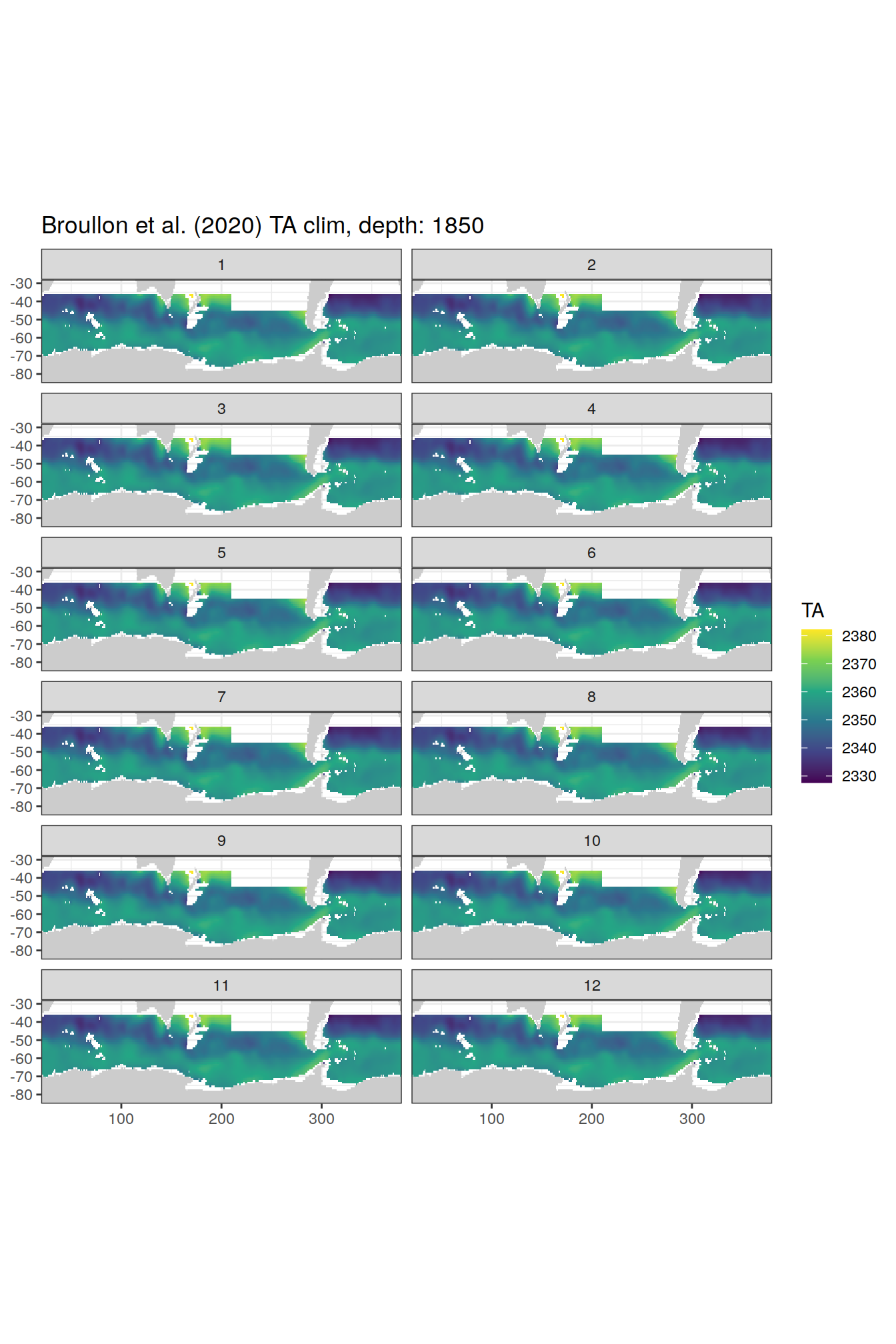
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[65]]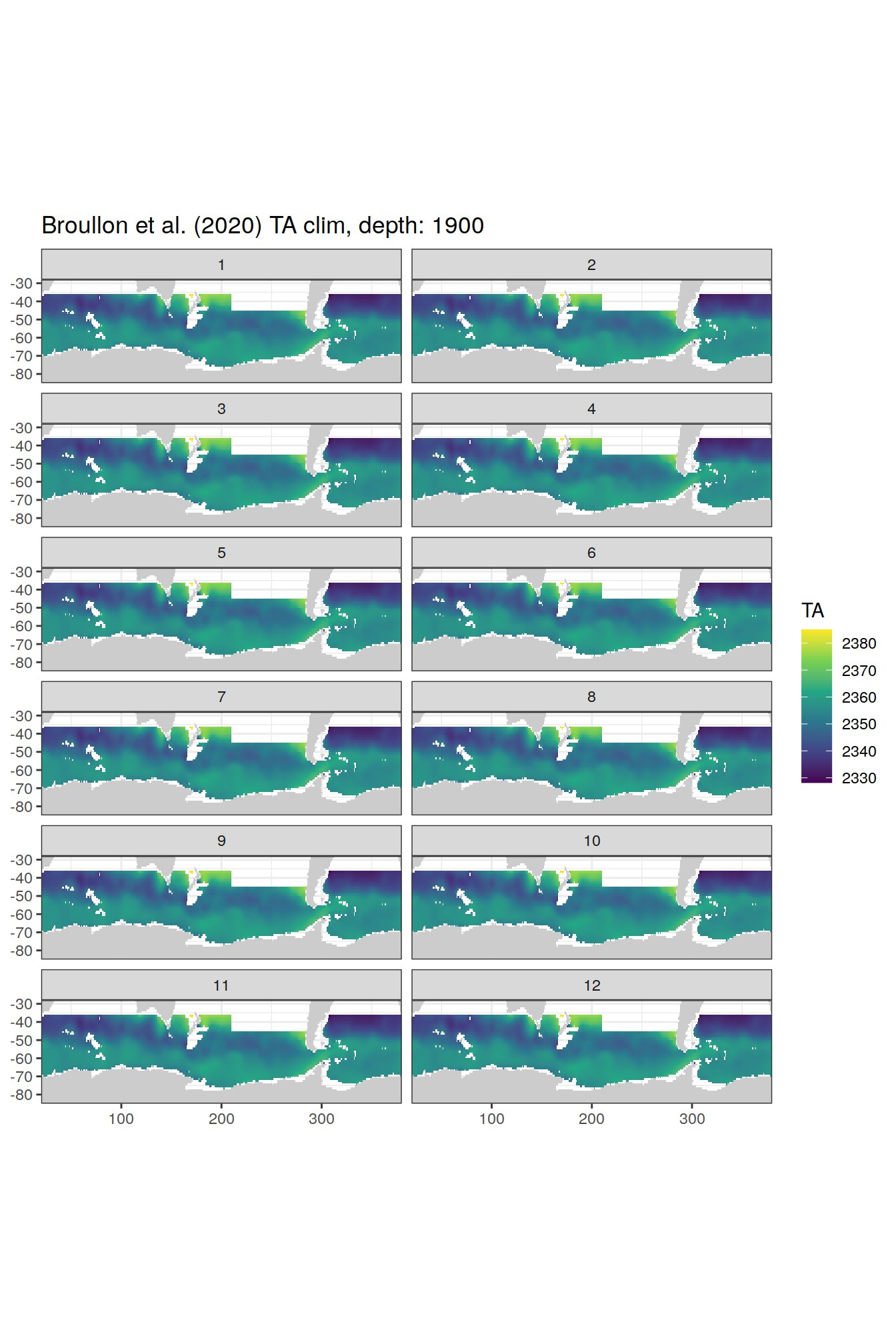
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[66]]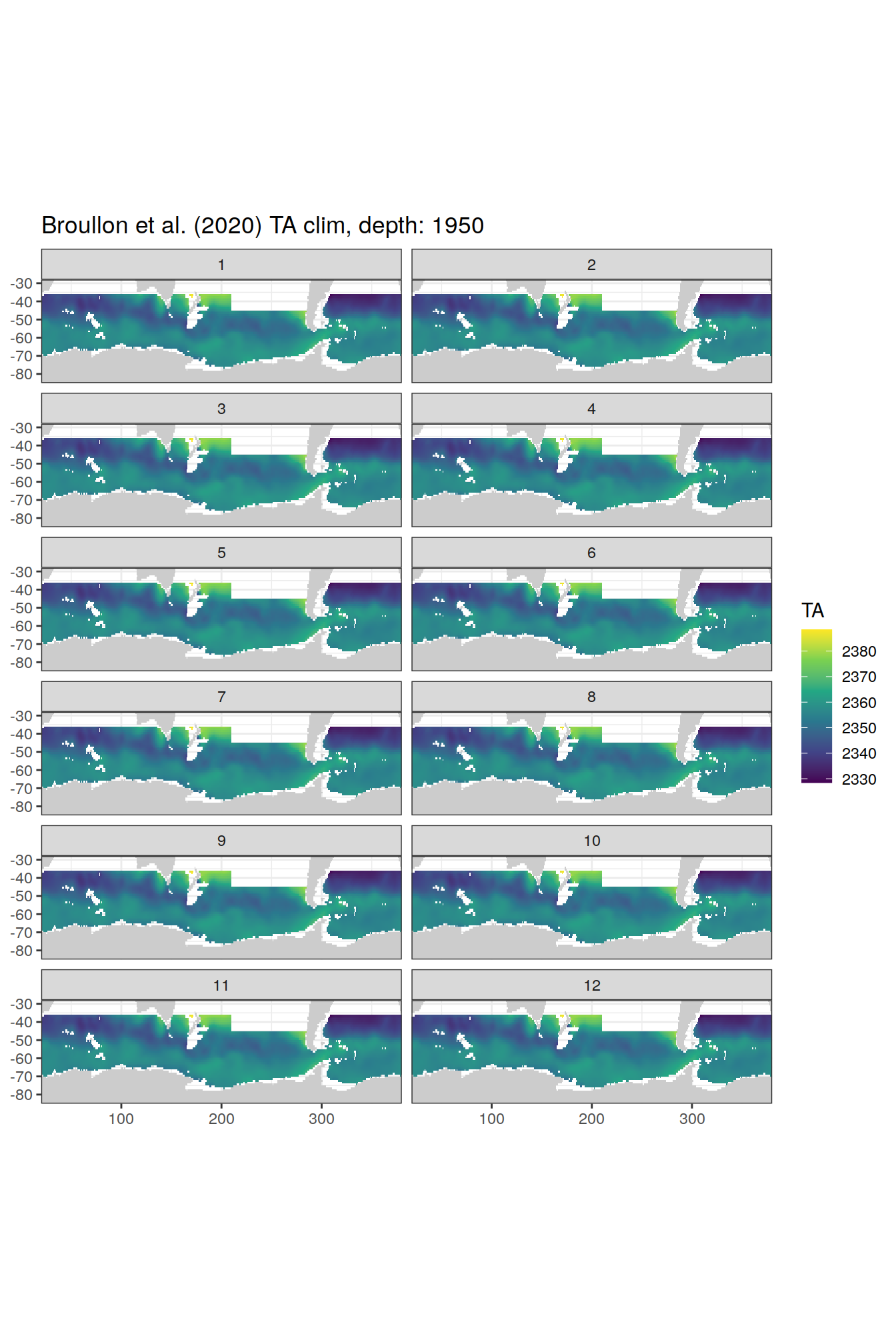
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[67]]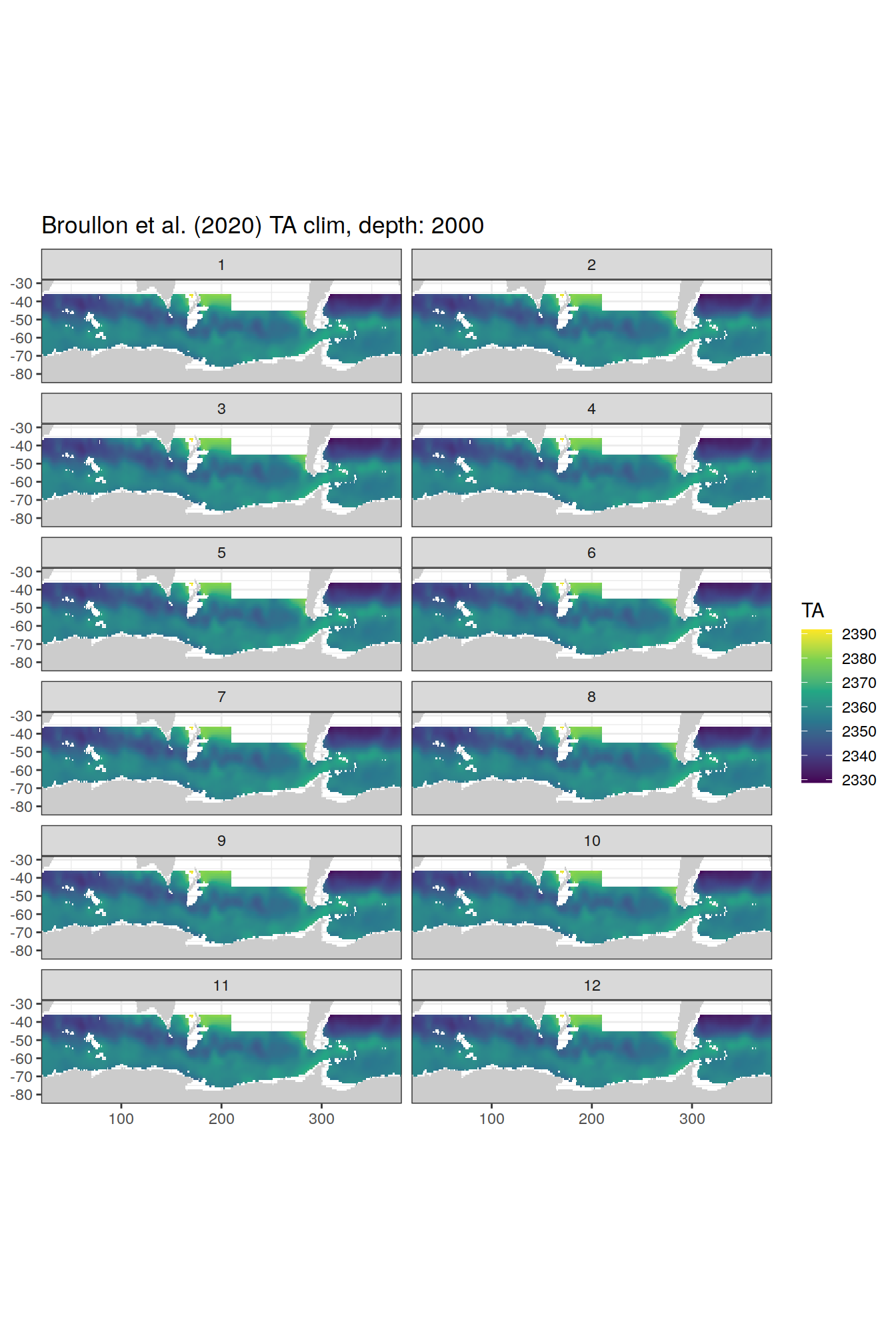
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[68]]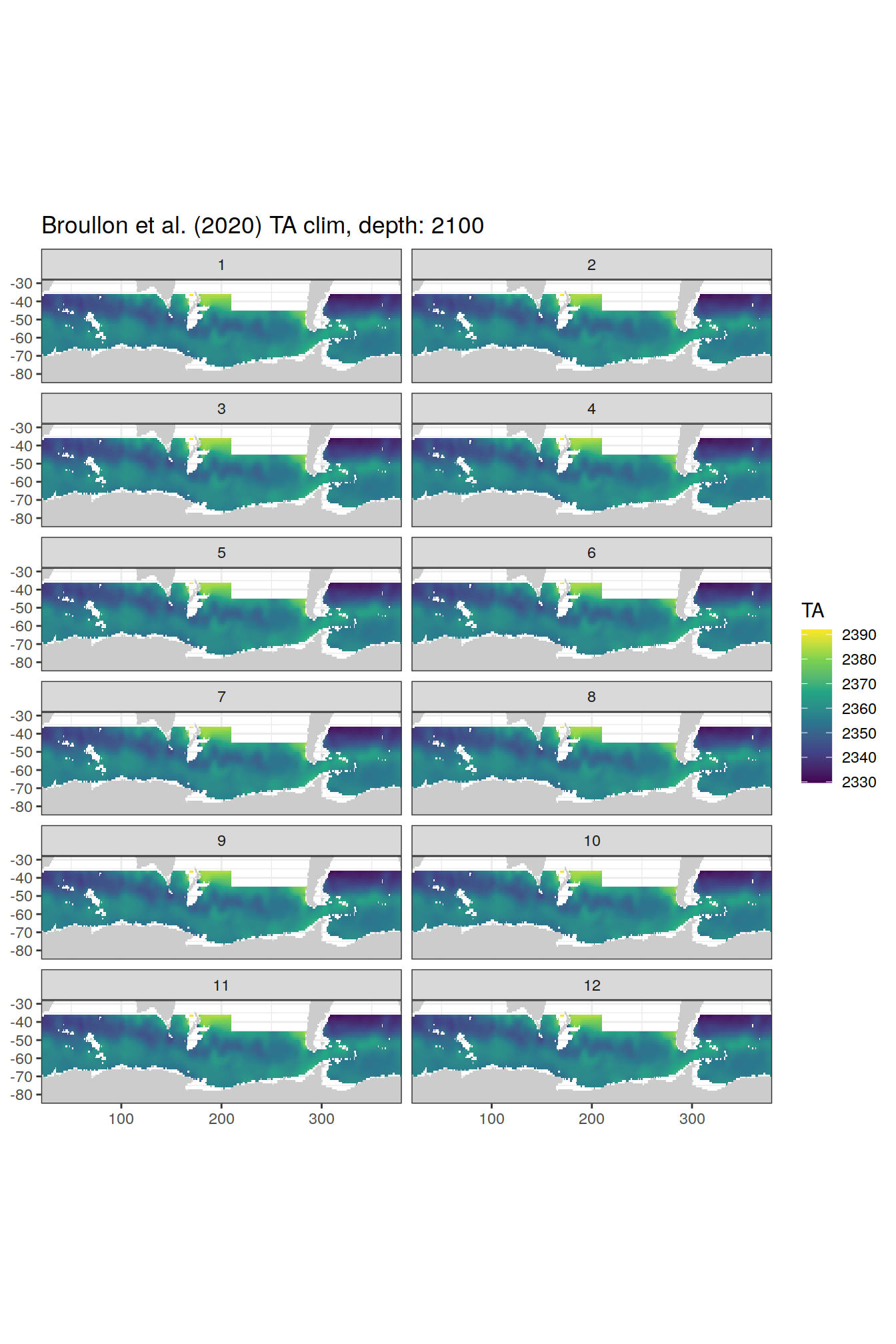
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[69]]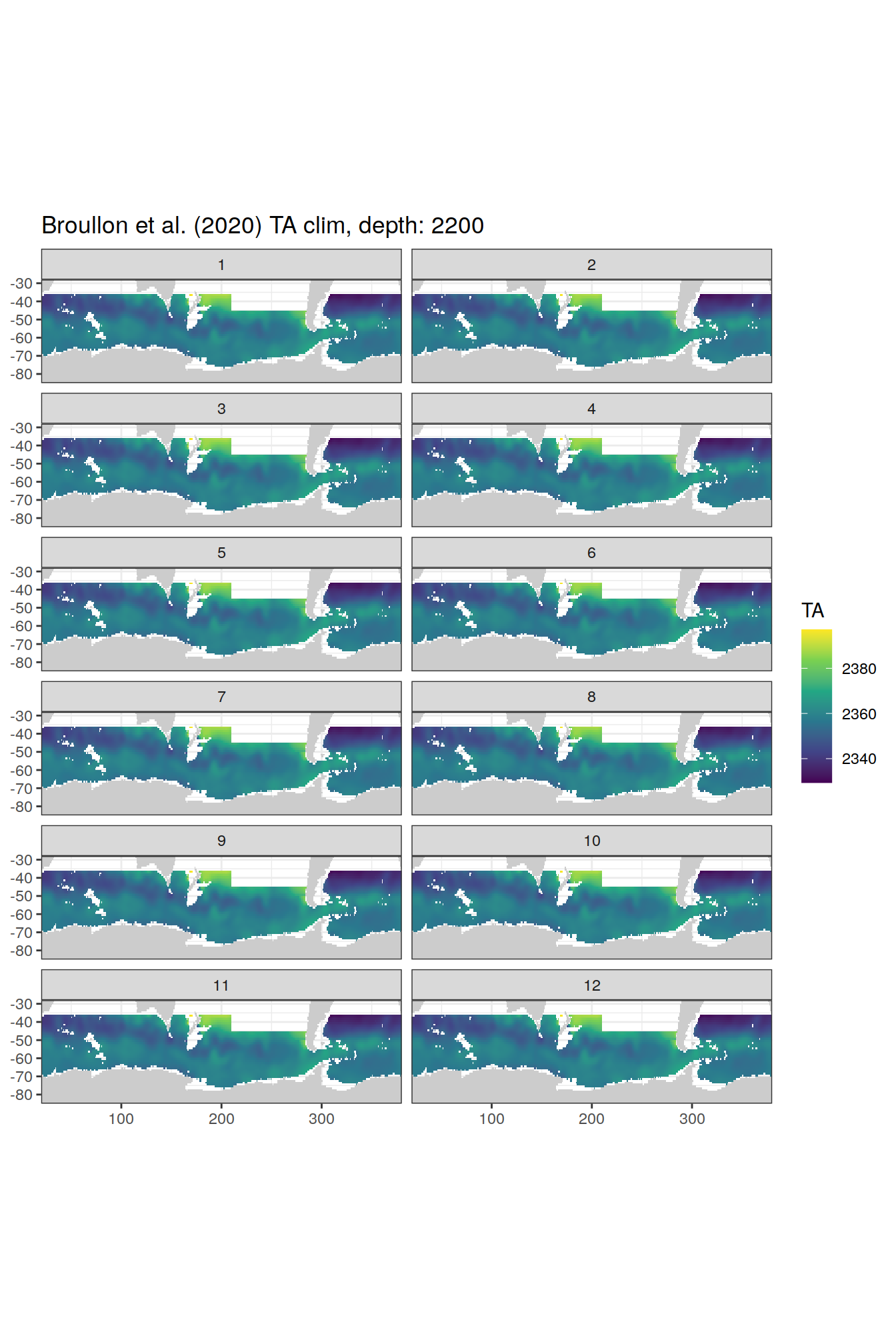
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[70]]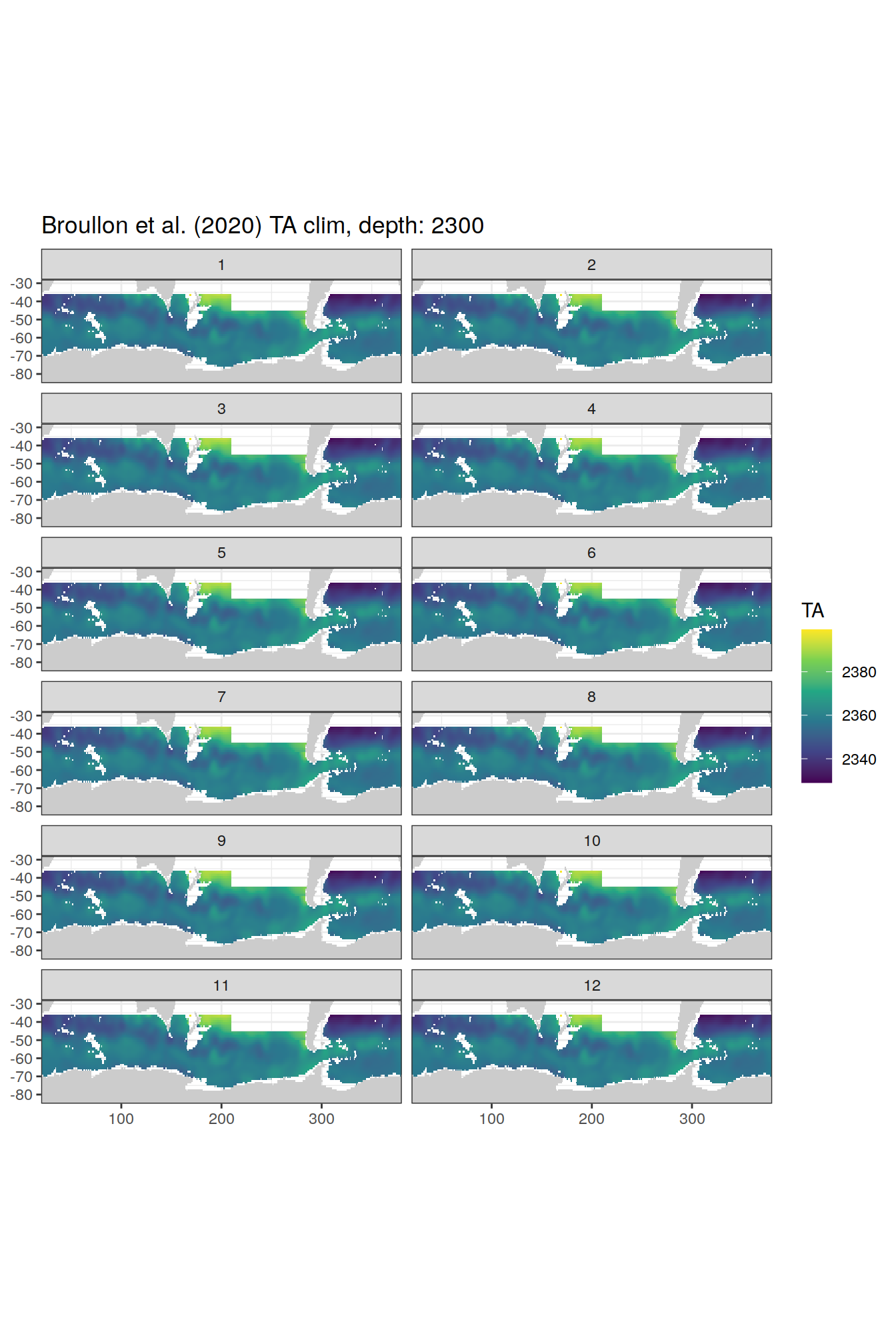
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[71]]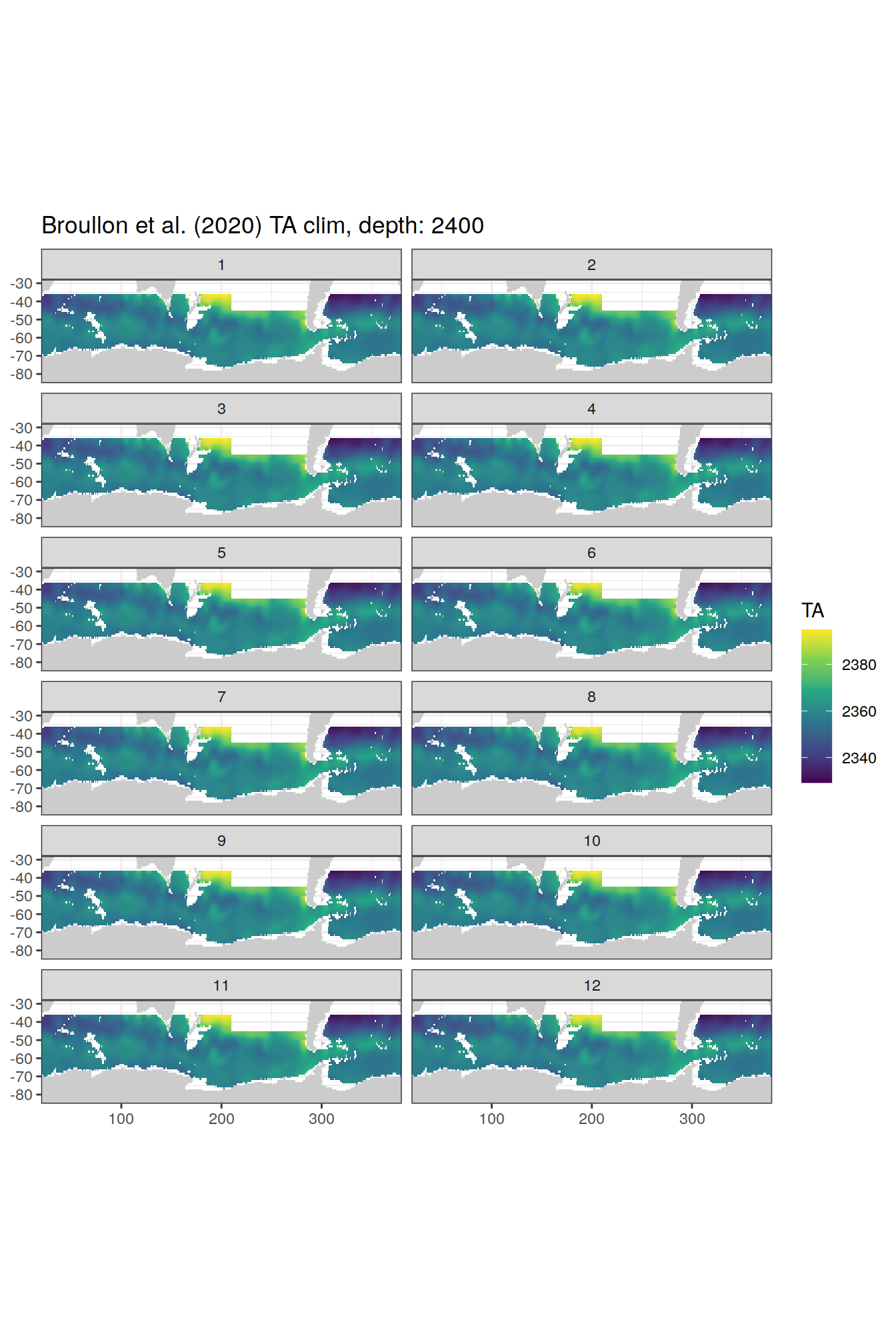
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[72]]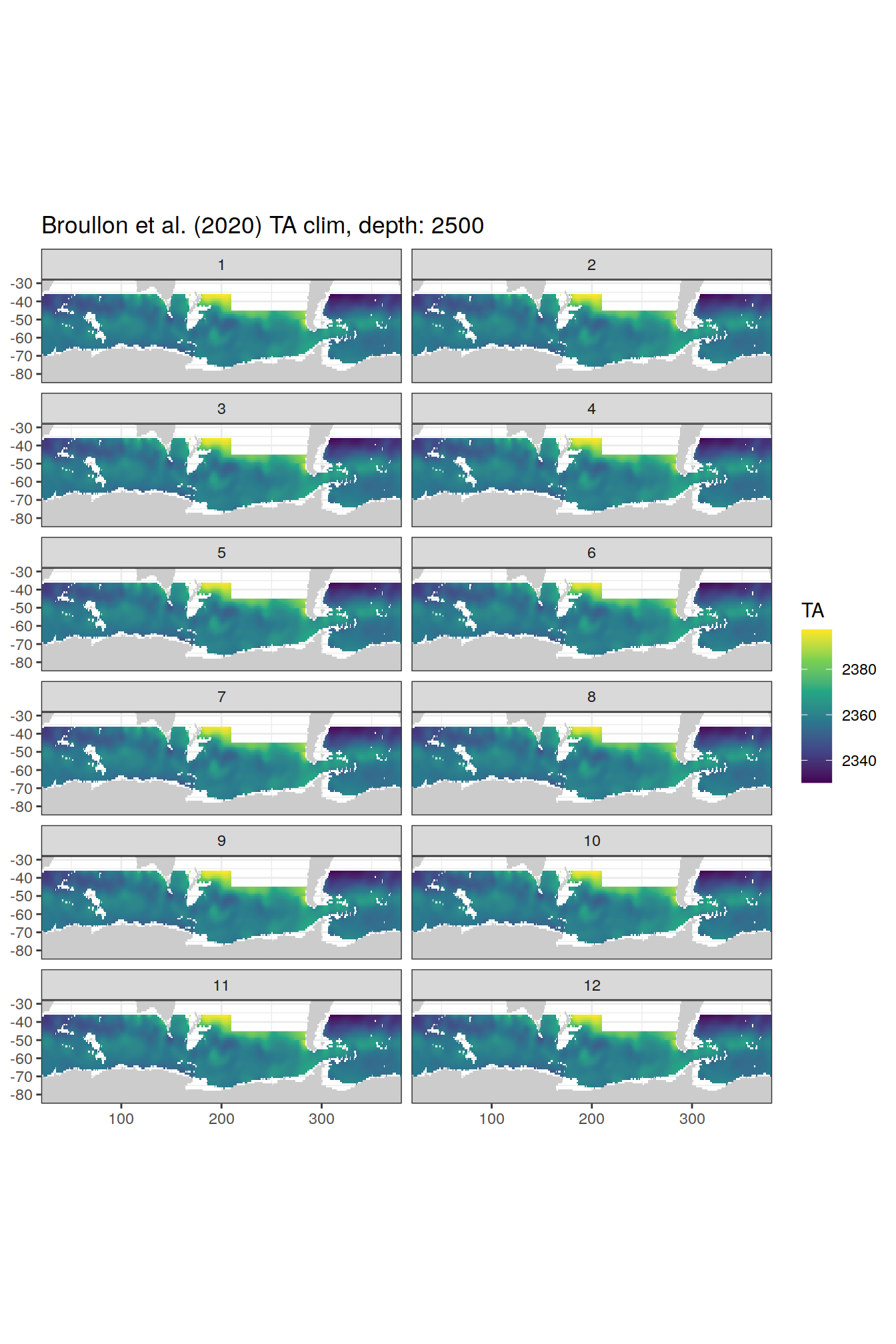
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[73]]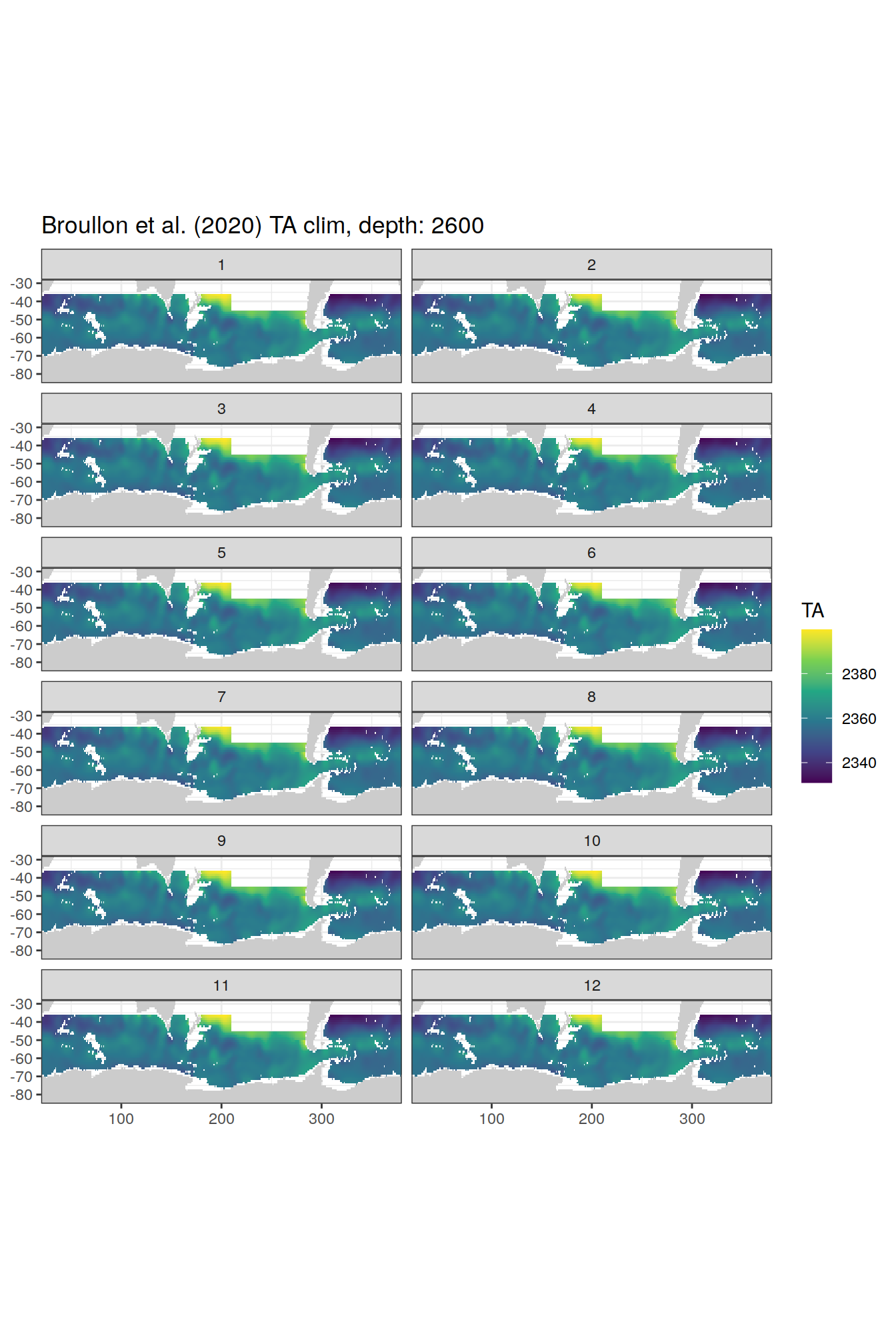
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[74]]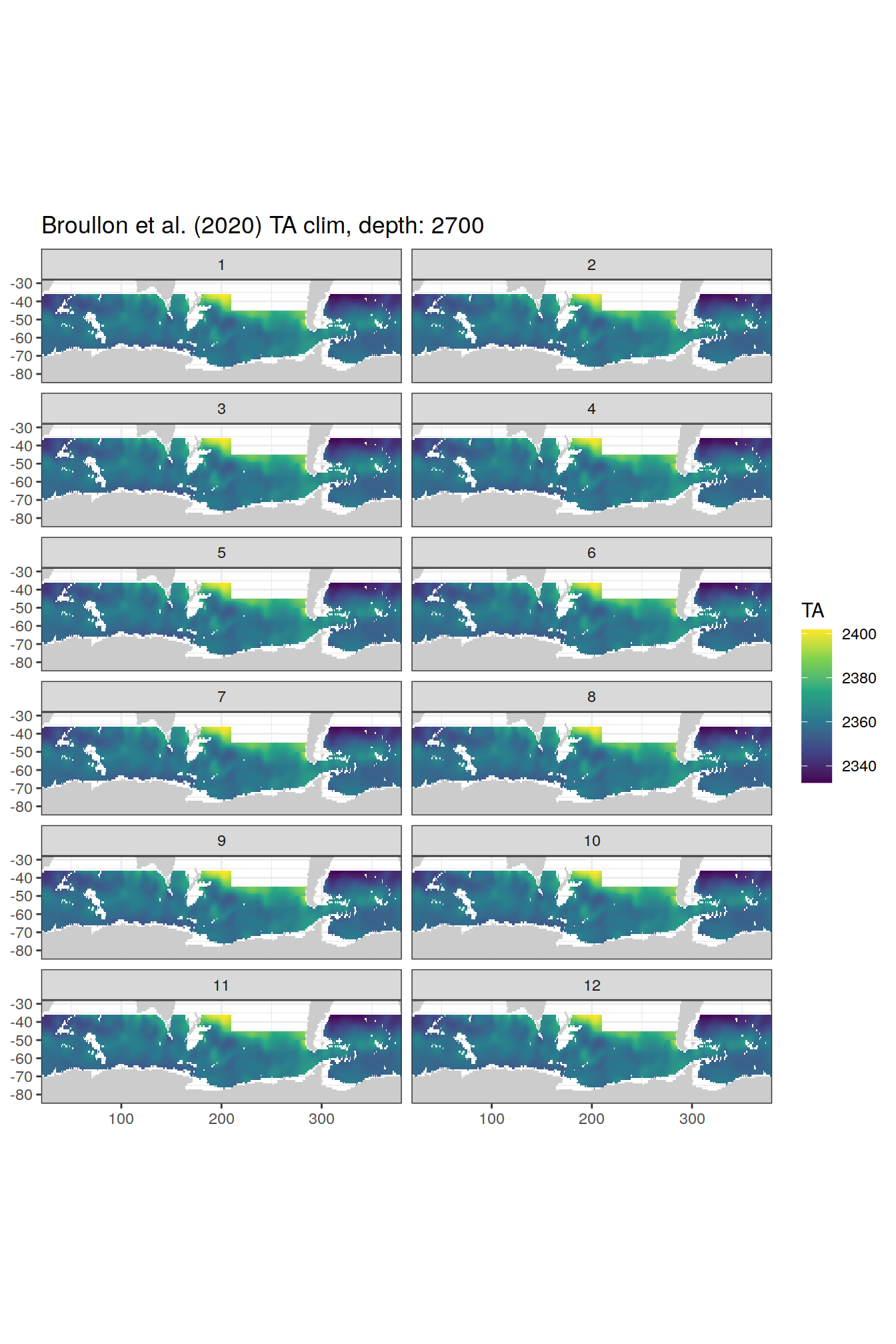
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[75]]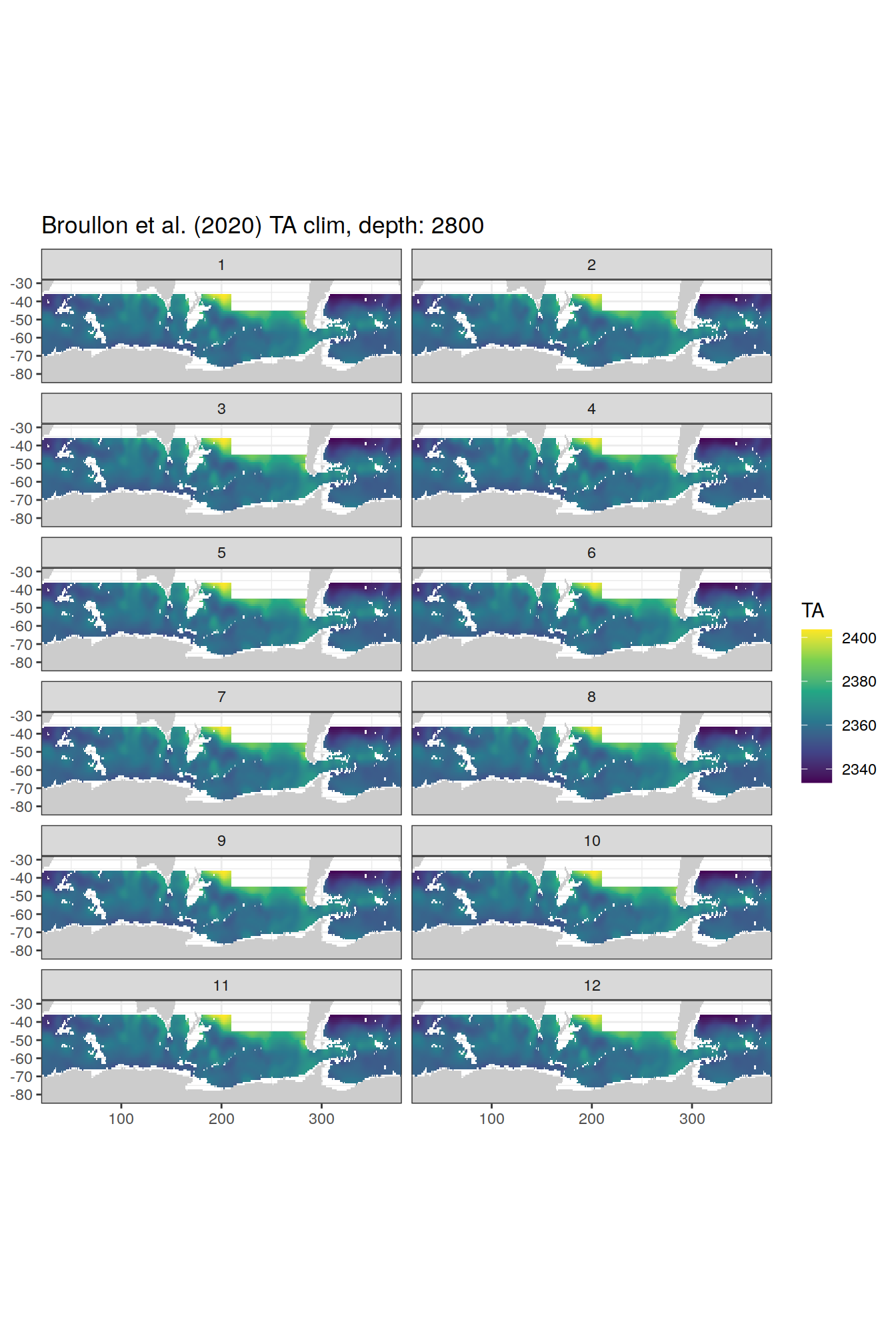
