Analysis
Tina Lasisi
August 11, 2022
Last updated: 2022-08-11
Checks: 6 1
Knit directory: HairManikin2022/
This reproducible R Markdown analysis was created with workflowr (version 1.7.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of
the R Markdown file created these results, you’ll want to first commit
it to the Git repo. If you’re still working on the analysis, you can
ignore this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20211024) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 9932591. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .Rproj.user/
Untracked files:
Untracked: data/current/March2022Sheet.xlsx
Untracked: data/current/reworked.csv
Untracked: data/current/reworked.xlsx
Untracked: data/current/reworkedUsable.csv
Untracked: data/current/reworkedUsable2.csv
Untracked: data/current/~$March2022Sheet.xlsx
Untracked: dataPrep.rmd
Unstaged changes:
Modified: analysis/analysis.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (analysis/analysis.Rmd) and HTML
(docs/analysis.html) files. If you’ve configured a remote
Git repository (see ?wflow_git_remote), click on the
hyperlinks in the table below to view the files as they were in that
past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| html | 6e70659 | Ben Zydney | 2022-08-06 | Updated Website |
| Rmd | 9db041e | Ben Zydney | 2022-08-06 | Reworked hair analysis calculations. |
| Rmd | dd0e722 | Tina Lasisi | 2022-03-13 | edited df creation |
| html | dd0e722 | Tina Lasisi | 2022-03-13 | edited df creation |
| Rmd | 210cee3 | Tina Lasisi | 2022-03-10 | updated figure size |
| html | 210cee3 | Tina Lasisi | 2022-03-10 | updated figure size |
| Rmd | bf62a07 | Tina Lasisi | 2022-03-10 | Updating analyses + figures |
| html | bf62a07 | Tina Lasisi | 2022-03-10 | Updating analyses + figures |
| html | 1a1f7bc | Tina Lasisi | 2022-03-07 | Build site. |
| Rmd | 7478e4c | Tina Lasisi | 2022-03-07 | updating analyses with models |
| html | c0ce5d2 | Tina Lasisi | 2022-03-06 | Build site. |
| Rmd | 4041aec | Tina Lasisi | 2022-03-06 | wflow_publish(files = "analysis/*", all = TRUE, republish = TRUE, |
