ebpower
Matthew Stephens
2025-03-09
Last updated: 2025-03-16
Checks: 7 0
Knit directory: misc/analysis/
This reproducible R Markdown analysis was created with workflowr (version 1.7.1). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(1) was run prior to running the
code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version fe667eb. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: analysis/.RData
Ignored: analysis/.Rhistory
Ignored: analysis/ALStruct_cache/
Ignored: data/.Rhistory
Ignored: data/methylation-data-for-matthew.rds
Ignored: data/pbmc/
Ignored: data/pbmc_purified.RData
Untracked files:
Untracked: .dropbox
Untracked: Icon
Untracked: analysis/GHstan.Rmd
Untracked: analysis/GTEX-cogaps.Rmd
Untracked: analysis/PACS.Rmd
Untracked: analysis/Rplot.png
Untracked: analysis/SPCAvRP.rmd
Untracked: analysis/abf_comparisons.Rmd
Untracked: analysis/admm_02.Rmd
Untracked: analysis/admm_03.Rmd
Untracked: analysis/bispca.Rmd
Untracked: analysis/cache/
Untracked: analysis/cholesky.Rmd
Untracked: analysis/compare-transformed-models.Rmd
Untracked: analysis/cormotif.Rmd
Untracked: analysis/cp_ash.Rmd
Untracked: analysis/eQTL.perm.rand.pdf
Untracked: analysis/eb_prepilot.Rmd
Untracked: analysis/eb_snmu.Rmd
Untracked: analysis/eb_var.Rmd
Untracked: analysis/ebpmf1.Rmd
Untracked: analysis/ebpmf_sla_text.Rmd
Untracked: analysis/ebspca_sims.Rmd
Untracked: analysis/explore_psvd.Rmd
Untracked: analysis/fa_check_identify.Rmd
Untracked: analysis/fa_iterative.Rmd
Untracked: analysis/flash_cov_overlapping_groups_init.Rmd
Untracked: analysis/flash_test_tree.Rmd
Untracked: analysis/flashier_newgroups.Rmd
Untracked: analysis/flashier_nmf_triples.Rmd
Untracked: analysis/flashier_pbmc.Rmd
Untracked: analysis/flashier_snn_shifted_prior.Rmd
Untracked: analysis/greedy_ebpmf_exploration_00.Rmd
Untracked: analysis/ieQTL.perm.rand.pdf
Untracked: analysis/lasso_em_03.Rmd
Untracked: analysis/m6amash.Rmd
Untracked: analysis/mash_bhat_z.Rmd
Untracked: analysis/mash_ieqtl_permutations.Rmd
Untracked: analysis/methylation_example.Rmd
Untracked: analysis/mixsqp.Rmd
Untracked: analysis/mr.ash_lasso_init.Rmd
Untracked: analysis/mr.mash.test.Rmd
Untracked: analysis/mr_ash_modular.Rmd
Untracked: analysis/mr_ash_parameterization.Rmd
Untracked: analysis/mr_ash_ridge.Rmd
Untracked: analysis/mv_gaussian_message_passing.Rmd
Untracked: analysis/nejm.Rmd
Untracked: analysis/nmf_bg.Rmd
Untracked: analysis/nonneg_underapprox.Rmd
Untracked: analysis/normal_conditional_on_r2.Rmd
Untracked: analysis/normalize.Rmd
Untracked: analysis/pbmc.Rmd
Untracked: analysis/pca_binary_weighted.Rmd
Untracked: analysis/pca_l1.Rmd
Untracked: analysis/poisson_nmf_approx.Rmd
Untracked: analysis/poisson_shrink.Rmd
Untracked: analysis/poisson_transform.Rmd
Untracked: analysis/qrnotes.txt
Untracked: analysis/ridge_iterative_02.Rmd
Untracked: analysis/ridge_iterative_splitting.Rmd
Untracked: analysis/samps/
Untracked: analysis/sc_bimodal.Rmd
Untracked: analysis/shrinkage_comparisons_changepoints.Rmd
Untracked: analysis/susie_cov.Rmd
Untracked: analysis/susie_en.Rmd
Untracked: analysis/susie_z_investigate.Rmd
Untracked: analysis/svd-timing.Rmd
Untracked: analysis/temp.RDS
Untracked: analysis/temp.Rmd
Untracked: analysis/test-figure/
Untracked: analysis/test.Rmd
Untracked: analysis/test.Rpres
Untracked: analysis/test.md
Untracked: analysis/test_qr.R
Untracked: analysis/test_sparse.Rmd
Untracked: analysis/tree_dist_top_eigenvector.Rmd
Untracked: analysis/z.txt
Untracked: code/multivariate_testfuncs.R
Untracked: code/rqb.hacked.R
Untracked: data/4matthew/
Untracked: data/4matthew2/
Untracked: data/E-MTAB-2805.processed.1/
Untracked: data/ENSG00000156738.Sim_Y2.RDS
Untracked: data/GDS5363_full.soft.gz
Untracked: data/GSE41265_allGenesTPM.txt
Untracked: data/Muscle_Skeletal.ACTN3.pm1Mb.RDS
Untracked: data/P.rds
Untracked: data/Thyroid.FMO2.pm1Mb.RDS
Untracked: data/bmass.HaemgenRBC2016.MAF01.Vs2.MergedDataSources.200kRanSubset.ChrBPMAFMarkerZScores.vs1.txt.gz
Untracked: data/bmass.HaemgenRBC2016.Vs2.NewSNPs.ZScores.hclust.vs1.txt
Untracked: data/bmass.HaemgenRBC2016.Vs2.PreviousSNPs.ZScores.hclust.vs1.txt
Untracked: data/eb_prepilot/
Untracked: data/finemap_data/fmo2.sim/b.txt
Untracked: data/finemap_data/fmo2.sim/dap_out.txt
Untracked: data/finemap_data/fmo2.sim/dap_out2.txt
Untracked: data/finemap_data/fmo2.sim/dap_out2_snp.txt
Untracked: data/finemap_data/fmo2.sim/dap_out_snp.txt
Untracked: data/finemap_data/fmo2.sim/data
Untracked: data/finemap_data/fmo2.sim/fmo2.sim.config
Untracked: data/finemap_data/fmo2.sim/fmo2.sim.k
Untracked: data/finemap_data/fmo2.sim/fmo2.sim.k4.config
Untracked: data/finemap_data/fmo2.sim/fmo2.sim.k4.snp
Untracked: data/finemap_data/fmo2.sim/fmo2.sim.ld
Untracked: data/finemap_data/fmo2.sim/fmo2.sim.snp
Untracked: data/finemap_data/fmo2.sim/fmo2.sim.z
Untracked: data/finemap_data/fmo2.sim/pos.txt
Untracked: data/logm.csv
Untracked: data/m.cd.RDS
Untracked: data/m.cdu.old.RDS
Untracked: data/m.new.cd.RDS
Untracked: data/m.old.cd.RDS
Untracked: data/mainbib.bib.old
Untracked: data/mat.csv
Untracked: data/mat.txt
Untracked: data/mat_new.csv
Untracked: data/matrix_lik.rds
Untracked: data/paintor_data/
Untracked: data/running_data_chris.csv
Untracked: data/running_data_matthew.csv
Untracked: data/temp.txt
Untracked: data/y.txt
Untracked: data/y_f.txt
Untracked: data/zscore_jointLCLs_m6AQTLs_susie_eQTLpruned.rds
Untracked: data/zscore_jointLCLs_random.rds
Untracked: explore_udi.R
Untracked: output/fit.k10.rds
Untracked: output/fit.nn.pbmc.purified.rds
Untracked: output/fit.nn.rds
Untracked: output/fit.nn.s.001.rds
Untracked: output/fit.nn.s.01.rds
Untracked: output/fit.nn.s.1.rds
Untracked: output/fit.nn.s.10.rds
Untracked: output/fit.snn.s.001.rds
Untracked: output/fit.snn.s.01.nninit.rds
Untracked: output/fit.snn.s.01.rds
Untracked: output/fit.varbvs.RDS
Untracked: output/fit2.nn.pbmc.purified.rds
Untracked: output/glmnet.fit.RDS
Untracked: output/snn07.txt
Untracked: output/snn34.txt
Untracked: output/test.bv.txt
Untracked: output/test.gamma.txt
Untracked: output/test.hyp.txt
Untracked: output/test.log.txt
Untracked: output/test.param.txt
Untracked: output/test2.bv.txt
Untracked: output/test2.gamma.txt
Untracked: output/test2.hyp.txt
Untracked: output/test2.log.txt
Untracked: output/test2.param.txt
Untracked: output/test3.bv.txt
Untracked: output/test3.gamma.txt
Untracked: output/test3.hyp.txt
Untracked: output/test3.log.txt
Untracked: output/test3.param.txt
Untracked: output/test4.bv.txt
Untracked: output/test4.gamma.txt
Untracked: output/test4.hyp.txt
Untracked: output/test4.log.txt
Untracked: output/test4.param.txt
Untracked: output/test5.bv.txt
Untracked: output/test5.gamma.txt
Untracked: output/test5.hyp.txt
Untracked: output/test5.log.txt
Untracked: output/test5.param.txt
Unstaged changes:
Modified: .gitignore
Modified: analysis/ebnm_binormal.Rmd
Modified: analysis/flashier_log1p.Rmd
Modified: analysis/flashier_sla_text.Rmd
Modified: analysis/logistic_z_scores.Rmd
Modified: analysis/mr_ash_pen.Rmd
Modified: analysis/nmu_em.Rmd
Modified: analysis/susie_flash.Rmd
Modified: misc.Rproj
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (analysis/ebpower.Rmd) and HTML
(docs/ebpower.html) files. If you’ve configured a remote
Git repository (see ?wflow_git_remote), click on the
hyperlinks in the table below to view the files as they were in that
past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | fe667eb | Matthew Stephens | 2025-03-16 | workflowr::wflow_publish("ebpower.Rmd") |
| html | 9006efb | Matthew Stephens | 2025-03-15 | Build site. |
| Rmd | 4c2850b | Matthew Stephens | 2025-03-15 | workflowr::wflow_publish("ebpower.Rmd") |
| html | 7c17b31 | Matthew Stephens | 2025-03-15 | Build site. |
| Rmd | 0608521 | Matthew Stephens | 2025-03-15 | workflowr::wflow_publish("ebpower.Rmd") |
| html | 2956a39 | Matthew Stephens | 2025-03-12 | Build site. |
| Rmd | 6aa4dfe | Matthew Stephens | 2025-03-12 | workflowr::wflow_publish("analysis/ebpower.Rmd") |
# load the ebnm library and define an ebnm_binormal function
library("ebnm")
dbinormal = function (x,s,s0,lambda,log=TRUE){
pi0 = 0.5
pi1 = 0.5
s2 = s^2
s02 = s0^2
l0 = dnorm(x,0,sqrt(lambda^2 * s02 + s2),log=TRUE)
l1 = dnorm(x,lambda,sqrt(lambda^2 * s02 + s2),log=TRUE)
logsum = log(pi0*exp(l0) + pi1*exp(l1))
m = pmax(l0,l1)
logsum = m + log(pi0*exp(l0-m) + pi1*exp(l1-m))
if (log) return(sum(logsum))
else return(exp(sum(logsum)))
}
ebnm_binormal = function(x,s){
x = drop(x)
s0 = 0.01
lambda = optimize(function(lambda){-dbinormal(x,s,s0,lambda,log=TRUE)},
lower = 0, upper = max(x))$minimum
g = ashr::normalmix(pi=c(0.5,0.5), mean=c(0,lambda), sd=c(lambda * s0,lambda * s0))
postmean = ashr::postmean(g,ashr::set_data(x,s))
postsd = ashr::postsd(g,ashr::set_data(x,s))
return(list(g = g, posterior = data.frame(mean=postmean,sd=postsd)))
}Introduction
I want to implement an EB version of the power update for a symmetric data matrix S. I haven’t implemented an update to the residual error yet
Rank 1
# This is just the regular power update, used for initialization
power_update_r1 = function(S,v){
newv = drop(S %*% v)
if(!all(newv==0))
v = newv/sqrt(sum(newv^2))
return(v)
}
eb_power_update_r1 = function(S,v,ebnm_fn,sigma){
newv = drop(S %*% v)
if(!all(newv==0))
v = ebnm_fn(newv, sigma)$posterior$mean
if(!all(v==0))
v = v/sqrt(sum(v^2))
return(v)
}Simulate data under a tree model (with very small errors)
set.seed(1)
n = 40
x = cbind(c(rep(1,n),rep(0,n)), c(rep(0,n),rep(1,n)), c(rep(1,n/2),rep(0,3*n/2)), c(rep(0,n/2), rep(1,n/2), rep(0,n)), c(rep(0,n),rep(1,n/2),rep(0,n/2)), c(rep(0,3*n/2),rep(1,n/2)))
E = matrix(0.01*rexp(2*n*2*n),nrow=2*n)
E = E+t(E) #symmetric errors
S = x %*% diag(c(1,1,1,1,1,1)) %*% t(x) + E
image(S)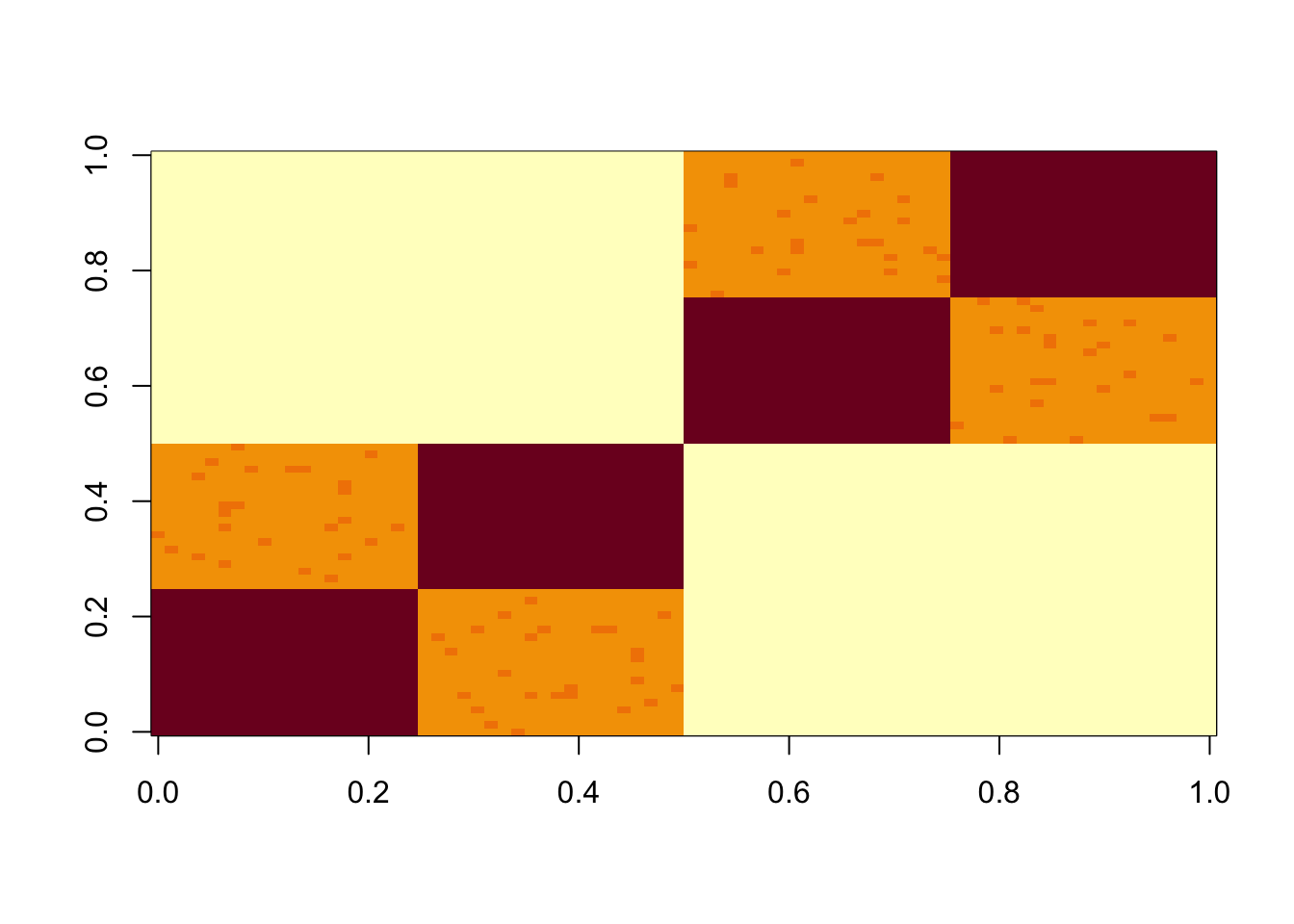
Check the first PC. It is essentially the constant vector.