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[76]]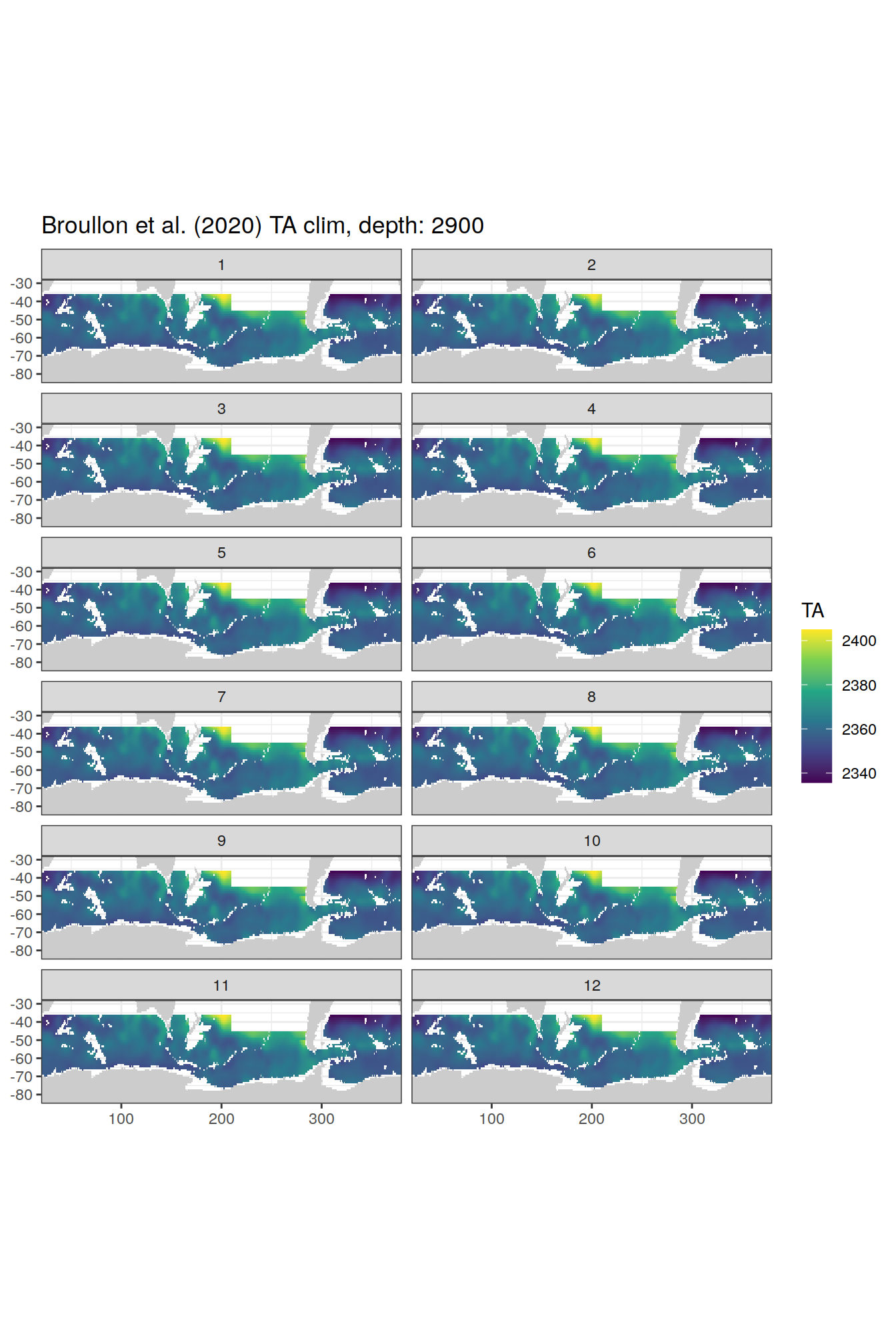
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[77]]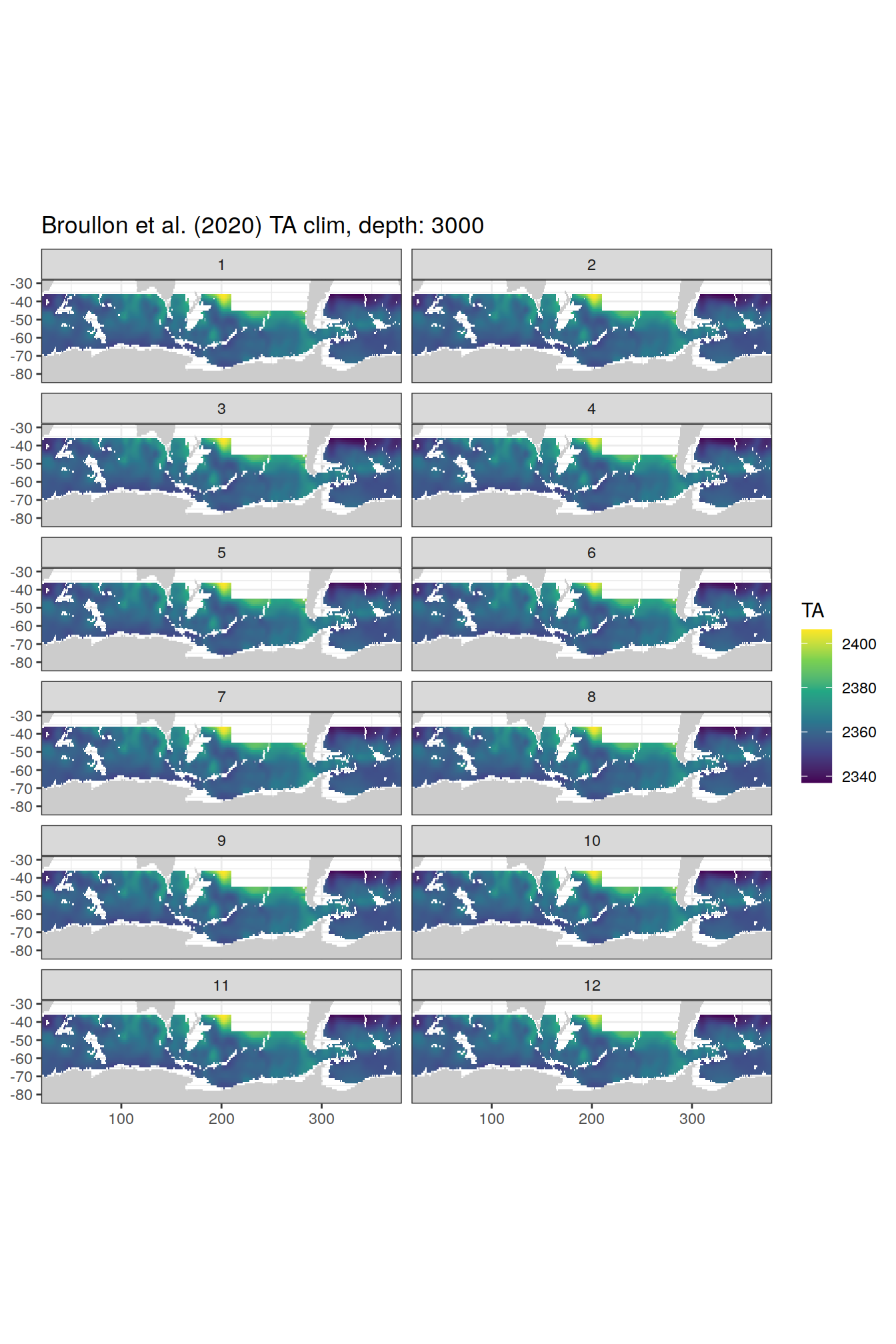
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[78]]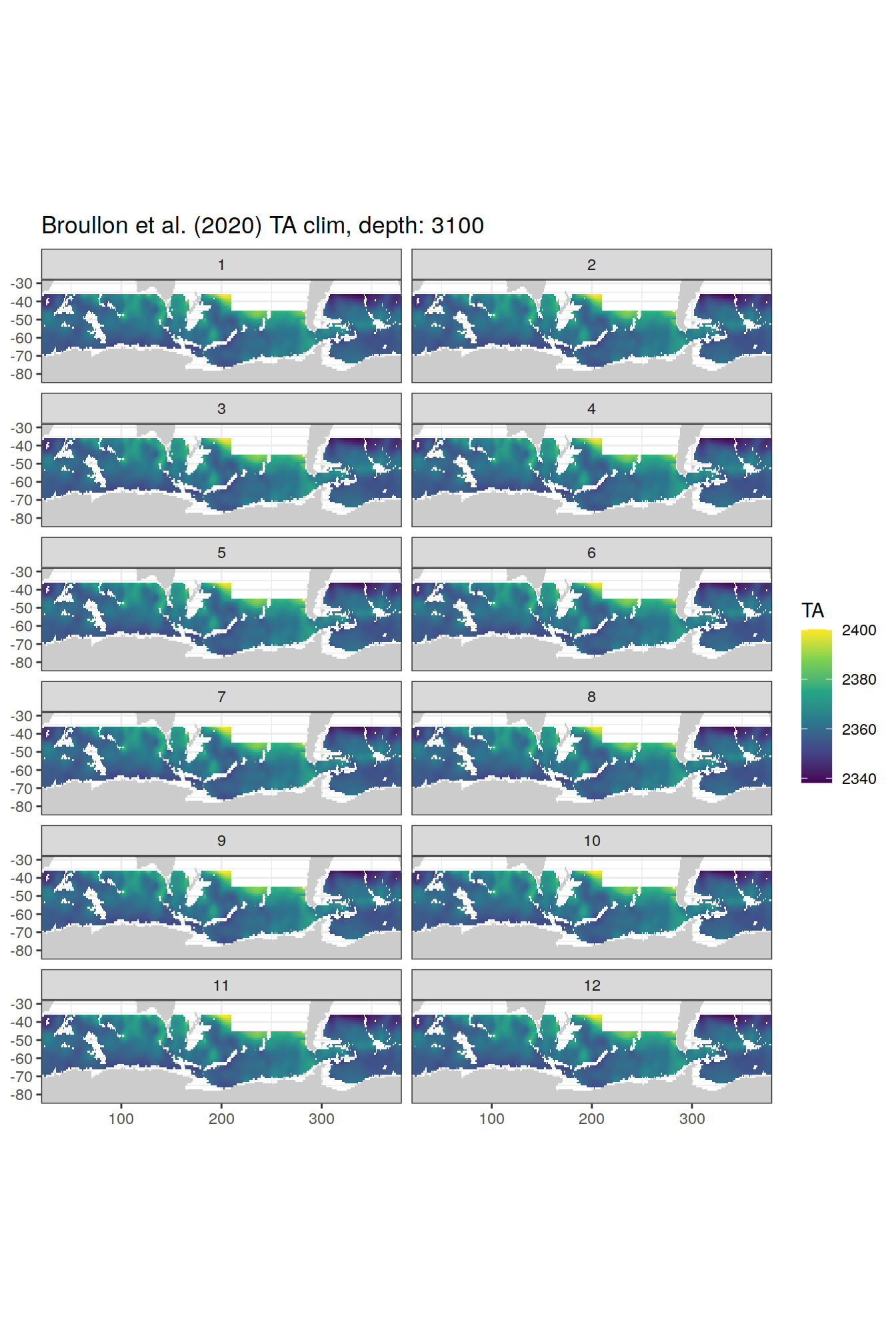
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[79]]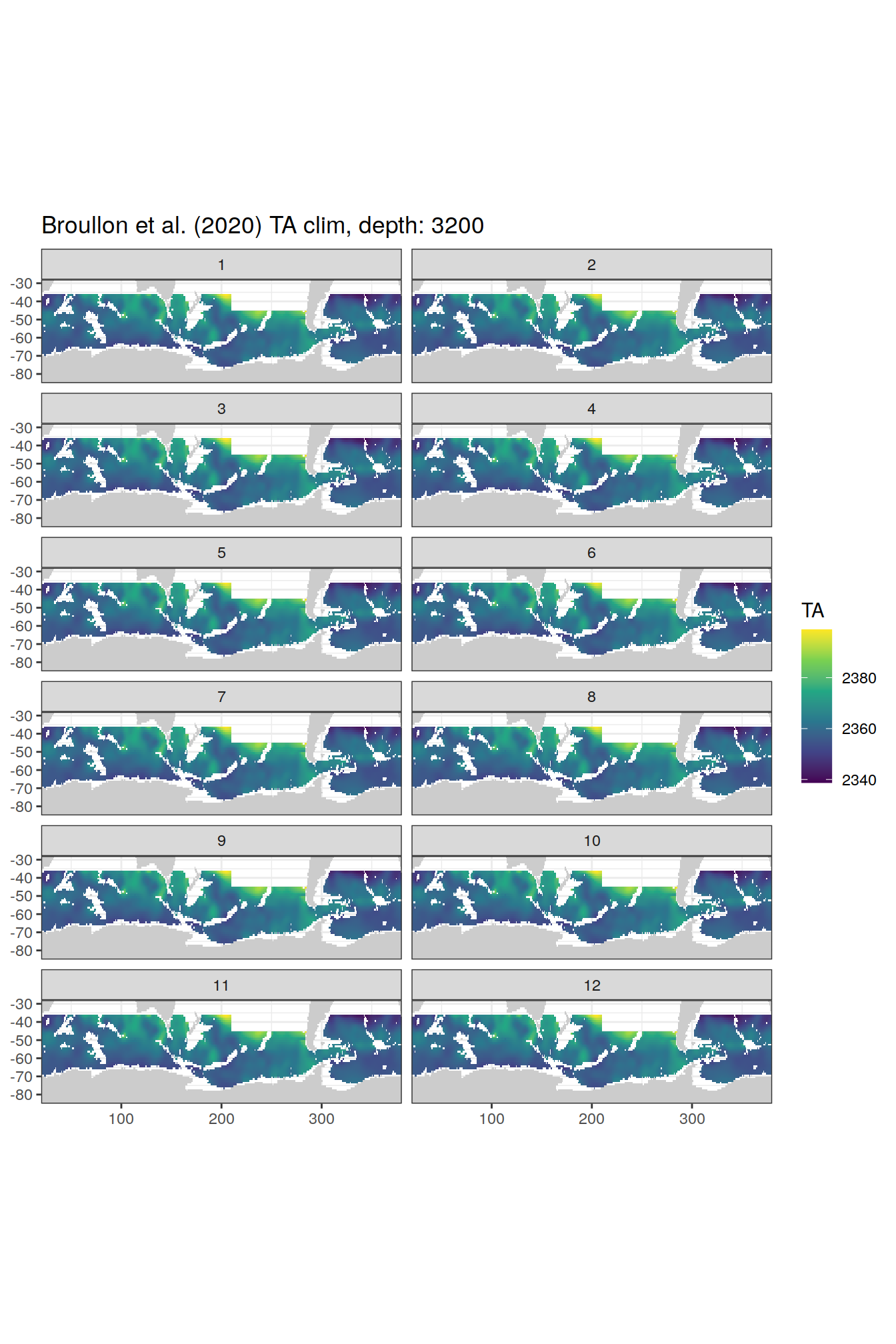
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[80]]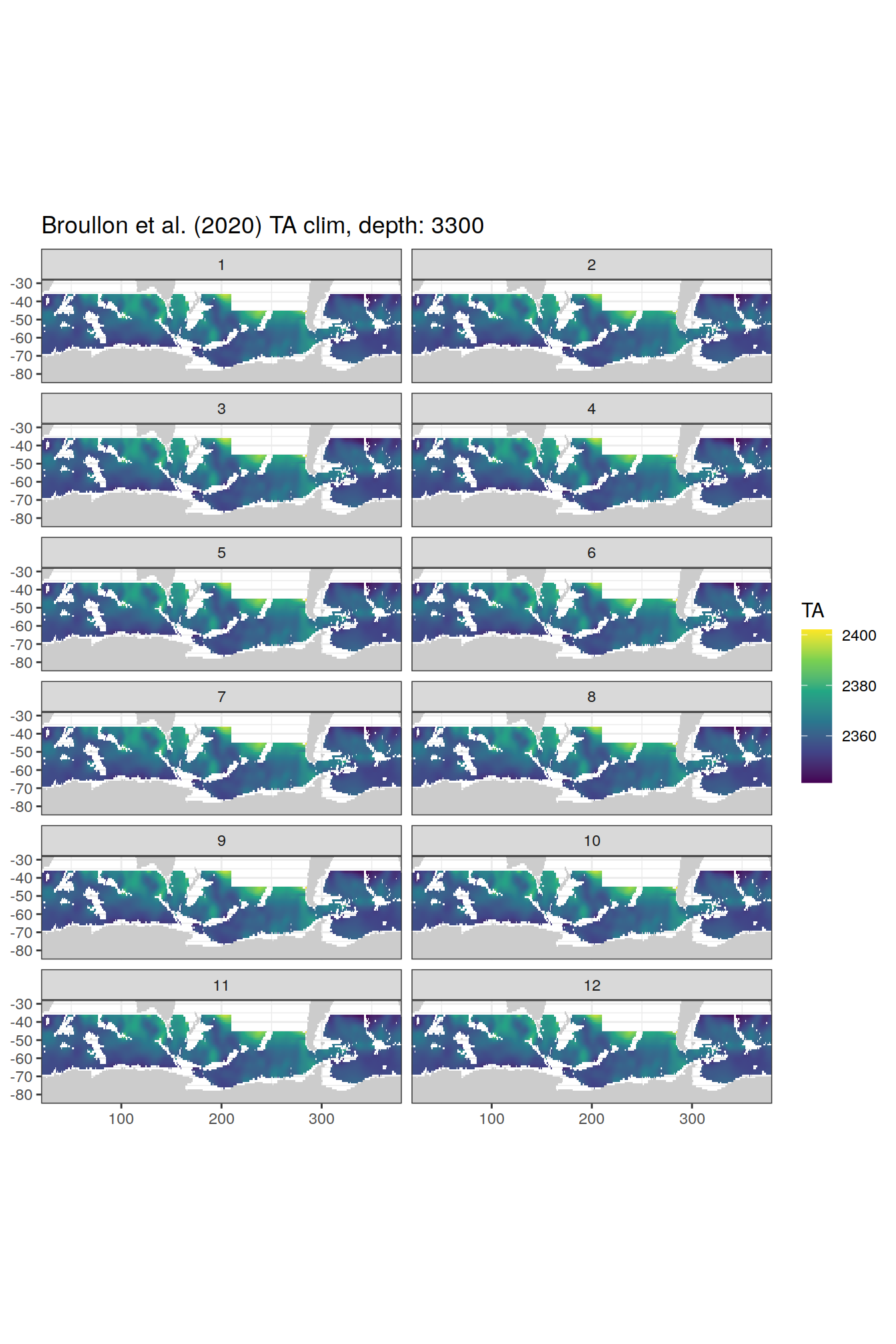
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[81]]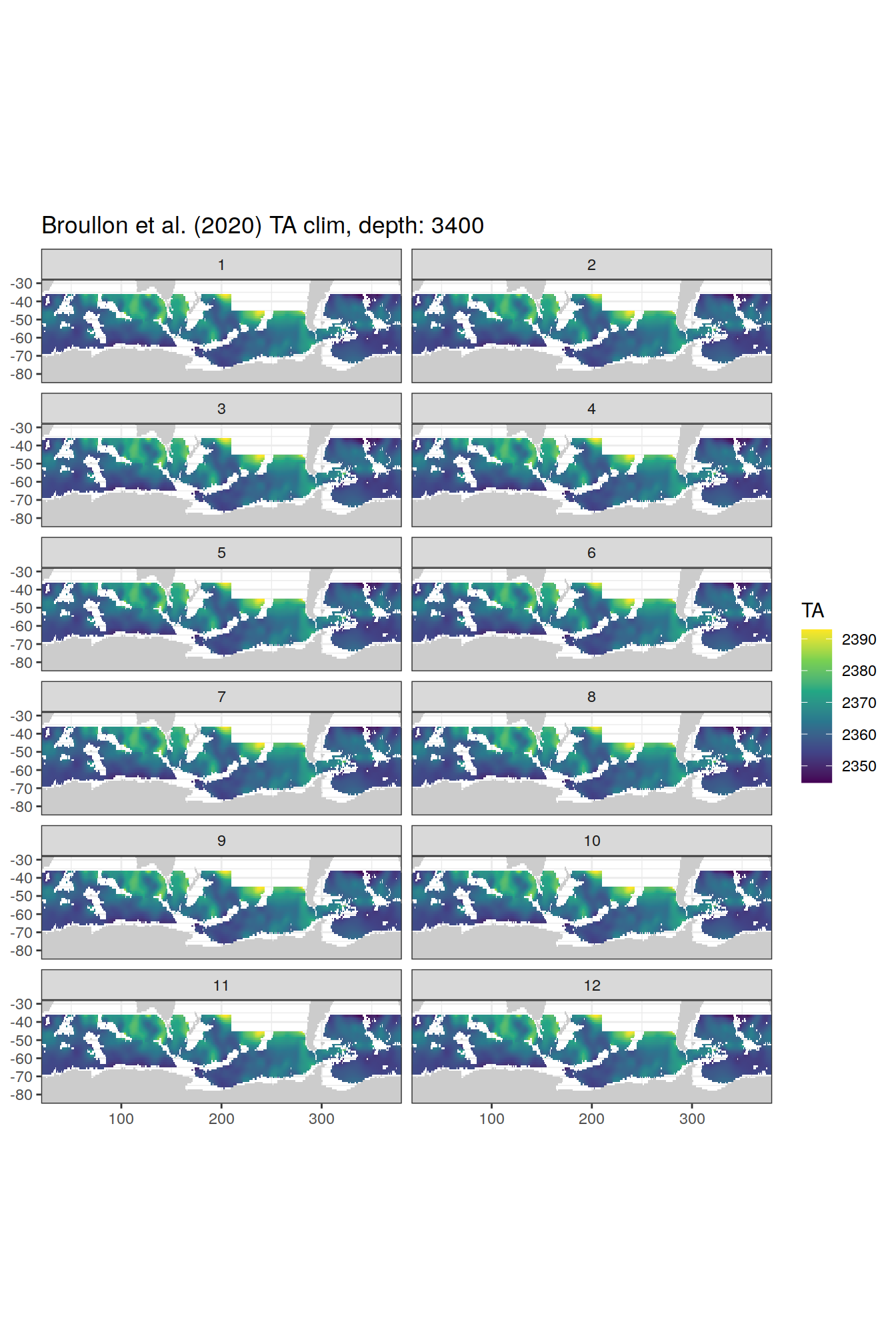
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[82]]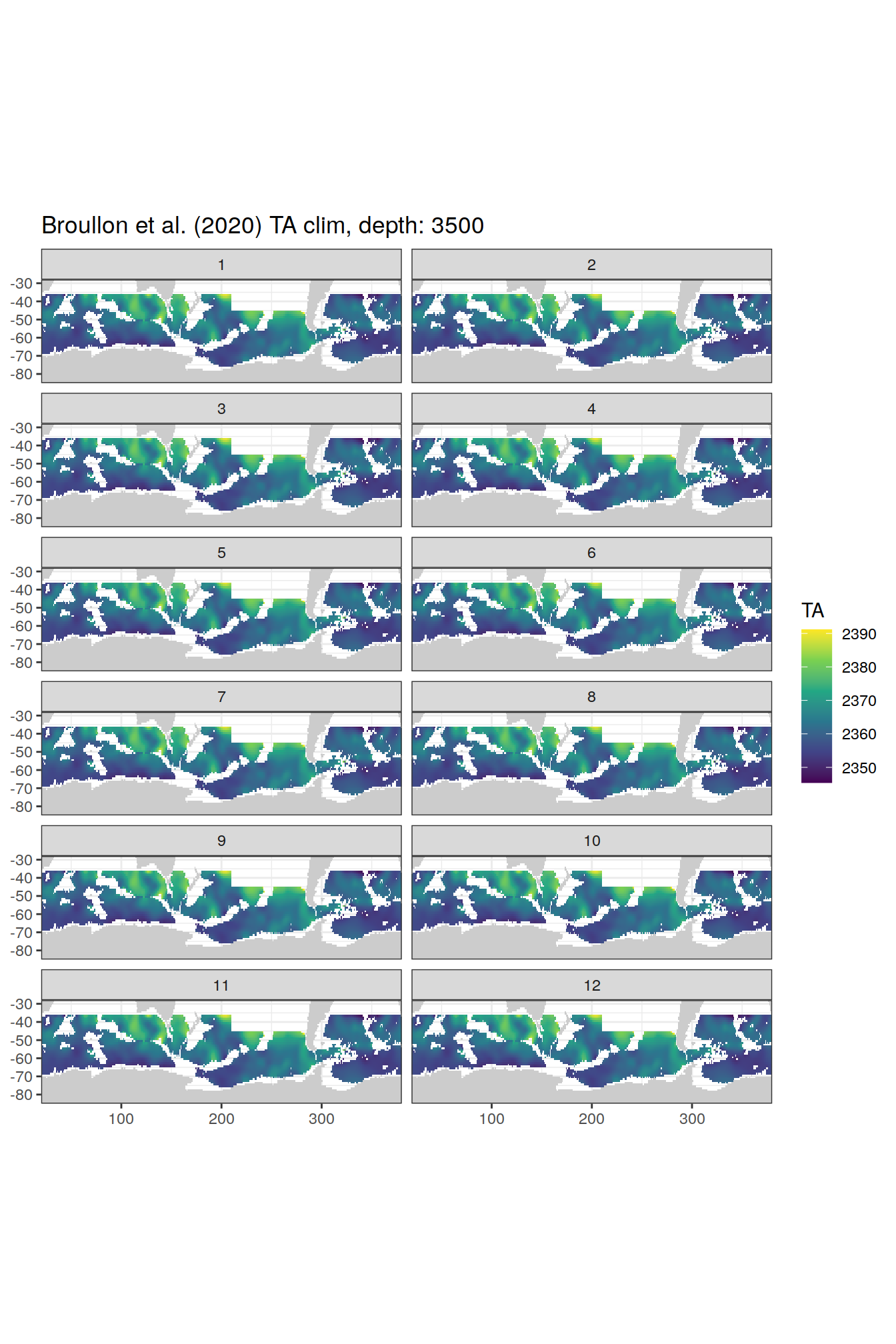
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[83]]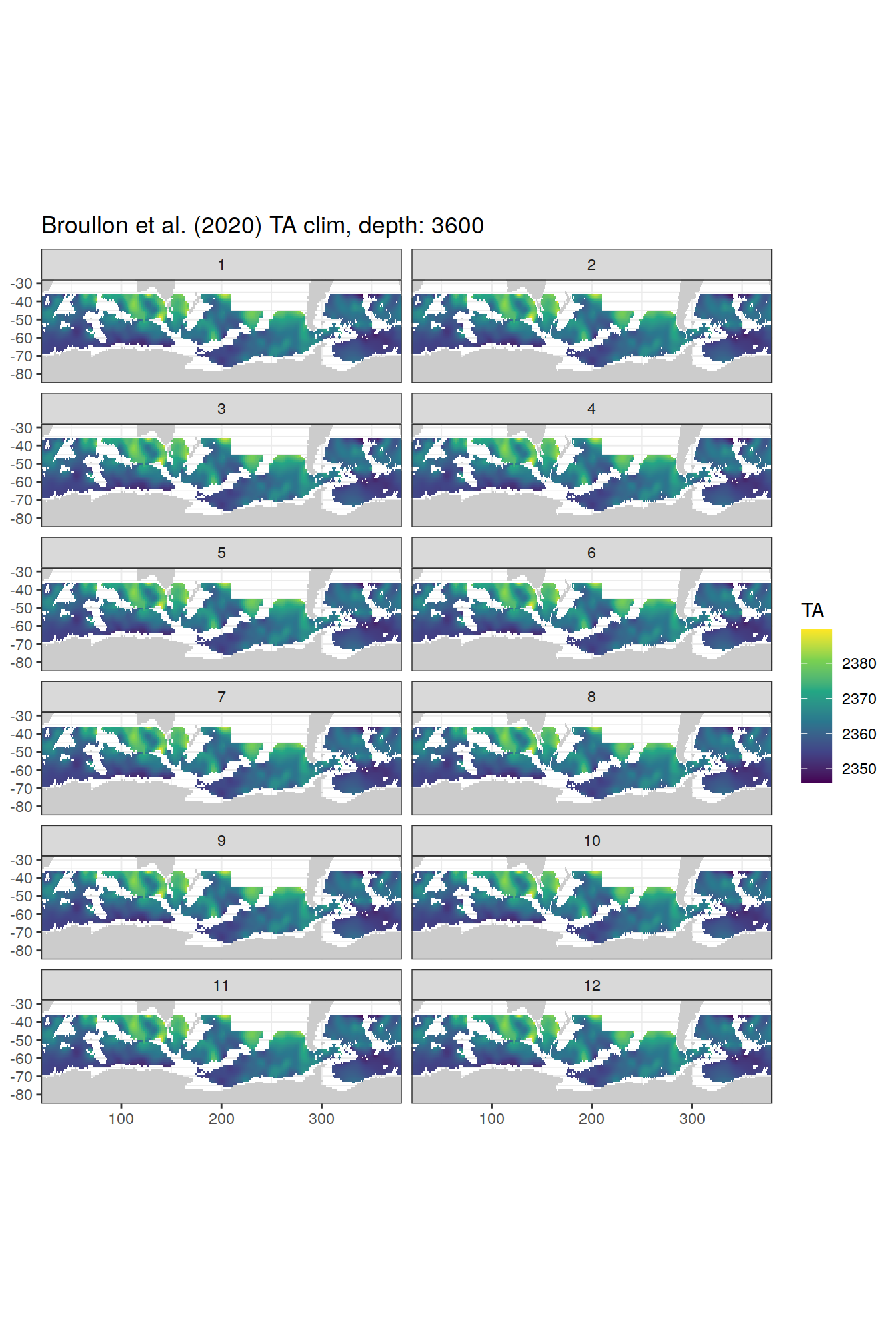
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[84]]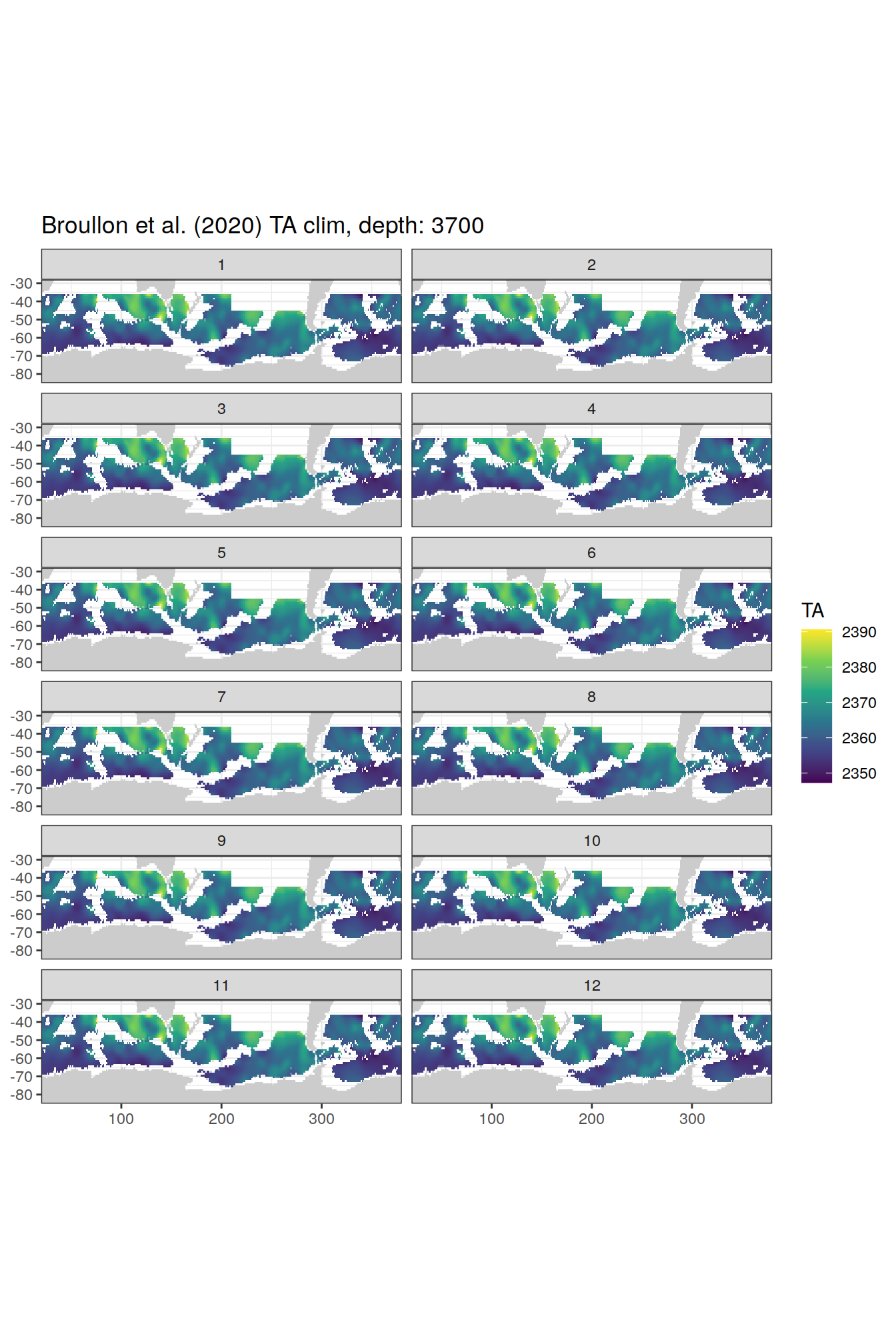
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[85]]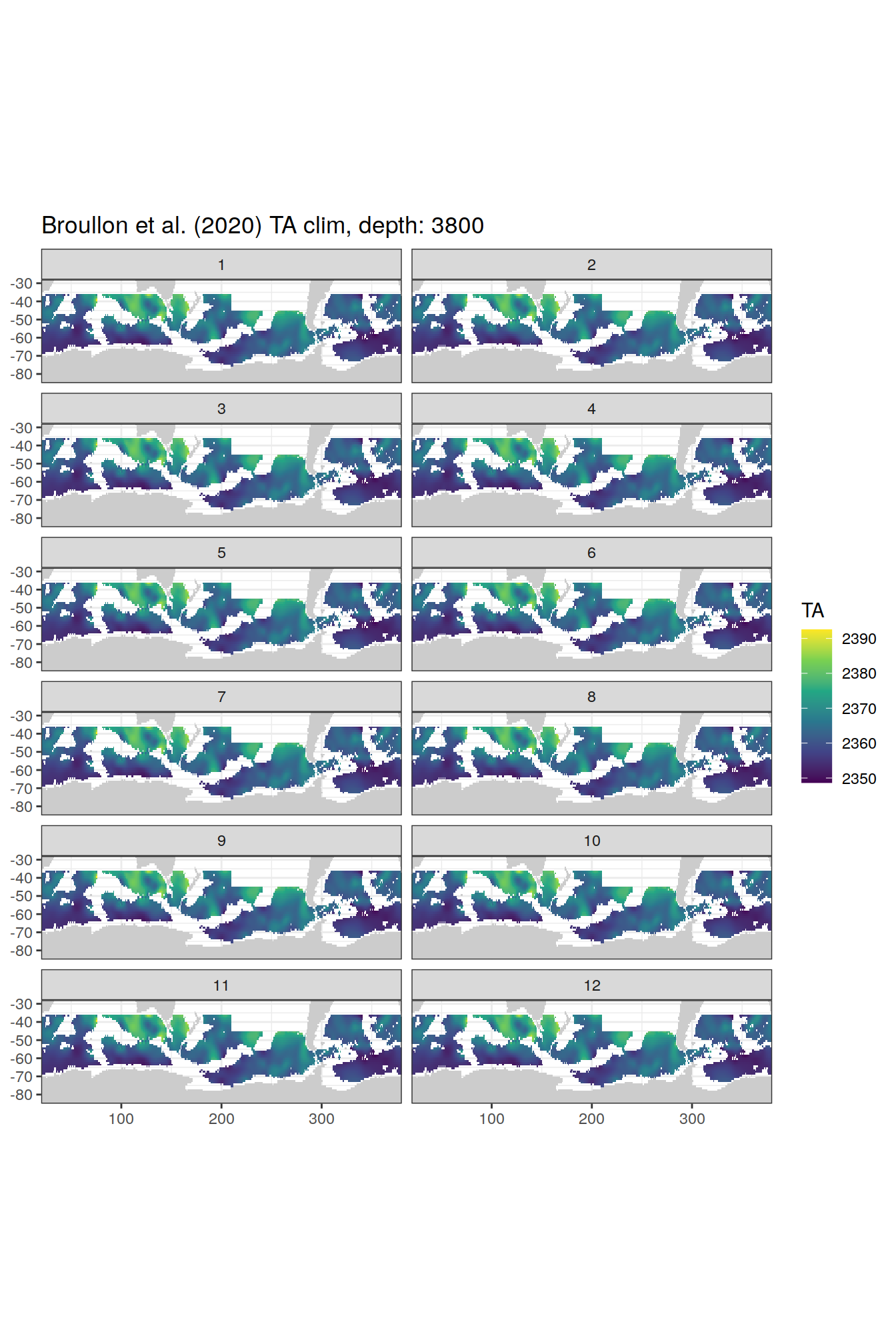
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[86]]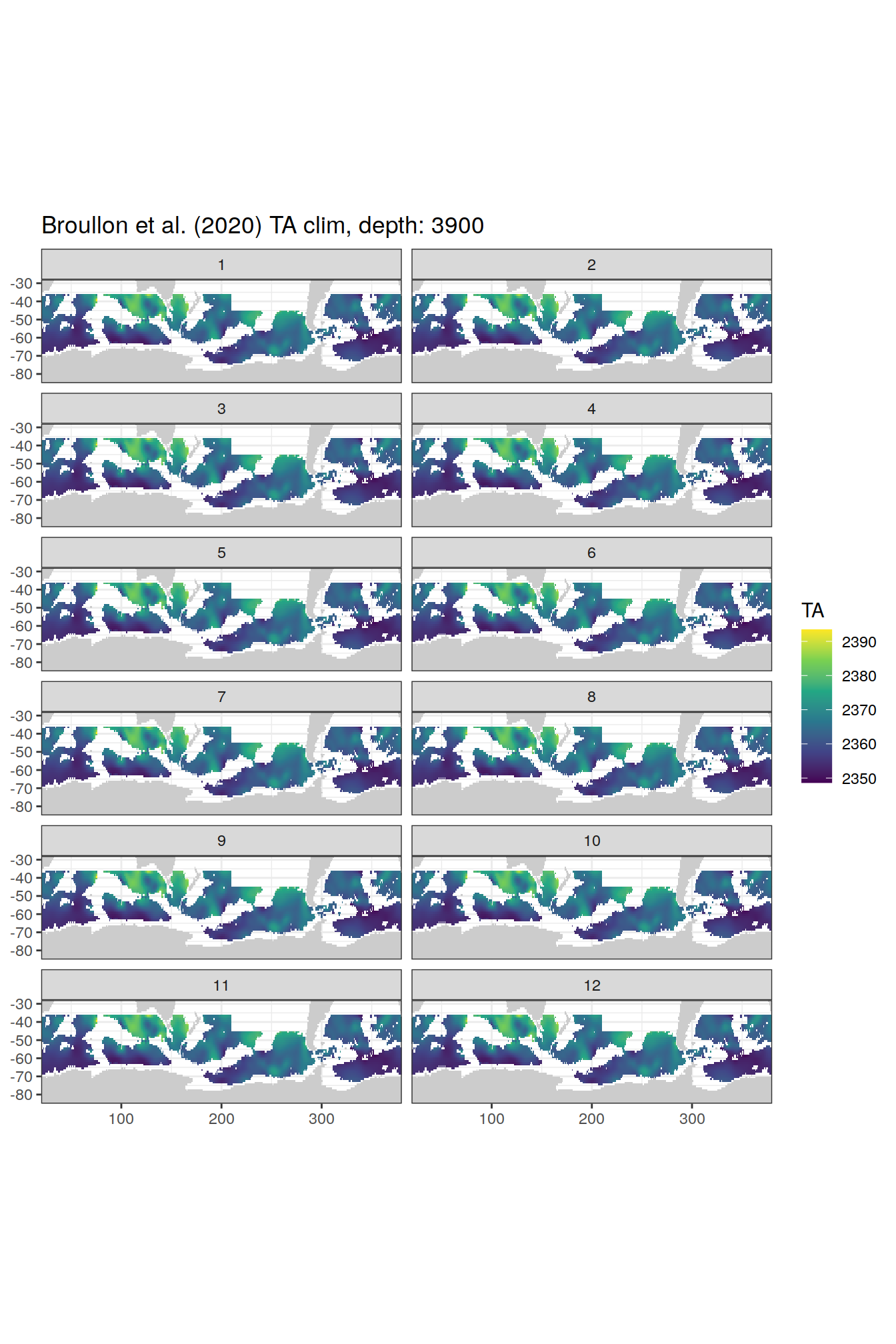
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[87]]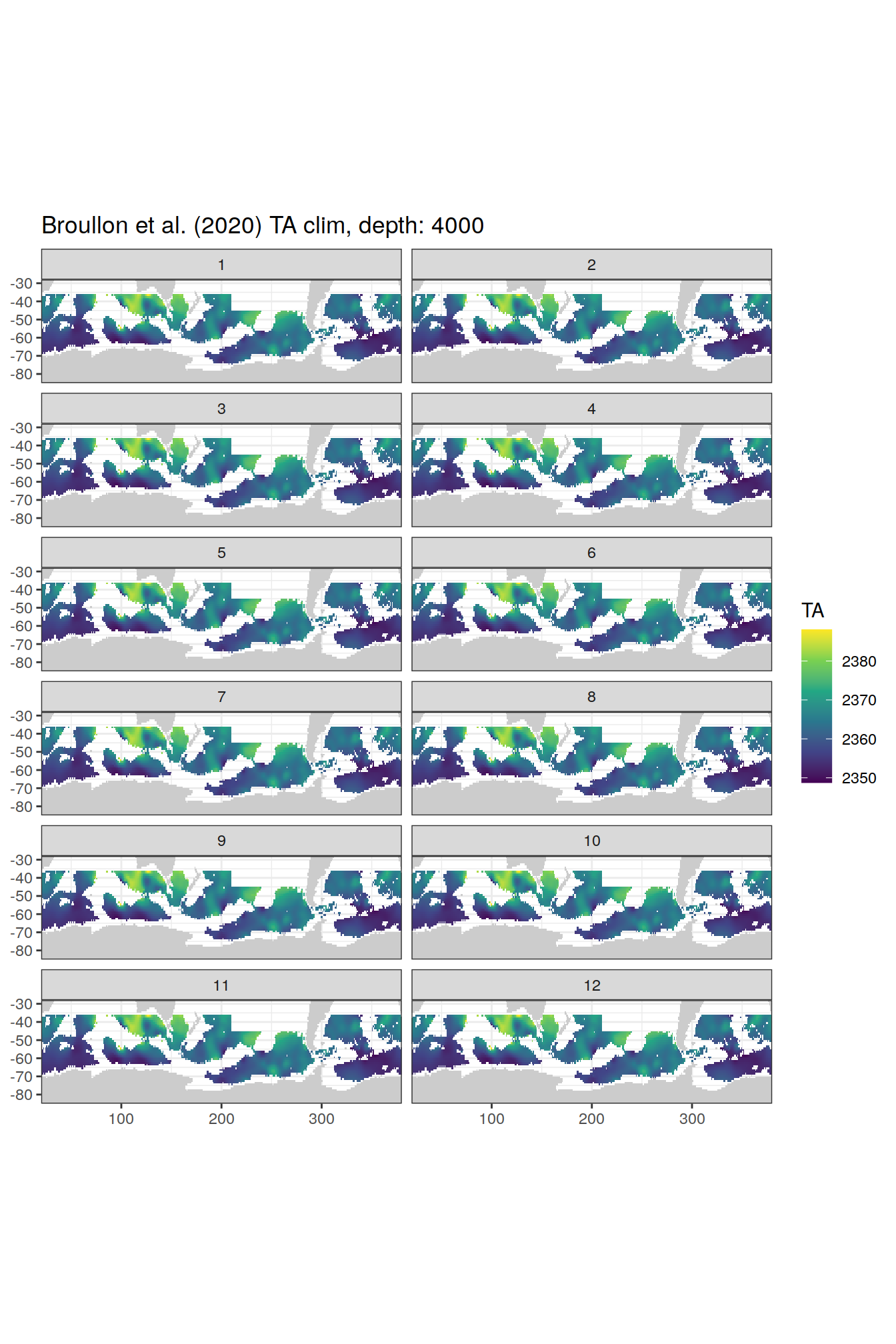