| html | 4041aec | Tina Lasisi | 2022-03-06 | wflow_publish(files = "analysis/*", all = TRUE, republish = TRUE, |
| Rmd | a796ceb | Ginawsy | 2022-02-21 | updated sum_manikin_df variable |
| Rmd | 520dcfc | GitHub | 2022-02-16 | Update analysis.Rmd |
| Rmd | 9c5b0d8 | Tina Lasisi | 2022-02-15 | Update analysis + figures |
| html | 9c5b0d8 | Tina Lasisi | 2022-02-15 | Update analysis + figures |
| Rmd | e9fa430 | GitHub | 2022-01-30 | Update analysis.Rmd |
| html | bb36720 | Tina Lasisi | 2022-01-19 | Build site. |
| Rmd | 05389ae | Tina Lasisi | 2022-01-19 | Added figures + analysis placeholders |
| html | 05389ae | Tina Lasisi | 2022-01-19 | Added figures + analysis placeholders |
| Rmd | bb99f1d | Tina Lasisi | 2022-01-08 | Update analysis.rmd + add data |
| html | bb99f1d | Tina Lasisi | 2022-01-08 | Update analysis.rmd + add data |
| Rmd | 0a50ef7 | Tina Lasisi | 2022-01-07 | Adding main analysis file |
1 Preparing Data
First, we import the data and label the variables.
# Preview data
head(df_wetdry)| wind | wig | wet_dry | heat_loss | skin_temp | resistance | clo | Mean Tsk (sel zones) | amb_temp | amb_rh | radiation | trial |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0.3 | Nude | wet | 90.9 | 34 | 4.36e-08 | 2.81e-07 | 34.2 | 34 | 45.8 | on | 1 |
| 0.3 | Nude | wet | 86.7 | 34 | -0.00121 | -0.00782 | 34.2 | 34.1 | 45.8 | on | 2 |
| 1 | Nude | wet | 227 | 34 | 0.000119 | 0.000767 | 34.2 | 34 | 46.3 | on | 1 |
| 2.5 | Nude | wet | 288 | 34 | 2.85e-05 | 0.000184 | 34.2 | 34 | 47.1 | on | 1 |
| 2.5 | Nude | wet | 289 | 34 | -1.25e-06 | -8.07e-06 | 34.2 | 34 | 47 | on | 2 |
| 0.3 | Straight | wet | 30 | 34 | -0.00422 | -0.0272 | 34.2 | 34.1 | 45.4 | on | 1 |
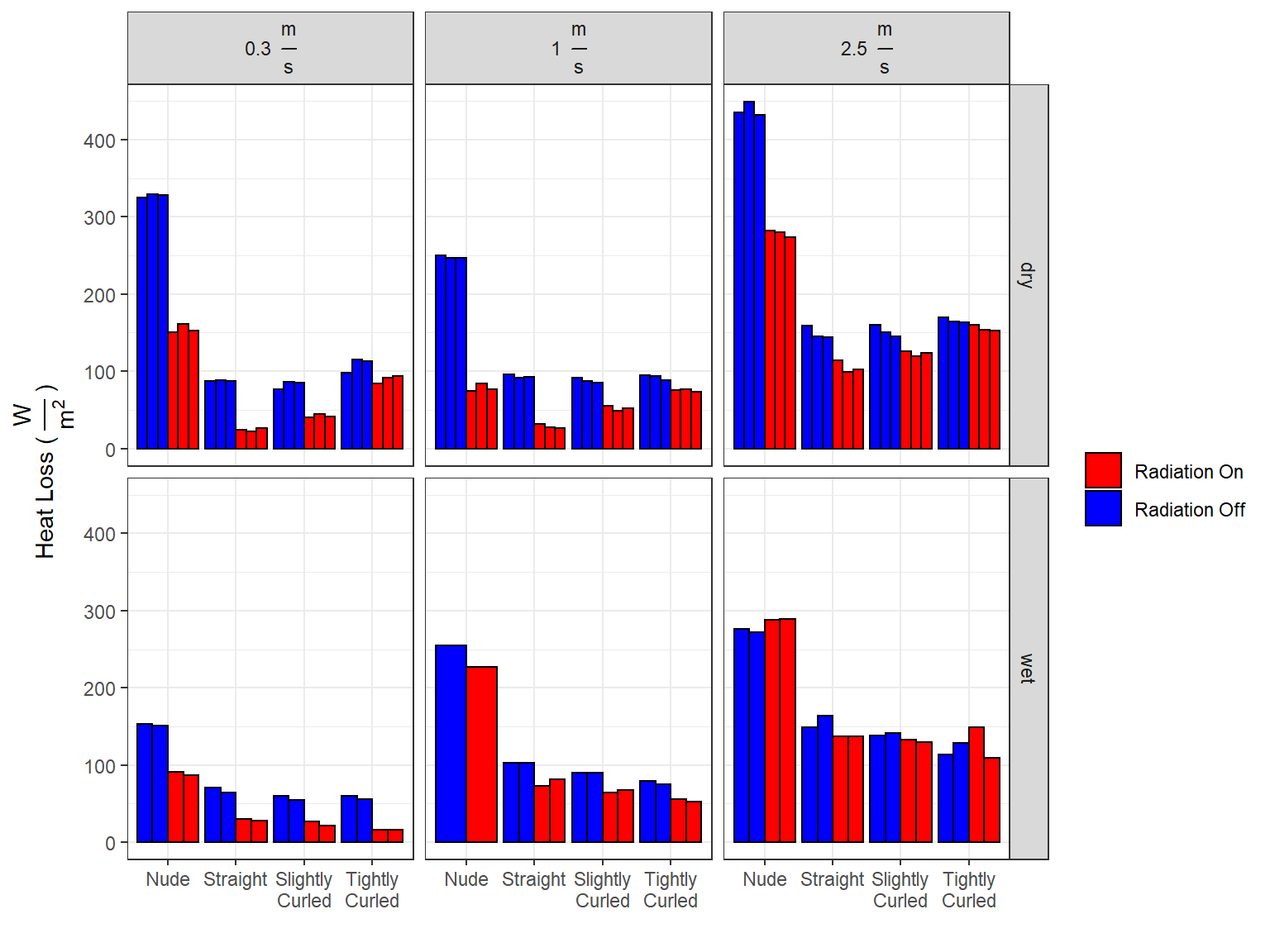
1.1 Removing Outlier
It was noticed that the 2nd trial conducted with wet, tightly curled hair, 2.5 m/s wind speed, and radiation on, had more heat loss than any of the trials with radiation off. With the understanding that radiation should always decrease heat loss, we elected to remove that data point.