pc1 = cbind(svd(S)$v[,1])
plot(-pc1,ylim=c(0,0.16))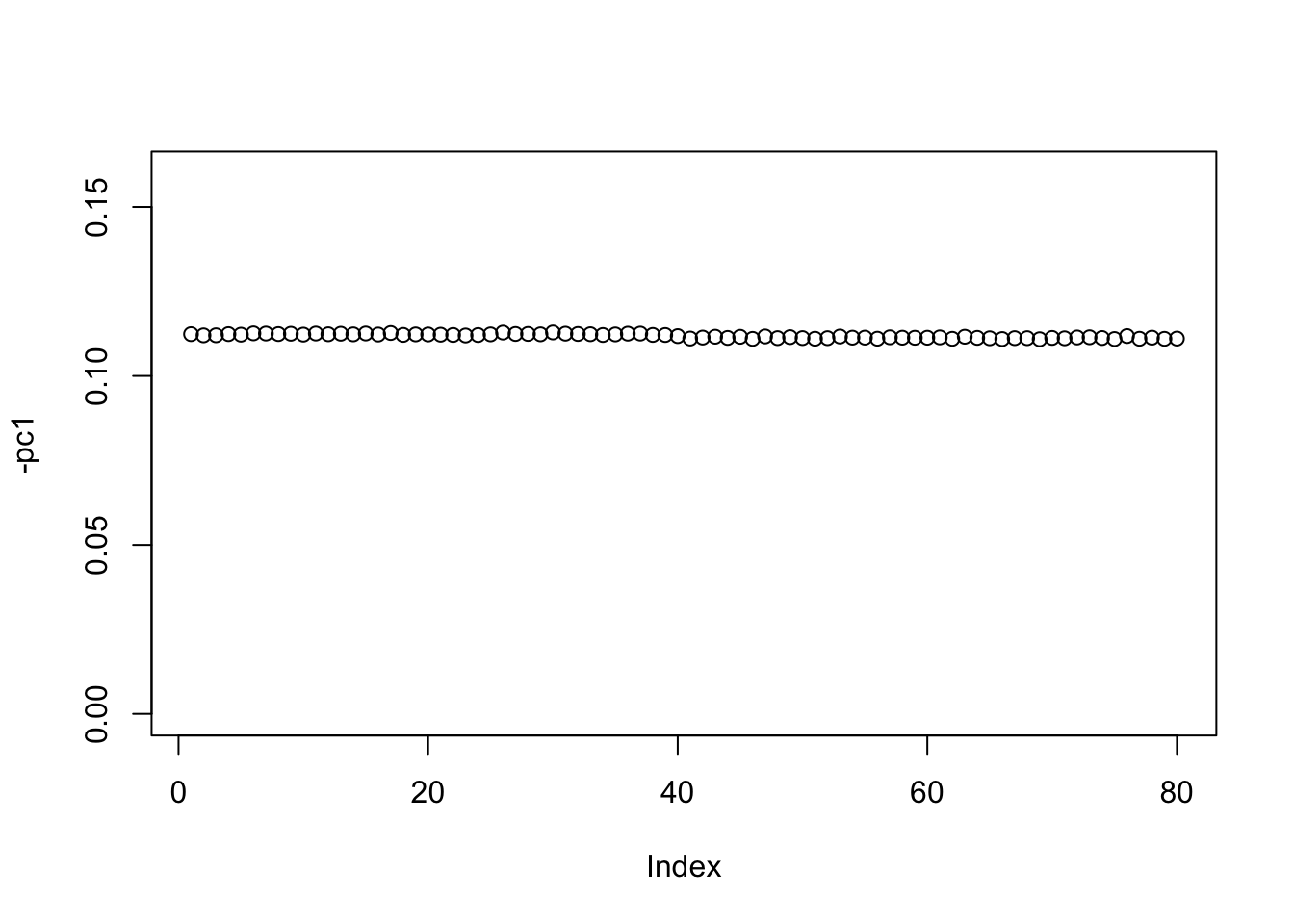
If one uses a completely random initialization then the eb method will often (but not always) zero things out completely on the first iteration:
set.seed(1)
v = cbind(rnorm(2*n))
v= v/sqrt(sum(v^2))
sigma = sqrt(mean(S^2))
v = eb_power_update_r1(S,v,ebnm_point_laplace,sigma)
plot(v,ylim=c(-0.25,0.25))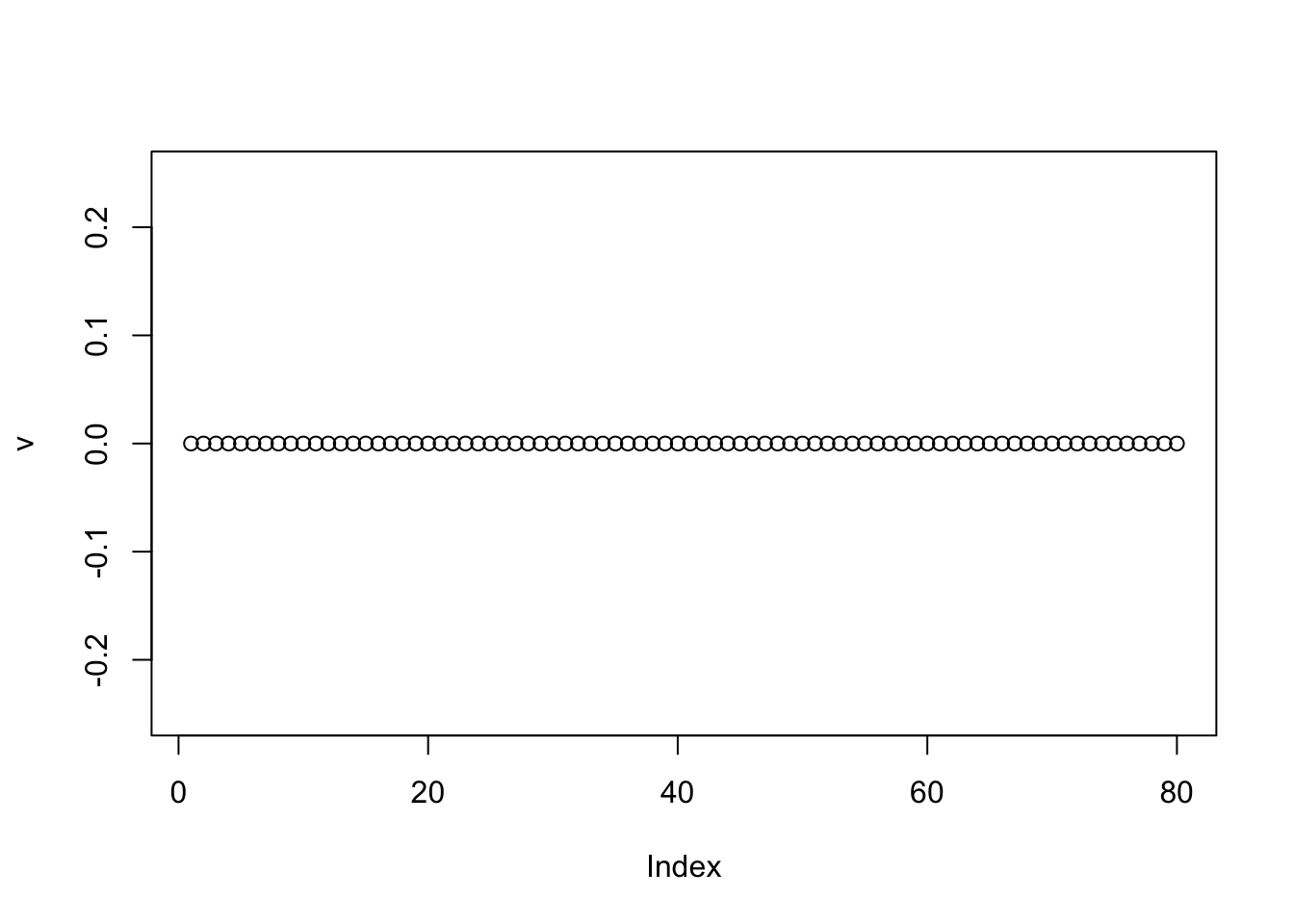
We can help avoid this here by doing a single iteration of the power method before moving to EB (because the matrix is very low rank the single iteration moves us quickly to the right subspace):
set.seed(1)
v = cbind(rnorm(2*n))
v= v/sqrt(sum(v^2))
sigma = sqrt(mean(S^2))
v = power_update_r1(S,v)