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[88]]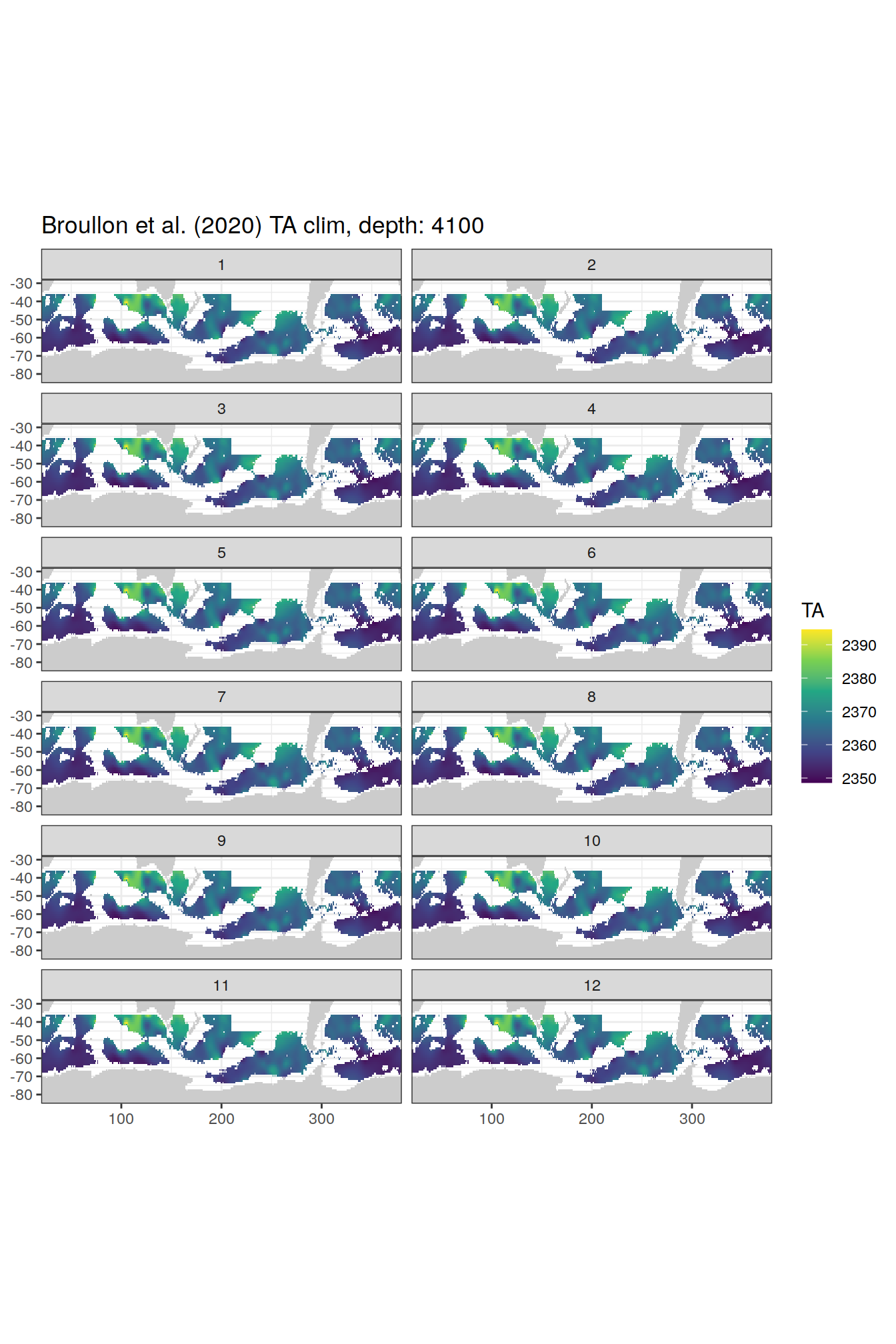
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[89]]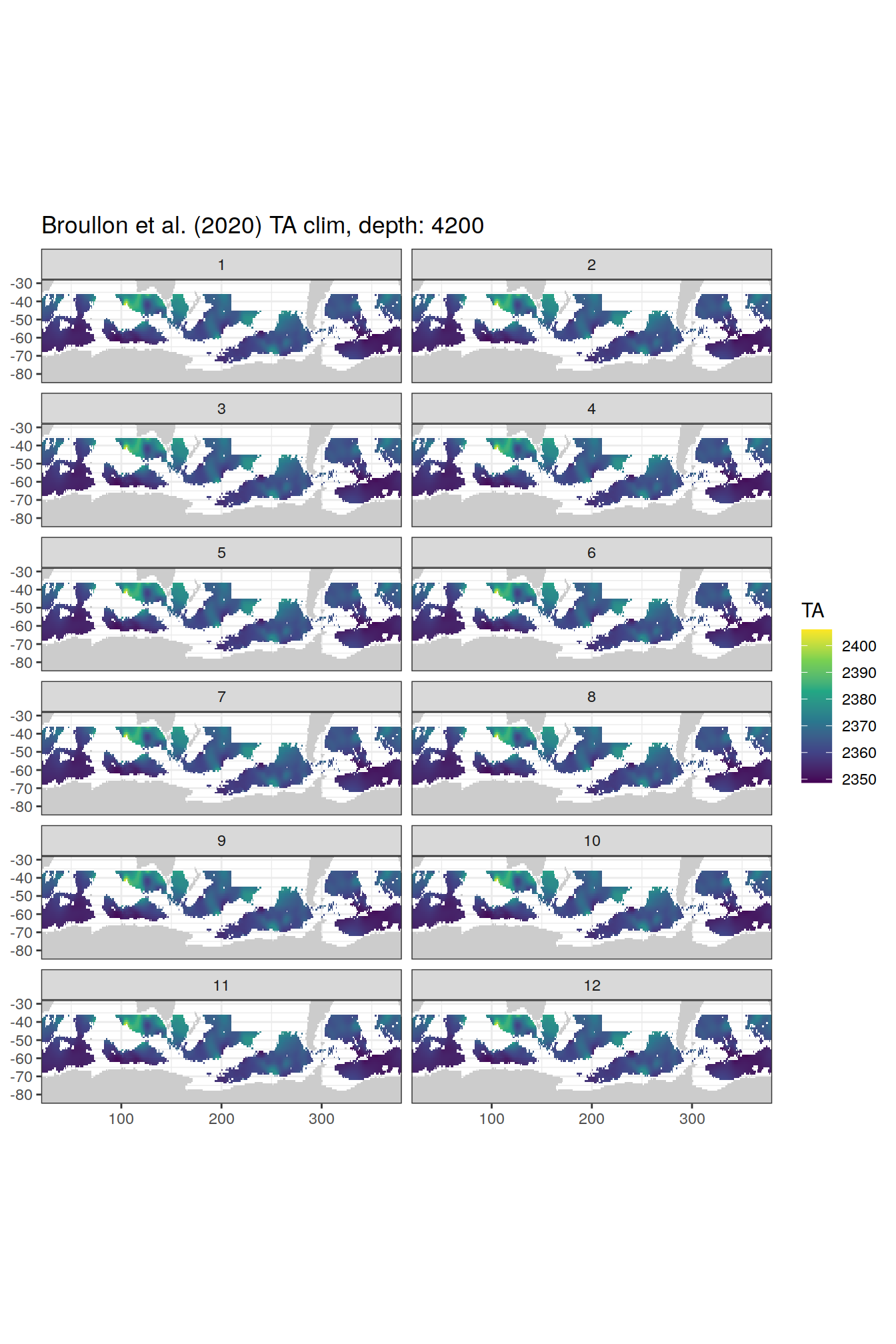
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[90]]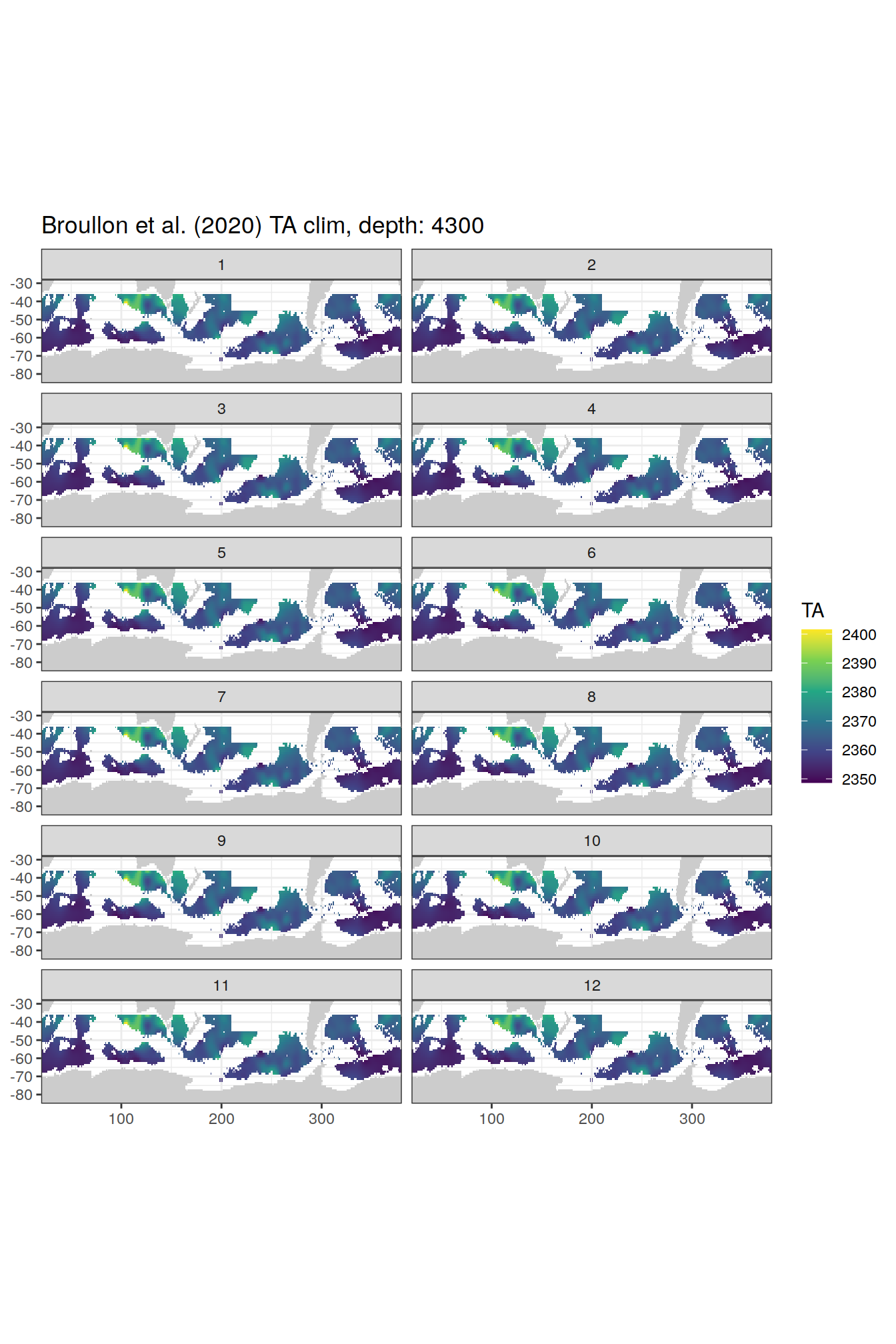
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[91]]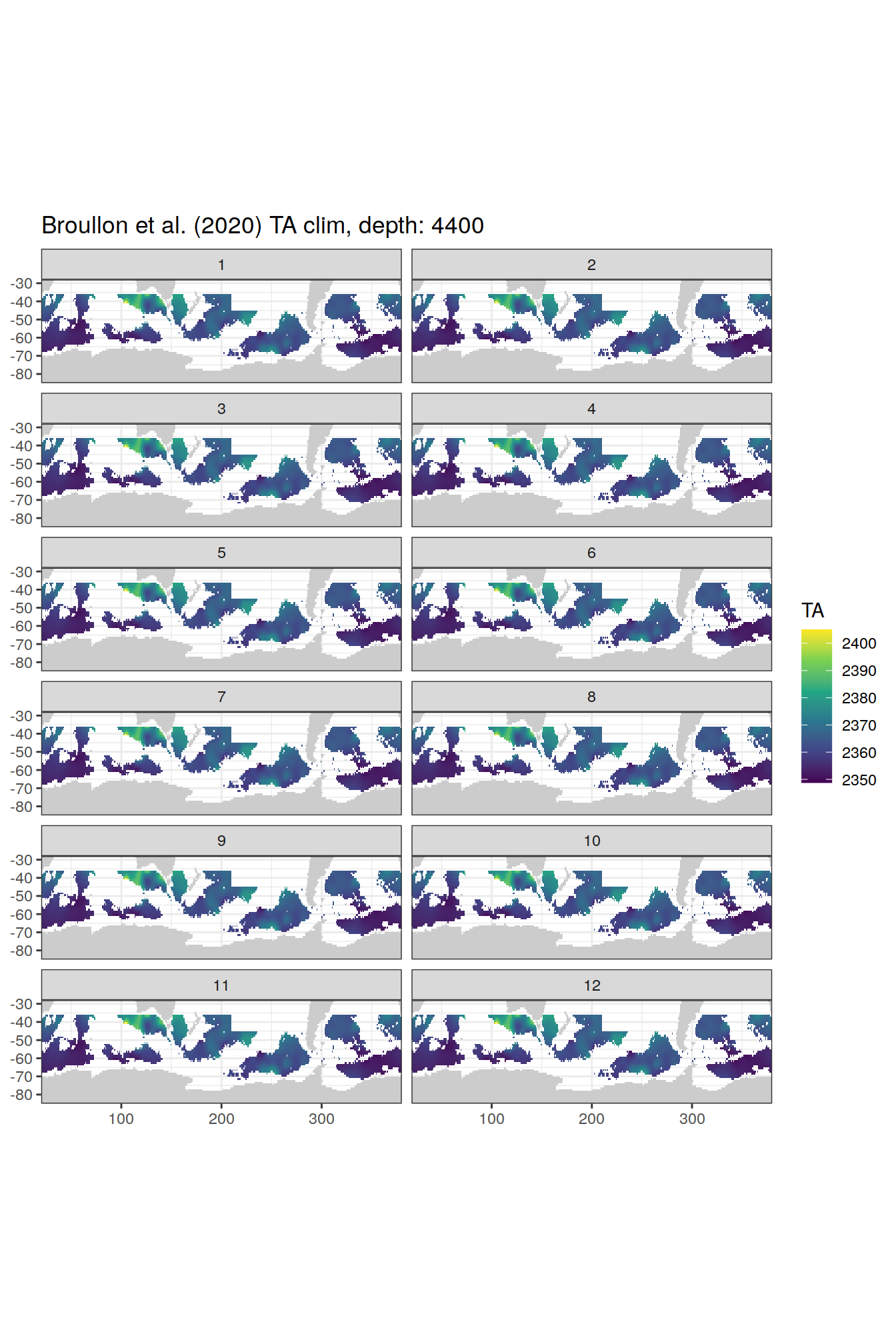
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[92]]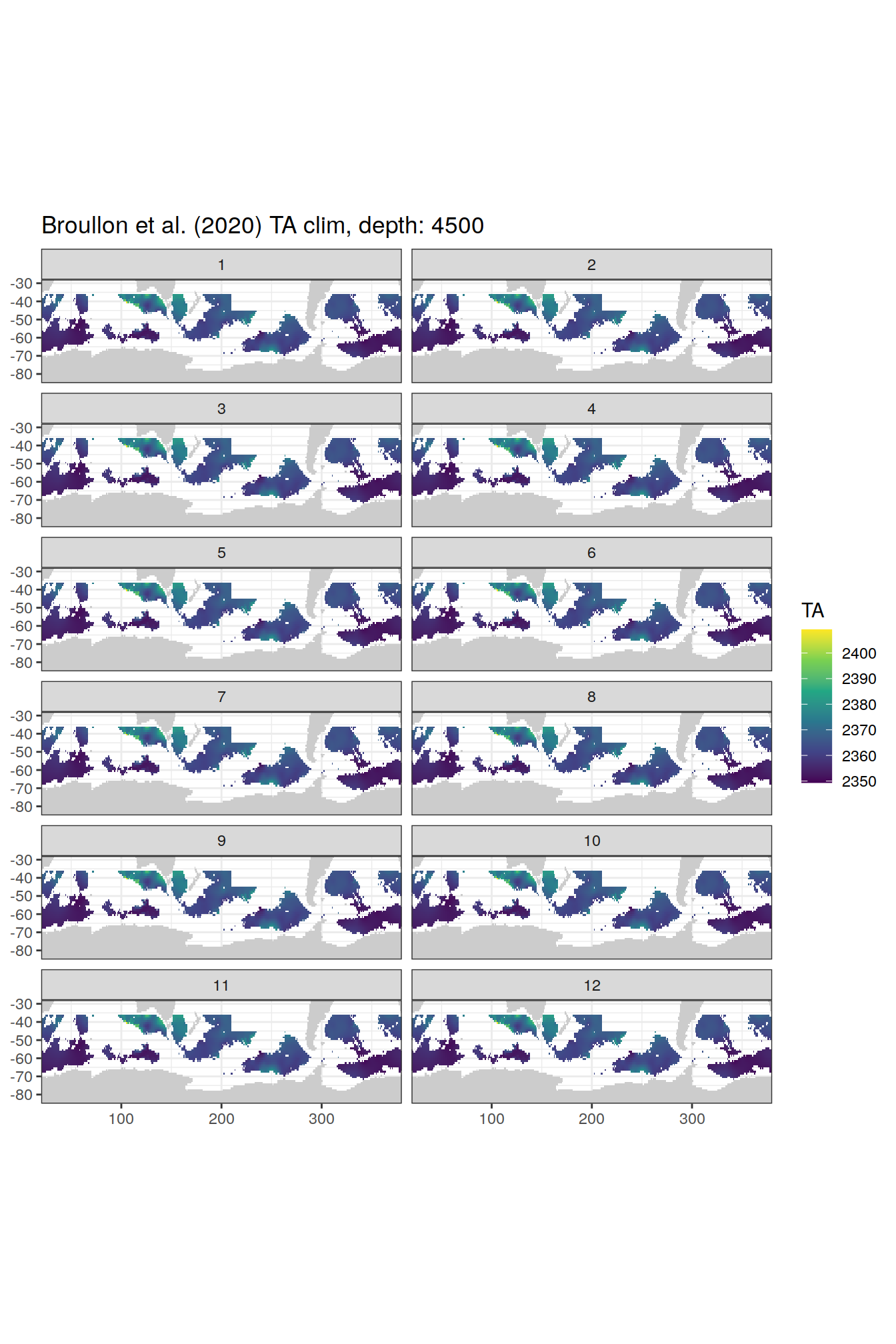
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[93]]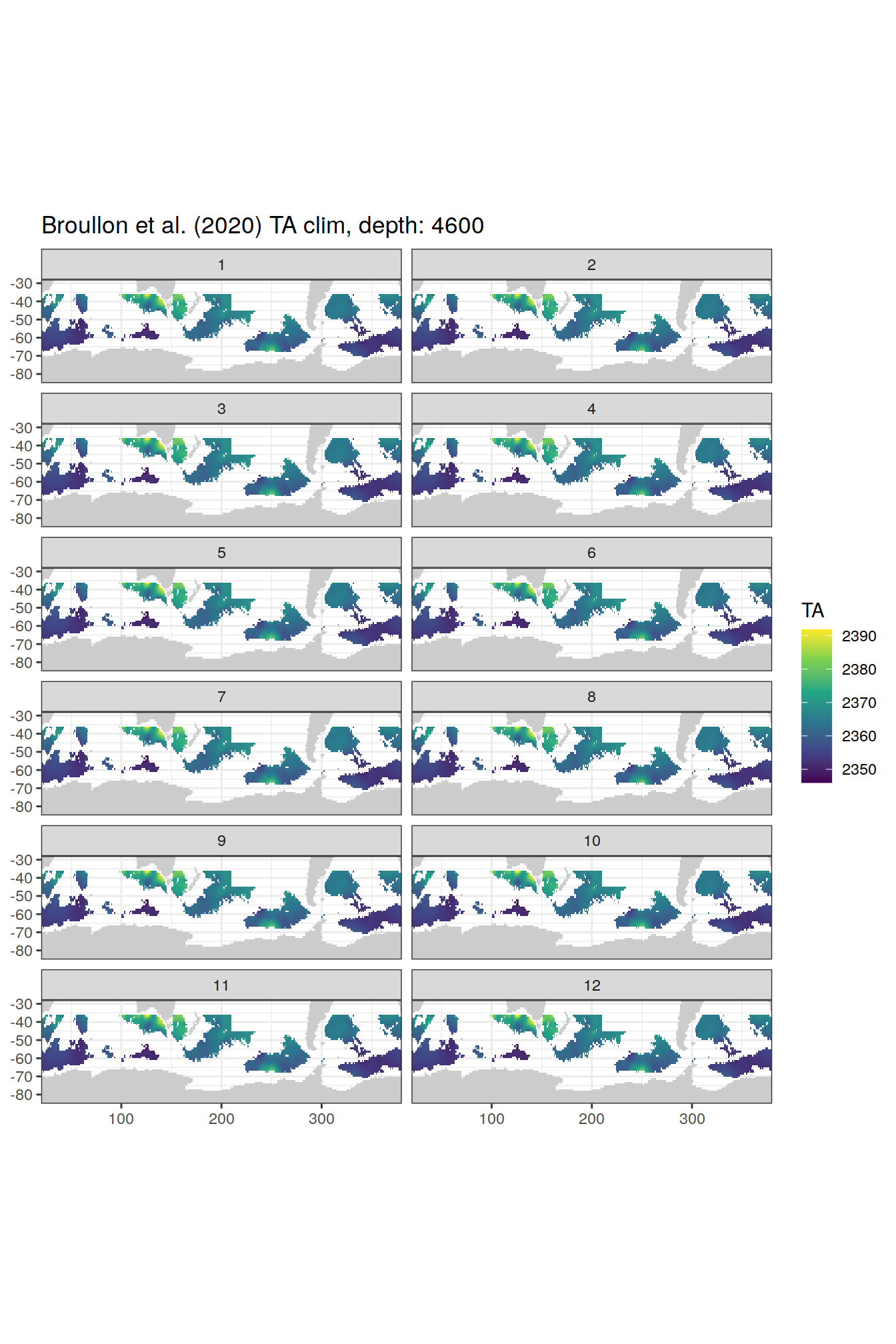
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[94]]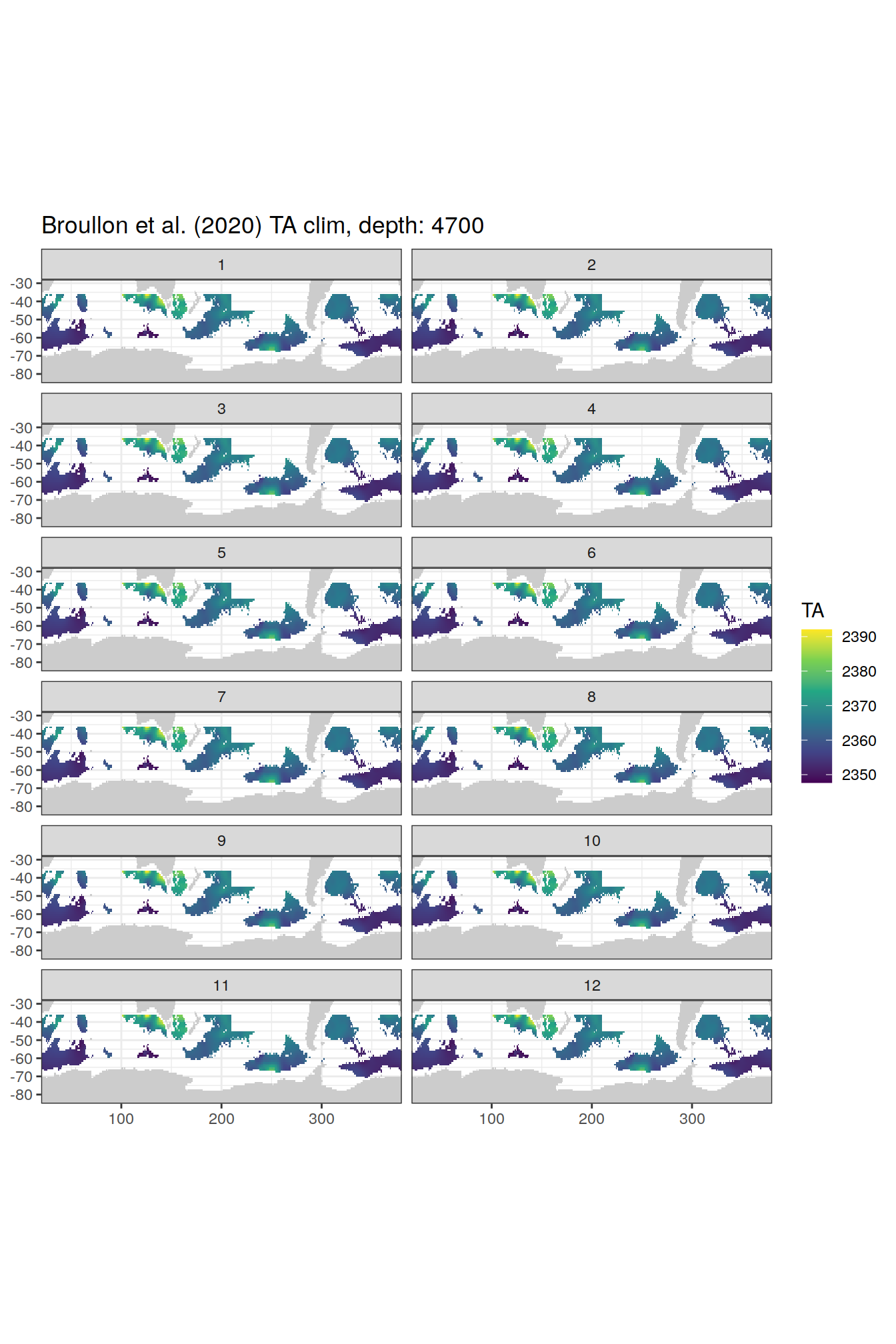
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[95]]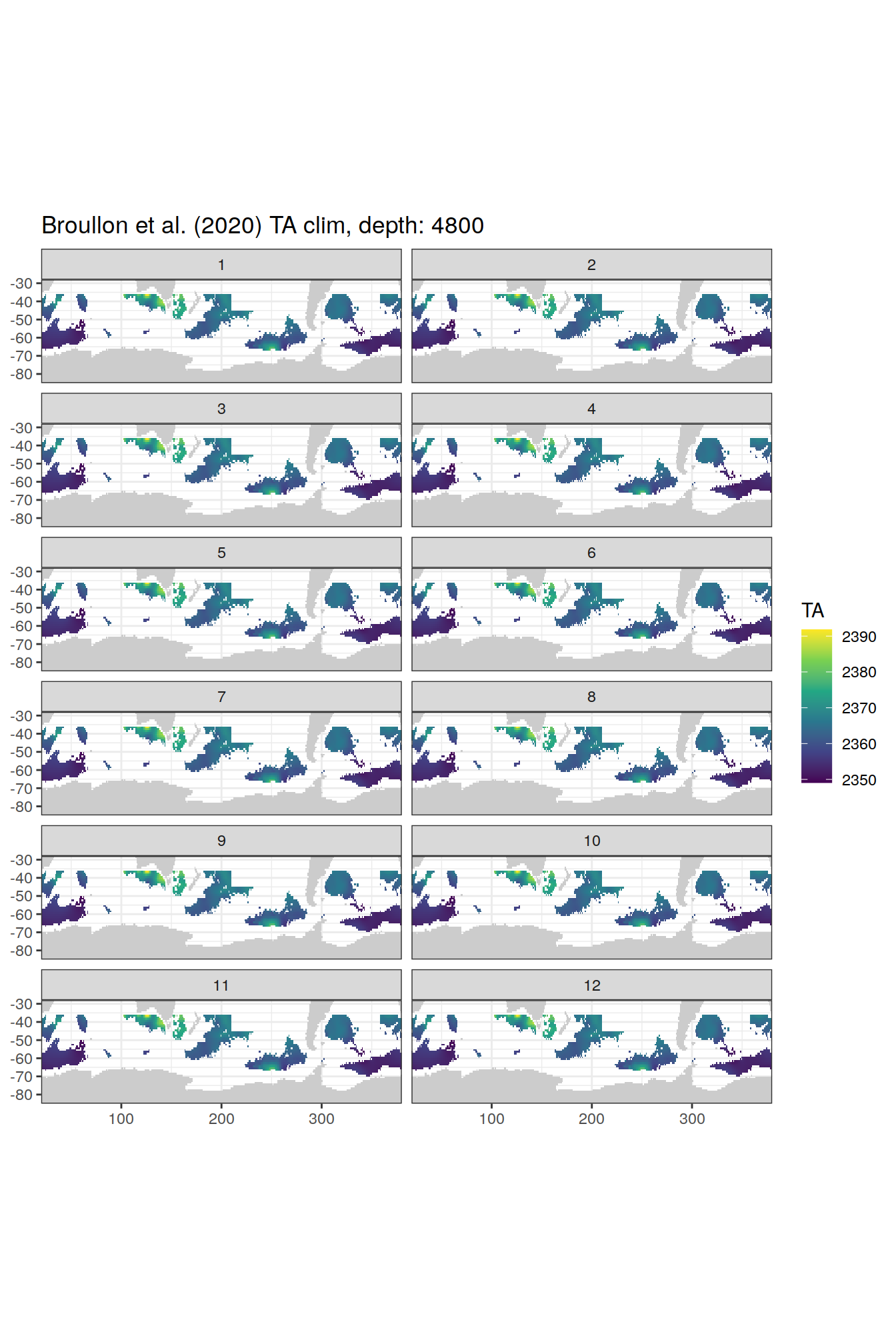
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[96]]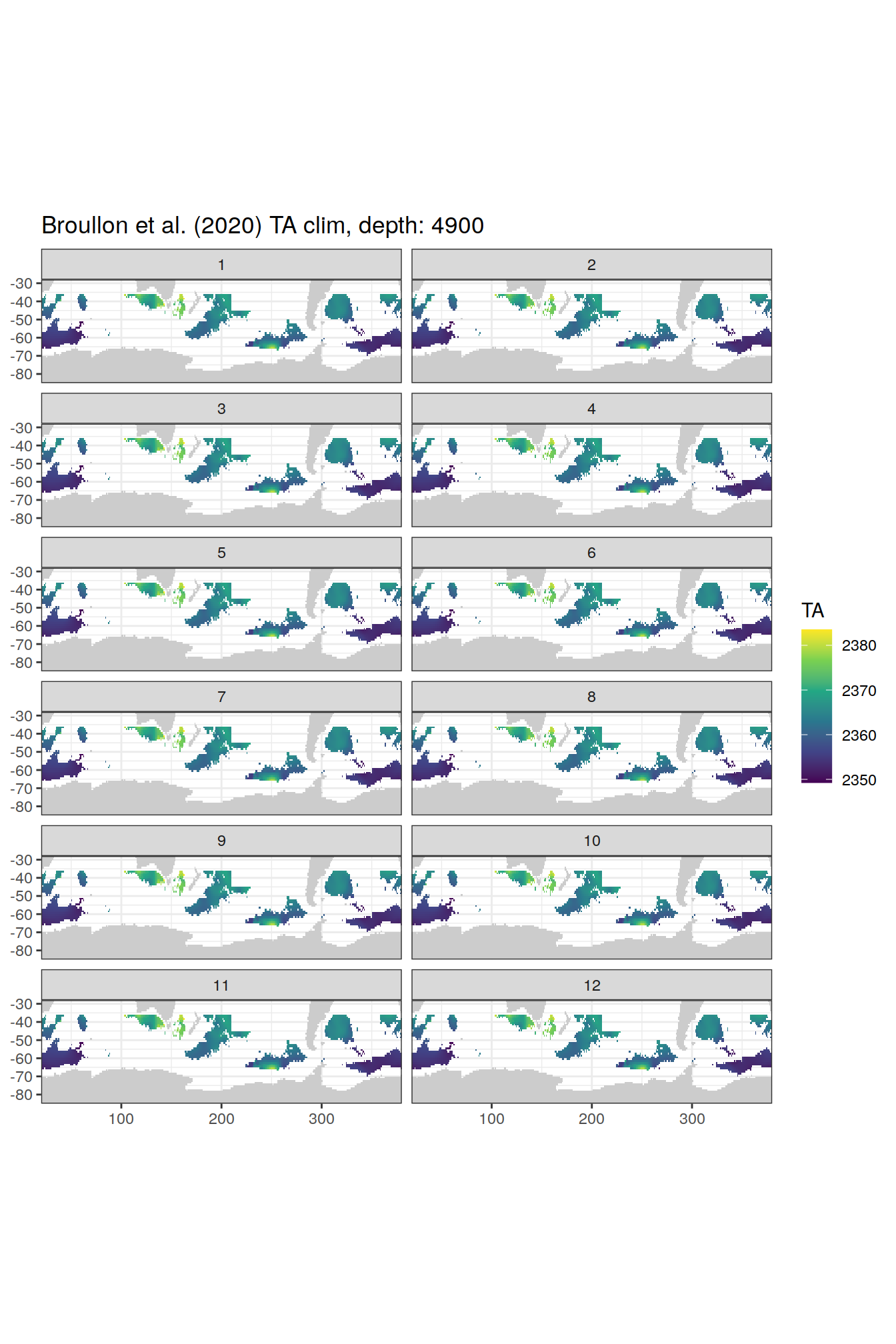
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[97]]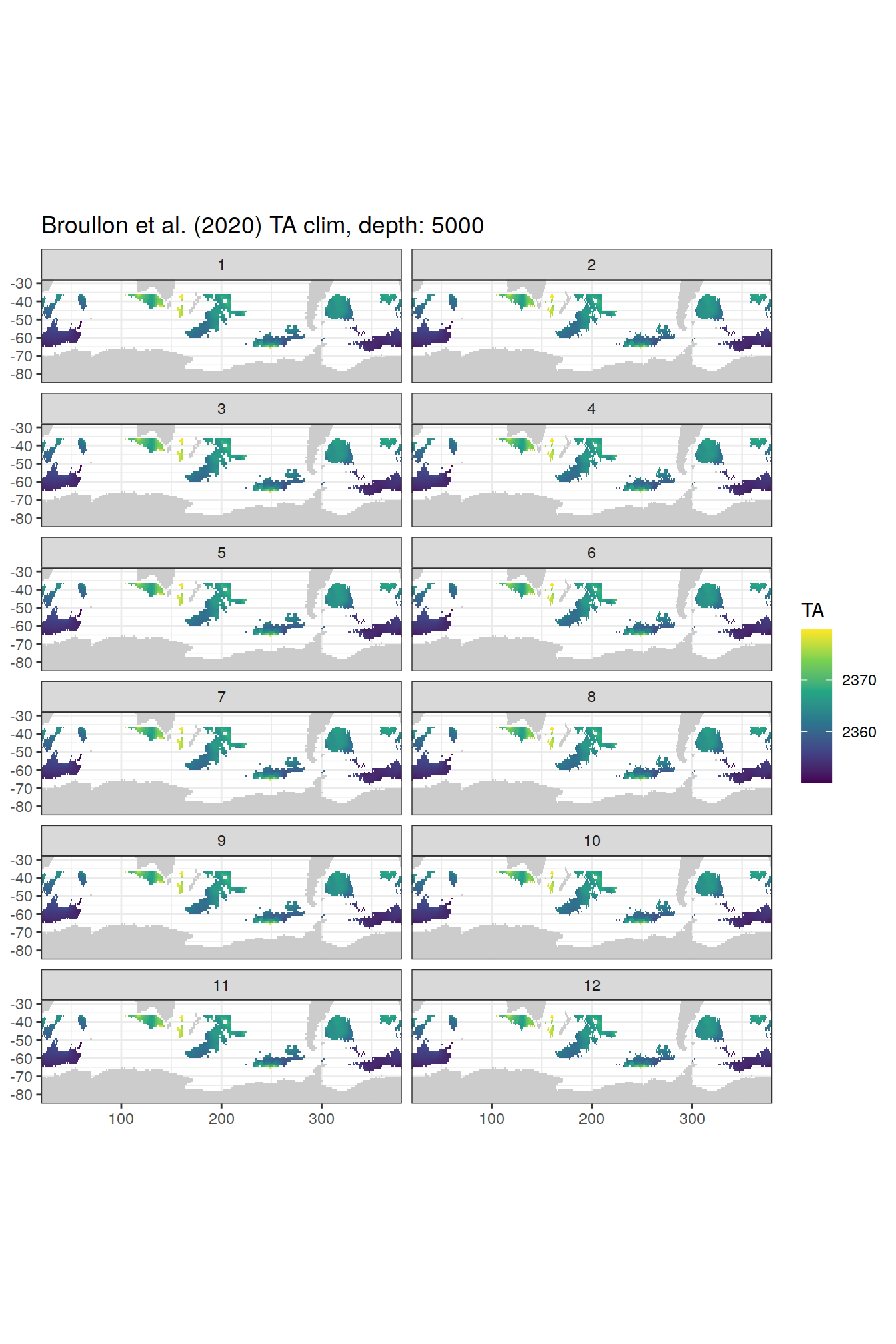
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[98]]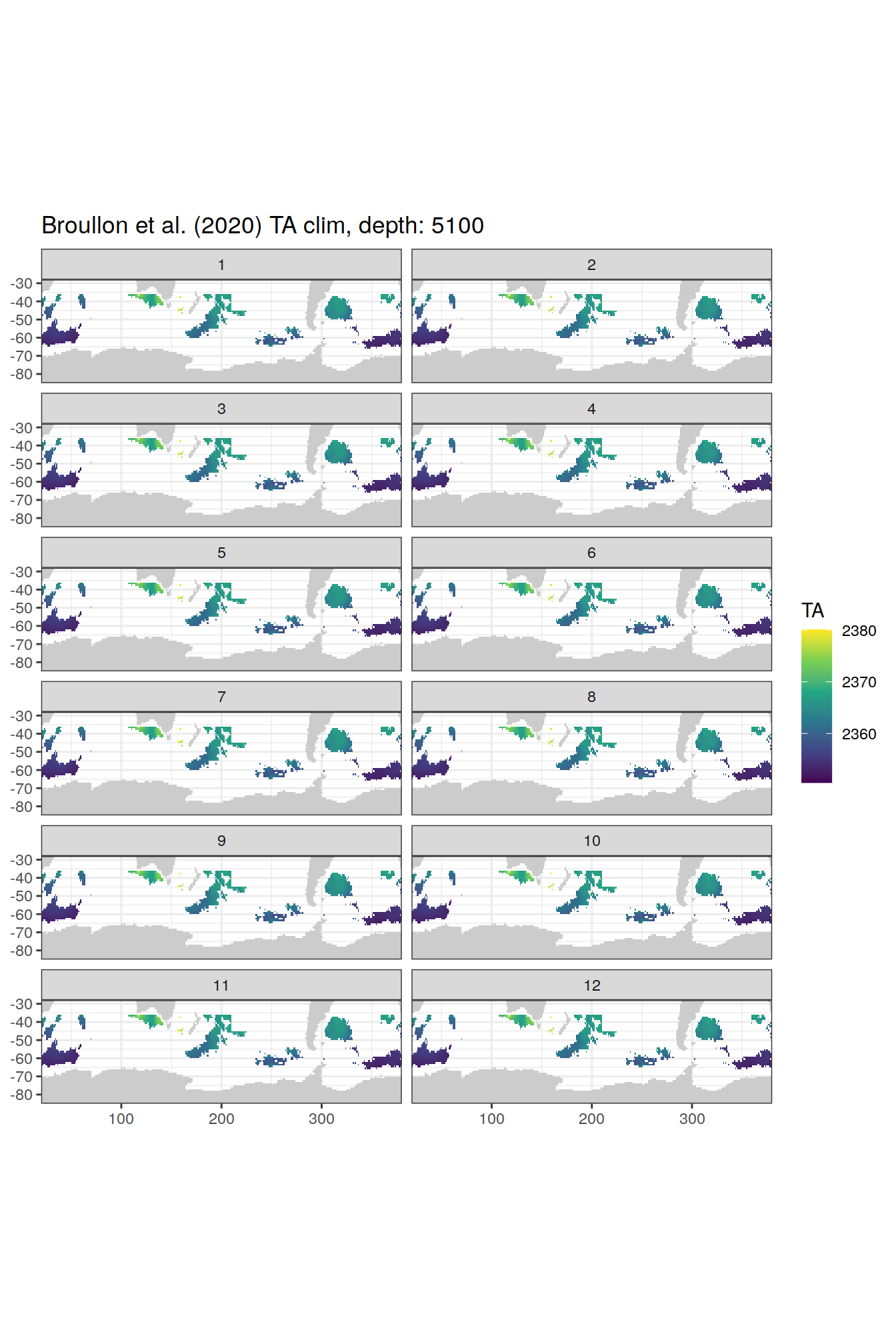
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[99]]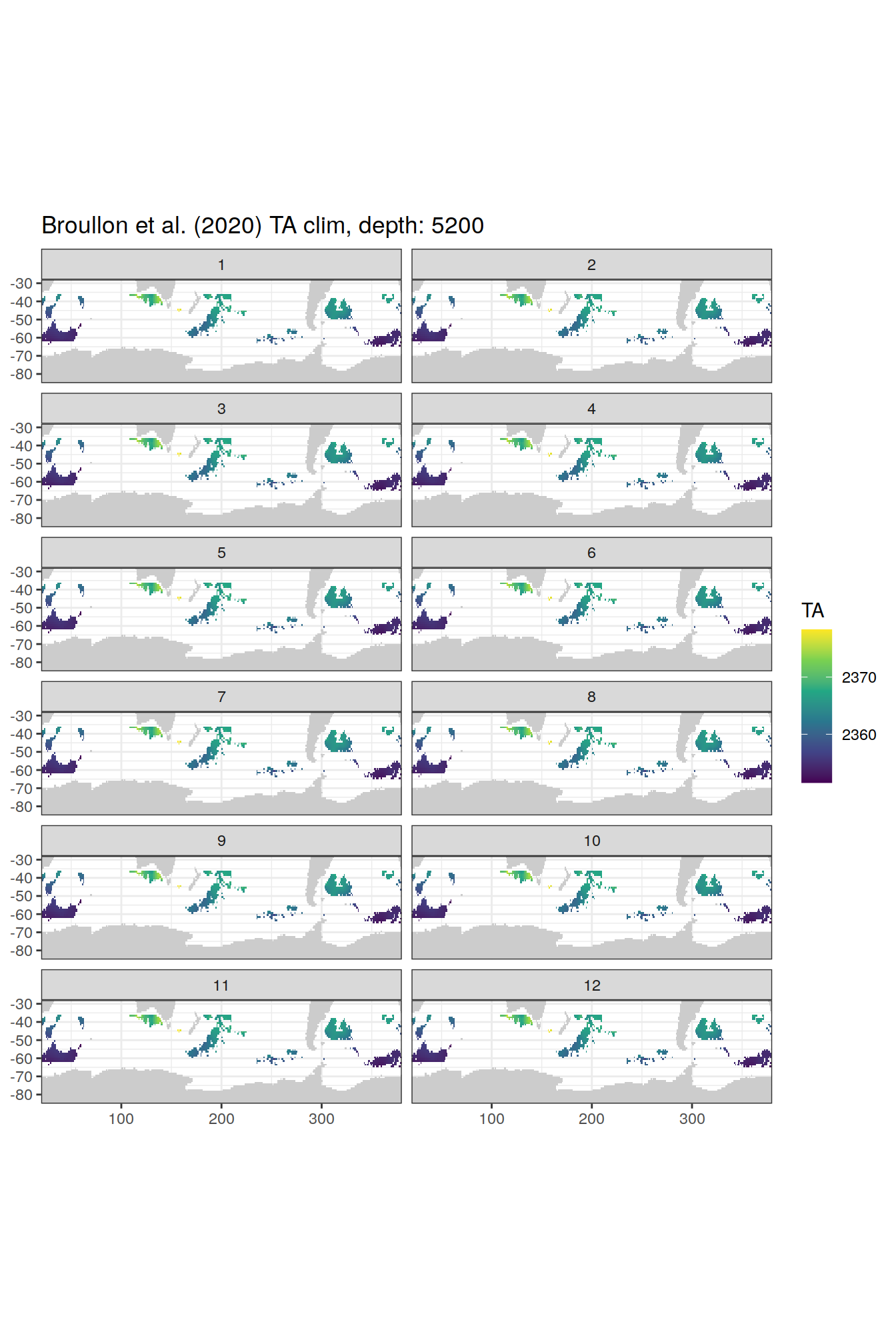