# Remove specific entry
df_wetdry %>% filter((wig == "Tightly\nCurled" & wind == 2.5 & radiation == "on" & wet_dry == "wet" & trial == "1"))| wind | wig | wet_dry | heat_loss | skin_temp | resistance | clo | Mean Tsk (sel zones) | amb_temp | amb_rh | radiation | trial |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 2.5 | Tightly Curled | wet | 149 | 34 | 5.4e-05 | 0.000348 | 34.2 | 34 | 46.9 | on | 1 |
df_wetdry <- df_wetdry %>% filter(!(wig == "Tightly\nCurled" & wind == 2.5 & radiation == "on" & wet_dry == "wet" & trial == "1"))2 Calculating Thermal Resistance
\[I_t = \frac{T_{Skin} - T_{Air}}{H_{Dry}}\]
df_wetdry['dry_heat_resistance'] <- (df_wetdry['skin_temp'] - df_wetdry['amb_temp']) / df_wetdry['heat_loss']
# For the dry data, leave this blank
df_wetdry <- df_wetdry %>% mutate(dry_heat_resistance = ifelse(wet_dry == 'wet', NaN, dry_heat_resistance))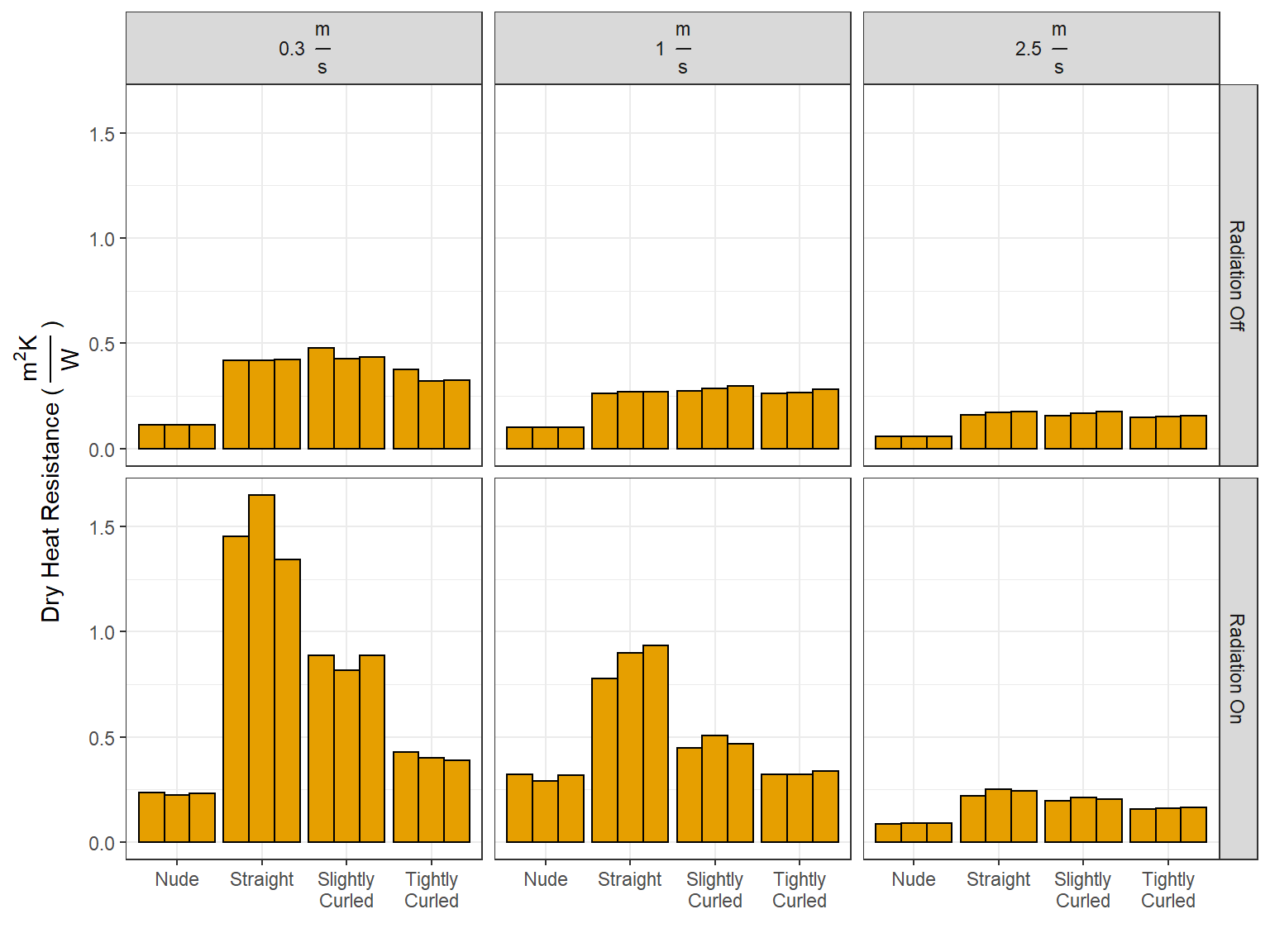
3 Calculating Net Solar Influx
\[I_{Dry} = H_{Dry} - H_{Dry}^{Solar}\] \[I_{Evap} = H_{Evap} - H_{Evap}^{Solar}\]