plot(v,ylim=c(-0.25,0.25))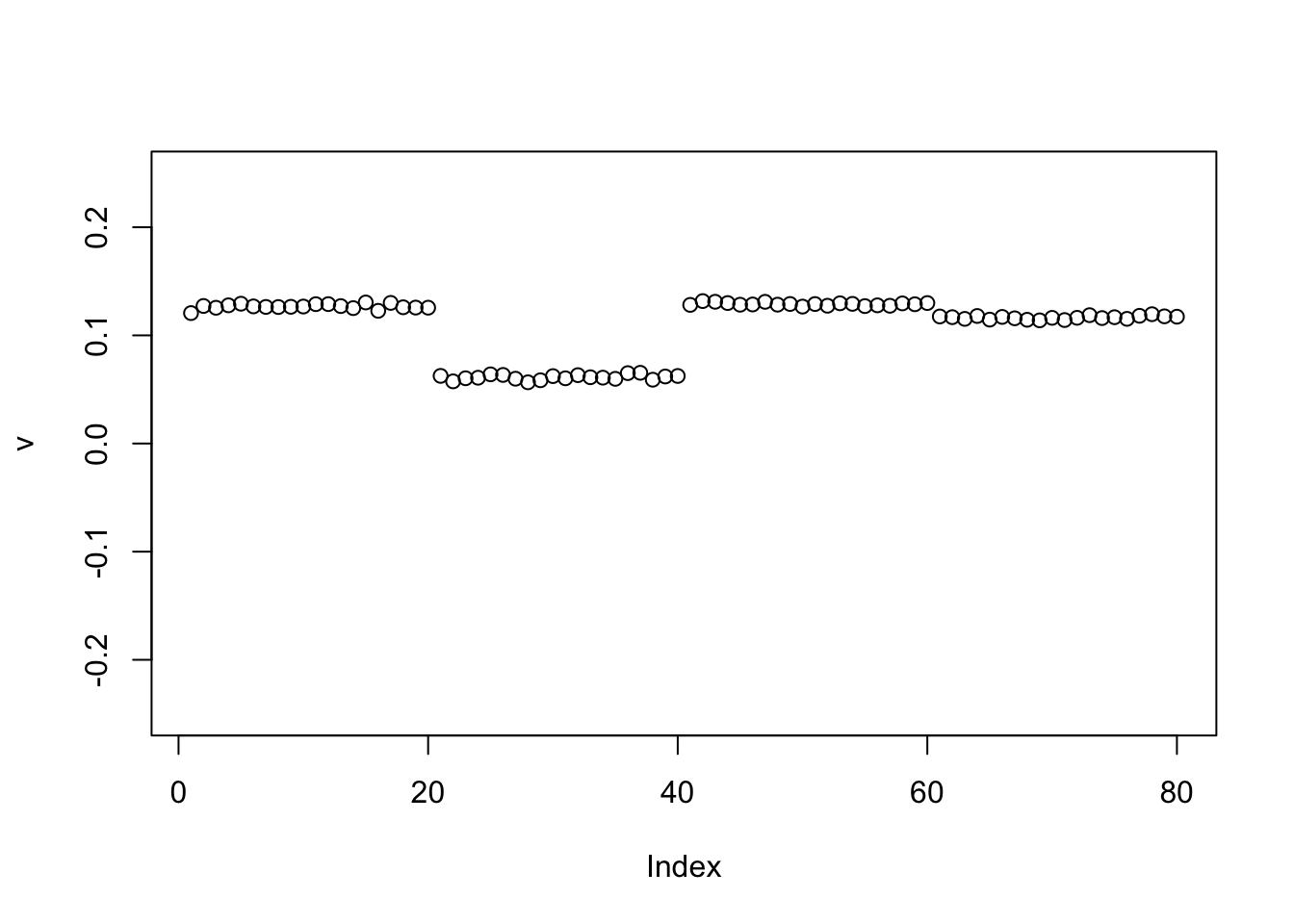
for(i in 1:100){
v = eb_power_update_r1(S,v,ebnm_point_laplace,sigma)
}
plot(v,ylim=c(-0.25,0.25))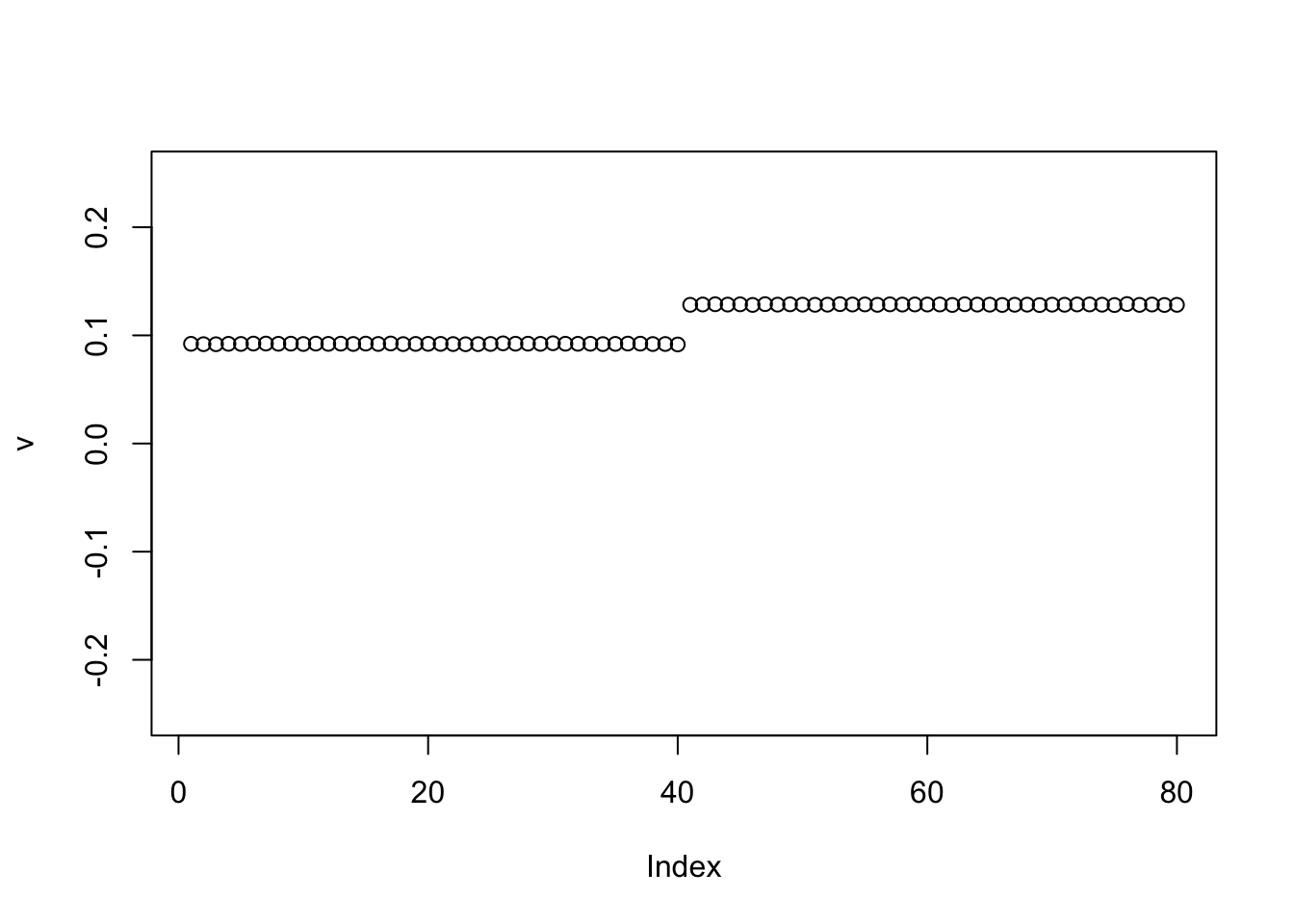
However, if we use a different random init, it can find the first split of the tree (here a 0-1 split rather than +-1 split).
set.seed(4)
v = cbind(rnorm(2*n))
v= v/sqrt(sum(v^2))
sigma = sqrt(mean(S^2))
v = power_update_r1(S,v)
plot(v,ylim=c(-0.25,0.25))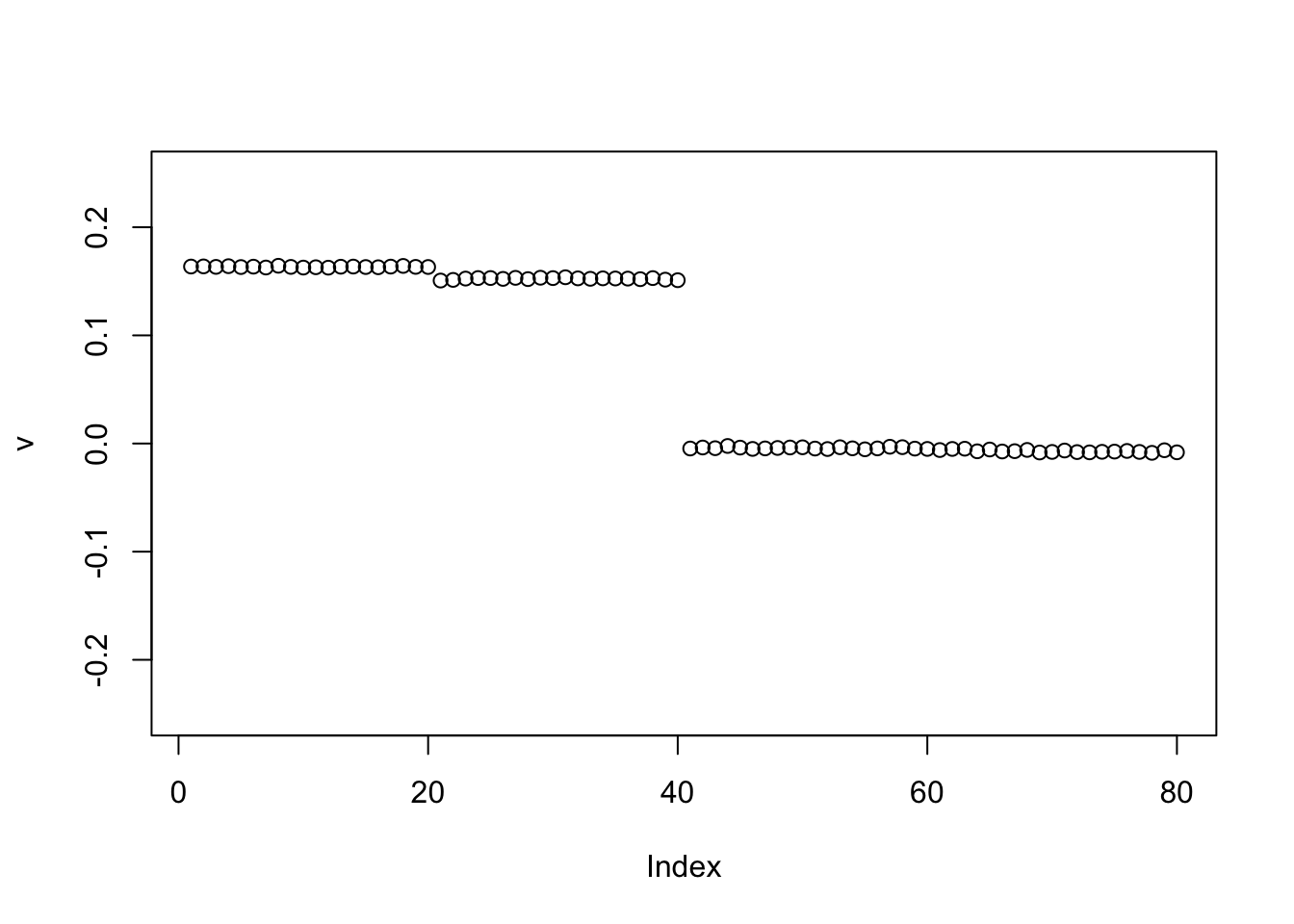
for(i in 1:100){
v = eb_power_update_r1(S,v,ebnm_point_laplace,sigma)
}
plot(v,ylim=c(-0.25,0.25))
abline(h=0)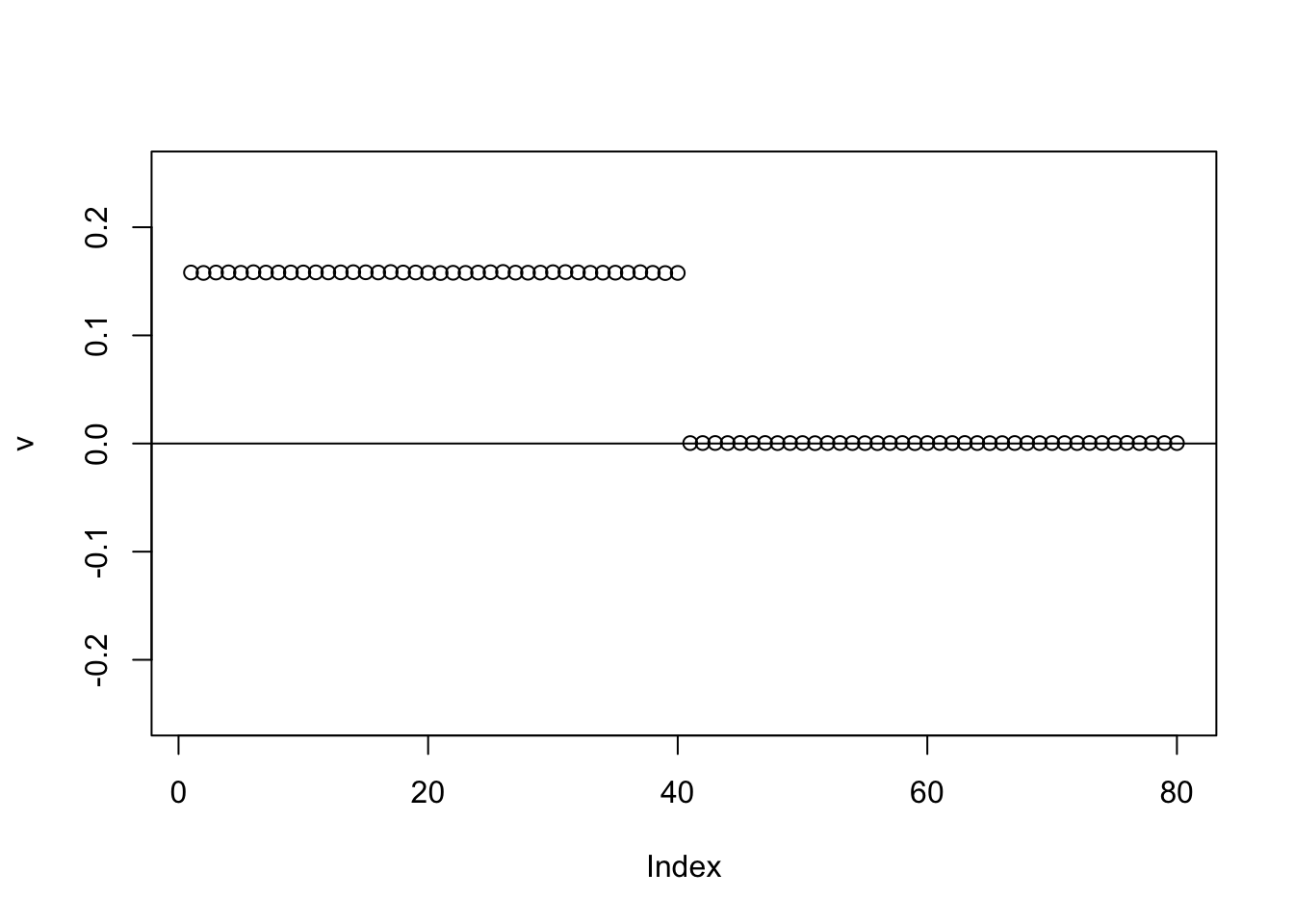
If we initialize to the first PC, it finds something close to the first PC (a bit more split).
