Last updated: 2026-02-17
Checks:
Knit directory: jovial-shockley/
This reproducible R Markdown
analysis was created with workflowr (version
1.7.1). The Checks tab describes the reproducibility checks
that were applied when the results were created. The Past
versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git
repository, you know the exact version of the code that produced these
results.
Great job! The global environment was empty. Objects defined in the
global environment can affect the analysis in your R Markdown file in
unknown ways. For reproduciblity it’s best to always run the code in an
empty environment.
The command set.seed(20260127) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Nice! There were no cached chunks for this analysis, so you can be
confident that you successfully produced the results during this
run.
Great job! Using relative paths to the files within your workflowr
project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development
and connecting the code version to the results is critical for
reproducibility.
The results in this page were generated with repository version
2410c8d .
See the Past versions tab to see a history of the changes made
to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .claude/
Ignored: analysis/figure/
Ignored: renv/library/
Ignored: renv/staging/
Untracked files:
Untracked: Data/laconia_messinia_cups_skulls.csv
Untracked: Data/period_dictionary.csv
Untracked: Data/tomb_type_dictionary.csv
Unstaged changes:
Deleted: data/laconia_messinia_cups_skulls - Sheet2.csv
Note that any generated files, e.g. HTML, png, CSS, etc., are not
included in this status report because it is ok for generated content to
have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown
(analysis/cups_and_skulls_analysis.Rmd) and HTML
(docs/cups_and_skulls_analysis.html) files. If you’ve
configured a remote Git repository (see ?wflow_git_remote),
click on the hyperlinks in the table below to view the files as they
were in that past version.
File
Version
Author
Date
Message
Rmd
2410c8d
tinalasisi
2026-02-17
Add dictionary loading template and switch to grouped bar plots
html
8d3e1c7
tinalasisi
2026-02-17
Build site.
Rmd
d689566
tinalasisi
2026-02-17
Add color customization and region-based faceted plots
html
2f9f6dc
tinalasisi
2026-02-17
Build site.
Rmd
c592bd3
tinalasisi
2026-02-17
Add data dictionary worksheet and chamber space analysis
Introduction
This analysis examines the relationship between drinking vessels and
human skeletal remains in Laconian and Messinian tholos and chamber
tombs. The data focuses on grave contexts where drinking vessels are
recorded alongside information about primary and secondary burials.
Load Libraries
library(readr)
library(dplyr)
library(ggplot2)
library(tidyr)
library(stringr)
library(janitor)
library(knitr)
Color Customization
Customize colors for the analyses. Edit the values below to use your
preferred colors:
# Set your color palette here
colors <- list(
laconia = "#1B9E77", # Laconia color (default: teal/green)
messenia = "#D95F02", # Messenia color (default: orange)
drinking_vessel = "#7570B3", # Drinking vessels (default: purple)
skulls = "#E7298A", # Skulls (default: pink)
background = "#FFFFCC" # Background/neutral (default: light yellow)
)
# These colors will be used throughout all plots
# Change the hex codes to your preferred colors
Load Data
# Load the CSV data
data_path <- here::here("data", "laconia_messinia_cups_skulls.csv")
cups_skulls <- read_csv(data_path)Rows: 296 Columns: 14
── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
chr (11): Grave ID, site, type, period, secondary, primary, dimensions, orie...
dbl (3): drinking vessel number, skulls number, area of chamber estimated
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.# Display dimensions
cat("Dataset dimensions:", nrow(cups_skulls), "graves ×", ncol(cups_skulls), "columns\n")Dataset dimensions: 296 graves × 14 columns# Show column names
glimpse(cups_skulls)Rows: 296
Columns: 14
$ `Grave ID` <chr> "vapheio_tholos", "arkynes_tholos_A", "ark…
$ site <chr> "Vapheio", "Arkynes", "Arkynes", "Arkynes"…
$ type <chr> "tholos", "tholos", "tholos", "tholos", "t…
$ period <chr> "LHIIA-LHIIIA1", "LHIIIA- LHIIIB", "LHIII"…
$ secondary <chr> "absent", "absent", "absent", "present", "…