# Average all trials with the same characteristics
df_averaged_trials <- df_wetdry %>%
group_by(wig, wind, radiation, wet_dry) %>%
drop_na(heat_loss) %>%
summarise(heat_loss = mean(heat_loss))
# Pivot the dataframe to incldue radiation on and off as part of same event
df_radiation_split <- df_averaged_trials %>%
pivot_wider(names_from = c(radiation), values_from = c(heat_loss)) %>%
rename(heat_loss_off = off) %>%
rename(heat_loss_on = on)
# Calculate the net influx
df_net_influx_plots <- df_radiation_split %>%
group_by(wig, wind) %>%
summarise(wet_dry = wet_dry,
net_influx = heat_loss_off - heat_loss_on)
df_net_influx <- df_net_influx_plots %>% spread(wet_dry, net_influx)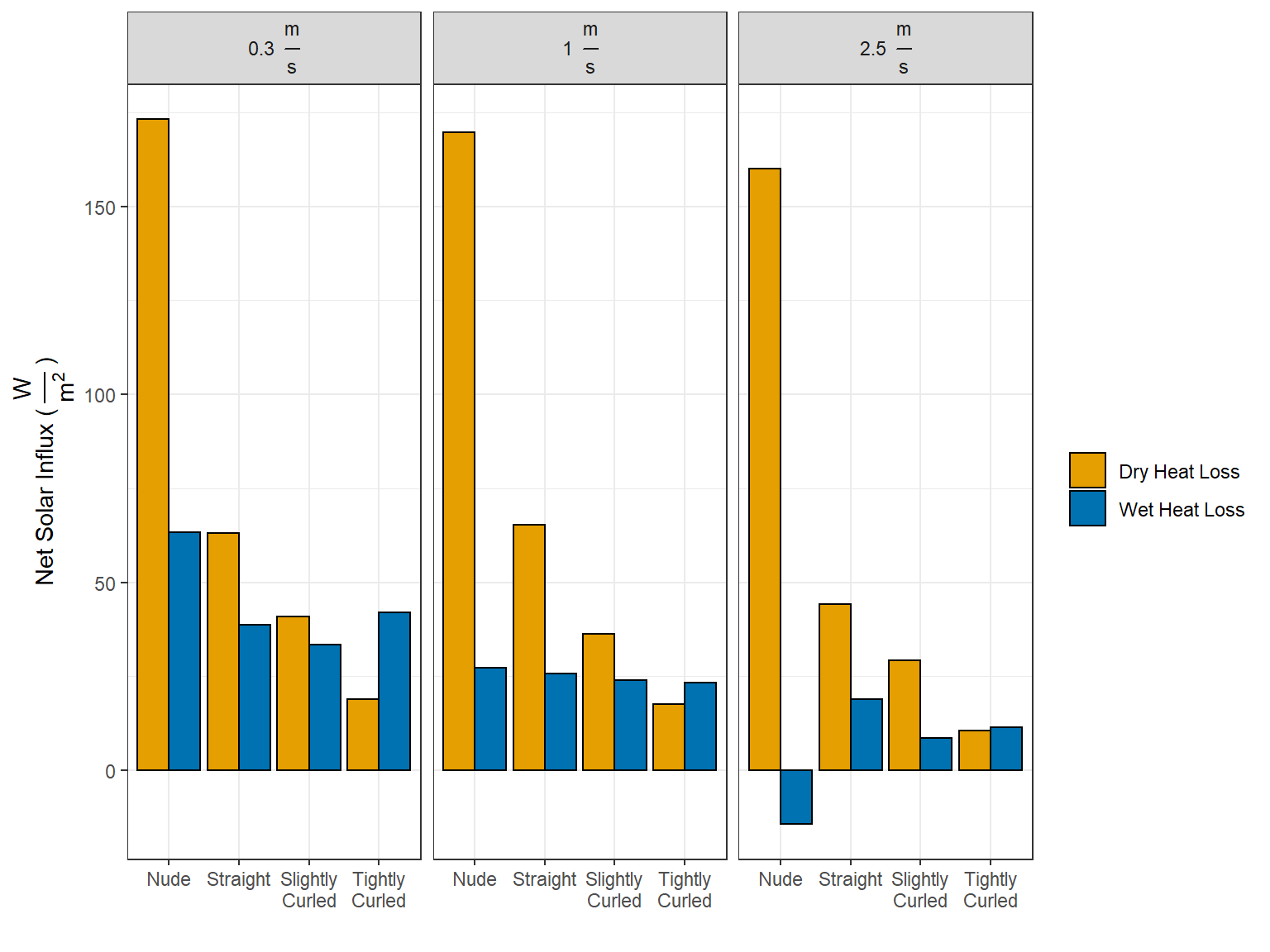
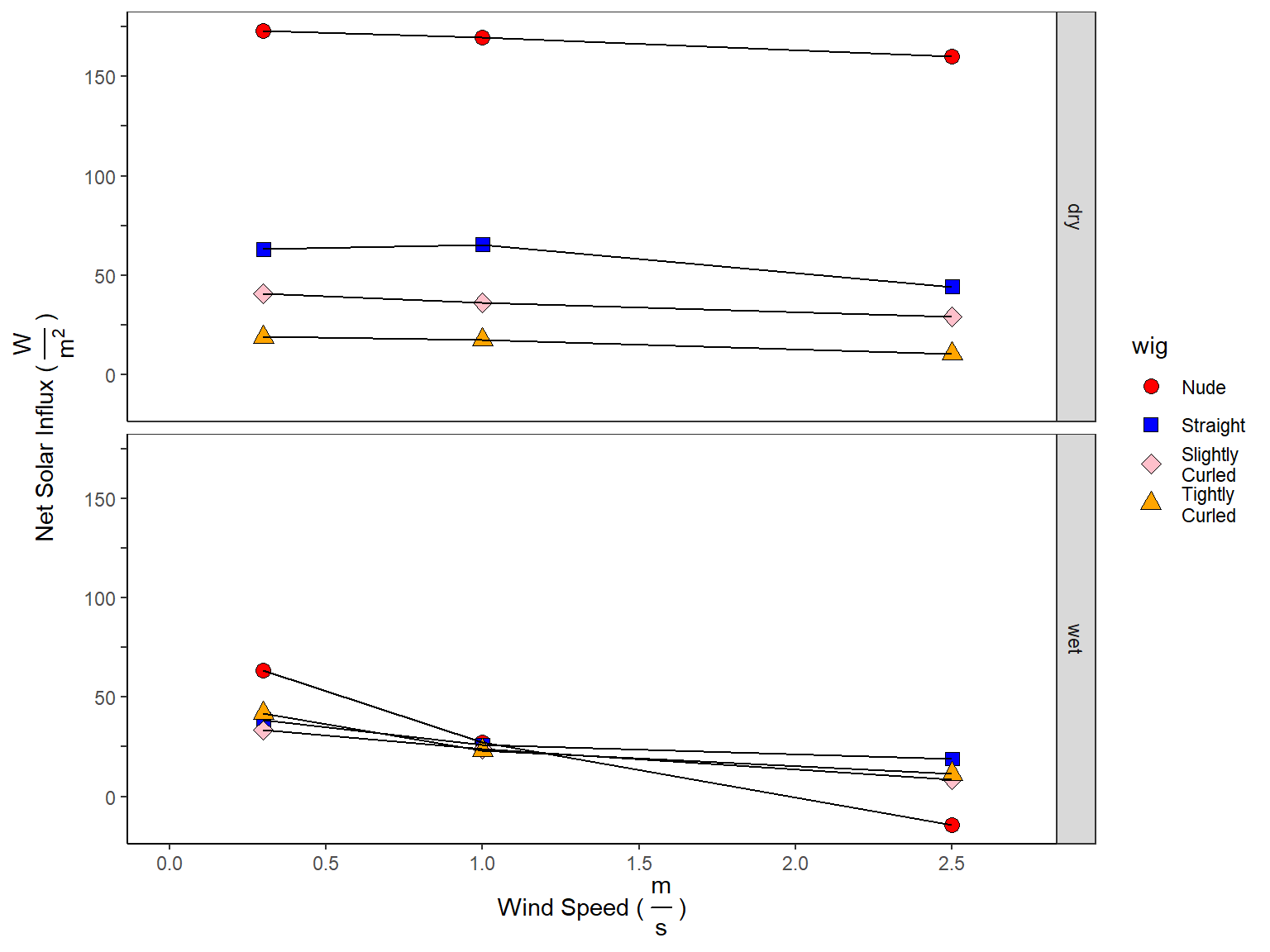
4 Adjusting Heat Losses to 30 Degrees Celsius
4.1 Dry Heat Loss
\[H_{Dry}^{30^\circ C} = \frac{35 -30}{I_t}\]
# Their calculation
df_wetdry['heat_30'] = (35 - 30) / df_wetdry['dry_heat_resistance']
# What I would expect
#df_wetdry['heat_30'] = (df_wetdry['skin_temp'] - 30) / df_wetdry['dry_heat_resistance']
