Visualize and interpret droplet and pulse-seq topics
Peter Carbonetto
Last updated: 2020-09-22
Checks: 7 0
Knit directory: single-cell-topics/analysis/
This reproducible R Markdown analysis was created with workflowr (version 1.6.2.9000). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(1) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 0877a1f. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: data/droplet.RData
Ignored: data/pbmc_68k.RData
Ignored: data/pbmc_purified.RData
Ignored: data/pulseseq.RData
Ignored: output/droplet/fits-droplet.RData
Ignored: output/droplet/rds/
Ignored: output/pbmc-68k/fits-pbmc-68k.RData
Ignored: output/pbmc-68k/rds/
Ignored: output/pbmc-purified/fits-pbmc-purified.RData
Ignored: output/pbmc-purified/rds/
Ignored: output/pulseseq/fits-pulseseq.RData
Ignored: output/pulseseq/rds/
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were made to the R Markdown (analysis/plots_tracheal_epithelium.Rmd) and HTML (docs/plots_tracheal_epithelium.html) files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 0877a1f | Peter Carbonetto | 2020-09-22 | workflowr::wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| Rmd | 5af46f1 | Peter Carbonetto | 2020-09-20 | Working on Structure plot for droplet data. |
| Rmd | c072577 | Peter Carbonetto | 2020-09-19 | Re-created structure plot for pulseseq data. |
| Rmd | 942486b | Peter Carbonetto | 2020-09-18 | Fixing merge issue. |
| Rmd | 8cf758e | Peter Carbonetto | 2020-09-12 | Working on improvements to clustering of pulse-seq data. |
| Rmd | a128de5 | Peter Carbonetto | 2020-09-12 | Revamping the analysis of the pulseseq data in plots_tracheal_epithelium. |
| Rmd | 3ab7da1 | Peter Carbonetto | 2020-08-25 | A few minor edits. |
| html | f2e0b23 | Peter Carbonetto | 2020-08-25 | Fixed dimensions of PCA plots in plots_tracheal_epithelium analysis. |
| Rmd | 7b59815 | Peter Carbonetto | 2020-08-25 | workflowr::wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| html | 5589611 | Peter Carbonetto | 2020-08-25 | Added PCA plots and structure plots from pulseseq data. |
| Rmd | b731a4a | Peter Carbonetto | 2020-08-25 | workflowr::wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| html | c3c1b12 | Peter Carbonetto | 2020-08-25 | Build site. |
| Rmd | 2defe6d | Peter Carbonetto | 2020-08-25 | Added crosstab plot to plots_tracheal_epithelium analysis. |
| html | 97c13c2 | Peter Carbonetto | 2020-08-25 | Build site. |
| Rmd | e11855b | Peter Carbonetto | 2020-08-25 | Working on revised analysis of droplet and pulse-seq data sets. |
| html | e11855b | Peter Carbonetto | 2020-08-25 | Working on revised analysis of droplet and pulse-seq data sets. |
| Rmd | bf23ca0 | Peter Carbonetto | 2020-08-20 | Added manual labeling of purified PBMC data to plots_pbmc analysis. |
| Rmd | 077d3d5 | Peter Carbonetto | 2020-08-20 | Added k=9 and k=11 pulseseq fits to plots_tracheal_epithelium analysis. |
| html | 0ce9604 | Peter Carbonetto | 2020-08-20 | Re-built plots_tracheal_epithelium with fastTopics 0.3-162. |
| Rmd | 961570e | Peter Carbonetto | 2020-08-20 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| html | b17bfa4 | Peter Carbonetto | 2020-08-19 | Added pulseseq PCA plots to plots_tracheal_epithelium analysis. |
| Rmd | 76dc0c6 | Peter Carbonetto | 2020-08-19 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| Rmd | c70612f | Peter Carbonetto | 2020-08-19 | Revised structure plot settings for abundant droplet samples in plots_tracheal_epithelium. |
| html | adda33f | Peter Carbonetto | 2020-08-19 | Fixed another structure plot in plots_tracheal_epithelium analysis. |