$ primary <chr> "absent", "absent", "present", "absent", "…
$ dimensions <chr> "83.28m^2. dromos: L 29.8m x w3.18-3.45, s…
$ orientation <chr> "NE-SW", "SE-NW", NA, NA, "E-W", "N-S", "W…
$ `drinking vessel` <chr> "yes", "no", "no", "no", "no", "no", "no",…
$ remains <chr> "yes", "yes", "yes", "yes", "yes", "no", "…
$ `drinking vessel number` <dbl> 6, NA, NA, NA, NA, NA, NA, 4, NA, NA, 5, N…
$ `skulls number` <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
$ `area of chamber estimated` <dbl> 84.130, 17.350, NA, 28.274, 7.793, NA, 18.…
$ region <chr> "laconia", "laconia", "laconia", "laconia"…
Data Cleaning
cups_skulls_clean <- cups_skulls %>%
# Clean column names
janitor::clean_names() %>%
# Trim whitespace from character columns
mutate(across(where(is.character), str_trim)) %>%
# Standardize yes/no values in drinking vessel column
mutate(
has_drinking_vessel = case_when(
tolower(drinking_vessel) == "yes" ~ TRUE,
tolower(drinking_vessel) == "no" ~ FALSE,
TRUE ~ NA
),
has_remains = case_when(
tolower(remains) == "yes" ~ TRUE,
tolower(remains) == "no" ~ FALSE,
TRUE ~ NA
),
primary_burial = case_when(
tolower(primary) == "present" ~ TRUE,
tolower(primary) == "absent" ~ FALSE,
TRUE ~ NA
),
secondary_burial = case_when(
tolower(secondary) == "present" ~ TRUE,
tolower(secondary) == "absent" ~ FALSE,
TRUE ~ NA
)
) %>%
# Create burial category
mutate(
burial_category = case_when(
primary_burial & secondary_burial ~ "Both Primary & Secondary",
primary_burial & !secondary_burial ~ "Primary Only",
!primary_burial & secondary_burial ~ "Secondary Only",
TRUE ~ "Unknown/Not Recorded"
)
) %>%
# Simplify grave type
mutate(
grave_type_simple = case_when(
str_detect(tolower(type), "tholos") ~ "Tholos",
str_detect(tolower(type), "chamber") ~ "Chamber Tomb",
str_detect(tolower(type), "cist") ~ "Cist Grave",
str_detect(tolower(type), "pit") ~ "Pit Grave",
str_detect(tolower(type), "dromos") ~ "Dromos Only",
TRUE ~ "Other"
)
)
# Show cleaned data structure
glimpse(cups_skulls_clean)Rows: 296
Columns: 20
$ grave_id <chr> "vapheio_tholos", "arkynes_tholos_A", "arkyn…
$ site <chr> "Vapheio", "Arkynes", "Arkynes", "Arkynes", …
$ type <chr> "tholos", "tholos", "tholos", "tholos", "tho…
$ period <chr> "LHIIA-LHIIIA1", "LHIIIA- LHIIIB", "LHIII", …
$ secondary <chr> "absent", "absent", "absent", "present", "pr…
$ primary <chr> "absent", "absent", "present", "absent", "pr…
$ dimensions <chr> "83.28m^2. dromos: L 29.8m x w3.18-3.45, sto…
$ orientation <chr> "NE-SW", "SE-NW", NA, NA, "E-W", "N-S", "W-E…
$ drinking_vessel <chr> "yes", "no", "no", "no", "no", "no", "no", "…
$ remains <chr> "yes", "yes", "yes", "yes", "yes", "no", "no…
$ drinking_vessel_number <dbl> 6, NA, NA, NA, NA, NA, NA, 4, NA, NA, 5, NA,…
$ skulls_number <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, …
$ area_of_chamber_estimated <dbl> 84.130, 17.350, NA, 28.274, 7.793, NA, 18.32…
$ region <chr> "laconia", "laconia", "laconia", "laconia", …
$ has_drinking_vessel <lgl> TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FAL…
$ has_remains <lgl> TRUE, TRUE, TRUE, TRUE, TRUE, FALSE, FALSE, …
$ primary_burial <lgl> FALSE, FALSE, TRUE, FALSE, TRUE, FALSE, FALS…
$ secondary_burial <lgl> FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALS…
$ burial_category <chr> "Unknown/Not Recorded", "Unknown/Not Recorde…
$ grave_type_simple <chr> "Tholos", "Tholos", "Tholos", "Tholos", "Tho…
Import and Process Data Dictionaries
After the student completes the dictionaries , this
section will load and apply ordinal rankings:
# NOTE: Set eval=TRUE once the student has completed the dictionaries
# Load period dictionary
period_dict <- read_csv(here::here("data", "period_dictionary.csv"))
# Load tomb type dictionary
tomb_dict <- read_csv(here::here("data", "tomb_type_dictionary.csv"))
# Join period dictionary and extract period components
cups_skulls_clean <- cups_skulls_clean %>%
left_join(
period_dict %>% select(period_code, chronological_order, earliest_component, latest_component),
by = c("period" = "period_code")
) %>%
# Get ordinal values for earliest component
left_join(
period_dict %>% select(period_code, chronological_order) %>% rename(start_order = chronological_order),
by = c("earliest_component" = "period_code")
) %>%
# Get ordinal values for latest component
left_join(