v = -pc1
sigma = sqrt(mean(S^2))
for(i in 1:100){
v = eb_power_update_r1(S,v,ebnm_point_laplace,sigma)
}
plot(v, ylim=c(-0.25,0.25))
If we initialize to the (positive version of) the first PC, point-exponential performs similarly (we use the negative because the first PC here is negative).
v = -pc1
sigma = sqrt(mean(S^2))
for(i in 1:100){
v = eb_power_update_r1(S,v,ebnm_point_exponential,sigma)
}
plot(v, ylim=c(-0.25,0.25))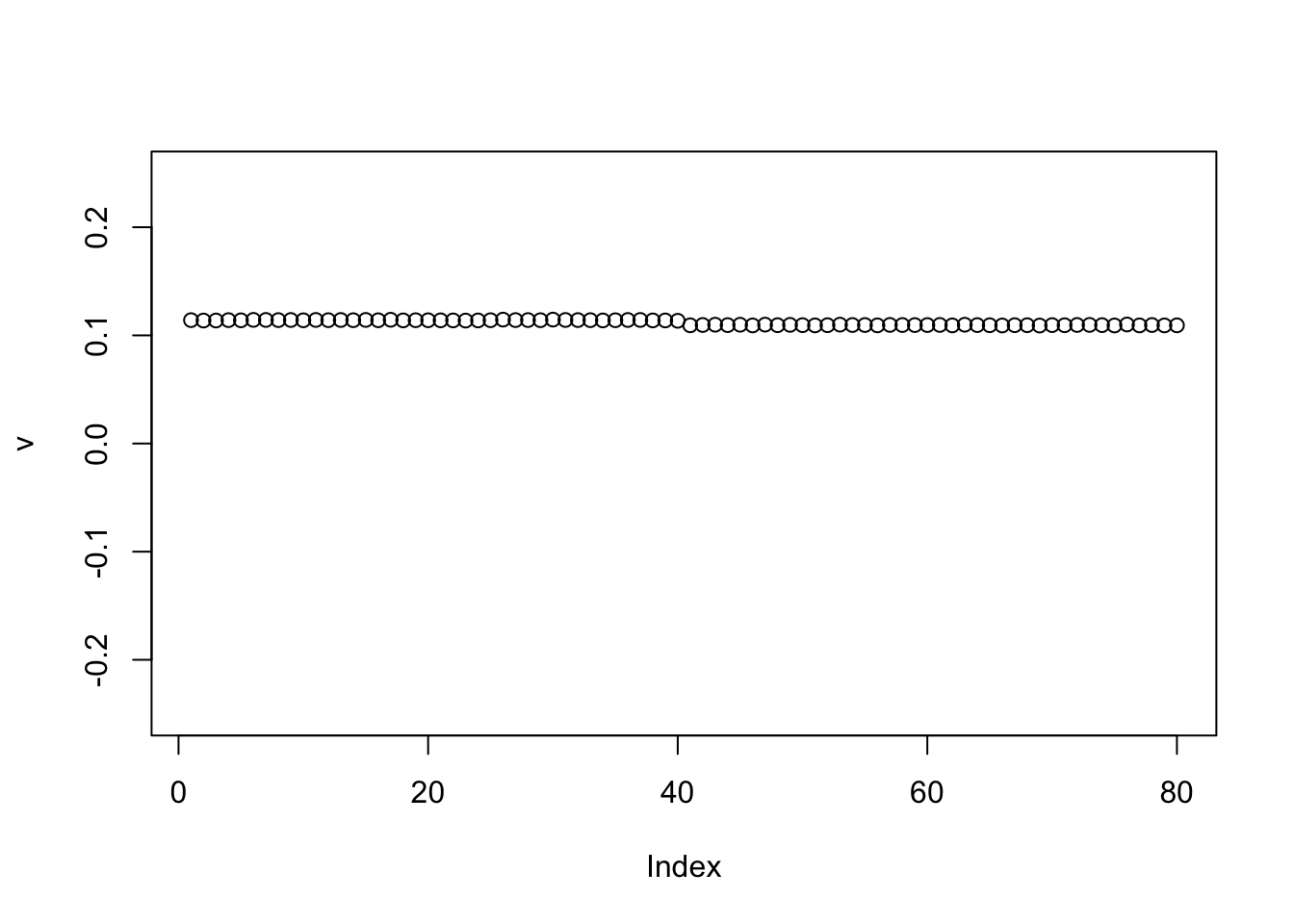
Try further running binormal - it levels things out.
for(i in 1:100){
v = eb_power_update_r1(S,v,ebnm_binormal,sigma)
}
plot(v, ylim=c(-0.25,0.25))
Rank K
# model is S \sim VDV' + E with eb prior on V
eb_power_update = function(S,v,d,ebnm_fn){
K = ncol(v)
sigma2=mean((S-v %*% diag(d,nrow=length(d)) %*% t(v))^2)
for(k in 1:K){
U = v[,-k,drop=FALSE]
D = diag(d[-k],nrow=length(d[-k]))
newv = (S %*% v[,k,drop=FALSE] - U %*% D %*% t(U) %*% v[,k,drop=FALSE] )
if(!all(newv==0)){
fit.ebnm = ebnm_fn(newv,sqrt(sigma2))
newv = fit.ebnm$posterior$mean
if(!all(newv==0)){
newv = newv/sqrt(sum(newv^2 + fit.ebnm$posterior$sd^2))
}
}
v[,k] = newv
d[k] = t(v[,k]) %*% S %*% v[,k] - t(v[,k]) %*% U %*% D %*% t(U) %*% v[,k]
}
return(list(v=v,d=d))
}
#helper function
compute_sqerr = function(S,fit){
sum((S-fit$v %*% diag(fit$d,nrow=length(fit$d)) %*% t(fit$v))^2)
}
# a random initialization
random_init = function(S,K,nonneg = FALSE){
n = nrow(S)
v = matrix(nrow=n,ncol=K)
for(k in 1:K){
v[,k] = cbind(rnorm(n)) # initialize v
if(nonneg)
v[,k] = pmax(v[,k],0)
v[,k] = v[,k]/sum(v[,k]^2)
}
d = rep(1e-8,K)
return(list(v=v,d=d))
}Tree structured data
Simulate data under a tree model (with very small errors)
set.seed(1)
n = 40
x = cbind(c(rep(1,n),rep(0,n)), c(rep(0,n),rep(1,n)), c(rep(1,n/2),rep(0,3*n/2)), c(rep(0,n/2), rep(1,n/2), rep(0,n)), c(rep(0,n),rep(1,n/2),rep(0,n/2)), c(rep(0,3*n/2),rep(1,n/2)))
E = matrix(0.01*rexp(2*n*2*n),nrow=2*n)
E = E+t(E) #symmetric errors
S = x %*% diag(c(1,1,1,1,1,1)) %*% t(x) + E
image(S)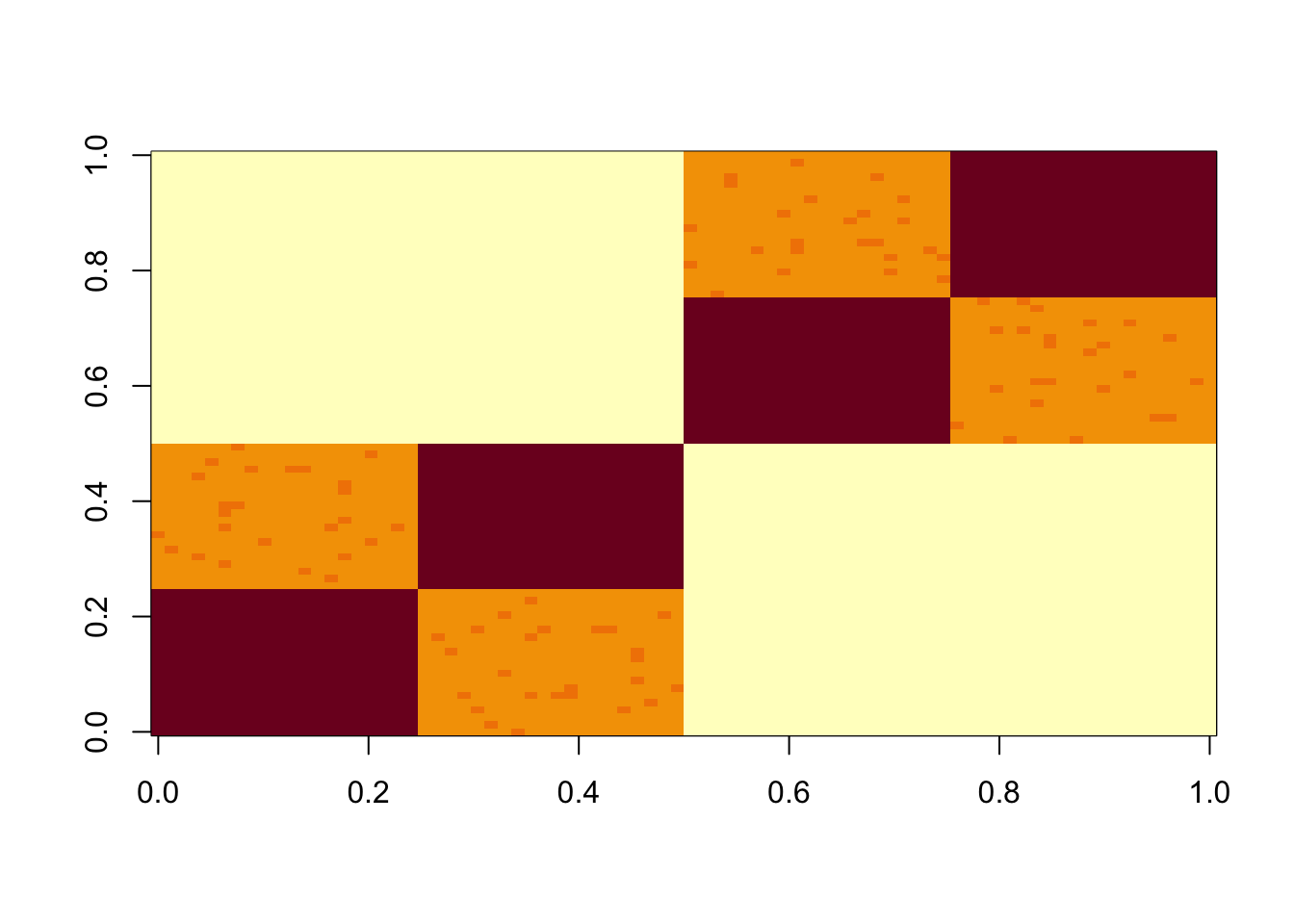
Run multiple factors
Here I run with \(K=9\) and point-exponential. It finds a rank 4 solution, essentially zeroing out the other 5. One can compare this with non-negative without the EB approach here.