# Recreate the radiation split dataframe to include heat_30
df_averaged_trials <- df_wetdry %>%
group_by(wig, wind, radiation, wet_dry) %>%
drop_na(heat_loss) %>%
summarise(heat_loss = mean(heat_loss),
heat_30 = mean(heat_30))
df_radiation_split <- df_averaged_trials %>%
pivot_wider(names_from = c(radiation), values_from = c(heat_loss, heat_30))4.2 Dry and Wet Heat Losses With Solar Radiation
\[H_{Dry}^{30^{\circ} C,\:Solar} = H_{Dry}^{30^{\circ} C} - I_{Dry}\] \[H_{Wet}^{30^{\circ} C,\:Solar} = H_{Evap + Dry}^{30^{\circ} C} = H_{Evap} + I_{Dry} + H_{Dry}^{30^{\circ} C,\:Solar}\]
dry_heat_30 = df_radiation_split[df_radiation_split$wet_dry == 'dry',]
heat_evap = df_radiation_split[df_radiation_split$wet_dry == 'wet',]
df_adjusted_solar <- data.frame(
dry_heat_loss <- dry_heat_30$heat_30_off - df_net_influx$dry,
wind <- dry_heat_30$wind,
wig <- dry_heat_30$wig
) %>% rename('dry_heat_loss' = 'dry_heat_loss....dry_heat_30.heat_30_off...df_net_influx.dry') %>%
rename('wind' = 'wind....dry_heat_30.wind') %>%
rename('wig' = 'wig....dry_heat_30.wig')