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[100]]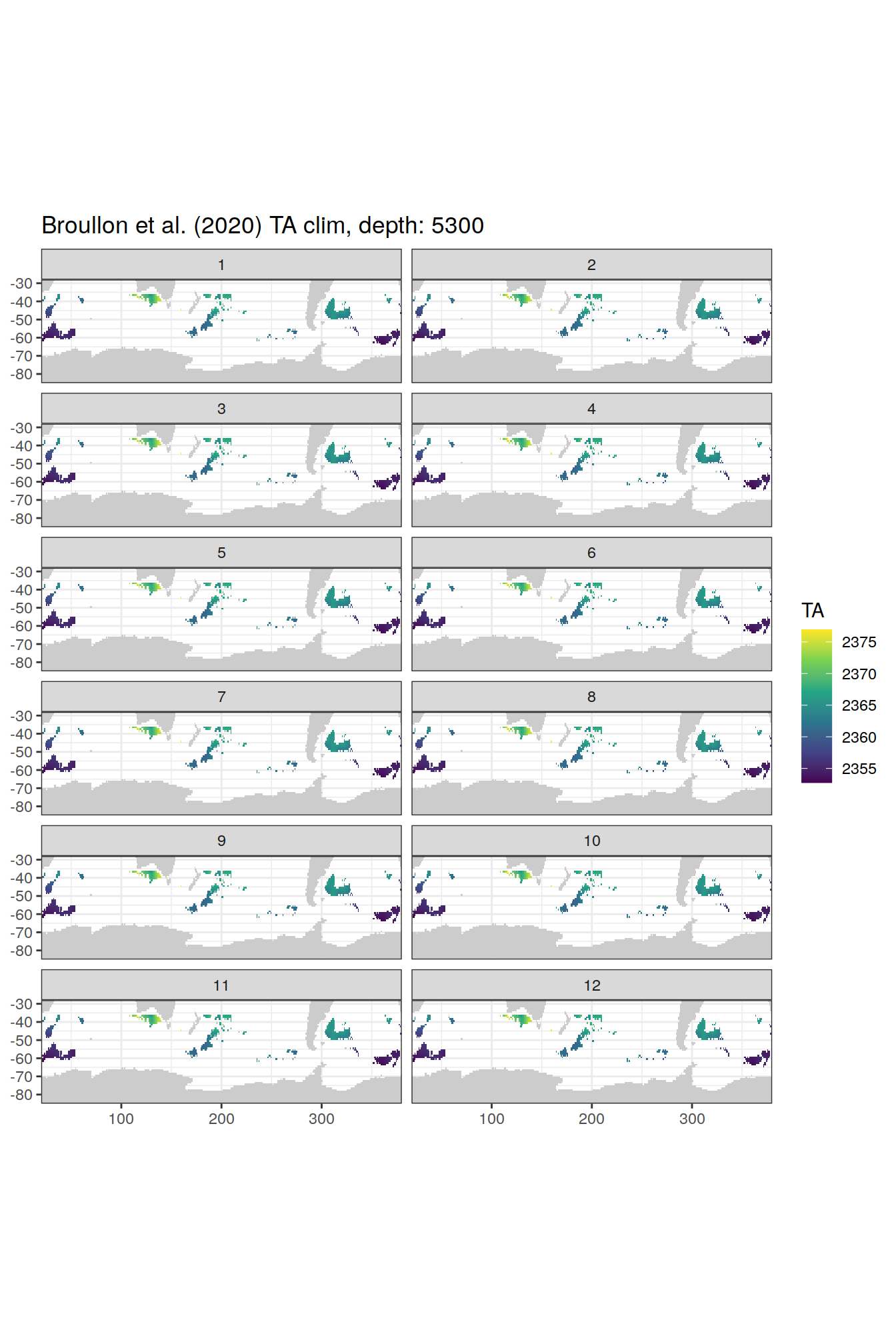
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[101]]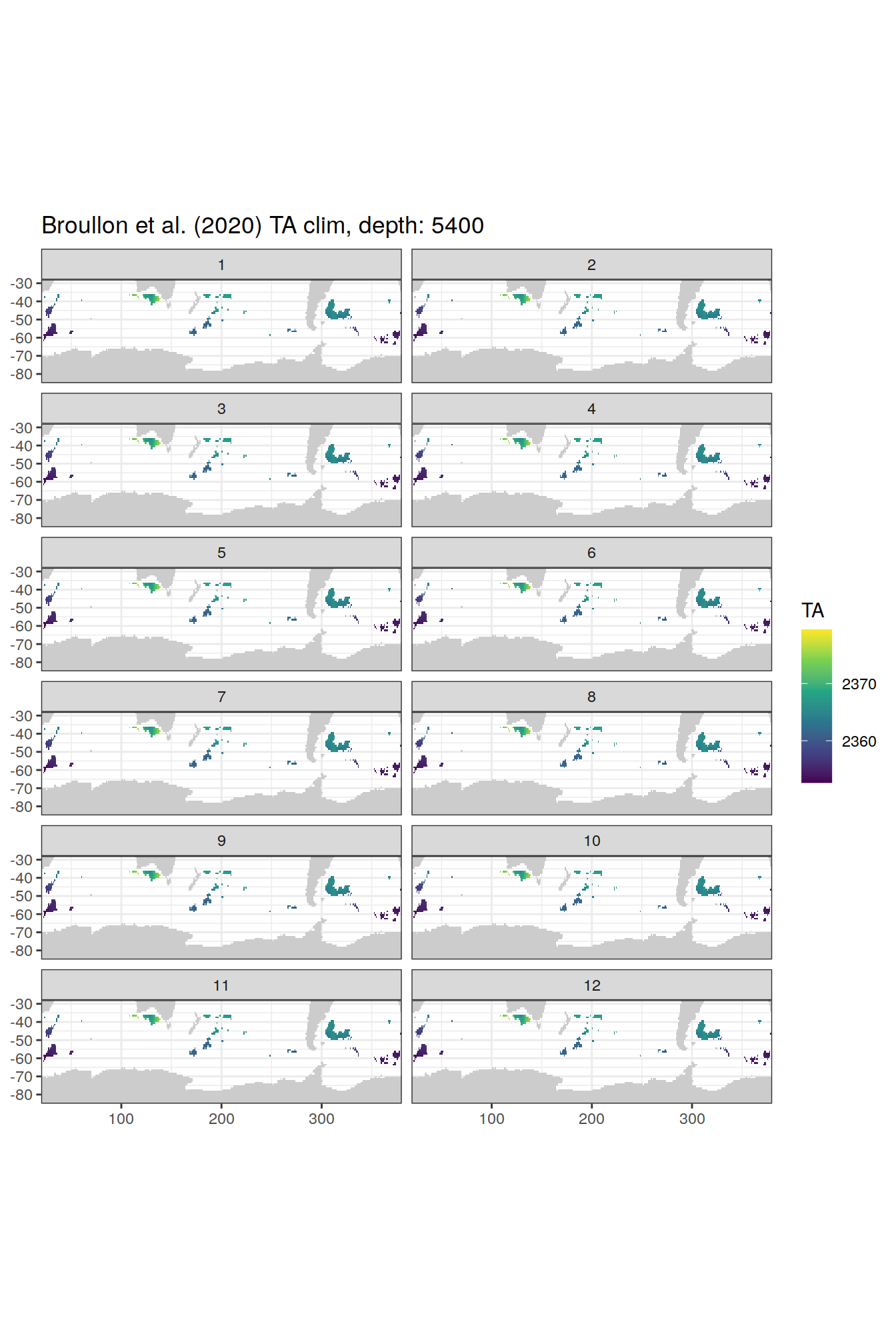
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[102]]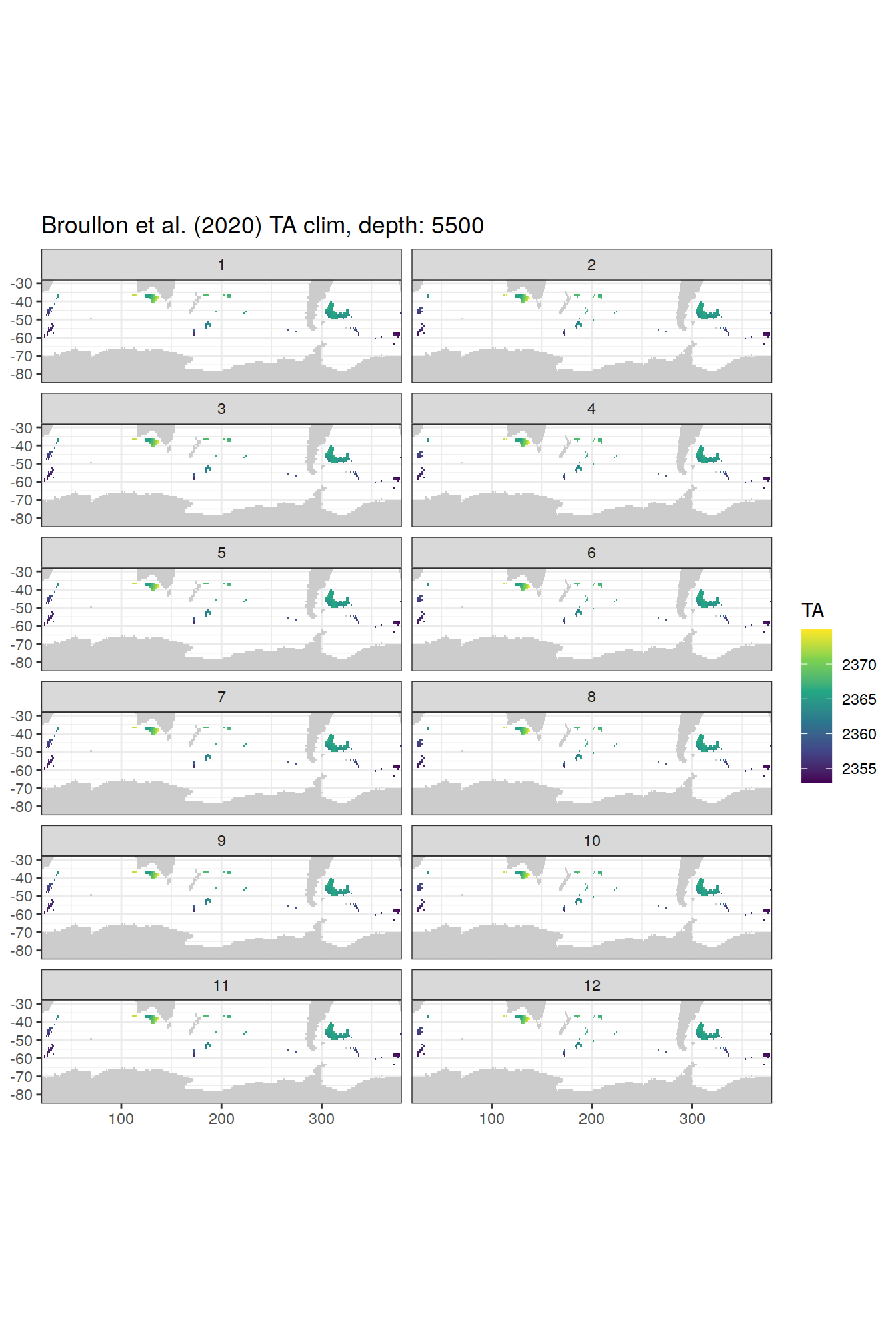
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
broullon_clim_SO %>%
# filter(depth <= 1500) %>%
group_split(month) %>%
map(
~ ggplot(data = .x,
aes(x = TA,
y = depth))+
geom_point(data = .x,
aes(x = TA,
y = depth),
size = 0.2,
pch = 1)+
scale_y_reverse()+
facet_grid(biome_name~basin_AIP)+
labs(title = paste0('Broullon et al. clim TA, month: ', unique(.x$month)))
)[[1]]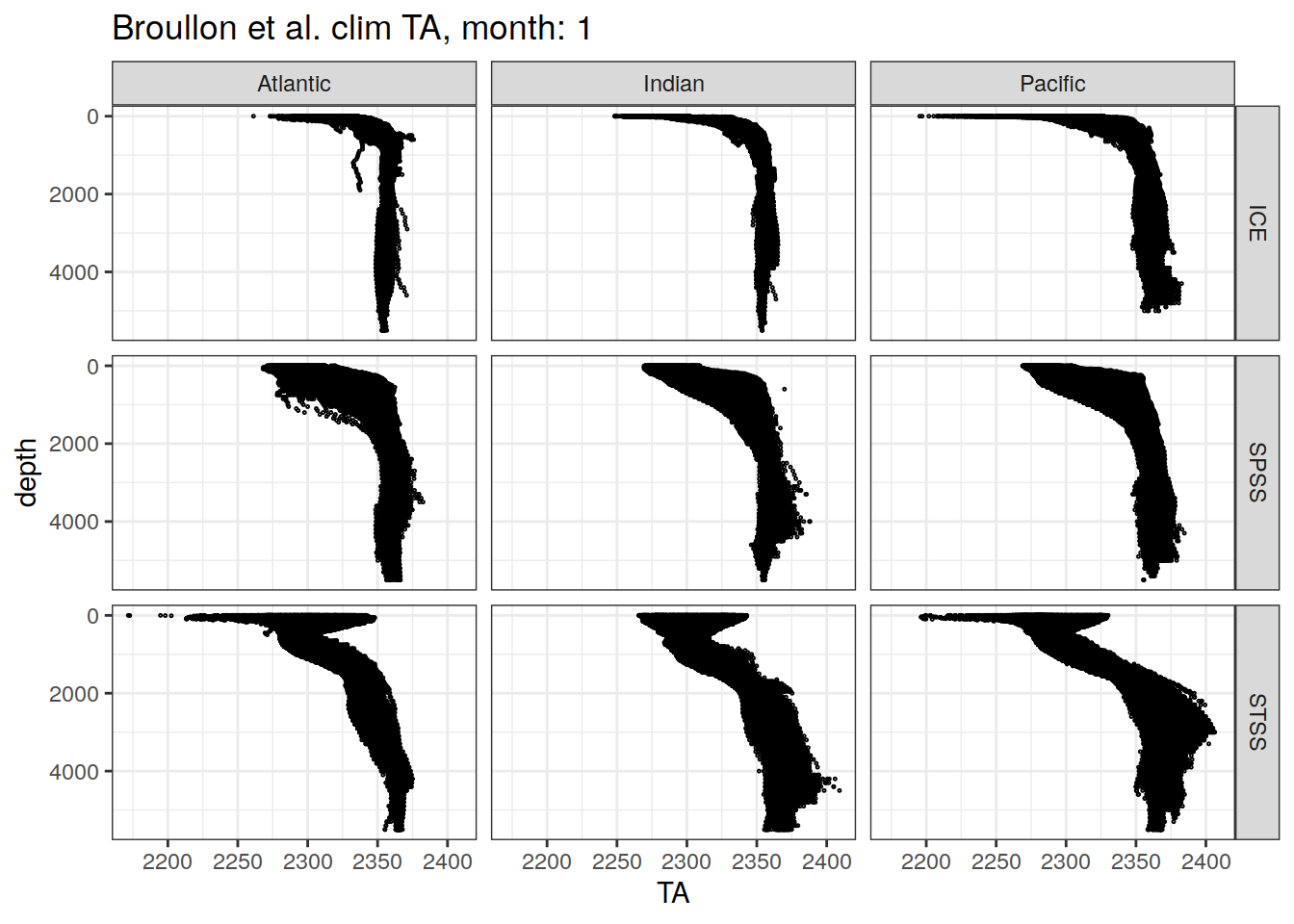
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[2]]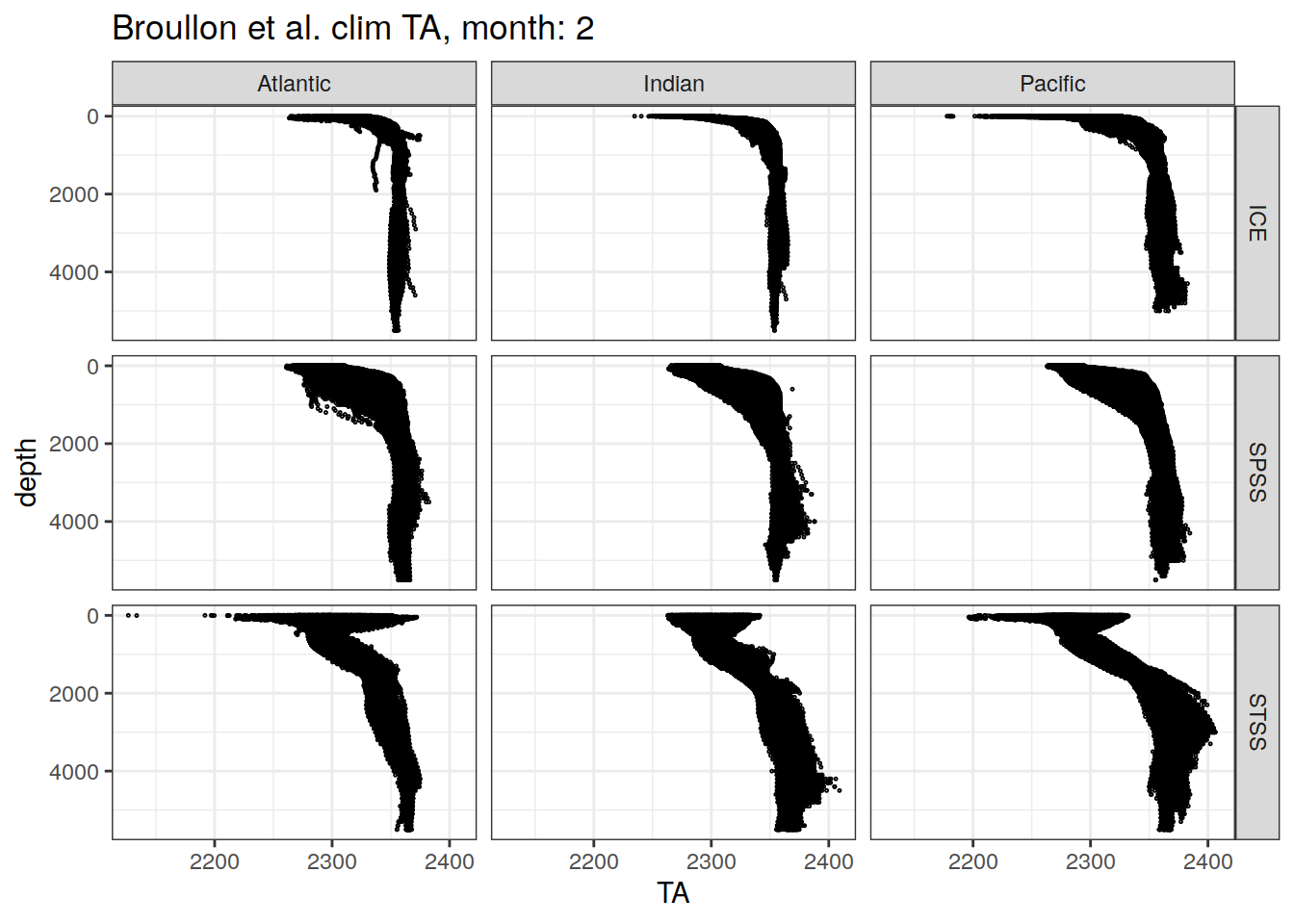
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[3]]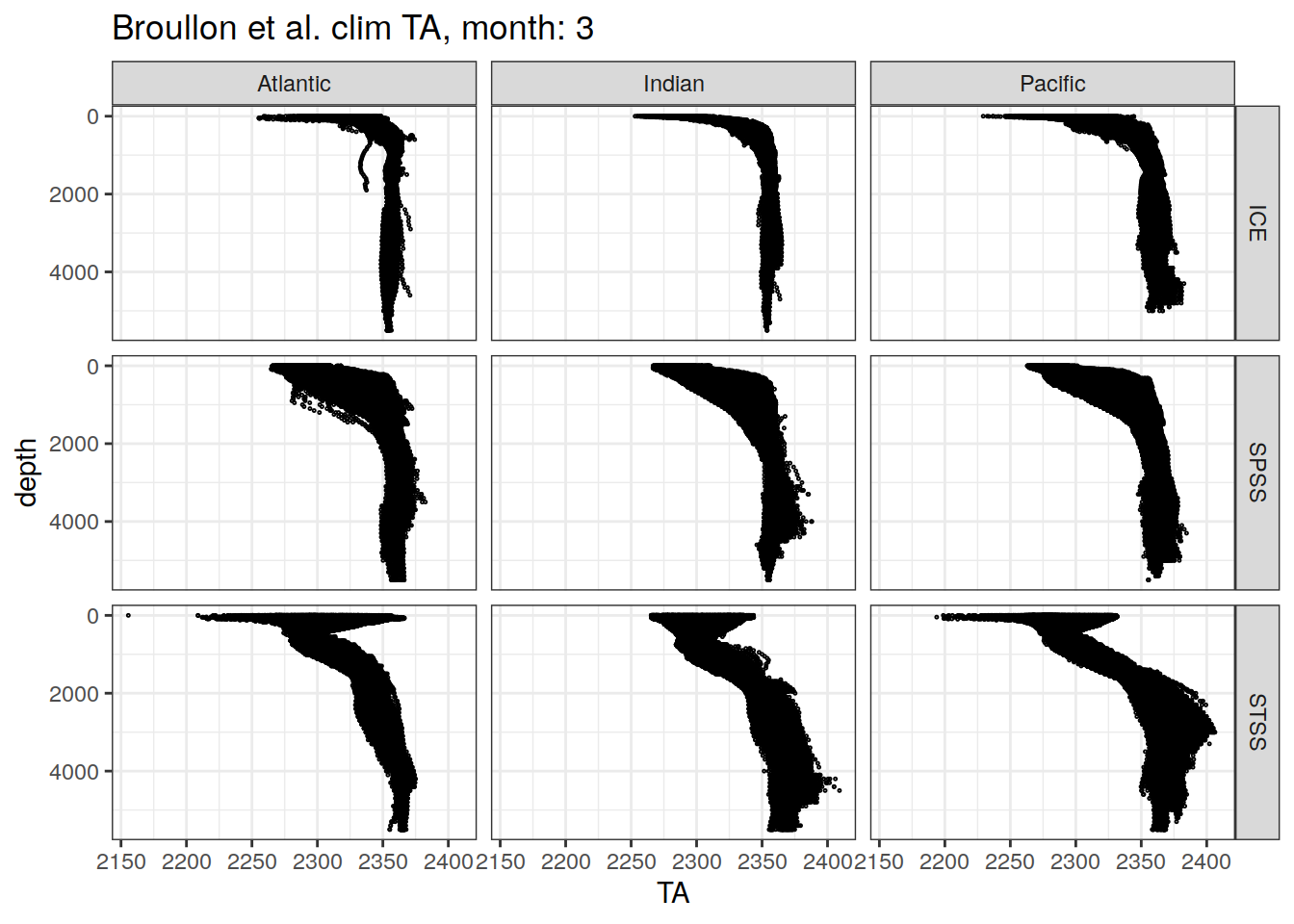
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[4]]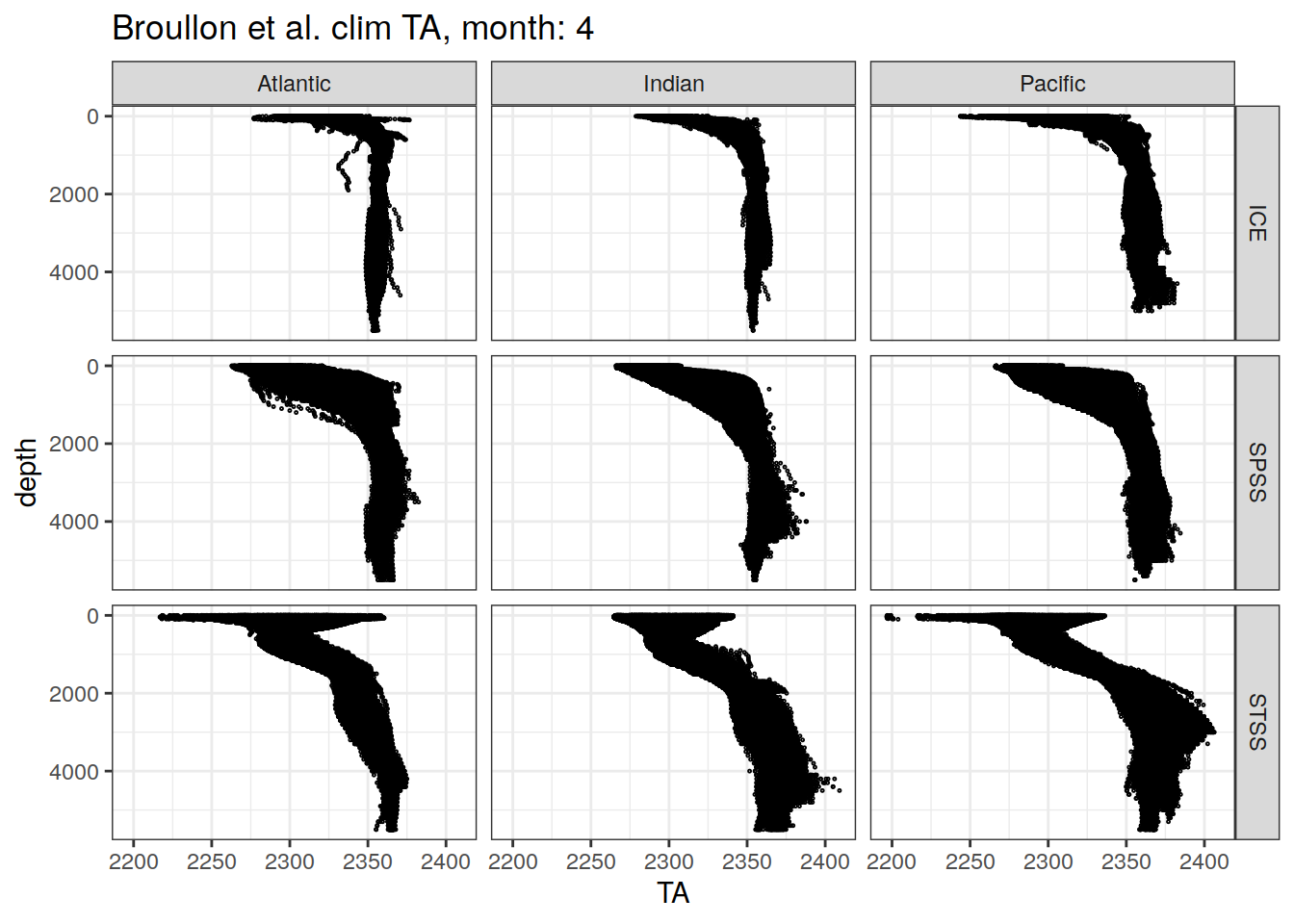
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[5]]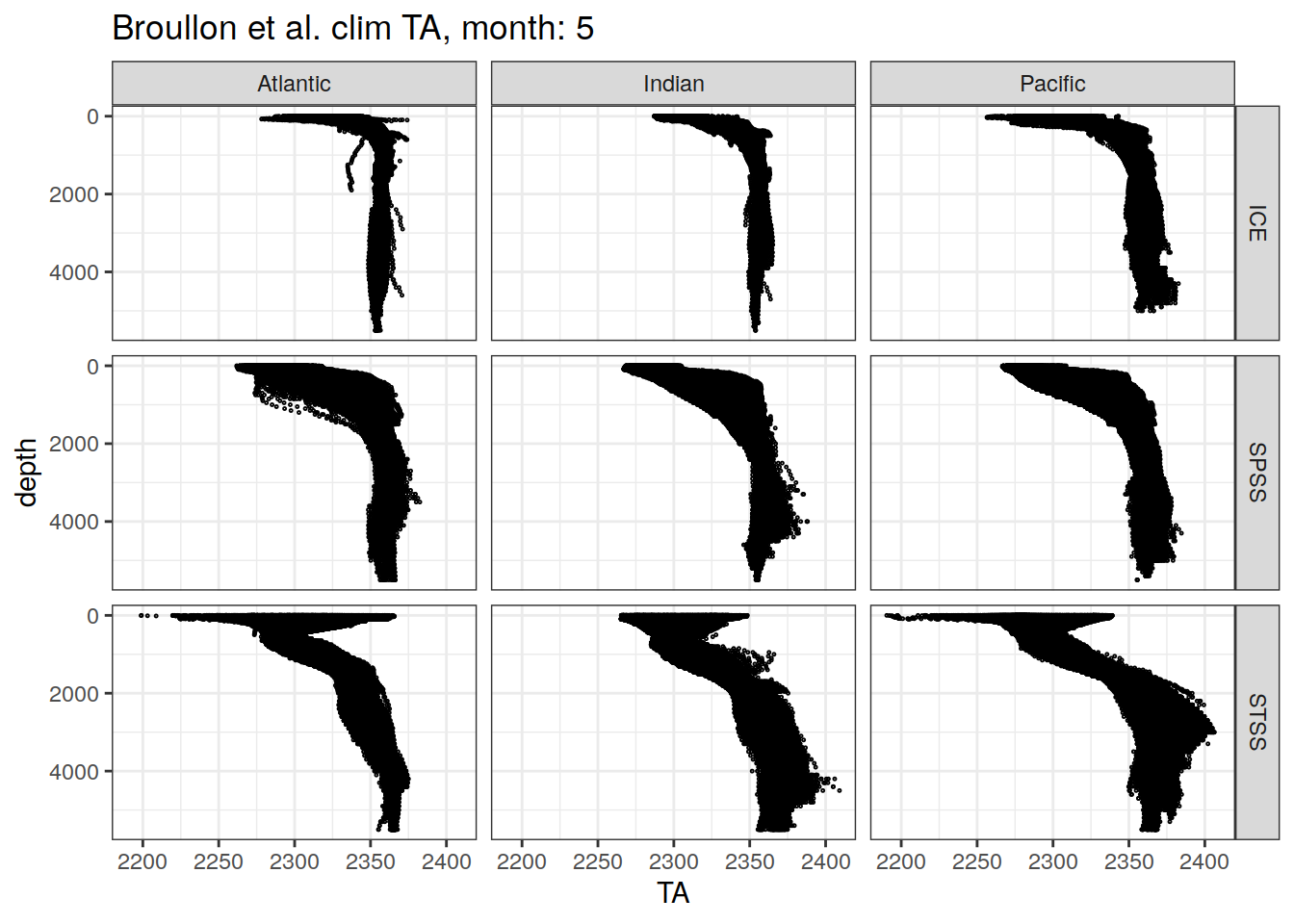
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[6]]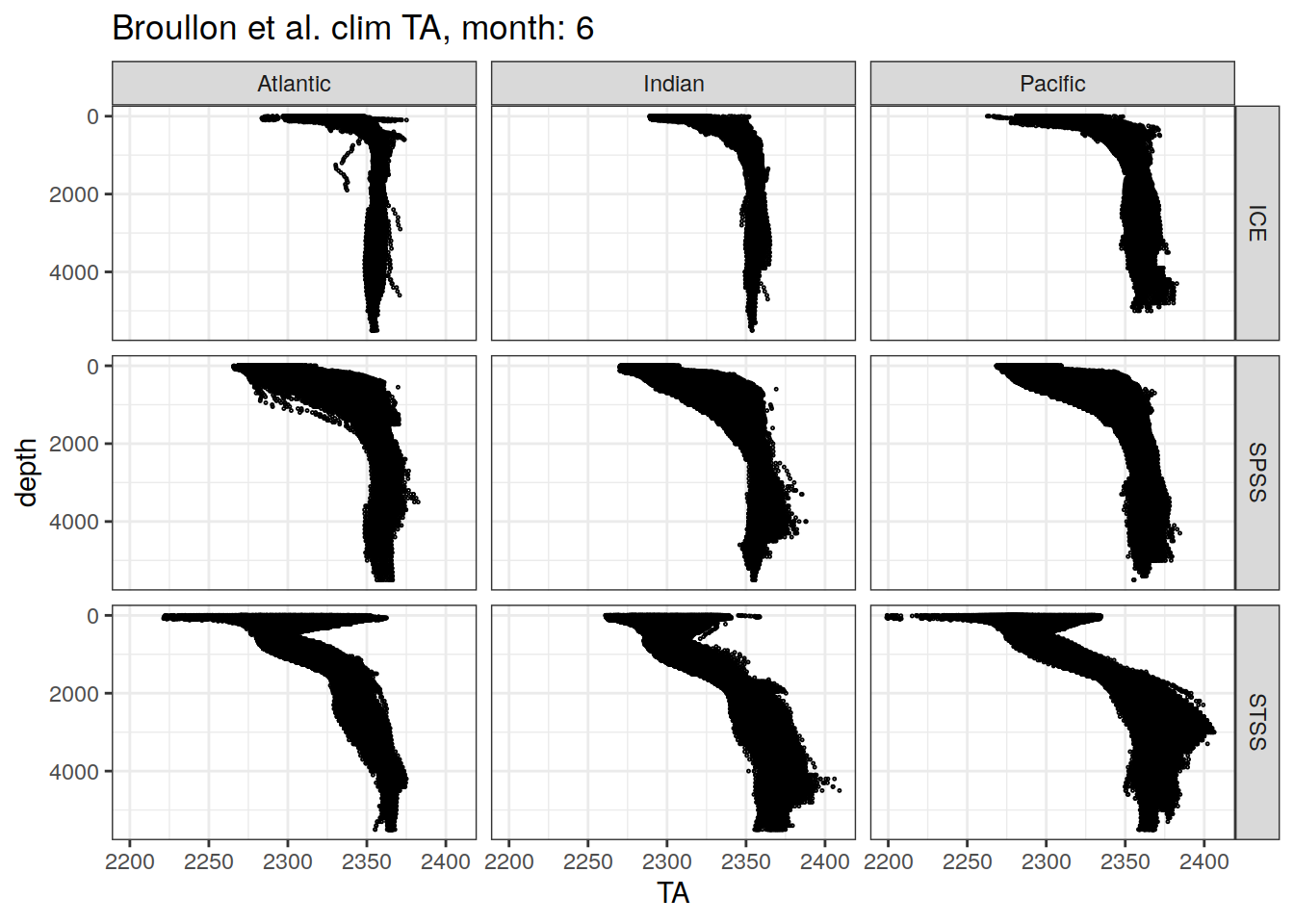