period_dict %>% select(period_code, chronological_order) %>% rename(end_order = chronological_order),
by = c("latest_component" = "period_code")
) %>%
# For single periods (where start = end), use that value
# For ranges, create midpoint
mutate(
period_start_order = start_order,
period_end_order = end_order,
period_midpoint_order = (start_order + end_order) / 2
)
# Join tomb type dictionary
cups_skulls_clean <- cups_skulls_clean %>%
left_join(
tomb_dict %>% select(tomb_type, tomb_type_order),
by = c("grave_type_simple" = "tomb_type")
) %>%
# Convert to ordered factors
mutate(
tomb_type_ordered = factor(
grave_type_simple,
levels = arrange(tomb_dict, tomb_type_order) %>% pull(tomb_type),
ordered = TRUE
),
period_ordered = factor(
period,
levels = arrange(period_dict, chronological_order) %>% pull(period_code),
ordered = TRUE
)
)
cat("✓ Dictionaries successfully loaded and applied!\n")
cat("New columns created:\n")
cat(" - period_start_order: chronological order of earliest period component\n")
cat(" - period_end_order: chronological order of latest period component\n")
cat(" - period_midpoint_order: midpoint between start and end\n")
cat(" - tomb_type_ordered: ordered factor for tomb types\n")
cat(" - period_ordered: ordered factor for periods\n")
Data Overview
Basic Statistics
cat("=== Dataset Summary ===\n\n")=== Dataset Summary ===cat("Total graves:", nrow(cups_skulls_clean), "\n")Total graves: 296 cat("Graves with drinking vessels recorded:", sum(cups_skulls_clean$has_drinking_vessel == TRUE, na.rm = TRUE), "\n")Graves with drinking vessels recorded: 80 cat("Graves with human remains:", sum(cups_skulls_clean$has_remains == TRUE, na.rm = TRUE), "\n")Graves with human remains: 226 cat("\n")# Missing data summary
cat("=== Missing Data ===\n")=== Missing Data ===cups_skulls_clean %>%
summarise(across(everything(), ~sum(is.na(.)))) %>%
pivot_longer(everything(), names_to = "column", values_to = "missing_count") %>%
filter(missing_count > 0) %>%
arrange(desc(missing_count)) %>%
kable()
orientation
262
drinking_vessel_number
222
skulls_number
213
area_of_chamber_estimated
197
dimensions
171
period
90
type
12
Sites Represented
sites_summary <- cups_skulls_clean %>%
count(site, region, name = "n_graves") %>%
arrange(desc(n_graves))
cat("Number of sites:", n_distinct(cups_skulls_clean$site), "\n")Number of sites: 50 cat("Sites represented:\n\n")Sites represented:print(sites_summary, n = 30)# A tibble: 50 × 3
site region n_graves
<chr> <chr> <int>
1 Agios Stephanos laconia 63
2 volimidhia messenia 35
3 Xerokambi: Agios Vasileios North Cemetery laconia 25
4 Ayos Ioannis Papoulia messenia 23
5 Koukounara Gouvalari messenia 14
6 Epidavros Limera:PAL1 laconia 10
7 Peristeria & kokorakou messenia 9
8 Voidhokoloa messenia 9
9 Nihoria messenia 8
10 Handhrinou Kissos messenia 7
11 Kaminia messenia 7
12 Pellana laconia 7
13 Englianos messenia 6
14 Skoura: malathria laconia 6
15 Pellana: pelekete and Palaiokastro laconia 5
16 Peristeri: Solakos laconia 5
17 Routsi messenia 4
18 Sparti: Polydendron laconia 4
19 Sykea laconia 4
20 Amyklai: Spelakia laconia 3
21 Amyklaion laconia 3
22 Arkynes laconia 3
23 Geraki: Acropolis laconia 3
24 Daphi: Louria laconia 2
25 Daphni: Louria 3 laconia 2
26 Koukounara Livadhiti to Fities messenia 2
27 Koukounara akones messenia 2
28 Psari Metsiki messenia 2
29 Tragana messenia 2
30 Agios Georgios Voion laconia 1
# ℹ 20 more rows
Grave Type Distribution
grave_type_summary <- cups_skulls_clean %>%
count(region, grave_type_simple, name = "count") %>%
arrange(region, desc(count))
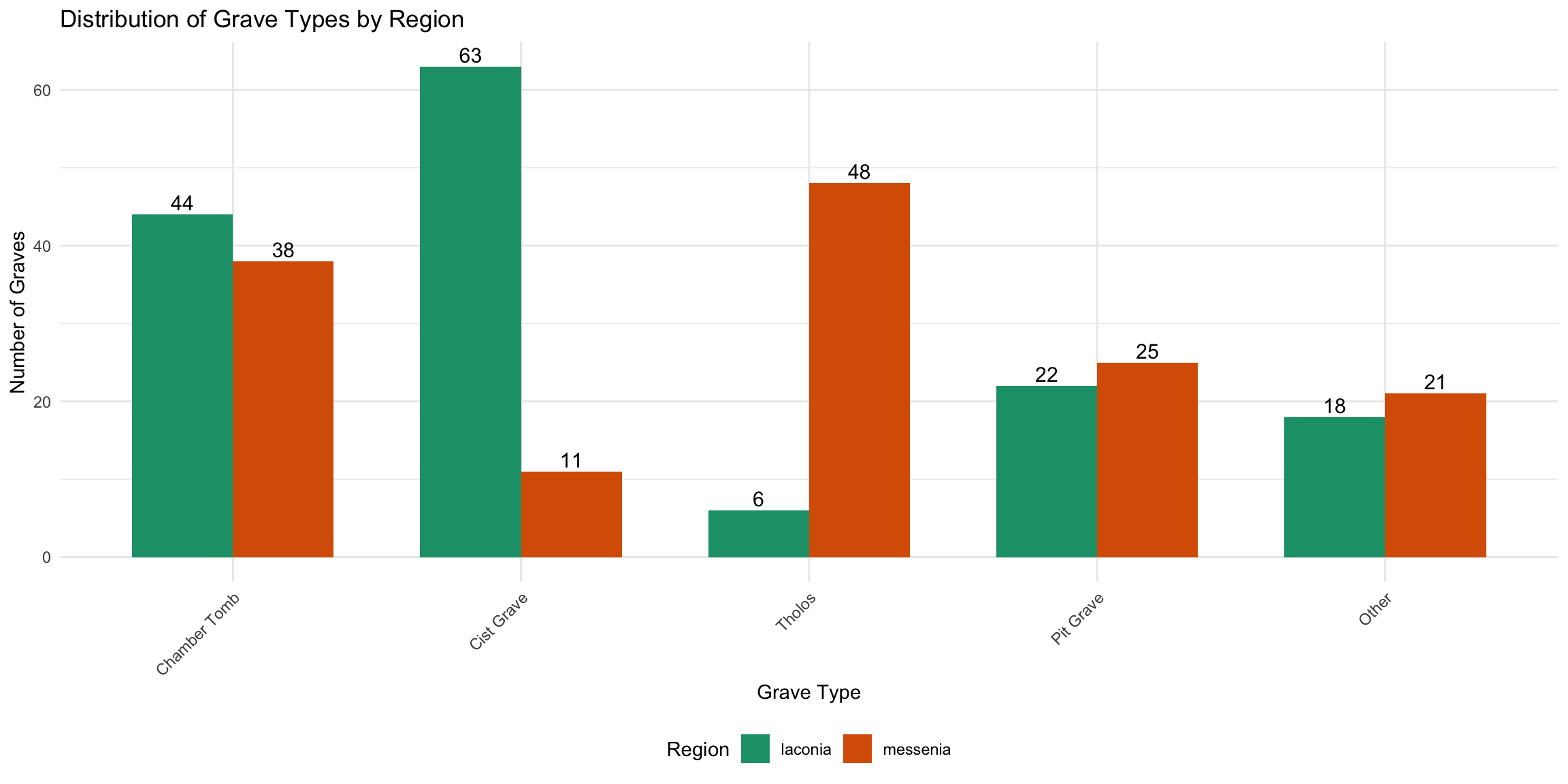
print(grave_type_summary)# A tibble: 10 × 3
region grave_type_simple count
<chr> <chr> <int>
1 laconia Cist Grave 63
2 laconia Chamber Tomb 44
3 laconia Pit Grave 22
4 laconia Other 18
5 laconia Tholos 6
6 messenia Tholos 48
7 messenia Chamber Tomb 38
8 messenia Pit Grave 25
9 messenia Other 21
10 messenia Cist Grave 11ggplot(grave_type_summary, aes(x = reorder(grave_type_simple, -count), y = count, fill = region)) +
geom_col(position = "dodge", width = 0.7) +
scale_fill_manual(values = c(laconia = colors$laconia, messenia = colors$messenia)) +
geom_text(aes(label = count), position = position_dodge(width = 0.7), vjust = -0.3, size = 4) +
labs(
title = "Distribution of Grave Types by Region",