df_adjusted_solar['wet_heat_loss'] <- + heat_evap$heat_loss_on + df_net_influx$dry + df_adjusted_solar$dry_heat_loss
df_adjusted_solar_plots <-df_adjusted_solar %>%
pivot_longer(cols = c('dry_heat_loss', 'wet_heat_loss'), names_to = 'wet_dry', values_to = 'heat_loss')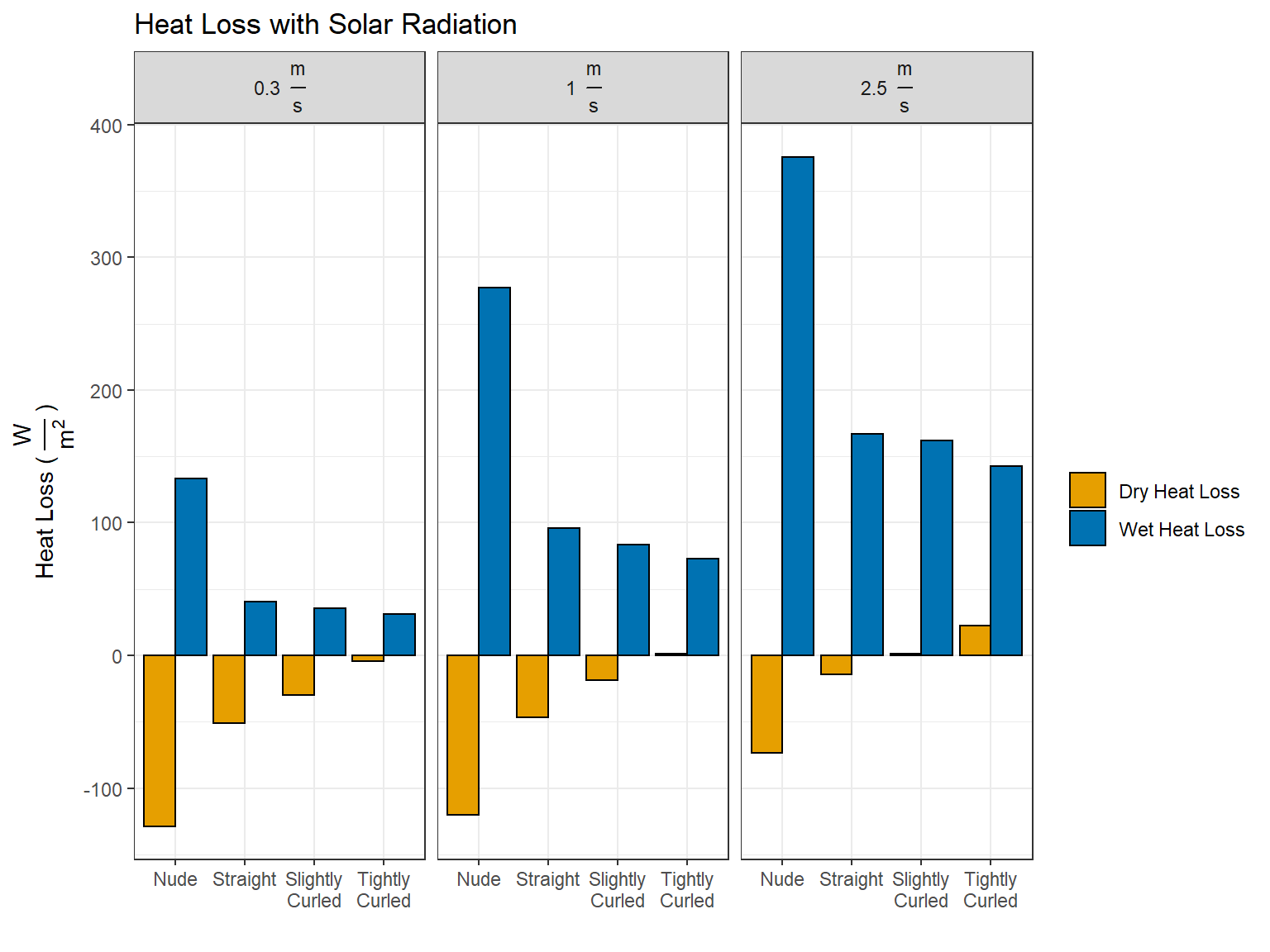
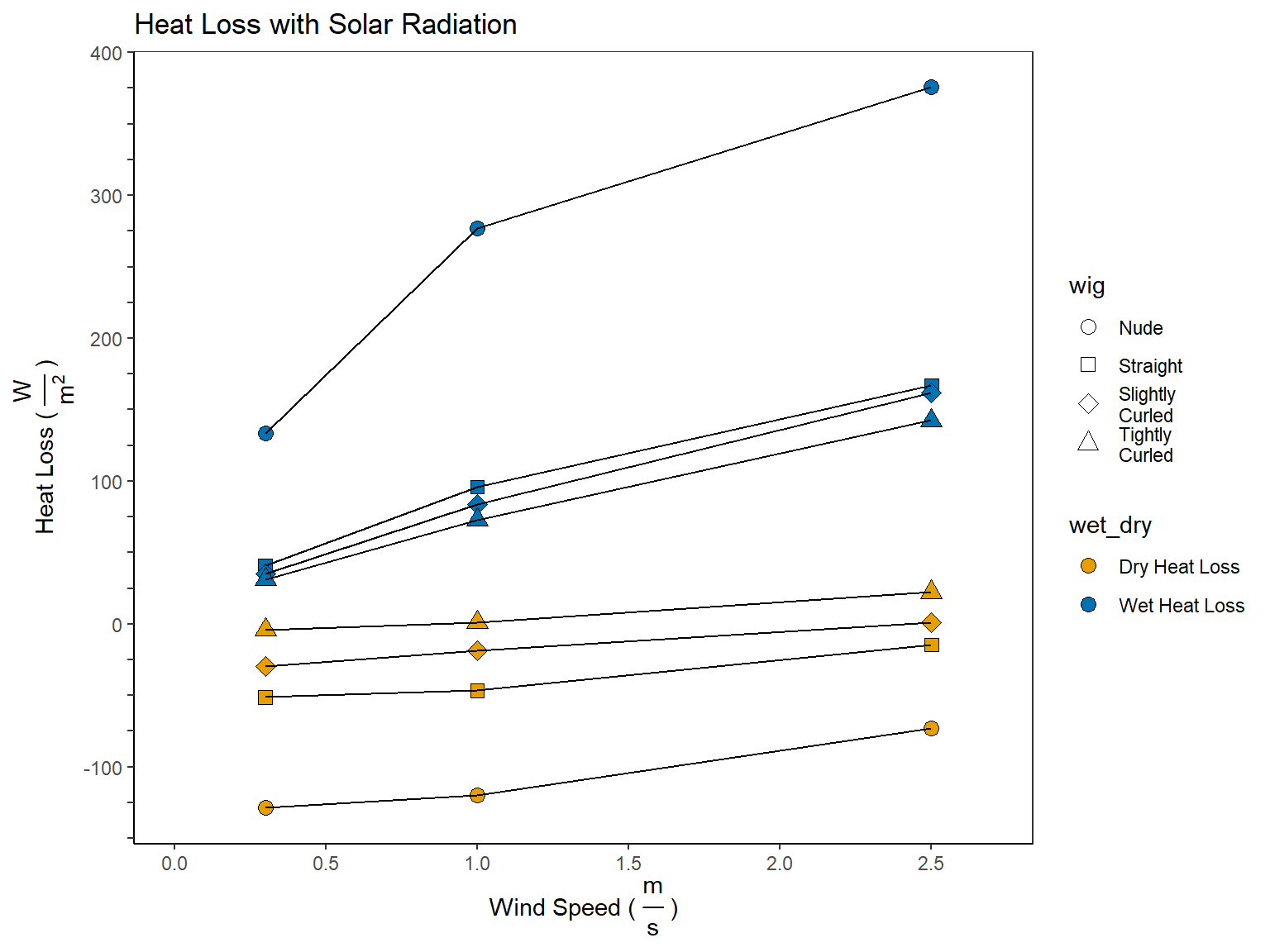
5 Calculating Evaporative Potential
\[H_{Max}^{30^{\circ} C,\:Solar} = H_{Wet}^{30^{\circ} C,\:Solar} - H_{Dry}^{30^{\circ} C,\:Solar}\]
df_evaporative_potential <- df_adjusted_solar$wet_heat_loss - df_adjusted_solar$dry_heat_loss6 Calculating Sweat Requirements
\[Sweat_{Max} = \frac{H_{Max}^{30^{\circ} C,\:Solar} * 3600}{2430}\]
\[ IF \; H_{Dry}^{30^{\circ} C,\:Solar} < 0, \; Sweat_{Zero} = -\frac{H_{Dry}^{30^{\circ} C,\:Solar} * 3600}{2430} \\ ELSE, \; Sweat_{Zero} = 0\]
# Create a new df with the sweat requirements
df_sweat_requirements <- data.frame(
sweat_max <- df_evaporative_potential * 3600 / 2430,
sweat_zero <- -3600 / 2430 * df_adjusted_solar['dry_heat_loss'],
wig <- df_adjusted_solar$wig,
wind <- df_adjusted_solar$wind
)
#Rename columns