| Rmd | 29a9258 | Peter Carbonetto | 2020-08-19 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| html | 0a16b60 | Peter Carbonetto | 2020-08-19 | Fixed structure plot in plots_tracheal_epithelium analysis. |
| Rmd | 3a7bd74 | Peter Carbonetto | 2020-08-19 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| html | f4bdf19 | Peter Carbonetto | 2020-08-19 | Added explanatory text and improved PC-based manual clustering of |
| Rmd | c7b77ee | Peter Carbonetto | 2020-08-19 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| Rmd | 70a4a60 | Peter Carbonetto | 2020-08-19 | Added note to plots_tracheal_epithelium.Rmd. |
| html | fb21b3b | Peter Carbonetto | 2020-08-19 | Added very initial Structure plots to plots_tracheal_epithelium analysis. |
| Rmd | d35cb03 | Peter Carbonetto | 2020-08-19 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| html | 368a74a | Peter Carbonetto | 2020-08-19 | Added some text to plots_tracheal_epithelium analysis. |
| Rmd | 223406b | Peter Carbonetto | 2020-08-19 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| html | aca46cc | Peter Carbonetto | 2020-08-19 | Added manual clustering of droplet samples based on PCs. |
| Rmd | 38f811b | Peter Carbonetto | 2020-08-19 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| Rmd | 343747e | Peter Carbonetto | 2020-08-19 | Small edit to figure dimensions. |
| html | 5a35bbd | Peter Carbonetto | 2020-08-19 | Added labeled PCA plot; adjusted plot dimensions in |
| Rmd | fb91075 | Peter Carbonetto | 2020-08-19 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| html | 8b9b528 | Peter Carbonetto | 2020-08-19 | Added more PCA plots to plots_tracheal_epithelium analysis. |
| Rmd | ee7cbf1 | Peter Carbonetto | 2020-08-19 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| html | c517ea2 | Peter Carbonetto | 2020-08-18 | Small fix to one of the PCA plots in plots_tracheal_epithelium. |
| Rmd | 8f5c210 | Peter Carbonetto | 2020-08-18 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| html | 01afbd2 | Peter Carbonetto | 2020-08-18 | Added some PC plots to the plots_tracheal_epithelium analysis. |
| Rmd | f1c7d02 | Peter Carbonetto | 2020-08-18 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| html | 0a04fc1 | Peter Carbonetto | 2020-08-18 | Added abundance plots to plots_tracheal_epithelium analysis. |
| Rmd | f914f7e | Peter Carbonetto | 2020-08-18 | wflow_publish(“plots_tracheal_epithelium.Rmd”) |
| Rmd | 61917ad | Peter Carbonetto | 2020-08-18 | Working on new analysis, plots_tracheal_epithelium.Rmd. |
Add introductory text here.
Load the packages used in the analysis below, as well as additional functions that we will use to generate some of the plots.
library(Matrix)
library(dplyr)
library(fastTopics)
library(ggplot2)
library(cowplot)
source("../code/plots.R")Load data and results
Load the smaller “droplet” data set, the \(k = 7\) Poisson NMF model fit for these data, and the 8 clusters identified in the clustering analysis.
load("../data/droplet.RData")
counts_droplet <- counts
samples_droplet <- readRDS("../output/droplet/clustering-droplet.rds")
fit_droplet <- readRDS("../output/droplet/rds/fit-droplet-scd-ex-k=7.rds")$fit
rm(samples,counts)For convenience, we show the Structure plot again, which summarizes the topic proportions in each of the 8 subsets:
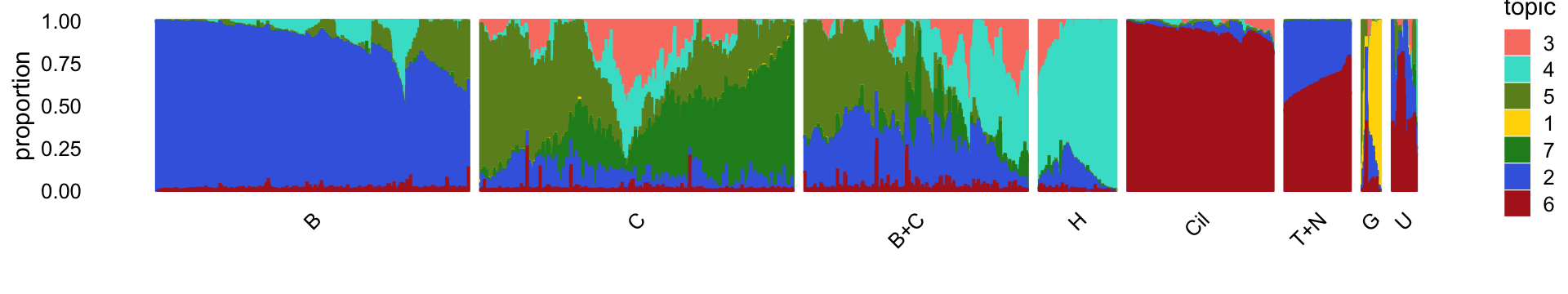
Next, load the larger “pulse-seq” data set, the \(k = 11\) Poisson NMF model fit for these data, and the 7 clusters identified in the clustering analysis.