set.seed(2)
fit = random_init(S,9,nonneg=TRUE)
err = rep(0,10)
err[1] = sum((S-fit$v %*% diag(fit$d) %*% t(fit$v))^2)
for(i in 2:100){
fit = eb_power_update(S,fit$v,fit$d,ebnm_point_exponential)
err[i] = sum((S-fit$v %*% diag(fit$d) %*% t(fit$v))^2)
}
plot(err)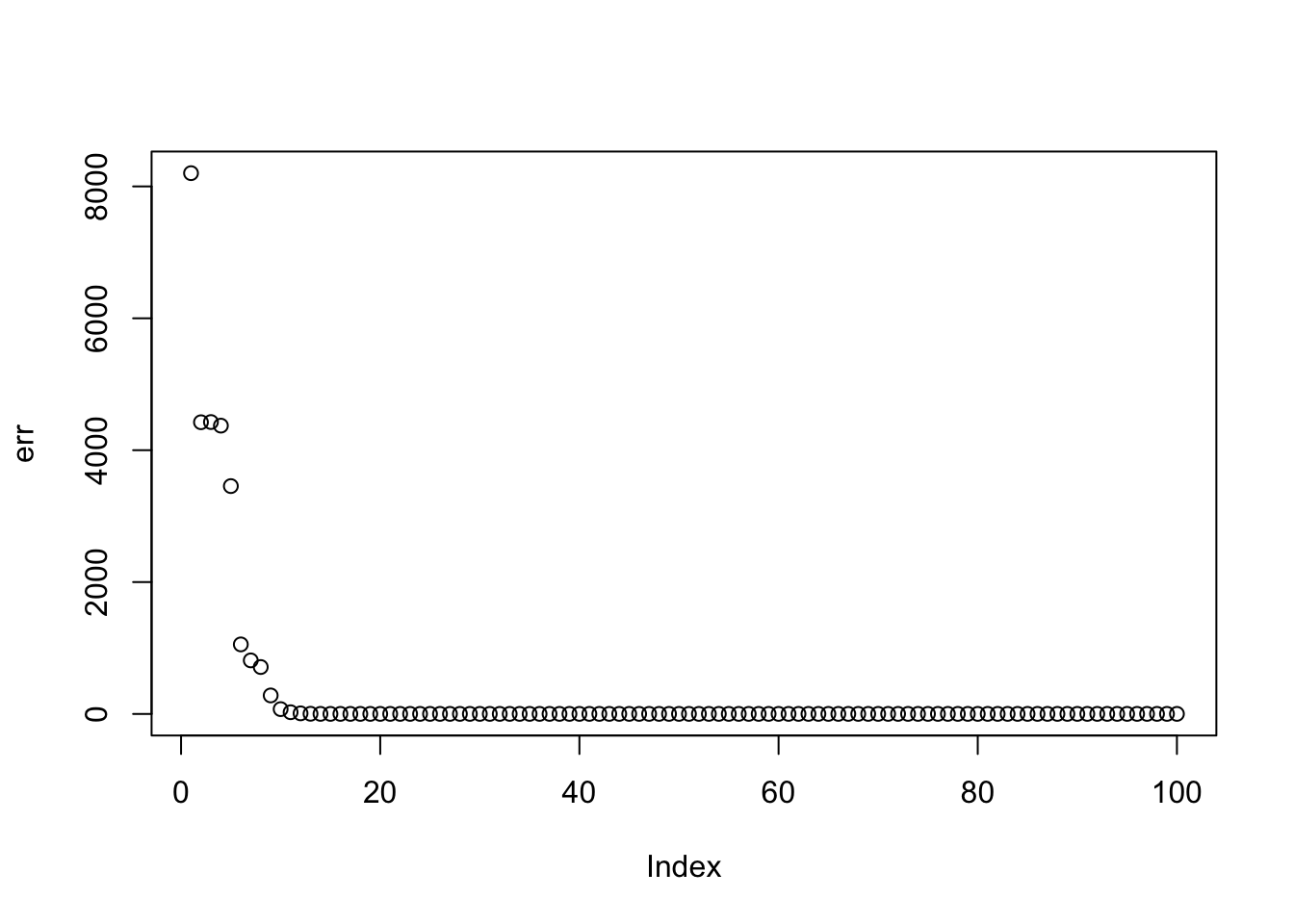
par(mfcol=c(3,3),mai=rep(0.3,4))
for(i in 1:9){plot(fit$v[,i],main=paste0(trunc(fit$d[i])))}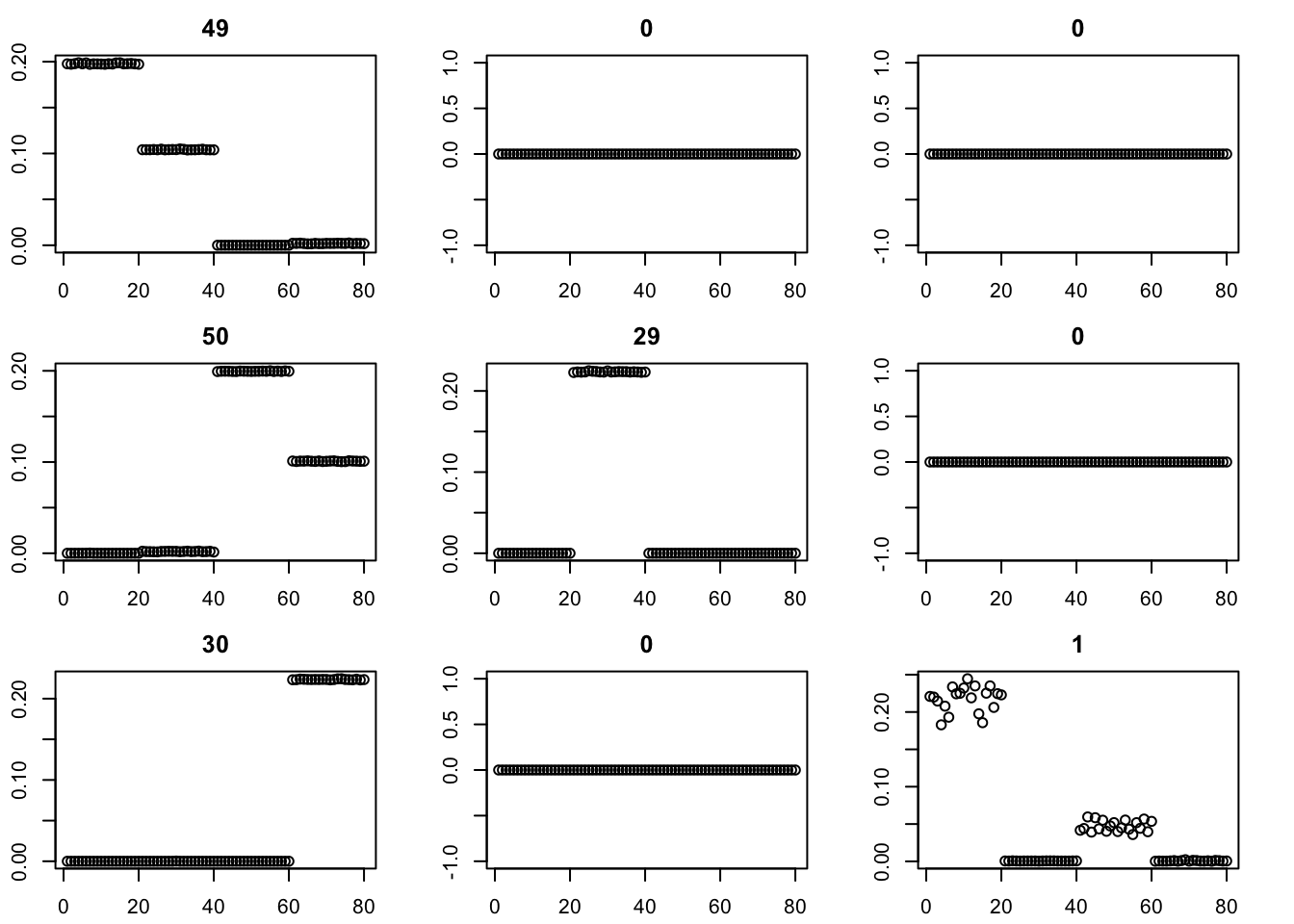
Here I try the generalized binary prior. So far I’m finding this does not work well, especially if started from random starting point. I debugged and found that what happens is that generally the v are bounded away from 0. So the gb prior puts all its weight on the non-null normal component and does not shrink anything. (Is it worth using a laplace for the non-null component?) The point exponential does not have that problem - it shrinks the smallest values towards 0, and eventually gets to a point where everything is 0. It seems clear that using the gb prior from random initialization is not going to work.
set.seed(2)
fit = random_init(S,9, nonneg=TRUE)
err = rep(0,10)
err[1] = sum((S-fit$v %*% diag(fit$d) %*% t(fit$v))^2)
for(i in 2:10){
fit = eb_power_update(S,fit$v,fit$d,ebnm_generalized_binary)
err[i] = sum((S-fit$v %*% diag(fit$d) %*% t(fit$v))^2)
}
plot(err)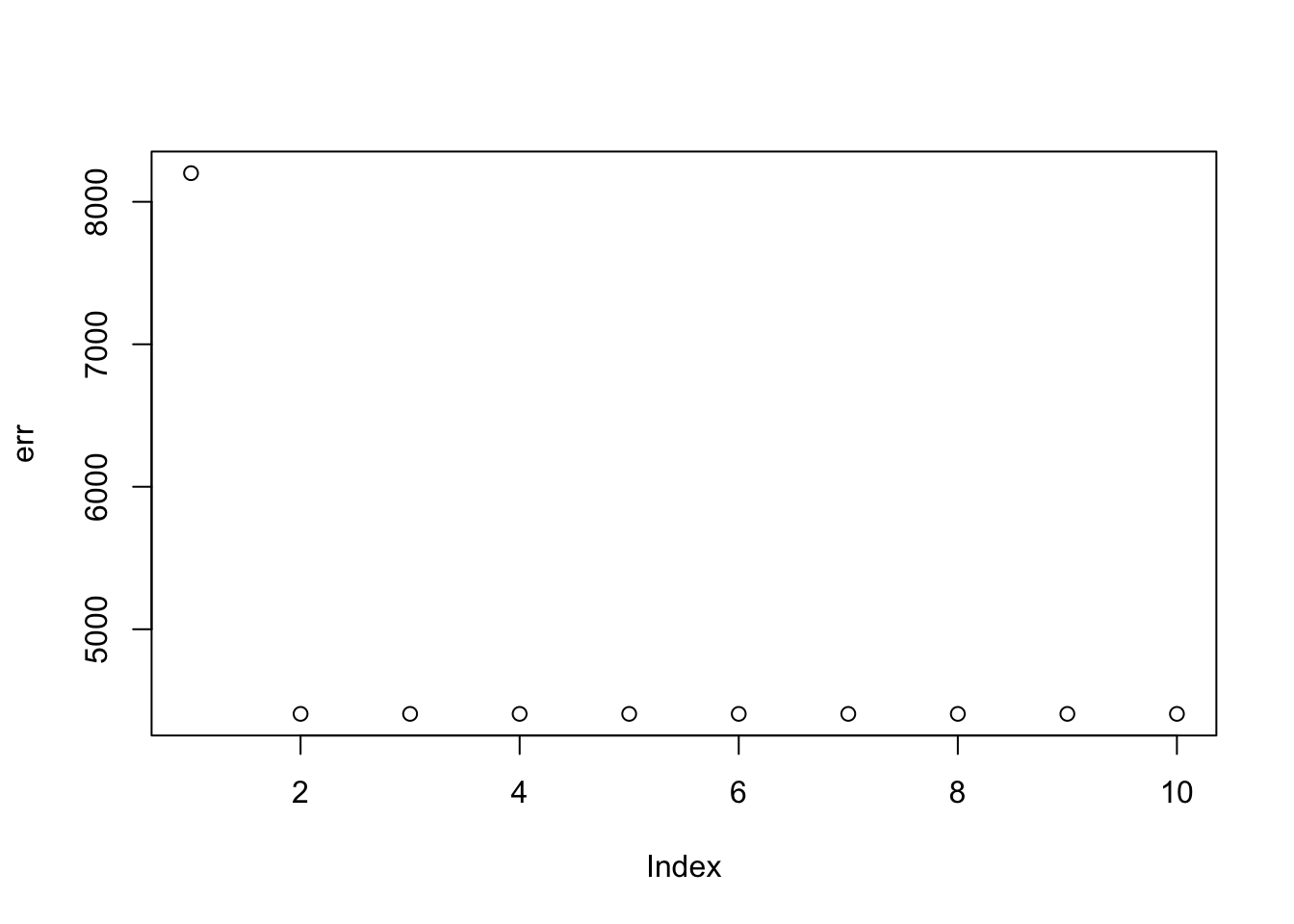
par(mfcol=c(3,3),mai=rep(0.3,4))
for(i in 1:9){plot(fit$v[,i],main=paste0(trunc(fit$d[i])))}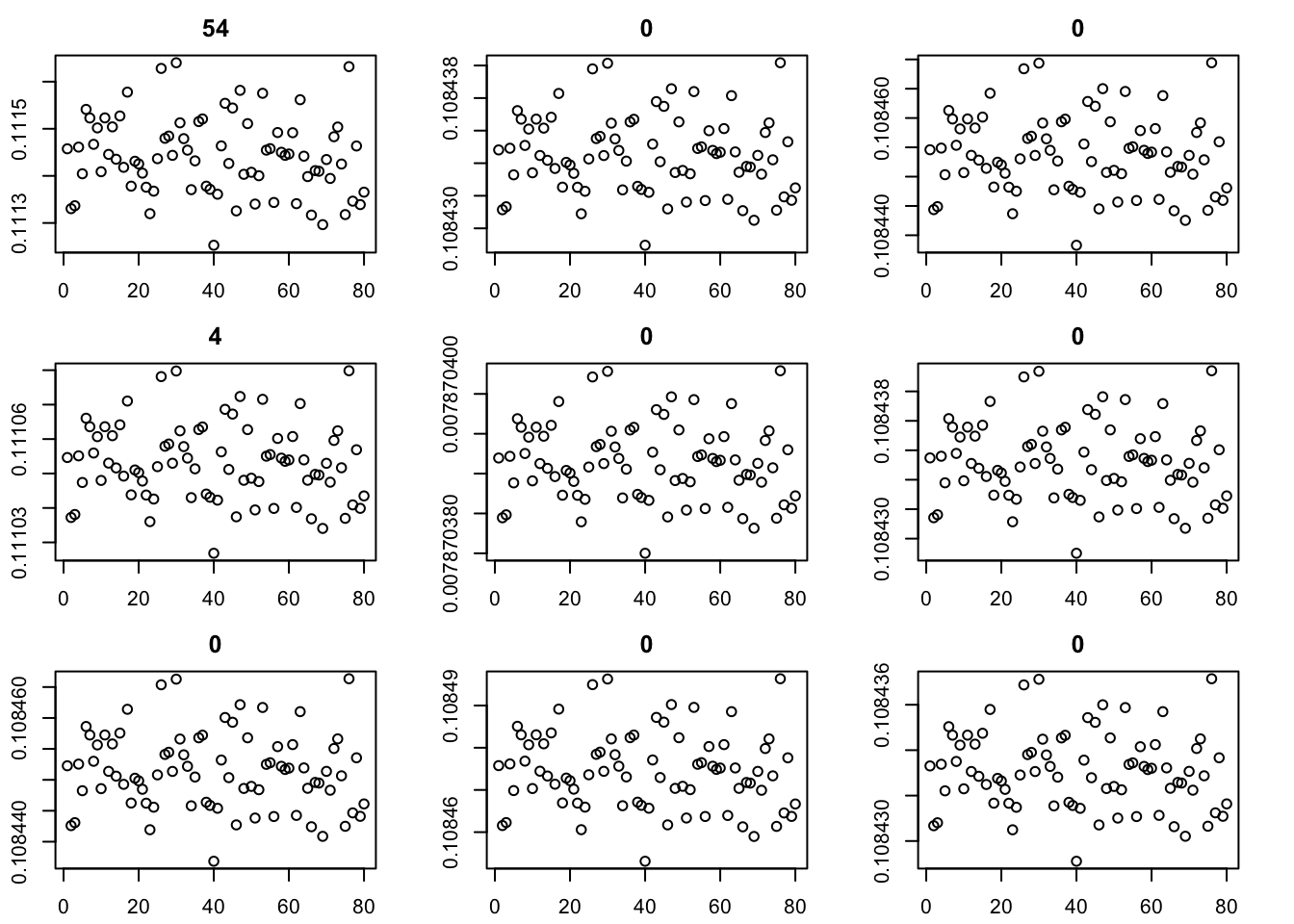
Here we try initializing GB with point-exponential. It only changes the fit very little.