colnames(df_sweat_requirements) <- c('sweat_max', 'sweat_zero')
#Replace all values less than 0 with 0 per formula
df_sweat_requirements['sweat_zero'][df_sweat_requirements['sweat_zero'] < 0] <- 0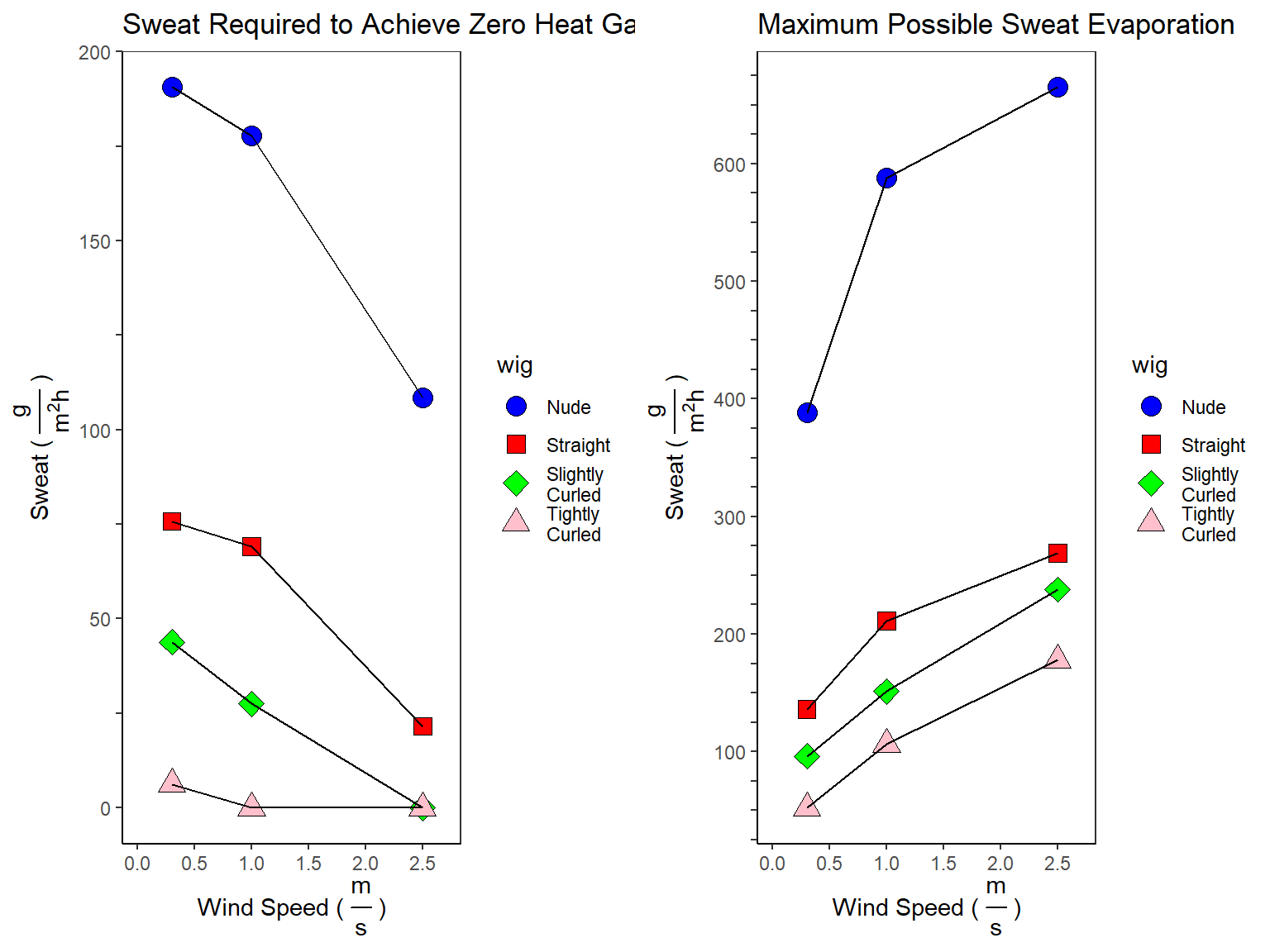
R version 4.2.1 (2022-06-23 ucrt)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 22000)
Matrix products: default
locale:
[1] LC_COLLATE=English_United States.utf8
[2] LC_CTYPE=English_United States.utf8
[3] LC_MONETARY=English_United States.utf8
[4] LC_NUMERIC=C
[5] LC_TIME=English_United States.utf8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] huxtable_5.5.0 broom.mixed_0.2.9.4 patchwork_1.1.1
[4] gridExtra_2.3 fs_1.5.2 knitr_1.39
[7] forcats_0.5.1 stringr_1.4.0 dplyr_1.0.9
[10] purrr_0.3.4 readr_2.1.2 tidyr_1.2.0
[13] tibble_3.1.8 ggplot2_3.3.6 tidyverse_1.3.2
loaded via a namespace (and not attached):
[1] nlme_3.1-157 bit64_4.0.5 lubridate_1.8.0
[4] httr_1.4.3 rprojroot_2.0.3 tools_4.2.1
[7] backports_1.4.1 bslib_0.4.0 utf8_1.2.2
[10] R6_2.5.1 DBI_1.1.3 colorspace_2.0-3
[13] withr_2.5.0 tidyselect_1.1.2 bit_4.0.4