x = "Grave Type",
y = "Number of Graves",
fill = "Region"
) +
theme_minimal() +
theme(legend.position = "bottom", axis.text.x = element_text(angle = 45, hjust = 1))
Past versions of grave-type-plot-1.png
Version
Author
Date
8d3e1c7
tinalasisi
2026-02-17
2f9f6dc
tinalasisi
2026-02-17
Burial Types
Primary vs Secondary Burials
burial_type_summary <- cups_skulls_clean %>%
summarise(
total = n(),
with_primary = sum(primary_burial == TRUE, na.rm = TRUE),
with_secondary = sum(secondary_burial == TRUE, na.rm = TRUE),
both = sum(primary_burial == TRUE & secondary_burial == TRUE, na.rm = TRUE),
neither = sum(primary_burial == FALSE & secondary_burial == FALSE, na.rm = TRUE),
unknown = sum(is.na(primary_burial) | is.na(secondary_burial))
) %>%
mutate(
pct_primary = round(with_primary / total * 100, 1),
pct_secondary = round(with_secondary / total * 100, 1),
pct_both = round(both / total * 100, 1)
)
print(burial_type_summary)# A tibble: 1 × 9
total with_primary with_secondary both neither unknown pct_primary
<int> <int> <int> <int> <int> <int> <dbl>
1 296 135 118 57 100 0 45.6
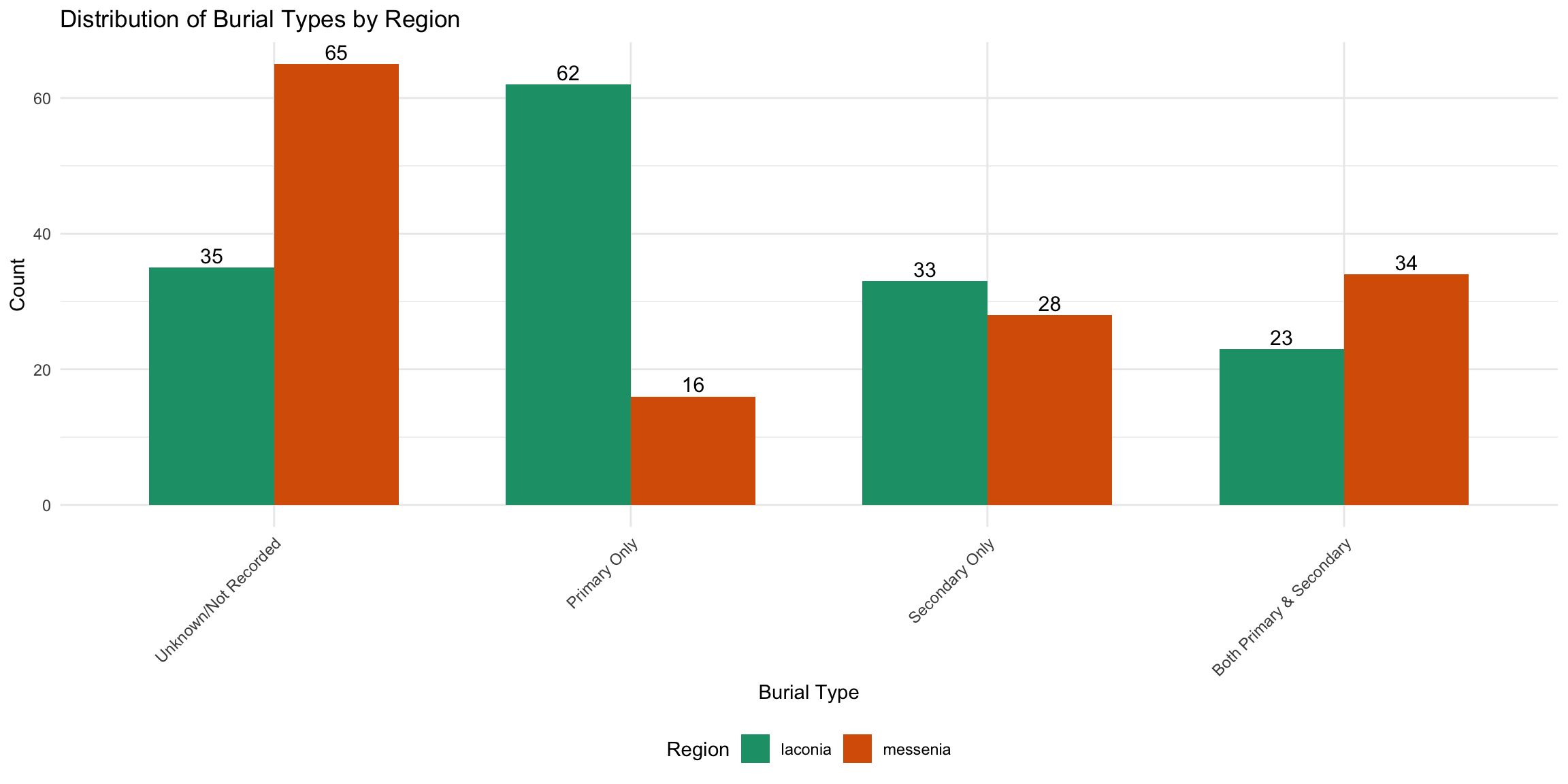
# ℹ 2 more variables: pct_secondary <dbl>, pct_both <dbl>burial_category_data_region <- cups_skulls_clean %>%
count(region, burial_category) %>%
arrange(region, desc(n))
ggplot(burial_category_data_region, aes(x = reorder(burial_category, -n), y = n, fill = region)) +
geom_col(position = "dodge", width = 0.7) +
scale_fill_manual(values = c(laconia = colors$laconia, messenia = colors$messenia)) +
geom_text(aes(label = n), position = position_dodge(width = 0.7), vjust = -0.3, size = 4) +
labs(
title = "Distribution of Burial Types by Region",
x = "Burial Type",
y = "Count",
fill = "Region"
) +
theme_minimal() +
theme(legend.position = "bottom", axis.text.x = element_text(angle = 45, hjust = 1))
Past versions of burial-plot-1.png
Version
Author
Date
8d3e1c7
tinalasisi
2026-02-17
2f9f6dc
tinalasisi
2026-02-17
Drinking Vessels Analysis
Overall Drinking Vessel Presence
dv_summary <- cups_skulls_clean %>%
summarise(
total_graves = n(),
with_vessels = sum(has_drinking_vessel == TRUE, na.rm = TRUE),
without_vessels = sum(has_drinking_vessel == FALSE, na.rm = TRUE),
unknown_vessel_status = sum(is.na(has_drinking_vessel))
) %>%
mutate(
pct_with = round(with_vessels / total_graves * 100, 1),
avg_vessels_recorded = round(mean(cups_skulls_clean$drinking_vessel_number, na.rm = TRUE), 2)
)
print(dv_summary)# A tibble: 1 × 6
total_graves with_vessels without_vessels unknown_vessel_status pct_with
<int> <int> <int> <int> <dbl>
1 296 80 216 0 27
# ℹ 1 more variable: avg_vessels_recorded <dbl>
Drinking Vessels by Grave Type
dv_by_type <- cups_skulls_clean %>%
group_by(grave_type_simple) %>%
summarise(
total = n(),
with_vessels = sum(has_drinking_vessel == TRUE, na.rm = TRUE),
pct_with_vessels = round(with_vessels / total * 100, 1),
avg_vessel_count = round(mean(drinking_vessel_number, na.rm = TRUE), 2)
) %>%
arrange(desc(with_vessels))
print(dv_by_type)# A tibble: 5 × 5
grave_type_simple total with_vessels pct_with_vessels avg_vessel_count
<chr> <int> <int> <dbl> <dbl>
1 Chamber Tomb 82 38 46.3 2.62
2 Tholos 54 20 37 2.74
3 Other 39 8 20.5 1.25
4 Pit Grave 47 8 17 1.14
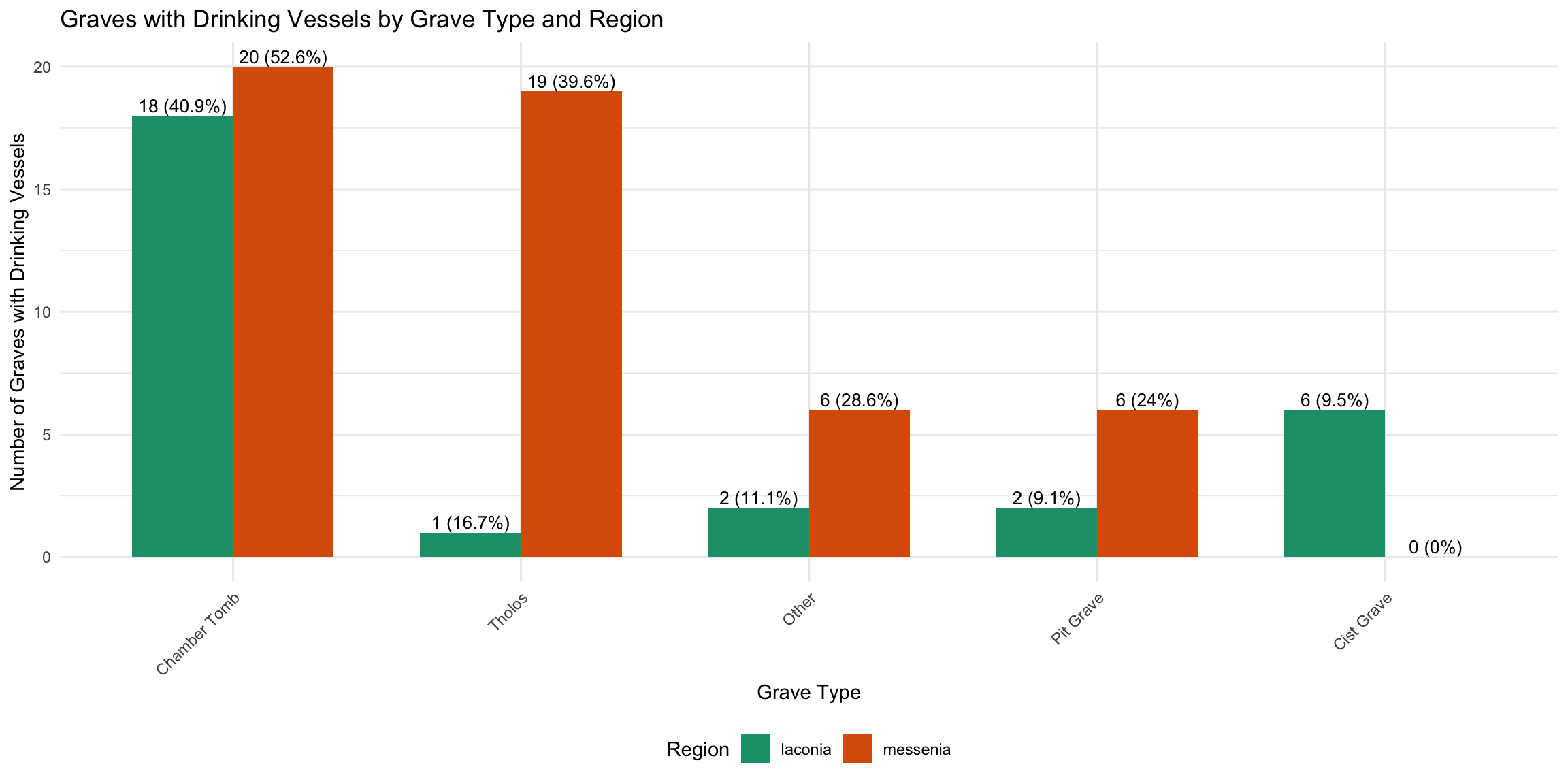
5 Cist Grave 74 6 8.1 1.33dv_by_type_region <- cups_skulls_clean %>%
group_by(region, grave_type_simple) %>%
summarise(
total = n(),
with_vessels = sum(has_drinking_vessel == TRUE, na.rm = TRUE),
pct_with_vessels = round(with_vessels / total * 100, 1),
avg_vessel_count = round(mean(drinking_vessel_number, na.rm = TRUE), 2),
.groups = "drop"
) %>%
arrange(region, desc(with_vessels))
ggplot(dv_by_type_region, aes(x = reorder(grave_type_simple, -with_vessels), y = with_vessels, fill = region)) +
geom_col(position = "dodge", width = 0.7) +
scale_fill_manual(values = c(laconia = colors$laconia, messenia = colors$messenia)) +
geom_text(aes(label = paste0(with_vessels, " (", pct_with_vessels, "%)")),
position = position_dodge(width = 0.7), vjust = -0.3, size = 3.5) +
labs(
title = "Graves with Drinking Vessels by Grave Type and Region",
x = "Grave Type",
y = "Number of Graves with Drinking Vessels",
fill = "Region"
) +
theme_minimal() +
theme(legend.position = "bottom", axis.text.x = element_text(angle = 45, hjust = 1))