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[7]]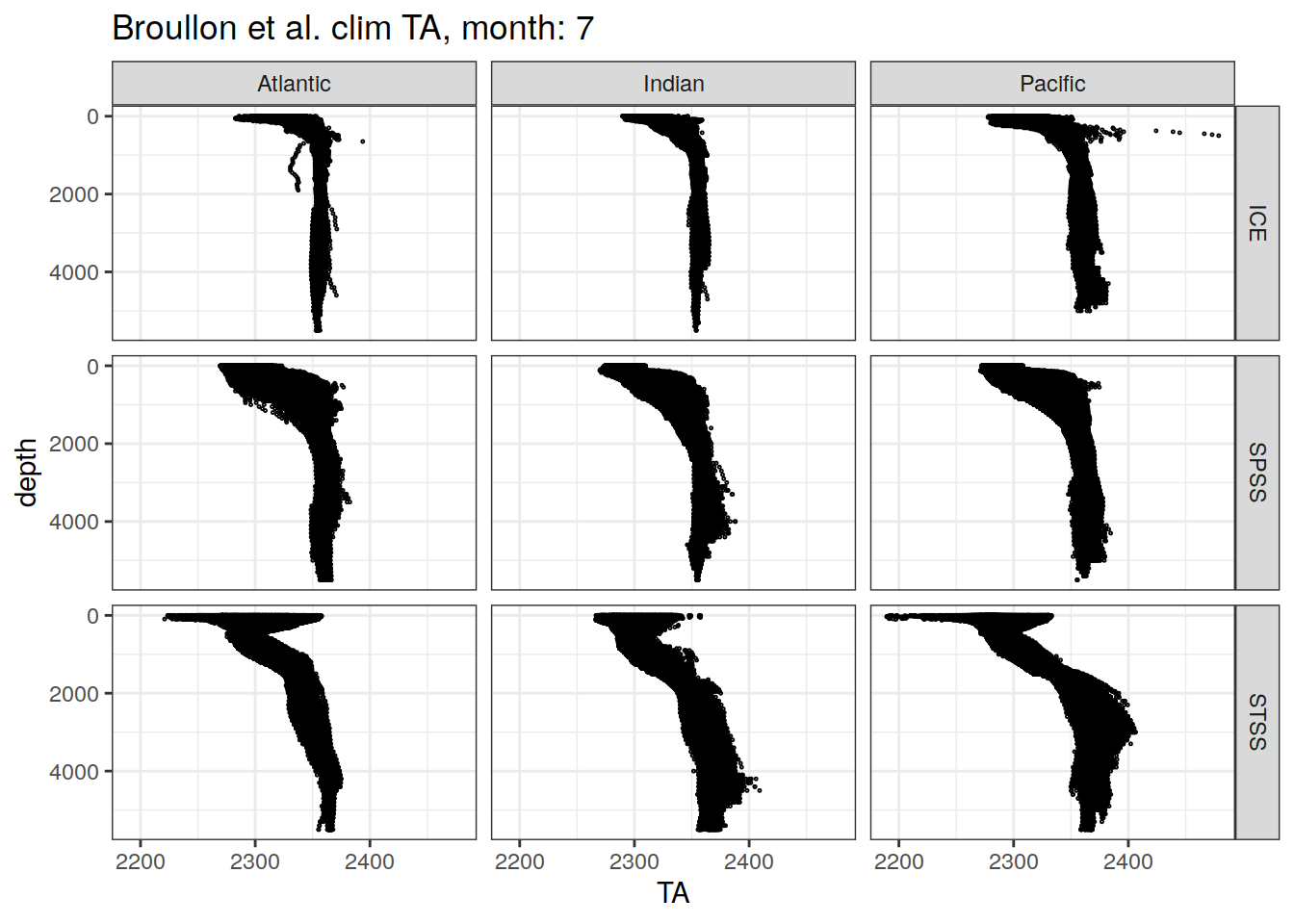
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[8]]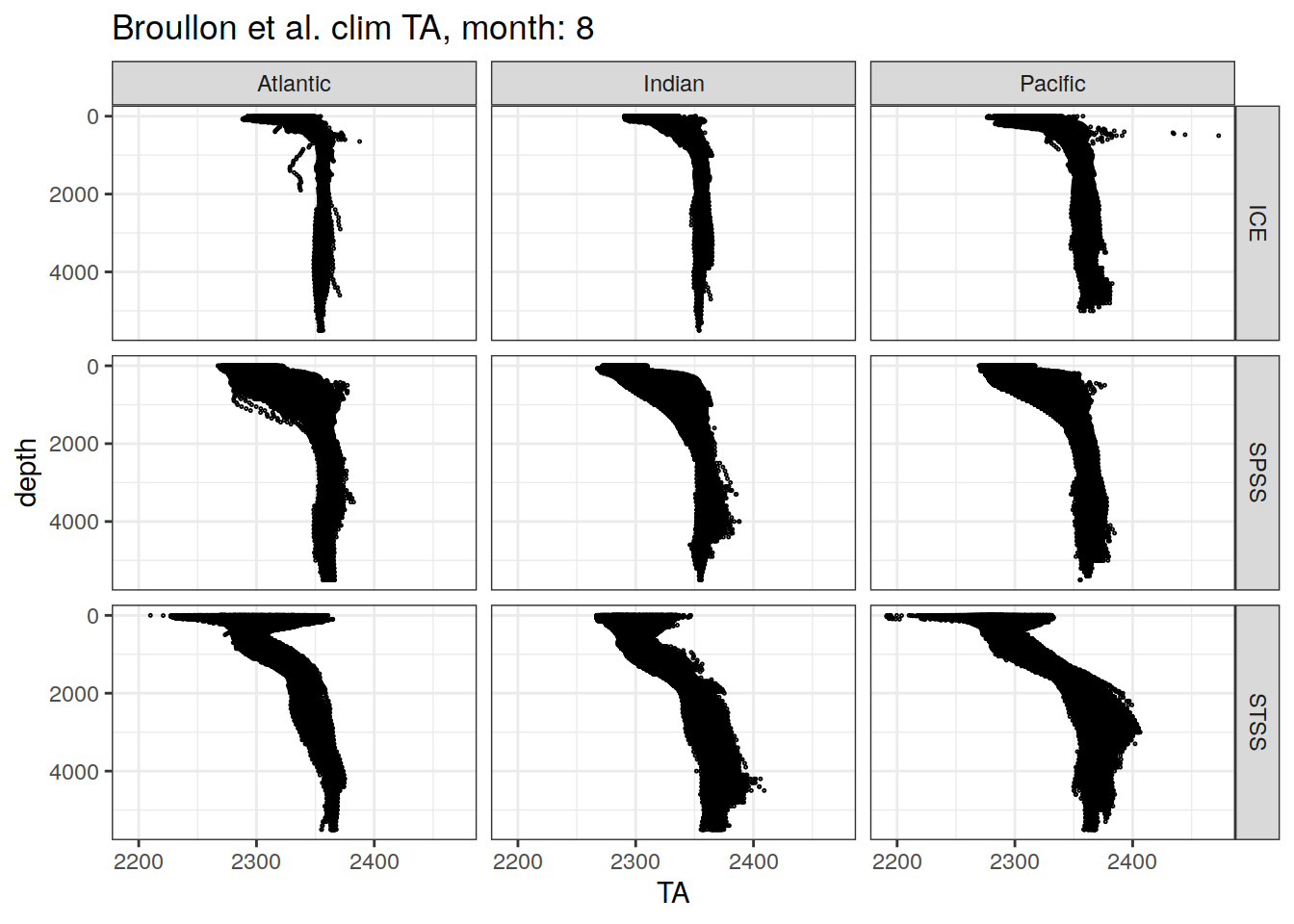
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[9]]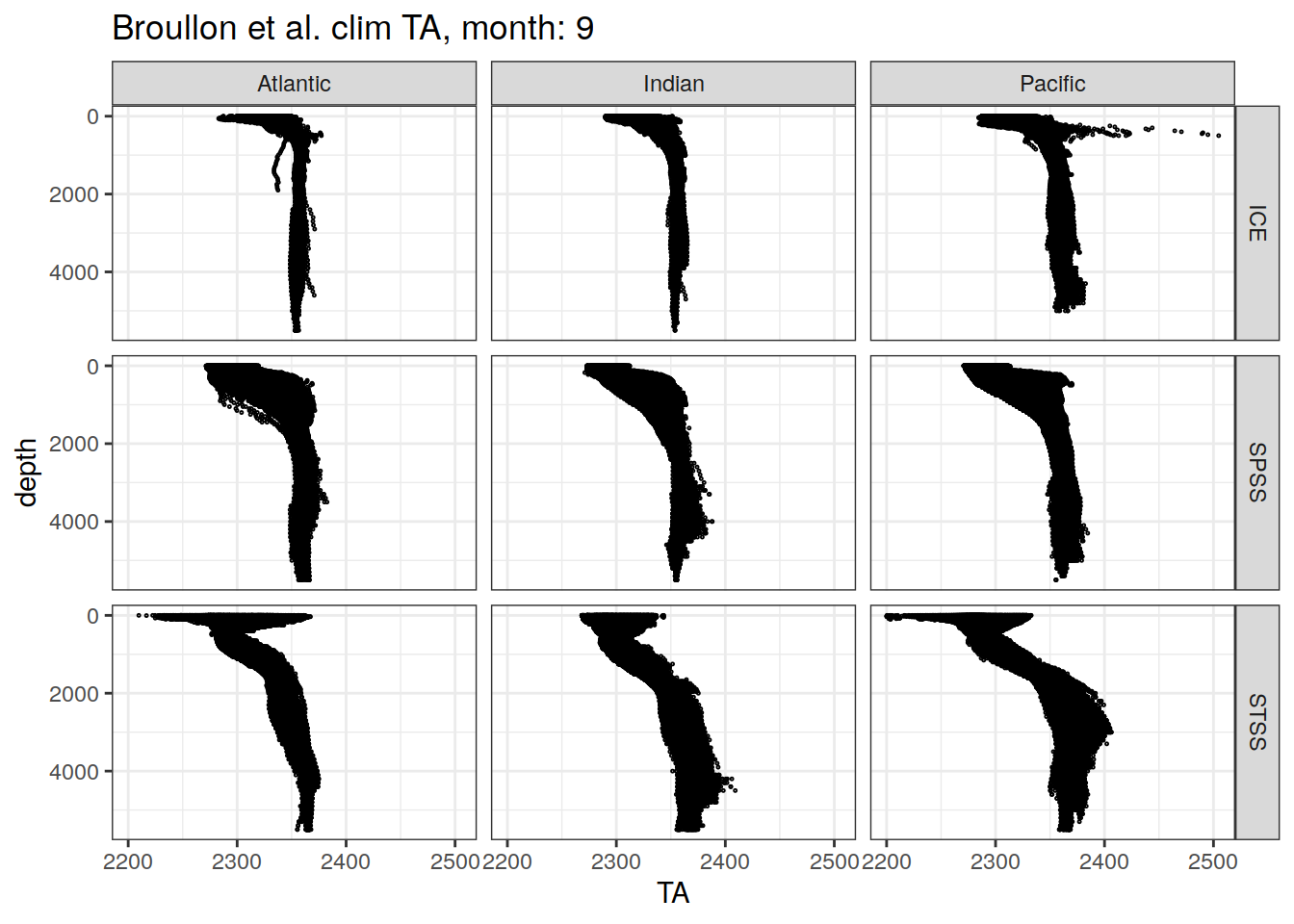
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[10]]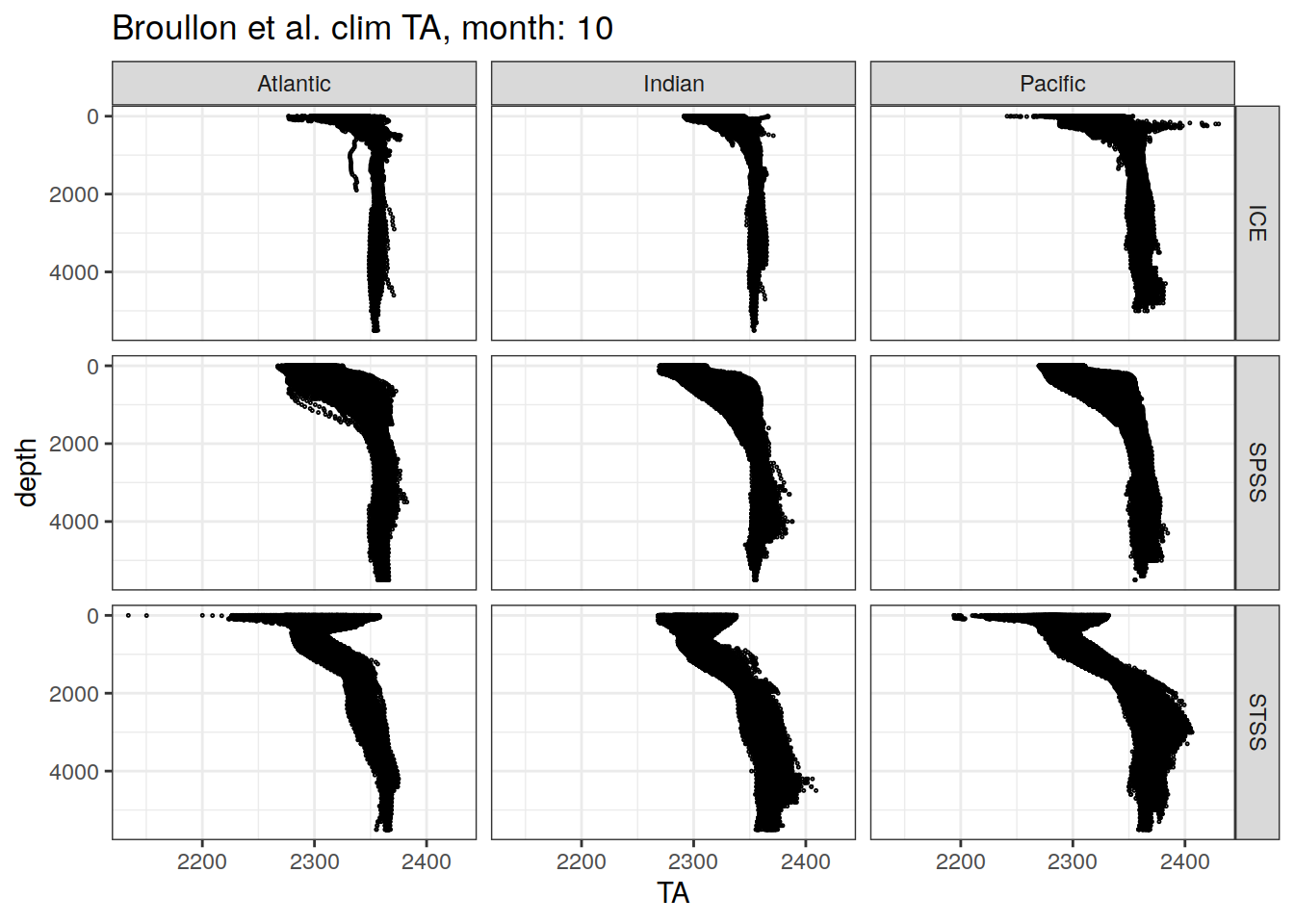
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[11]]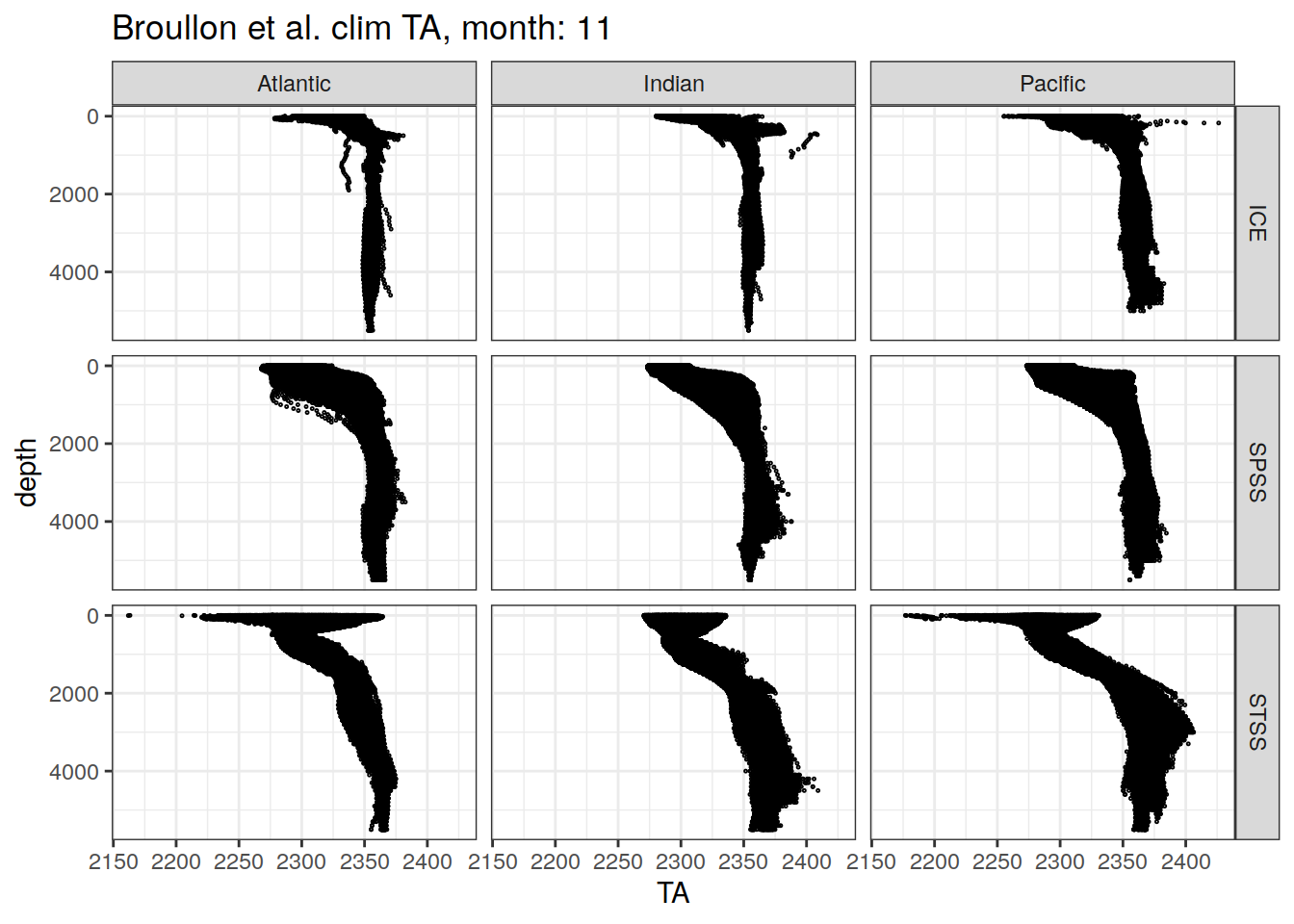
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
[[12]]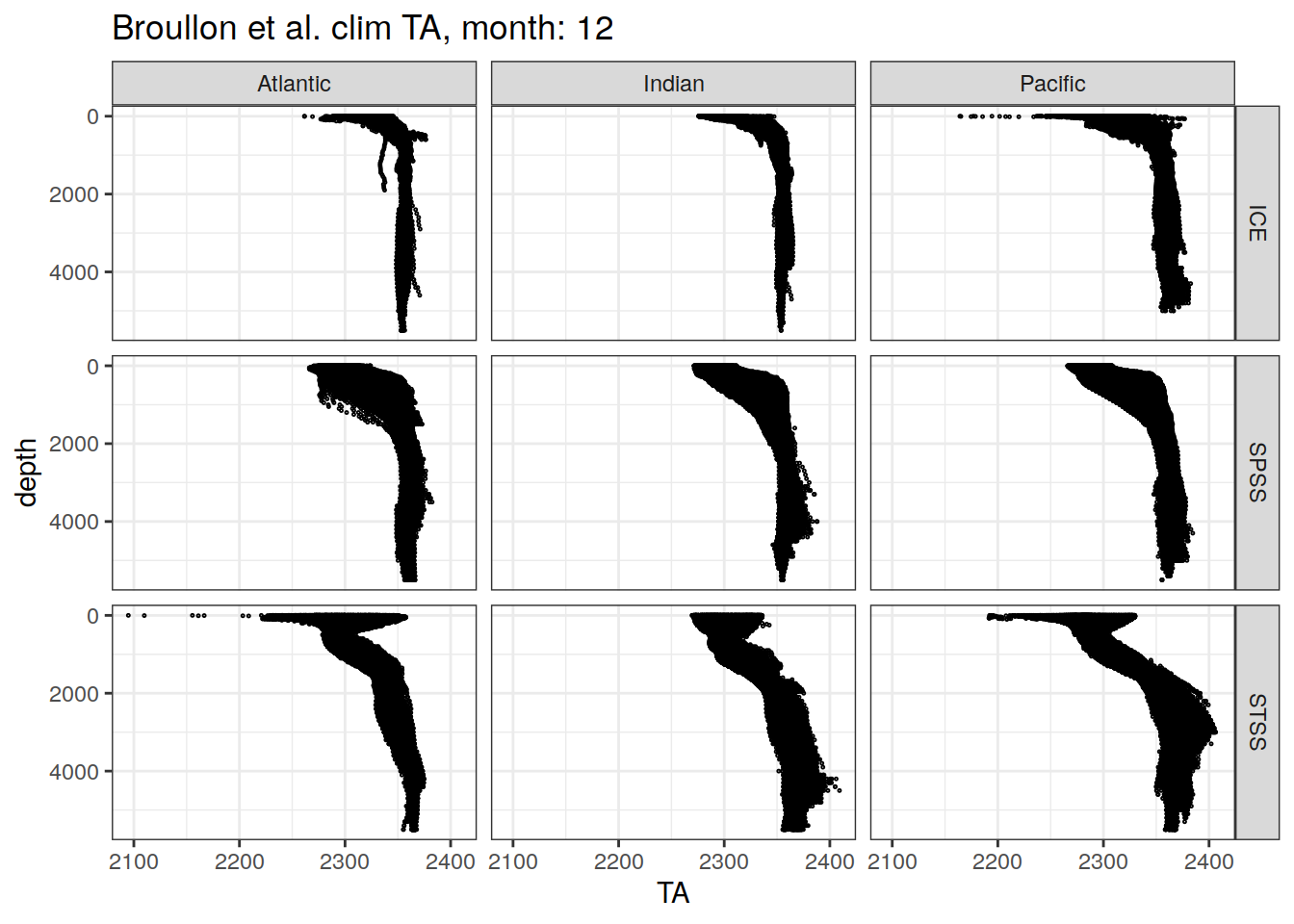
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
pCO2
map+
geom_tile(data = broullon_clim_SO %>% filter(depth_level == 1),
aes(x = lon,
y = lat,
fill = pco2))+
scale_fill_viridis_c()+
lims(y = c(-85, -28))+
facet_wrap(~month, ncol = 2)+
labs(title = 'Broullon et al. pCO2 clim')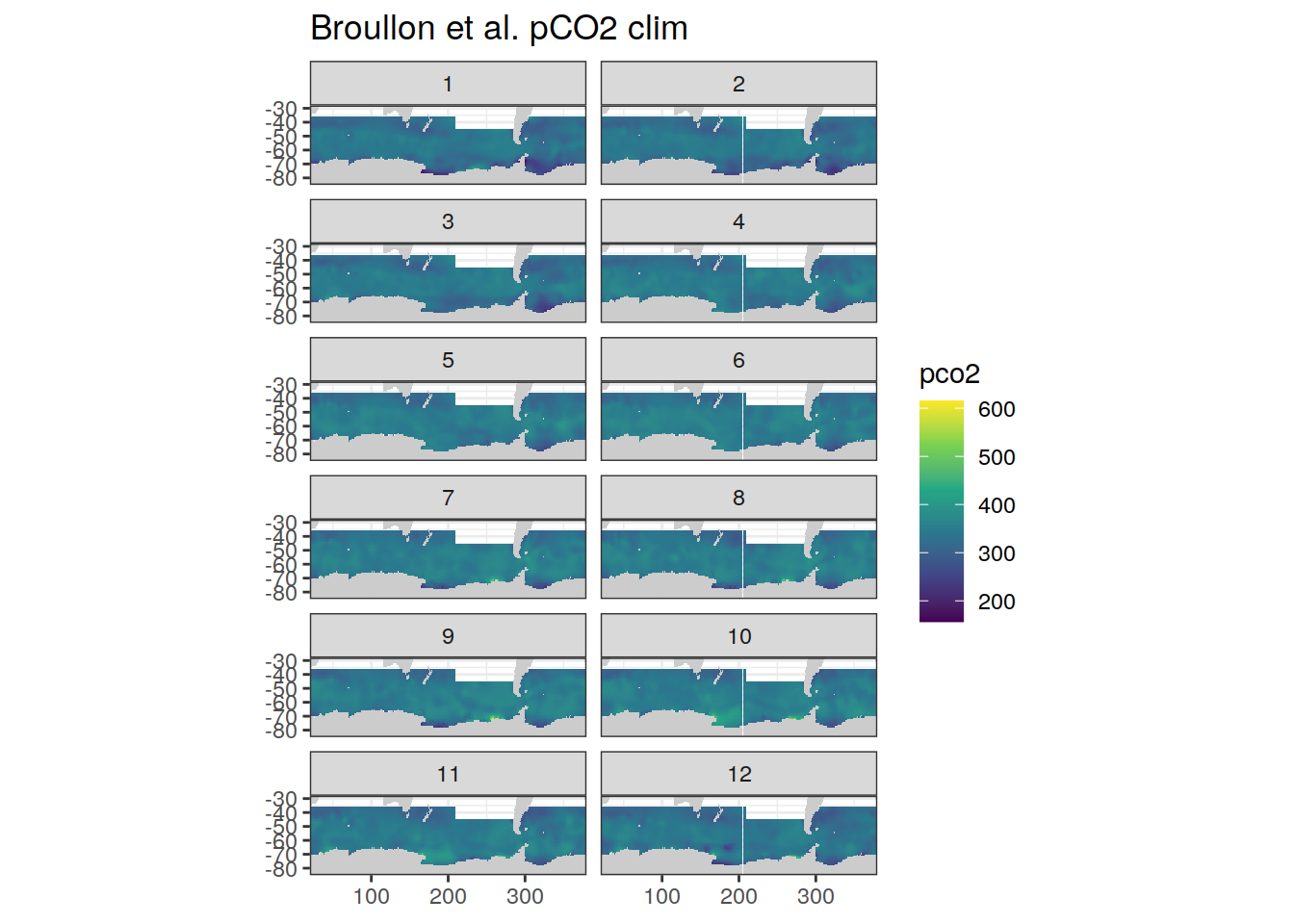
| Version | Author | Date |
|---|---|---|
| 2d44f8a | pasqualina-vonlanthendinenna | 2022-04-29 |
Oxygen
broullon_clim_SO %>%
group_split(depth) %>%
map(
~map +
geom_tile(data = .x,
aes(x = lon,
y = lat,