[16] compiler_4.2.1 git2r_0.30.1 cli_3.3.0
[19] rvest_1.0.2 xml2_1.3.3 labeling_0.4.2
[22] sass_0.4.2 scales_1.2.0 commonmark_1.8.0
[25] digest_0.6.29 rmarkdown_2.14 pkgconfig_2.0.3
[28] htmltools_0.5.3 parallelly_1.32.1 highr_0.9
[31] dbplyr_2.2.1 fastmap_1.1.0 rlang_1.0.4
[34] readxl_1.4.0 rstudioapi_0.13 farver_2.1.1
[37] jquerylib_0.1.4 generics_0.1.3 jsonlite_1.8.0
[40] vroom_1.5.7 googlesheets4_1.0.0 magrittr_2.0.3
[43] Rcpp_1.0.9 munsell_0.5.0 fansi_1.0.3
[46] lifecycle_1.0.1 furrr_0.3.0 stringi_1.7.8
[49] whisker_0.4 yaml_2.3.5 grid_4.2.1
[52] paletteer_1.4.0 parallel_4.2.1 listenv_0.8.0
[55] promises_1.2.0.1 crayon_1.5.1 lattice_0.20-45
[58] haven_2.5.0 splines_4.2.1 hms_1.1.1
[61] pillar_1.8.0 codetools_0.2-18 reprex_2.0.1
[64] glue_1.6.2 evaluate_0.15 modelr_0.1.8
[67] vctrs_0.4.1 tzdb_0.3.0 httpuv_1.6.5
[70] cellranger_1.1.0 gtable_0.3.0 rematch2_2.1.2
[73] future_1.27.0 assertthat_0.2.1 cachem_1.0.6
[76] xfun_0.31 broom_1.0.0 later_1.3.0
[79] googledrive_2.0.0 gargle_1.2.0 workflowr_1.7.0
[82] globals_0.15.1 ellipsis_0.3.2