set.seed(2)
fit = random_init(S,9,nonneg=TRUE)
for(i in 1:100){
fit = eb_power_update(S,fit$v,fit$d,ebnm_point_exponential)
}
par(mfcol=c(3,3),mai=rep(0.3,4))
for(i in 1:9){plot(fit$v[,i],main=paste0(trunc(fit$d[i])))}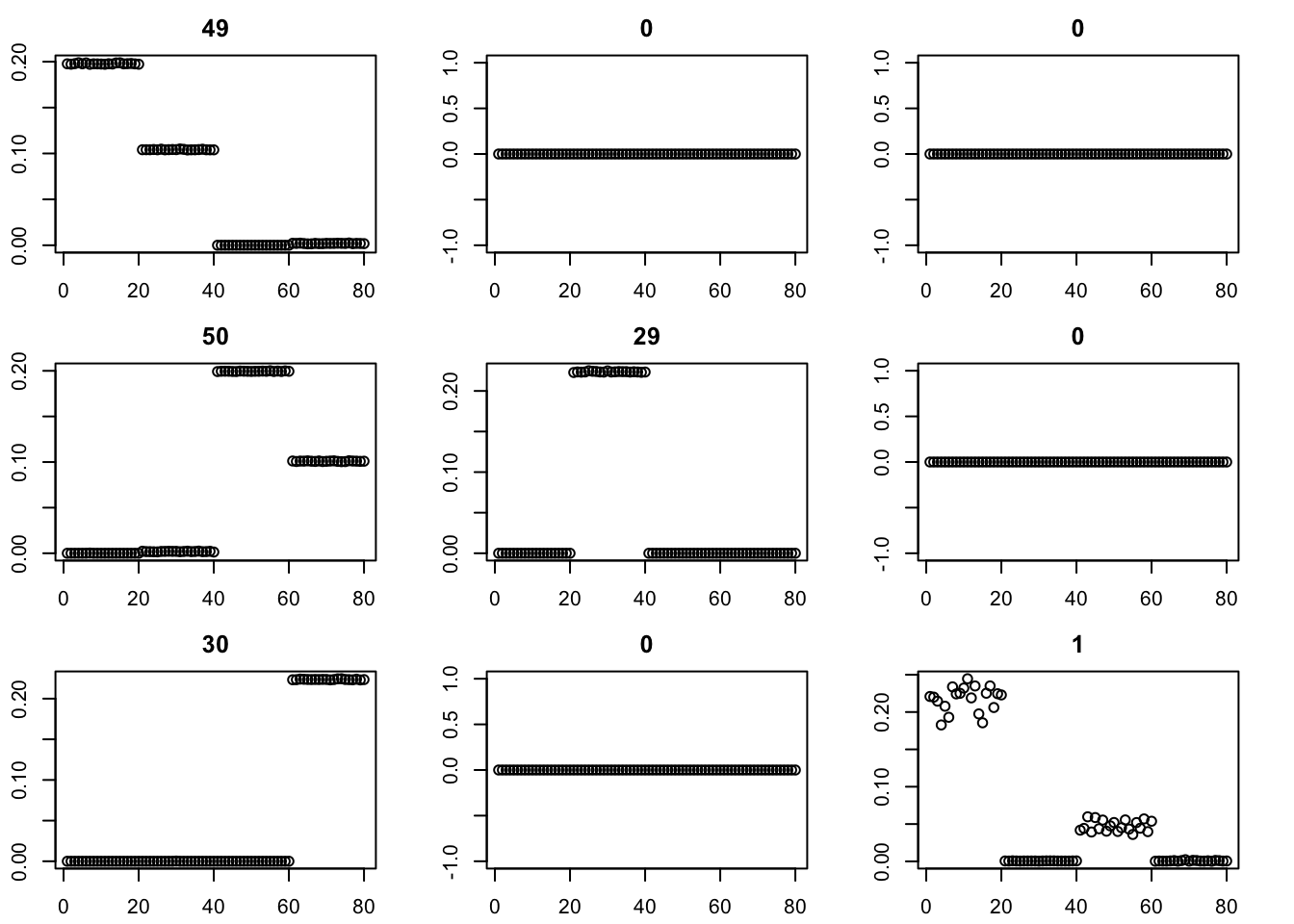
for(i in 1:100){
fit = eb_power_update(S,fit$v,fit$d,ebnm_generalized_binary)
}
par(mfcol=c(3,3),mai=rep(0.3,4))
for(i in 1:9){plot(fit$v[,i],main=paste0(trunc(fit$d[i])))}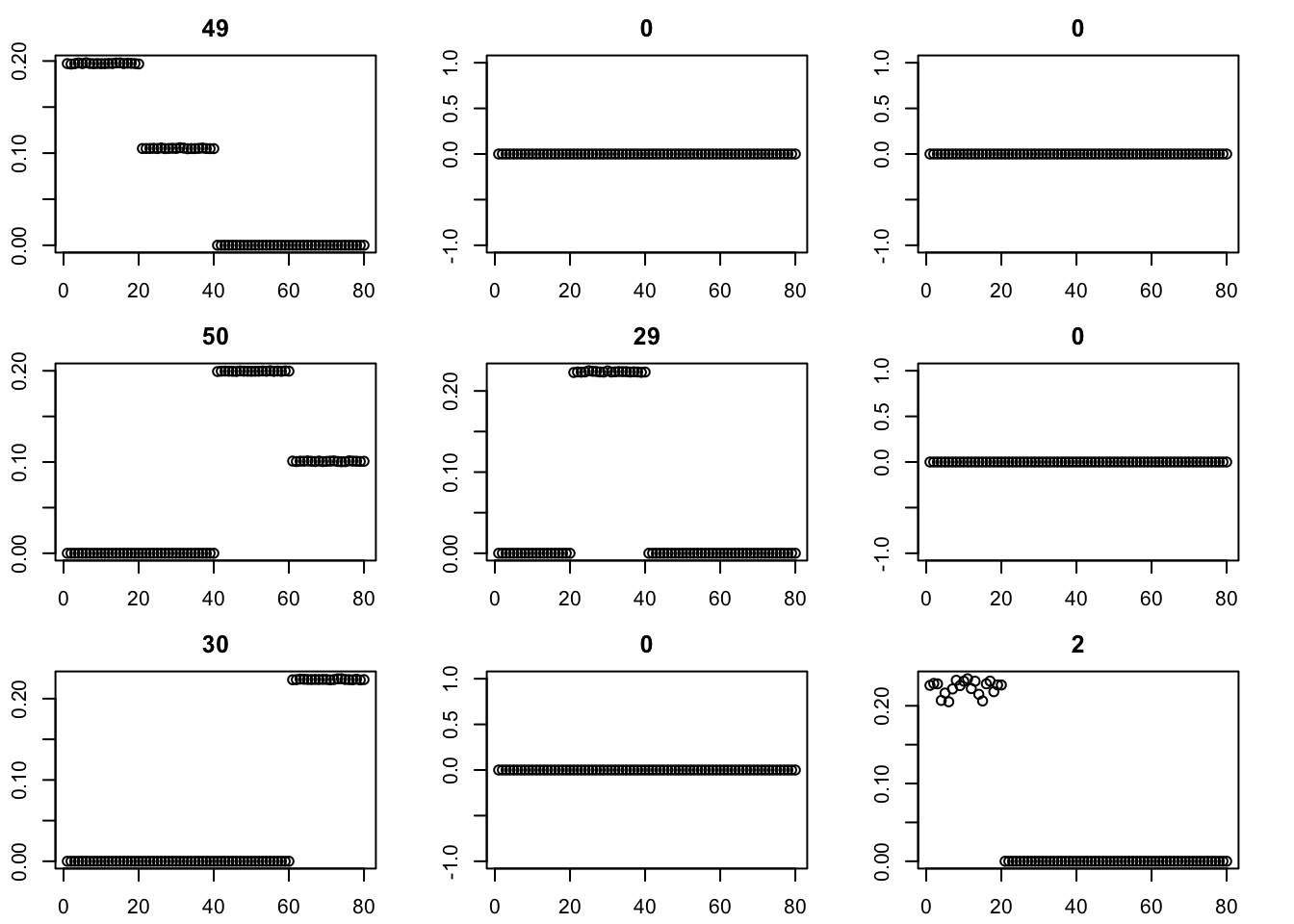
| Version | Author | Date |
|---|---|---|
| 9006efb | Matthew Stephens | 2025-03-15 |
The binormal prior provides a more bimodal solution. (Note: although not seen here, I have seen issues with it including a factor that puts two non-neighboring populations together.)