fill = oxygen))+
scale_fill_viridis_c()+
lims(y = c(-85, -28))+
facet_wrap(~month, ncol = 2)+
labs(title = paste0('Broullon et al. (2020) oxygen clim, depth: ', unique(.x$depth)))
)broullon_clim_SO %>%
# filter(depth <= 1500) %>%
group_split(month) %>%
map(
~ ggplot(data = .x,
aes(x = oxygen,
y = depth))+
geom_point(data = .x,
aes(x = oxygen,
y = depth),
size = 0.2,
pch = 1)+
scale_y_reverse()+
facet_grid(biome_name~basin_AIP)+
labs(title = paste0('Broullon et al. clim oxygen, month: ', unique(.x$month)))
)Nitrate
broullon_clim_SO %>%
group_split(depth) %>%
map(
~map +
geom_tile(data = .x,
aes(x = lon,
y = lat,
fill = nitrate))+
scale_fill_viridis_c()+
lims(y = c(-85, -28))+
facet_wrap(~month, ncol = 2)+
labs(title = paste0('Broullon et al. (2020) nitrate clim, depth: ', unique(.x$depth)))
)broullon_clim_SO %>%
# filter(depth <= 1500) %>%
group_split(month) %>%
map(
~ ggplot(data = .x,
aes(x = nitrate,
y = depth))+
geom_point(data = .x,
aes(x = nitrate,
y = depth),
size = 0.2,
pch = 1)+
scale_y_reverse()+
facet_grid(biome_name~basin_AIP)+
labs(title = paste0('Broullon et al. clim nitrate, month: ', unique(.x$month)))
)Phosphate
broullon_clim_SO %>%