Past versions of dv-by-type-plot-1.png
Version
Author
Date
8d3e1c7
tinalasisi
2026-02-17
2f9f6dc
tinalasisi
2026-02-17
Drinking Vessels by Burial Type
dv_by_burial <- cups_skulls_clean %>%
group_by(burial_category) %>%
summarise(
total = n(),
with_vessels = sum(has_drinking_vessel == TRUE, na.rm = TRUE),
pct_with_vessels = round(with_vessels / total * 100, 1),
avg_vessel_count = round(mean(drinking_vessel_number, na.rm = TRUE), 2)
) %>%
arrange(desc(with_vessels))
print(dv_by_burial)# A tibble: 4 × 5
burial_category total with_vessels pct_with_vessels avg_vessel_count
<chr> <int> <int> <dbl> <dbl>
1 Both Primary & Secondary 57 32 56.1 3.1
2 Secondary Only 61 20 32.8 1.83
3 Unknown/Not Recorded 100 19 19 1.88
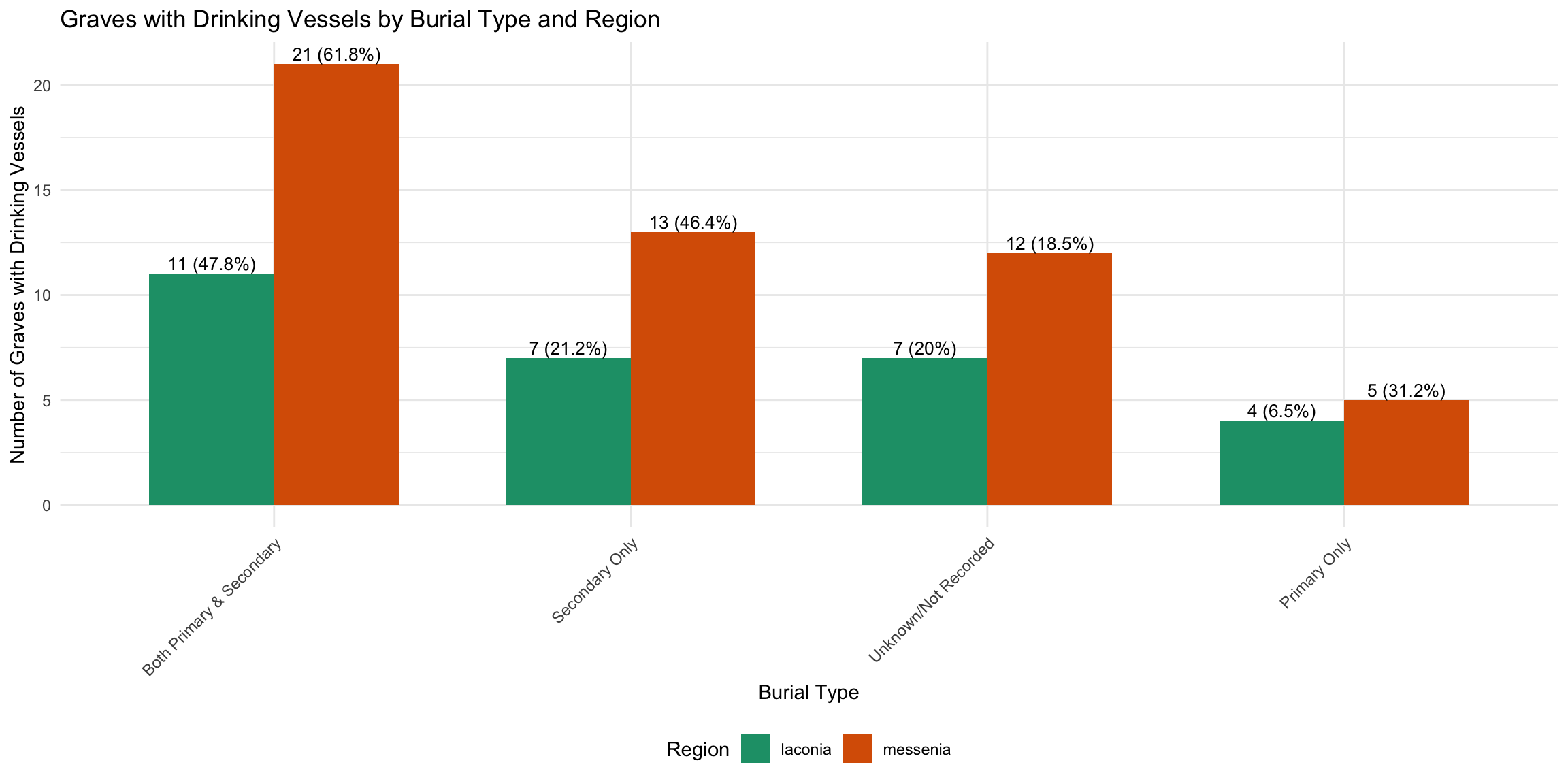
4 Primary Only 78 9 11.5 1 dv_by_burial_region <- cups_skulls_clean %>%
group_by(region, burial_category) %>%
summarise(
total = n(),
with_vessels = sum(has_drinking_vessel == TRUE, na.rm = TRUE),
pct_with_vessels = round(with_vessels / total * 100, 1),
avg_vessel_count = round(mean(drinking_vessel_number, na.rm = TRUE), 2),
.groups = "drop"
) %>%
arrange(region, desc(with_vessels))
ggplot(dv_by_burial_region, aes(x = reorder(burial_category, -with_vessels), y = with_vessels, fill = region)) +
geom_col(position = "dodge", width = 0.7) +
scale_fill_manual(values = c(laconia = colors$laconia, messenia = colors$messenia)) +
geom_text(aes(label = paste0(with_vessels, " (", pct_with_vessels, "%)")),
position = position_dodge(width = 0.7), vjust = -0.3, size = 3.5) +
labs(
title = "Graves with Drinking Vessels by Burial Type and Region",
x = "Burial Type",
y = "Number of Graves with Drinking Vessels",
fill = "Region"
) +
theme_minimal() +
theme(legend.position = "bottom", axis.text.x = element_text(angle = 45, hjust = 1))
Past versions of dv-by-burial-plot-1.png
Version
Author
Date
8d3e1c7
tinalasisi
2026-02-17
2f9f6dc
tinalasisi
2026-02-17
Skeletal Remains Analysis
Remains Distribution
remains_summary <- cups_skulls_clean %>%
summarise(
total_graves = n(),
with_remains = sum(has_remains == TRUE, na.rm = TRUE),
without_remains = sum(has_remains == FALSE, na.rm = TRUE),
unknown_remains = sum(is.na(has_remains))
) %>%
mutate(pct_with_remains = round(with_remains / total_graves * 100, 1))
print(remains_summary)# A tibble: 1 × 5
total_graves with_remains without_remains unknown_remains pct_with_remains
<int> <int> <int> <int> <dbl>
1 296 226 70 0 76.4
Skulls Recorded
skulls_stats <- cups_skulls_clean %>%
filter(!is.na(skulls_number)) %>%
summarise(
graves_with_skull_count = n(),
total_skulls = sum(skulls_number, na.rm = TRUE),
avg_skulls = round(mean(skulls_number, na.rm = TRUE), 2),
min_skulls = min(skulls_number, na.rm = TRUE),
max_skulls = max(skulls_number, na.rm = TRUE)
)
print(skulls_stats)# A tibble: 1 × 5
graves_with_skull_count total_skulls avg_skulls min_skulls max_skulls
<int> <dbl> <dbl> <dbl> <dbl>
1 83 350 4.22 1 47skulls_by_count <- cups_skulls_clean %>%
filter(!is.na(skulls_number)) %>%
count(skulls_number, name = "graves") %>%
arrange(skulls_number)
print(skulls_by_count)# A tibble: 16 × 2
skulls_number graves
<dbl> <int>
1 1 48
2 2 10
3 3 5
4 4 2
5 5 1
6 6 3
7 7 1
8 9 2
9 10 2
10 12 2
11 14 1
12 15 1
13 20 2
14 24 1
15 27 1
16 47 1
Cross-Analysis: Drinking Vessels and Remains
Do graves with drinking vessels tend to have remains?
vessels_remains_cross <- cups_skulls_clean %>%
filter(!is.na(has_drinking_vessel) & !is.na(has_remains)) %>%
count(has_drinking_vessel, has_remains) %>%
pivot_wider(names_from = has_remains, values_from = n, values_fill = 0) %>%
rename(no_remains = `FALSE`, with_remains = `TRUE`)
rownames(vessels_remains_cross) <- c("No Drinking Vessels", "With Drinking Vessels")Warning: Setting row names on a tibble is deprecated.print(vessels_remains_cross)# A tibble: 2 × 3
has_drinking_vessel no_remains with_remains
* <lgl> <int> <int>
1 FALSE 55 161
2 TRUE 15 65# Calculate cross-tabulation with percentages
cross_analysis <- cups_skulls_clean %>%
filter(!is.na(has_drinking_vessel) & !is.na(has_remains)) %>%
group_by(has_drinking_vessel) %>%
summarise(
total = n(),
with_remains = sum(has_remains == TRUE, na.rm = TRUE),
pct_with_remains = round(with_remains / total * 100, 1)
) %>%
mutate(vessel_status = ifelse(has_drinking_vessel, "With Drinking Vessels", "Without Drinking Vessels"))