for(i in 1:100){
fit = eb_power_update(S,fit$v,fit$d,ebnm_binormal)
err[i] = compute_sqerr(S,fit)
}
par(mfcol=c(3,3),mai=rep(0.3,4))
for(i in 1:9){plot(fit$v[,i],main=paste0(trunc(fit$d[i])))}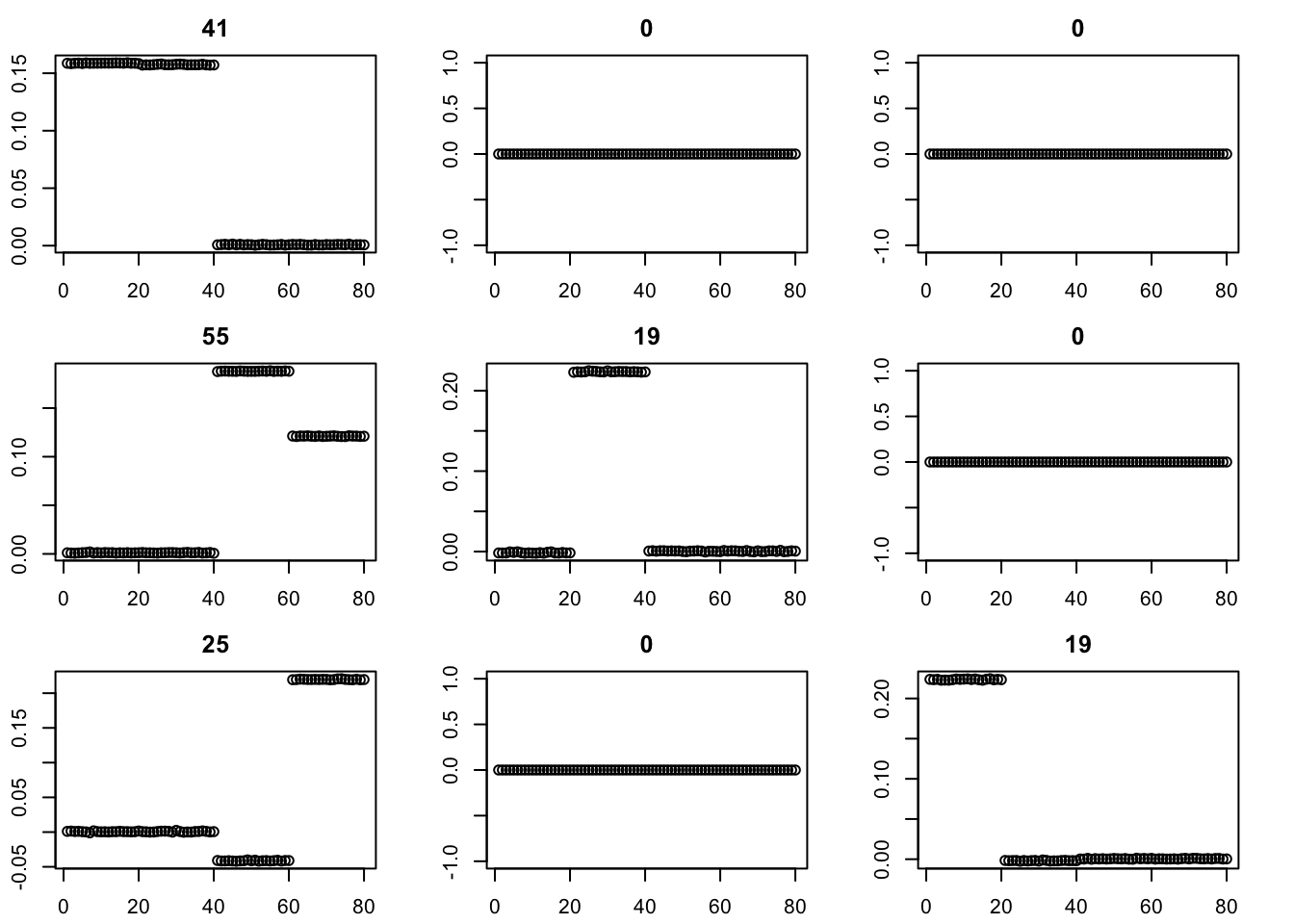
| Version | Author | Date |
|---|---|---|
| 7c17b31 | Matthew Stephens | 2025-03-15 |
for(i in 1:100){
fit = eb_power_update(S,fit$v,fit$d,ebnm_binormal)
}
par(mfcol=c(3,3),mai=rep(0.3,4))
for(i in 1:9){plot(fit$v[,i],main=paste0(trunc(fit$d[i])))}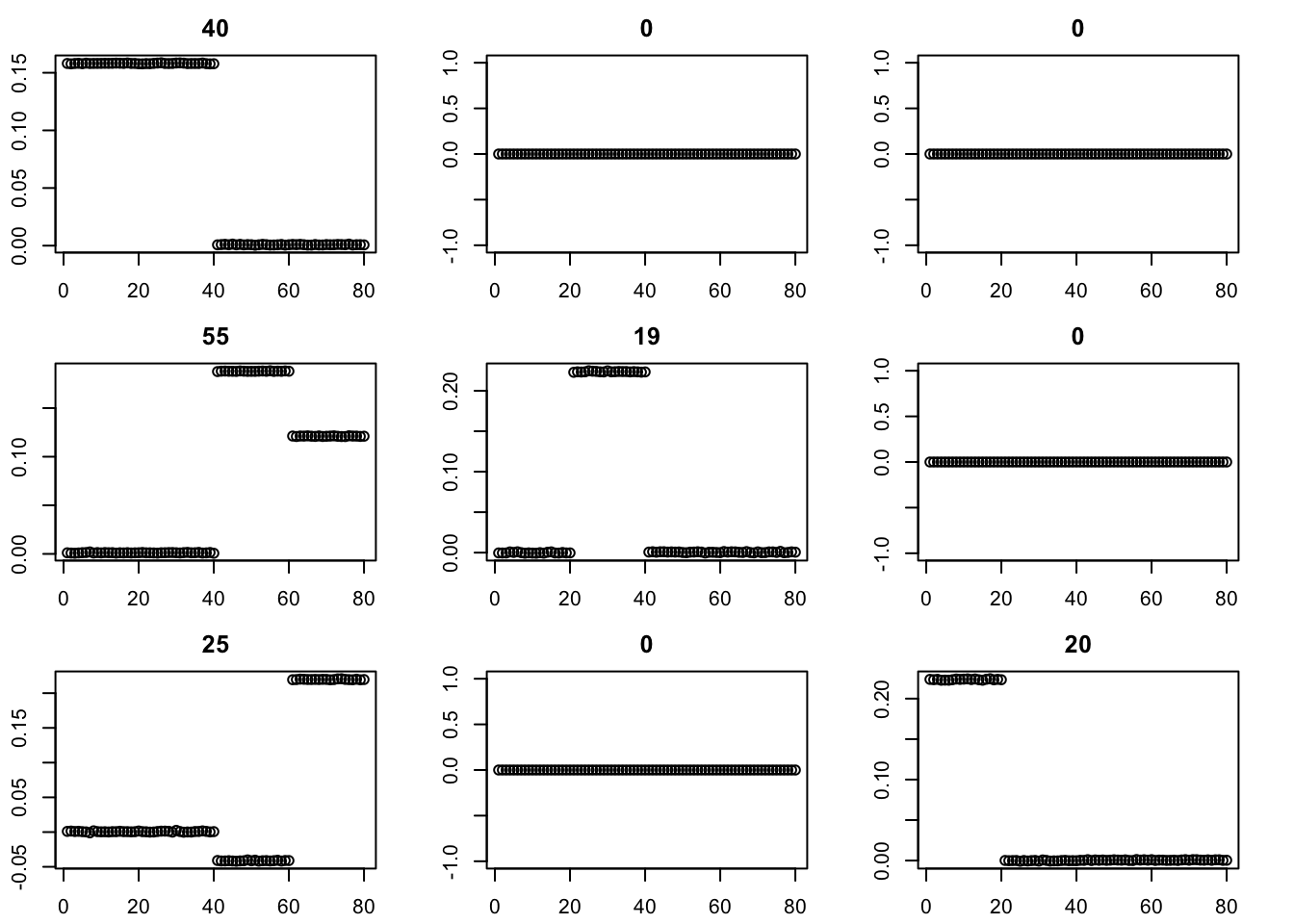
SVD initialization
I’m going to try initializing with SVD, then running point-laplace, then running GB, similar to the strategy in the GBCD paper. (Note: here I initialize with the svd values for d; an alternative is to set these to be very small and just use the v from svd to initialize.)
set.seed(2)
S.svd = svd(S)
fit = list(v=S.svd$u[,1:4],d=S.svd$d[1:4]) #rep(1e-8,4)) #init d to be very small
err = rep(0,10)
err[1] = compute_sqerr(S,fit)
for(i in 2:100){
fit = eb_power_update(S,fit$v,fit$d,ebnm_point_laplace)
err[i] = compute_sqerr(S,fit)
}
par(mfcol=c(3,3),mai=rep(0.3,4))
for(i in 1:4){plot(fit$v[,i],main=paste0(trunc(fit$d[i])))}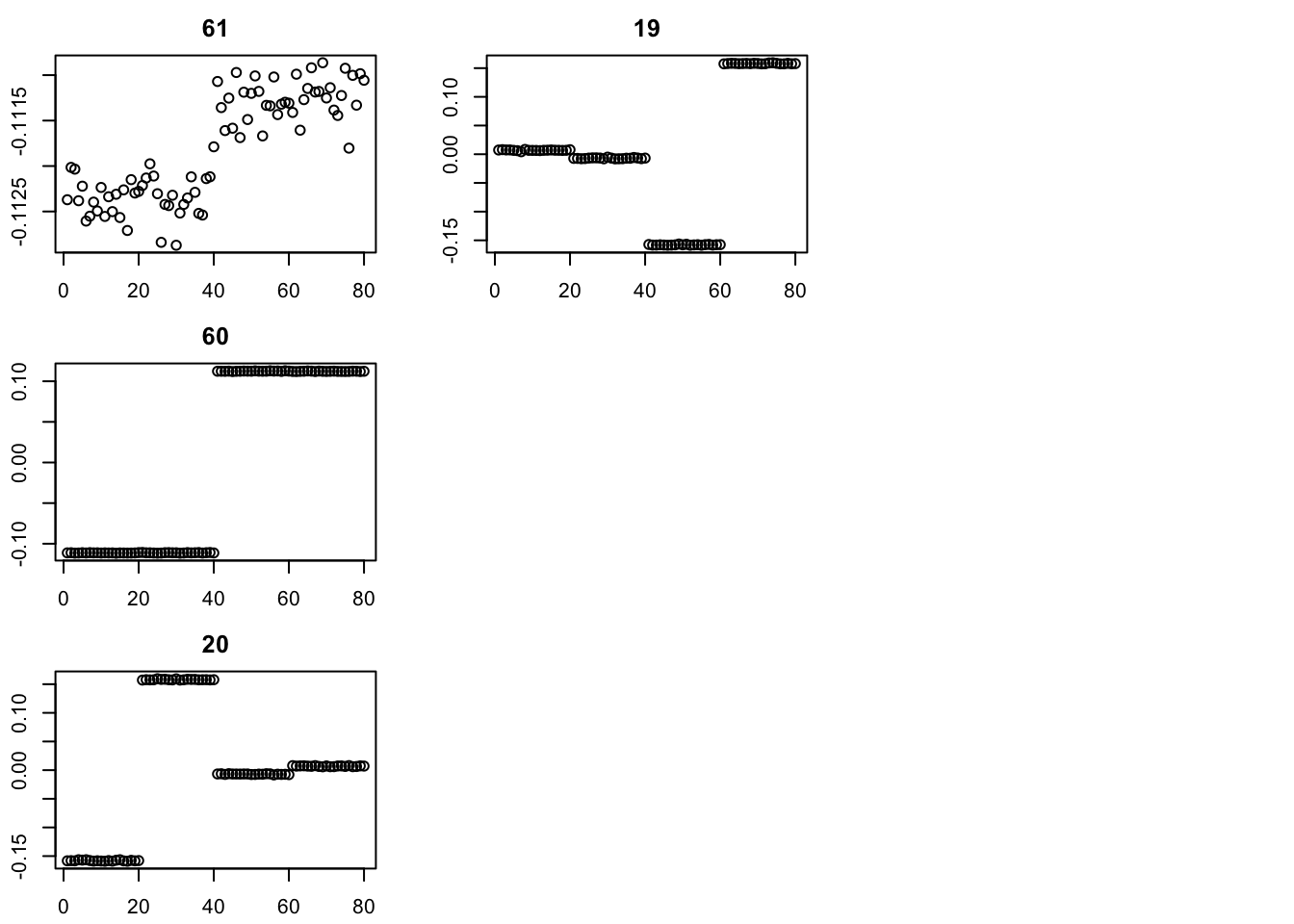
| Version | Author | Date |
|---|---|---|
| 7c17b31 | Matthew Stephens | 2025-03-15 |
split_v = function(v){
v = cbind(pmax(v,0),pmax(-v,0))
}
fit$v = split_v(fit$v)
fit$d= rep(fit$d/2,2)
for(i in 2:100){
fit = eb_power_update(S,fit$v,fit$d,ebnm_point_exponential)
err[i] = compute_sqerr(S,fit)
}
par(mfcol=c(3,3),mai=rep(0.3,4))
for(i in 1:8){plot(fit$v[,i],main=paste0(trunc(fit$d[i])))}
fit.pe = fit # save for later use
for(i in 2:200){
fit = eb_power_update(S,fit$v,fit$d,ebnm_generalized_binary)
err[i] = compute_sqerr(S,fit)
}