group_split(depth) %>%
map(
~map +
geom_tile(data = .x,
aes(x = lon,
y = lat,
fill = phosphate))+
scale_fill_viridis_c()+
lims(y = c(-85, -28))+
facet_wrap(~month, ncol = 2)+
labs(title = paste0('Broullon et al. (2020) phosphate clim, depth: ', unique(.x$depth)))
)broullon_clim_SO %>%
# filter(depth <= 1500) %>%
group_split(month) %>%
map(
~ ggplot(data = .x,
aes(x = phosphate,
y = depth))+
geom_point(data = .x,
aes(x = phosphate,
y = depth),
size = 0.2,
pch = 1)+
scale_y_reverse()+
facet_grid(biome_name~basin_AIP)+
labs(title = paste0('Broullon et al. clim phosphate, month: ', unique(.x$month)))
)
sessionInfo()R version 4.1.2 (2021-11-01)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: openSUSE Leap 15.3
Matrix products: default
BLAS: /usr/local/R-4.1.2/lib64/R/lib/libRblas.so
LAPACK: /usr/local/R-4.1.2/lib64/R/lib/libRlapack.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] stars_0.5-5 sf_1.0-5 abind_1.4-5 lubridate_1.8.0
[5] forcats_0.5.1 stringr_1.4.0 dplyr_1.0.7 purrr_0.3.4
[9] readr_2.1.1 tidyr_1.1.4 tibble_3.1.6 ggplot2_3.3.5
[13] tidyverse_1.3.1 workflowr_1.7.0
loaded via a namespace (and not attached):