kable(select(cross_analysis, vessel_status, total, with_remains, pct_with_remains))
Without Drinking Vessels
216
161
74.5
With Drinking Vessels
80
65
81.2
Drinking Vessels and Skull Count
vessels_skulls_data <- cups_skulls_clean %>%
filter(!is.na(drinking_vessel_number) & !is.na(skulls_number)) %>%
mutate(has_vessels_label = ifelse(has_drinking_vessel, "With Vessels", "No Vessels"))
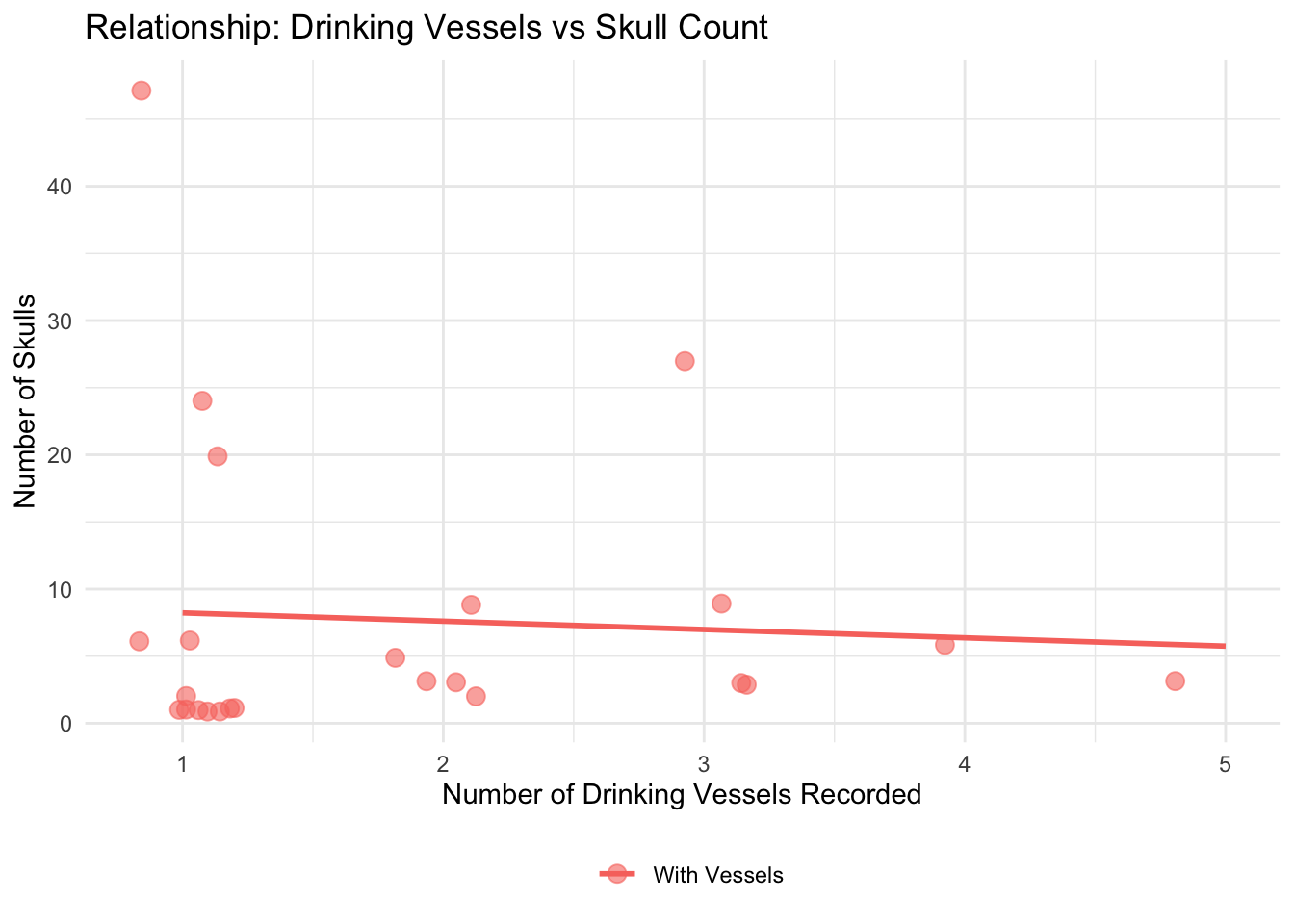
ggplot(vessels_skulls_data, aes(x = drinking_vessel_number, y = skulls_number, color = has_vessels_label)) +
geom_jitter(size = 3, alpha = 0.6, width = 0.2, height = 0.2) +
geom_smooth(method = "lm", se = FALSE, alpha = 0.2) +
labs(
title = "Relationship: Drinking Vessels vs Skull Count",
x = "Number of Drinking Vessels Recorded",
y = "Number of Skulls",
color = ""
) +
theme_minimal() +
theme(legend.position = "bottom")`geom_smooth()` using formula = 'y ~ x'
Past versions of vessels-skulls-scatter-1.png
Version
Author
Date
2f9f6dc
tinalasisi
2026-02-17
Regional Analysis
Laconia vs Other Regions
regional_summary <- cups_skulls_clean %>%
group_by(region) %>%
summarise(
total_graves = n(),
with_drinking_vessels = sum(has_drinking_vessel == TRUE, na.rm = TRUE),
pct_with_vessels = round(with_drinking_vessels / total_graves * 100, 1),
with_remains = sum(has_remains == TRUE, na.rm = TRUE),
pct_with_remains = round(with_remains / total_graves * 100, 1),
avg_skulls = round(mean(skulls_number, na.rm = TRUE), 2)
)
print(regional_summary)# A tibble: 2 × 7
region total_graves with_drinking_vessels pct_with_vessels with_remains
<chr> <int> <int> <dbl> <int>
1 laconia 153 29 19 127
2 messenia 143 51 35.7 99
# ℹ 2 more variables: pct_with_remains <dbl>, avg_skulls <dbl>
Chronological Overview
Periods Represented
period_summary <- cups_skulls_clean %>%
mutate(
period_category = case_when(
str_detect(tolower(period), "mh") ~ "Middle Helladic",
str_detect(tolower(period), "lh\\s*i(?!i)") | str_detect(tolower(period), "lh i-") ~ "LH I-II",
str_detect(tolower(period), "lh\\s*ii") & !str_detect(tolower(period), "lh\\s*ii") ~ "LH II",
str_detect(tolower(period), "lh\\s*iii") ~ "LH III",
str_detect(tolower(period), "lh") ~ "Late Helladic (general)",
is.na(period) | period == "" ~ "Unrecorded",
TRUE ~ "Other"
)
) %>%
count(period_category, name = "graves") %>%
arrange(desc(graves))
print(period_summary)# A tibble: 5 × 2
period_category graves
<chr> <int>
1 Unrecorded 90
2 LH III 88
3 LH I-II 45
4 Middle Helladic 39
5 Late Helladic (general) 34
Key Findings Summary
cat("=== KEY FINDINGS ===\n\n")=== KEY FINDINGS ===cat("1. Drinking Vessel Prevalence:\n")1. Drinking Vessel Prevalence:cat(" -", sum(cups_skulls_clean$has_drinking_vessel == TRUE, na.rm = TRUE),
"graves (", round(sum(cups_skulls_clean$has_drinking_vessel == TRUE, na.rm = TRUE) / nrow(cups_skulls_clean) * 100, 1),
"%) have drinking vessels recorded\n\n") - 80 graves ( 27 %) have drinking vessels recordedcat("2. Skeletal Remains:\n")2. Skeletal Remains:cat(" -", sum(cups_skulls_clean$has_remains == TRUE, na.rm = TRUE),
"graves (", round(sum(cups_skulls_clean$has_remains == TRUE, na.rm = TRUE) / nrow(cups_skulls_clean) * 100, 1),
"%) have human remains\n") - 226 graves ( 76.4 %) have human remainscat(" - Skull counts recorded in", sum(!is.na(cups_skulls_clean$skulls_number)), "graves\n\n") - Skull counts recorded in 83 gravescat("3. Burial Type Association:\n")3. Burial Type Association:burial_counts <- cups_skulls_clean %>%
count(burial_category) %>%
arrange(desc(n))
for (i in 1:nrow(burial_counts)) {
cat(" -", burial_counts$burial_category[i], ":", burial_counts$n[i], "graves\n")
} - Unknown/Not Recorded : 100 graves
- Primary Only : 78 graves
- Secondary Only : 61 graves
- Both Primary & Secondary : 57 graves
Chamber Space and Burial Goods
Do larger chambers have more skulls and drinking vessels?
This section explores whether the estimated chamber area relates to
the number of skulls and drinking vessels found.
Data Availability
space_analysis_data <- cups_skulls_clean %>%
filter(!is.na(area_of_chamber_estimated))
cat("Graves with chamber area estimated:", nrow(space_analysis_data), "\n")Graves with chamber area estimated: 99 cat("Graves with area AND skull count:", sum(!is.na(space_analysis_data$skulls_number)), "\n")Graves with area AND skull count: 22 cat("Graves with area AND vessel count:", sum(!is.na(space_analysis_data$drinking_vessel_number)), "\n\n")Graves with area AND vessel count: 41 # Summary statistics for chamber area