load("../data/pulseseq.RData")
counts_pulseseq <- counts
samples_pulseseq <- readRDS("../output/pulseseq/clustering-pulseseq.rds")
fit_pulseseq <- readRDS("../output/pulseseq/rds/fit-pulseseq-scd-ex-k=11.rds")$fit
rm(samples,counts)For convenience, we show the Structure plot again, which summarizes the topic proportions in each of the 7 subsets:
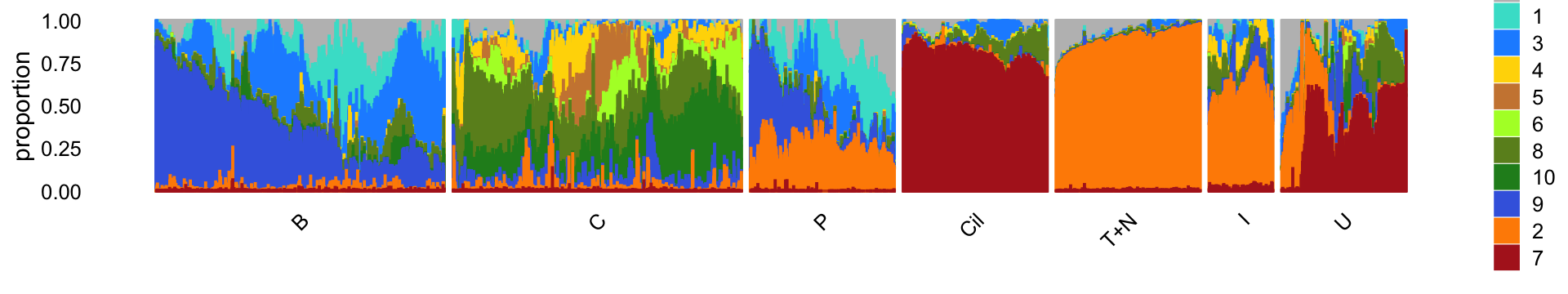
Differential expression analysis
sessionInfo()
# R version 3.6.2 (2019-12-12)
# Platform: x86_64-apple-darwin15.6.0 (64-bit)
# Running under: macOS Catalina 10.15.6
#
# Matrix products: default
# BLAS: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRblas.0.dylib
# LAPACK: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRlapack.dylib
#
# locale:
# [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
#
# attached base packages:
# [1] stats graphics grDevices utils datasets methods base
#
# other attached packages:
# [1] cowplot_1.0.0 ggplot2_3.3.0 fastTopics_0.3-175 dplyr_0.8.3
# [5] Matrix_1.2-18
#
# loaded via a namespace (and not attached):
# [1] progress_1.2.2 tidyselect_0.2.5 xfun_0.11
# [4] purrr_0.3.3 lattice_0.20-38 vctrs_0.2.1
# [7] colorspace_1.4-1 viridisLite_0.3.0 htmltools_0.4.0
# [10] yaml_2.2.0 MCMCpack_1.4-5 plotly_4.9.2
# [13] rlang_0.4.5 later_1.0.0 pillar_1.4.3
# [16] withr_2.1.2 glue_1.3.1 lifecycle_0.1.0
# [19] stringr_1.4.0 MatrixModels_0.4-1 munsell_0.5.0
# [22] gtable_0.3.0 workflowr_1.6.2.9000 htmlwidgets_1.5.1
# [25] coda_0.19-3 evaluate_0.14 knitr_1.26
# [28] SparseM_1.78 httpuv_1.5.2 quantreg_5.54
# [31] irlba_2.3.3 Rcpp_1.0.5 promises_1.1.0
# [34] backports_1.1.5 scales_1.1.0 jsonlite_1.6
# [37] RcppParallel_4.4.2 fs_1.3.1 mcmc_0.9-6
# [40] hms_0.5.2 digest_0.6.23 stringi_1.4.3
# [43] Rtsne_0.15 ggrepel_0.9.0 grid_3.6.2
# [46] rprojroot_1.3-2 quadprog_1.5-8 tools_3.6.2
# [49] magrittr_1.5 lazyeval_0.2.2 tibble_2.1.3
# [52] zeallot_0.1.0 tidyr_1.0.0 crayon_1.3.4
# [55] whisker_0.4 pkgconfig_2.0.3 MASS_7.3-51.4
# [58] prettyunits_1.1.1 data.table_1.12.8 assertthat_0.2.1
# [61] rmarkdown_2.3 httr_1.4.1 R6_2.4.1
# [64] git2r_0.26.1 compiler_3.6.2