[1] fs_1.5.2 bit64_4.0.5 httr_1.4.2 rprojroot_2.0.2
[5] tools_4.1.2 backports_1.4.1 bslib_0.3.1 utf8_1.2.2
[9] R6_2.5.1 KernSmooth_2.23-20 DBI_1.1.2 colorspace_2.0-2
[13] withr_2.4.3 tidyselect_1.1.1 processx_3.5.2 bit_4.0.4
[17] compiler_4.1.2 git2r_0.29.0 cli_3.1.1 rvest_1.0.2
[21] RNetCDF_2.5-2 xml2_1.3.3 labeling_0.4.2 sass_0.4.0
[25] scales_1.1.1 classInt_0.4-3 callr_3.7.0 proxy_0.4-26
[29] digest_0.6.29 rmarkdown_2.11 pkgconfig_2.0.3 htmltools_0.5.2
[33] highr_0.9 dbplyr_2.1.1 fastmap_1.1.0 rlang_1.0.2
[37] tidync_0.2.4 readxl_1.3.1 rstudioapi_0.13 farver_2.1.0
[41] jquerylib_0.1.4 generics_0.1.1 jsonlite_1.7.3 vroom_1.5.7
[45] magrittr_2.0.1 ncmeta_0.3.0 Rcpp_1.0.8 munsell_0.5.0
[49] fansi_1.0.2 lifecycle_1.0.1 stringi_1.7.6 whisker_0.4
[53] yaml_2.2.1 grid_4.1.2 parallel_4.1.2 promises_1.2.0.1
[57] crayon_1.4.2 haven_2.4.3 hms_1.1.1 knitr_1.37
[61] ps_1.6.0 pillar_1.6.4 reprex_2.0.1 glue_1.6.0
[65] evaluate_0.14 getPass_0.2-2 modelr_0.1.8 vctrs_0.3.8
[69] tzdb_0.2.0 httpuv_1.6.5 cellranger_1.1.0 gtable_0.3.0
[73] assertthat_0.2.1 xfun_0.29 lwgeom_0.2-8 broom_0.7.11
[77] e1071_1.7-9 later_1.3.0 class_7.3-20 ncdf4_1.19
[81] viridisLite_0.4.0 units_0.7-2 ellipsis_0.3.2