cat("Chamber Area Summary (m²):\n")Chamber Area Summary (m²):print(space_analysis_data %>%
pull(area_of_chamber_estimated) %>%
summary()) Min. 1st Qu. Median Mean 3rd Qu. Max.
1.419 7.069 12.566 19.056 23.330 88.250
Skulls vs Chamber Area
skulls_area_data <- cups_skulls_clean %>%
filter(!is.na(area_of_chamber_estimated) & !is.na(skulls_number))
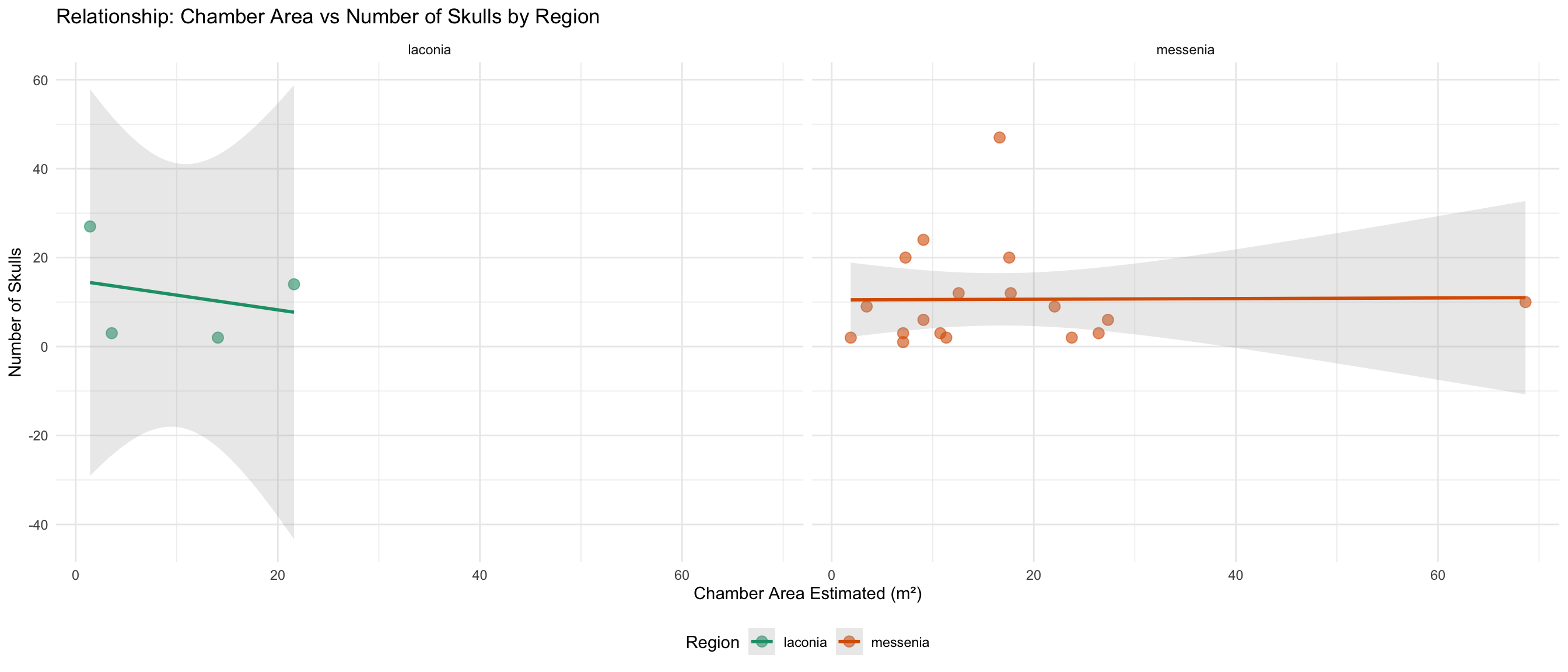
ggplot(skulls_area_data, aes(x = area_of_chamber_estimated, y = skulls_number, color = region)) +
geom_point(size = 3, alpha = 0.6) +
geom_smooth(method = "lm", se = TRUE, alpha = 0.2) +
scale_color_manual(values = c(laconia = colors$laconia, messenia = colors$messenia)) +
facet_wrap(~region) +
labs(
title = "Relationship: Chamber Area vs Number of Skulls by Region",
x = "Chamber Area Estimated (m²)",
y = "Number of Skulls",
color = "Region"
) +
theme_minimal() +
theme(legend.position = "bottom")`geom_smooth()` using formula = 'y ~ x'
Past versions of skulls-area-scatter-1.png
Version
Author
Date
8d3e1c7
tinalasisi
2026-02-17
2f9f6dc
tinalasisi
2026-02-17
# By region
for (reg in unique(skulls_area_data$region)) {
region_data <- skulls_area_data %>% filter(region == reg)
cor_val <- cor(region_data$area_of_chamber_estimated, region_data$skulls_number, use = "complete.obs")
lm_result <- lm(skulls_number ~ area_of_chamber_estimated, data = region_data)
cat("\n===", toupper(reg), "===\n")
cat("Pearson correlation (r):", round(cor_val, 3), "\n")
print(summary(lm_result))
}
=== LACONIA ===
Pearson correlation (r): -0.267
Call:
lm(formula = skulls_number ~ area_of_chamber_estimated, data = region_data)
Residuals:
1 2 3 4
6.281 -8.208 -10.682 12.609
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 14.8606 11.0108 1.350 0.310
area_of_chamber_estimated -0.3307 0.8450 -0.391 0.733
Residual standard error: 13.78 on 2 degrees of freedom
Multiple R-squared: 0.07112, Adjusted R-squared: -0.3933
F-statistic: 0.1531 on 1 and 2 DF, p-value: 0.7333
=== MESSENIA ===
Pearson correlation (r): 0.009
Call:
lm(formula = skulls_number ~ area_of_chamber_estimated, data = region_data)
Residuals:
Min 1Q Median 3Q Max
-9.542 -7.653 -3.103 1.409 36.389
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 10.491027 4.206518 2.494 0.024 *
area_of_chamber_estimated 0.007211 0.189755 0.038 0.970
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 11.78 on 16 degrees of freedom
Multiple R-squared: 9.024e-05, Adjusted R-squared: -0.0624
F-statistic: 0.001444 on 1 and 16 DF, p-value: 0.9702
Drinking Vessels vs Chamber Area
vessels_area_data <- cups_skulls_clean %>%
filter(!is.na(area_of_chamber_estimated) & !is.na(drinking_vessel_number))
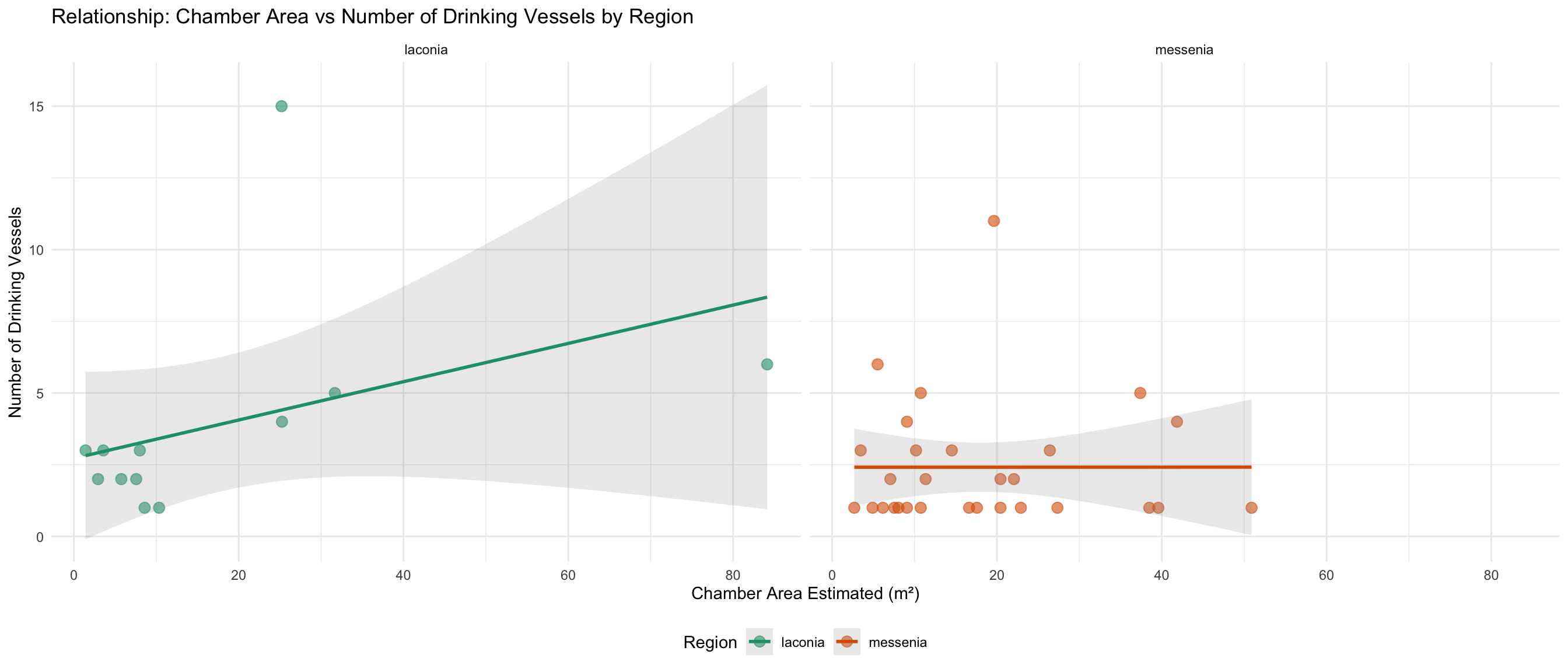
ggplot(vessels_area_data, aes(x = area_of_chamber_estimated, y = drinking_vessel_number, color = region)) +
geom_point(size = 3, alpha = 0.6) +
geom_smooth(method = "lm", se = TRUE, alpha = 0.2) +
scale_color_manual(values = c(laconia = colors$laconia, messenia = colors$messenia)) +
facet_wrap(~region) +
labs(
title = "Relationship: Chamber Area vs Number of Drinking Vessels by Region",
x = "Chamber Area Estimated (m²)",
y = "Number of Drinking Vessels",
color = "Region"
) +
theme_minimal() +
theme(legend.position = "bottom")`geom_smooth()` using formula = 'y ~ x'
Past versions of vessels-area-scatter-1.png
Version
Author
Date
8d3e1c7
tinalasisi
2026-02-17
2f9f6dc
tinalasisi
2026-02-17
# By region
for (reg in unique(vessels_area_data$region)) {
region_data <- vessels_area_data %>% filter(region == reg)
cor_val <- cor(region_data$area_of_chamber_estimated, region_data$drinking_vessel_number, use = "complete.obs")
lm_result <- lm(drinking_vessel_number ~ area_of_chamber_estimated, data = region_data)
cat("\n===", toupper(reg), "===\n")
cat("Pearson correlation (r):", round(cor_val, 3), "\n")
print(summary(lm_result))
}
=== LACONIA ===
Pearson correlation (r): 0.406
Call:
lm(formula = drinking_vessel_number ~ area_of_chamber_estimated,
data = region_data)
Residuals:
Min 1Q Median 3Q Max
-2.4141 -1.4954 -0.6643 0.0692 10.5935
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 2.72350 1.35122 2.016 0.0715 .