par(mfcol=c(3,3),mai=rep(0.3,4))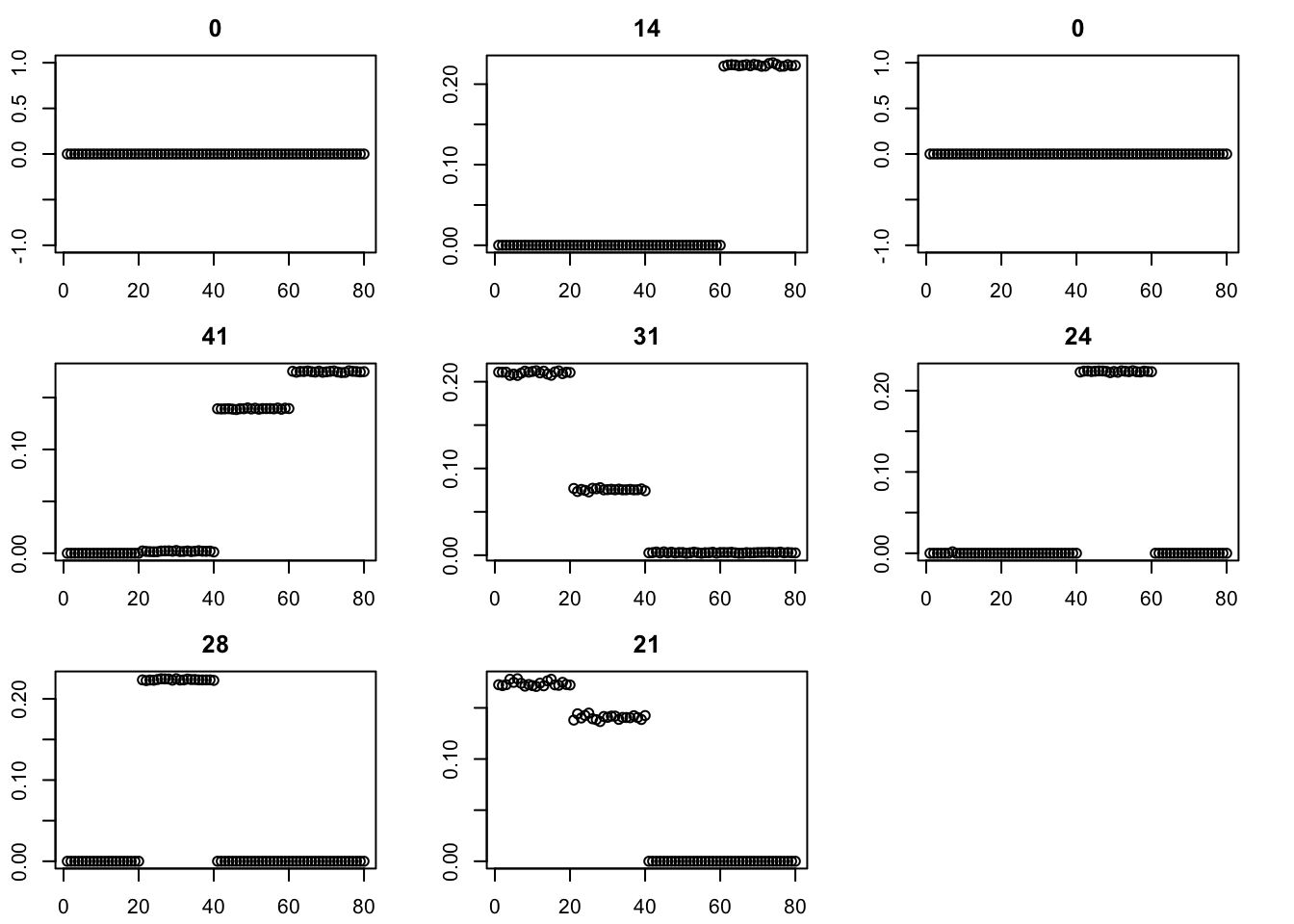
| Version | Author | Date |
|---|---|---|
| 7c17b31 | Matthew Stephens | 2025-03-15 |
for(i in 1:8){plot(fit$v[,i],main=paste0(trunc(fit$d[i])))}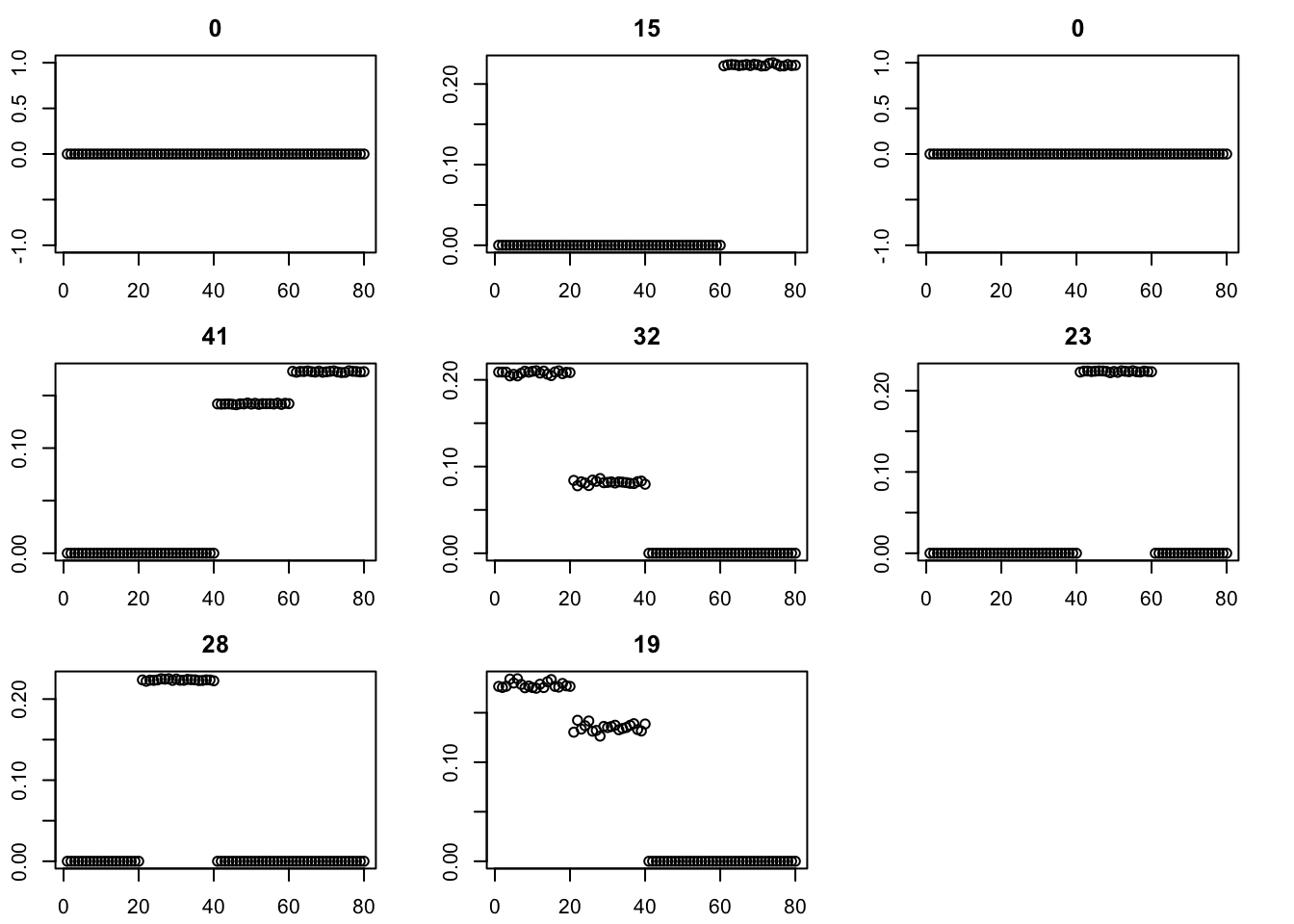
Try binormal instead, initialized from point-exp. This works better.
fit = fit.pe
for(i in 2:200){
fit = eb_power_update(S,fit$v,fit$d,ebnm_binormal)
err[i] = compute_sqerr(S,fit)
}
par(mfcol=c(3,3),mai=rep(0.3,4))
for(i in 1:8){plot(fit$v[,i],main=paste0(trunc(fit$d[i])))}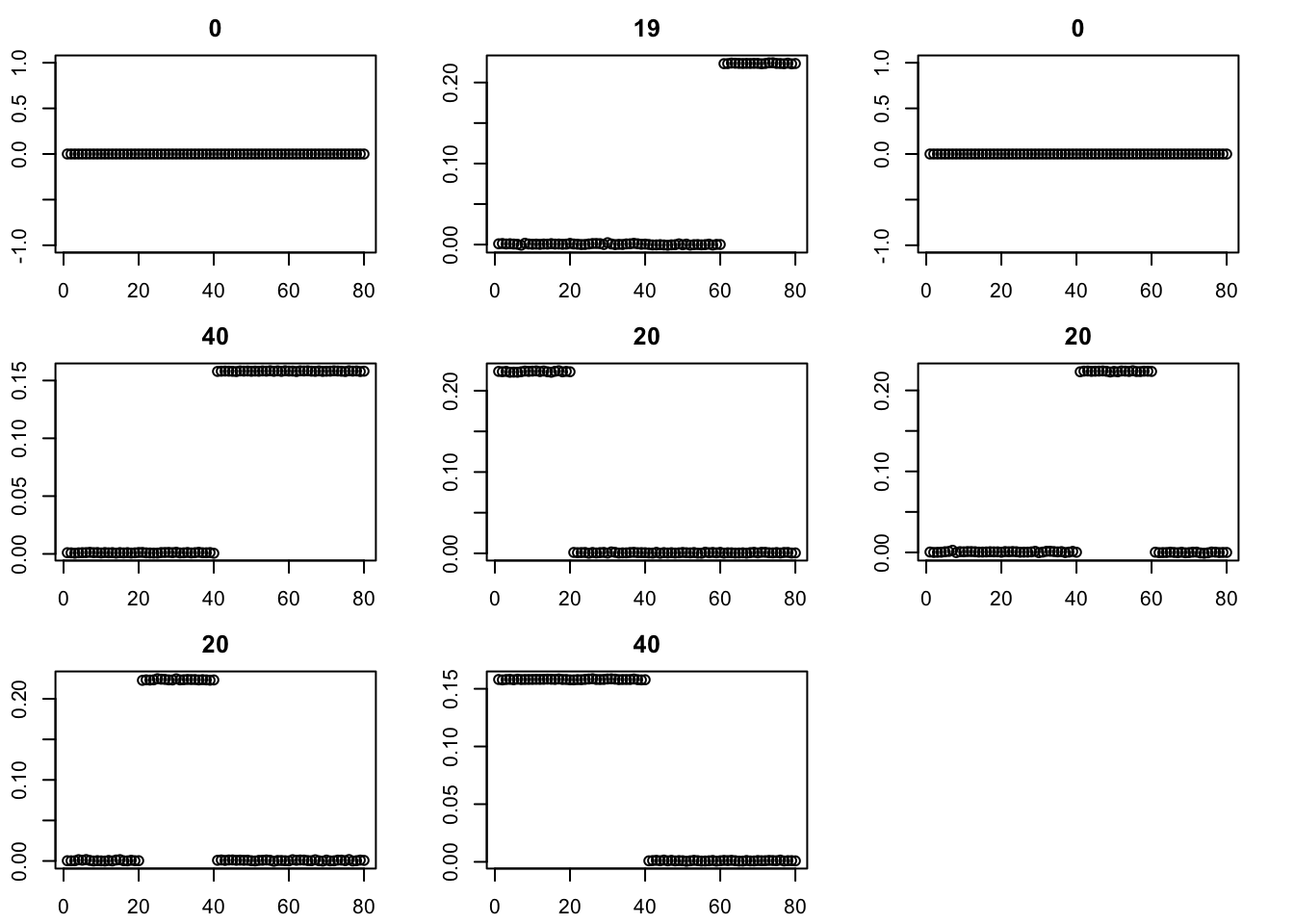
sessionInfo()R version 4.4.2 (2024-10-31)
Platform: aarch64-apple-darwin20
Running under: macOS Sequoia 15.3.1
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
time zone: America/Chicago
tzcode source: internal
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] ebnm_1.1-2
loaded via a namespace (and not attached):
[1] sass_0.4.9 generics_0.1.3 ashr_2.2-63 stringi_1.8.4
[5] lattice_0.22-6 digest_0.6.37 magrittr_2.0.3 evaluate_1.0.3
[9] grid_4.4.2 fastmap_1.2.0 rprojroot_2.0.4 workflowr_1.7.1
[13] jsonlite_1.8.9 Matrix_1.7-2 whisker_0.4.1 mixsqp_0.3-54
[17] promises_1.3.2 scales_1.3.0 truncnorm_1.0-9 invgamma_1.1
[21] jquerylib_0.1.4 cli_3.6.3 rlang_1.1.5 deconvolveR_1.2-1
[25] munsell_0.5.1 splines_4.4.2 cachem_1.1.0 yaml_2.3.10
[29] tools_4.4.2 SQUAREM_2021.1 dplyr_1.1.4 colorspace_2.1-1
[33] ggplot2_3.5.1 httpuv_1.6.15 vctrs_0.6.5 R6_2.5.1
[37] lifecycle_1.0.4 git2r_0.35.0 stringr_1.5.1 fs_1.6.5
[41] trust_0.1-8 irlba_2.3.5.1 pkgconfig_2.0.3 pillar_1.10.1
[45] bslib_0.9.0 later_1.4.1 gtable_0.3.6 glue_1.8.0
[49] Rcpp_1.0.14 xfun_0.50 tibble_3.2.1 tidyselect_1.2.1
[53] rstudioapi_0.17.1 knitr_1.49 htmltools_0.5.8.1 rmarkdown_2.29
[57] compiler_4.4.2 horseshoe_0.2.0