area_of_chamber_estimated 0.06679 0.04750 1.406 0.1900
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 3.643 on 10 degrees of freedom
Multiple R-squared: 0.1651, Adjusted R-squared: 0.08157
F-statistic: 1.977 on 1 and 10 DF, p-value: 0.19
=== MESSENIA ===
Pearson correlation (r): 0
Call:
lm(formula = drinking_vessel_number ~ area_of_chamber_estimated,
data = region_data)
Residuals:
Min 1Q Median 3Q Max
-1.4161 -1.4137 -1.4127 0.5868 8.5861
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 2.413e+00 7.226e-01 3.339 0.00247 **
area_of_chamber_estimated 7.082e-05 3.268e-02 0.002 0.99829
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 2.253 on 27 degrees of freedom
Multiple R-squared: 1.739e-07, Adjusted R-squared: -0.03704
F-statistic: 4.695e-06 on 1 and 27 DF, p-value: 0.9983
Both Skulls and Vessels vs Chamber Area
both_area_data <- cups_skulls_clean %>%
filter(!is.na(area_of_chamber_estimated) & !is.na(skulls_number) & !is.na(drinking_vessel_number))
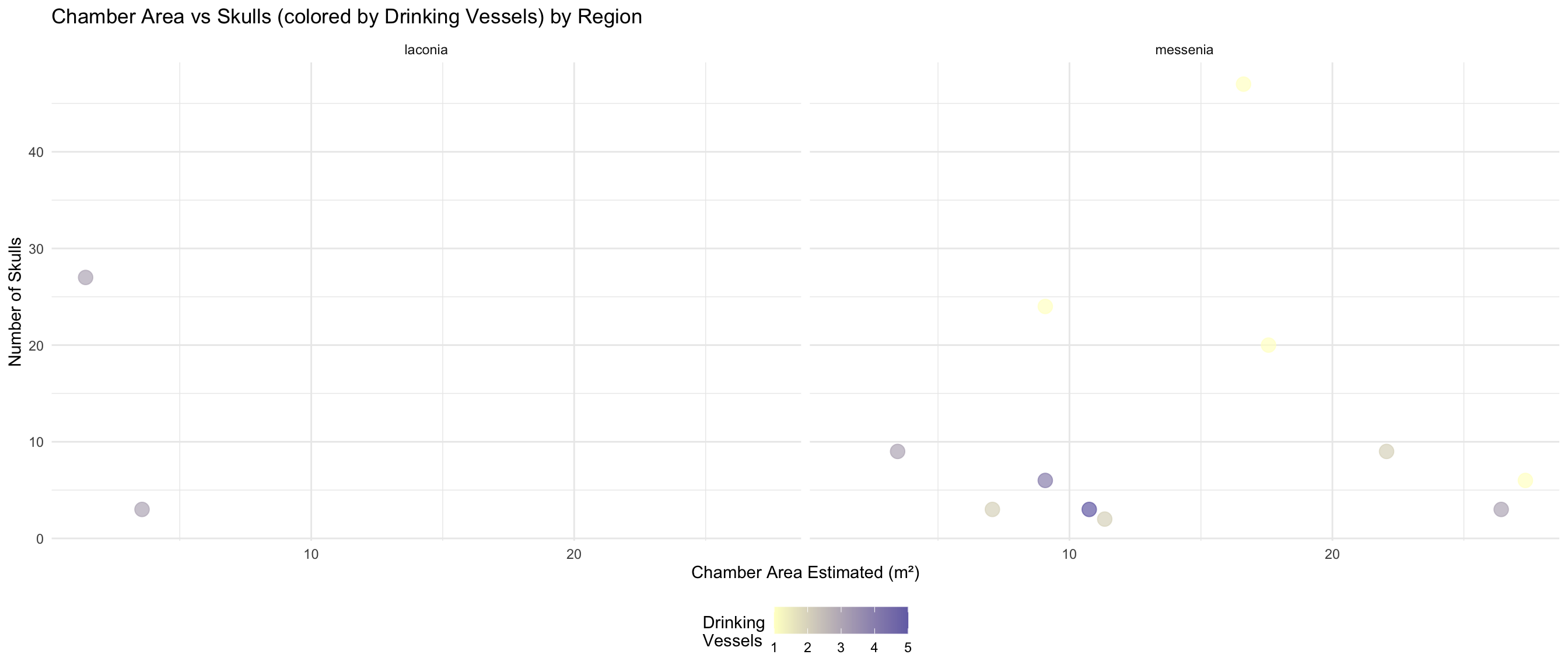
ggplot(both_area_data, aes(x = area_of_chamber_estimated, y = skulls_number)) +
geom_point(aes(color = drinking_vessel_number), size = 4, alpha = 0.7) +
scale_color_gradient(name = "Drinking\nVessels", low = colors$background, high = colors$drinking_vessel) +
facet_wrap(~region) +
labs(
title = "Chamber Area vs Skulls (colored by Drinking Vessels) by Region",
x = "Chamber Area Estimated (m²)",
y = "Number of Skulls"
) +
theme_minimal() +
theme(legend.position = "bottom")
Past versions of both-area-scatter-1.png
Version
Author
Date
8d3e1c7
tinalasisi
2026-02-17
2f9f6dc
tinalasisi
2026-02-17
Session Info
sessionInfo()R version 4.5.2 (2025-10-31)
Platform: aarch64-apple-darwin20
Running under: macOS Tahoe 26.2
Matrix products: default
BLAS: /System/Library/Frameworks/Accelerate.framework/Versions/A/Frameworks/vecLib.framework/Versions/A/libBLAS.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.1
locale:
[1] en_US/en_US/en_US/C/en_US/en_US
time zone: America/Detroit
tzcode source: internal
attached base packages:
[1] stats graphics grDevices datasets utils methods base
other attached packages:
[1] knitr_1.50 janitor_2.2.1 tidyr_1.3.1 ggplot2_4.0.1 dplyr_1.1.4
[6] readr_2.1.5 stringr_1.6.0
loaded via a namespace (and not attached):
[1] utf8_1.2.6 sass_0.4.10 generics_0.1.4 renv_1.1.6
[5] lattice_0.22-7 stringi_1.8.7 hms_1.1.3 digest_0.6.37
[9] magrittr_2.0.4 evaluate_1.0.3 grid_4.5.2 timechange_0.3.0
[13] RColorBrewer_1.1-3 fastmap_1.2.0 Matrix_1.7-4 rprojroot_2.0.4
[17] workflowr_1.7.1 jsonlite_2.0.0 whisker_0.4.1 promises_1.3.3
[21] mgcv_1.9-3 purrr_1.2.0 scales_1.4.0 jquerylib_0.1.4
[25] cli_3.6.5 crayon_1.5.3 rlang_1.1.7 splines_4.5.2
[29] bit64_4.6.0-1 withr_3.0.2 cachem_1.1.0 yaml_2.3.10
[33] parallel_4.5.2 tools_4.5.2 tzdb_0.5.0 httpuv_1.6.16
[37] here_1.0.1 vctrs_0.7.1 R6_2.6.1 lifecycle_1.0.5
[41] lubridate_1.9.4 git2r_0.36.2 snakecase_0.11.1 bit_4.6.0
[45] fs_1.6.6 vroom_1.6.5 pkgconfig_2.0.3 pillar_1.11.1
[49] bslib_0.9.0 later_1.4.2 gtable_0.3.6 glue_1.8.0
[53] Rcpp_1.1.1 xfun_0.54 tibble_3.3.1 tidyselect_1.2.1
[57] dichromat_2.0-0.1 farver_2.1.2 nlme_3.1-168 htmltools_0.5.8.1
[61] labeling_0.4.3 rmarkdown_2.29 compiler_4.5.2 S7_0.2.0
sessionInfo()R version 4.5.2 (2025-10-31)
Platform: aarch64-apple-darwin20
Running under: macOS Tahoe 26.2
Matrix products: default
BLAS: /System/Library/Frameworks/Accelerate.framework/Versions/A/Frameworks/vecLib.framework/Versions/A/libBLAS.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.1
locale:
[1] en_US/en_US/en_US/C/en_US/en_US
time zone: America/Detroit
tzcode source: internal
attached base packages:
[1] stats graphics grDevices datasets utils methods base
other attached packages:
[1] knitr_1.50 janitor_2.2.1 tidyr_1.3.1 ggplot2_4.0.1 dplyr_1.1.4
[6] readr_2.1.5 stringr_1.6.0
loaded via a namespace (and not attached):
[1] utf8_1.2.6 sass_0.4.10 generics_0.1.4 renv_1.1.6
[5] lattice_0.22-7 stringi_1.8.7 hms_1.1.3 digest_0.6.37
[9] magrittr_2.0.4 evaluate_1.0.3 grid_4.5.2 timechange_0.3.0
[13] RColorBrewer_1.1-3 fastmap_1.2.0 Matrix_1.7-4 rprojroot_2.0.4
[17] workflowr_1.7.1 jsonlite_2.0.0 whisker_0.4.1 promises_1.3.3
[21] mgcv_1.9-3 purrr_1.2.0 scales_1.4.0 jquerylib_0.1.4
[25] cli_3.6.5 crayon_1.5.3 rlang_1.1.7 splines_4.5.2
[29] bit64_4.6.0-1 withr_3.0.2 cachem_1.1.0 yaml_2.3.10
[33] parallel_4.5.2 tools_4.5.2 tzdb_0.5.0 httpuv_1.6.16
[37] here_1.0.1 vctrs_0.7.1 R6_2.6.1 lifecycle_1.0.5
[41] lubridate_1.9.4 git2r_0.36.2 snakecase_0.11.1 bit_4.6.0
[45] fs_1.6.6 vroom_1.6.5 pkgconfig_2.0.3 pillar_1.11.1
[49] bslib_0.9.0 later_1.4.2 gtable_0.3.6 glue_1.8.0
[53] Rcpp_1.1.1 xfun_0.54 tibble_3.3.1 tidyselect_1.2.1
[57] dichromat_2.0-0.1 farver_2.1.2 nlme_3.1-168 htmltools_0.5.8.1
[61] labeling_0.4.3 rmarkdown_2.29 compiler_4.5.2 S7_0.2.0