Last updated: 2026-04-07
Checks: 6 1
Knit directory:
Integrating-nir-genomic-kernel/
This reproducible R Markdown analysis was created with workflowr (version 1.7.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of
the R Markdown file created these results, you’ll want to first commit
it to the Git repo. If you’re still working on the analysis, you can
ignore this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20250829) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version d4d1044. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: analysis/analysis.Rmd
Ignored: analysis/run_cv0_cv00_only.R
Ignored: analysis/run_cv1_cv2_only.R
Ignored: data/Article_documents/
Ignored: data/Maize-NIRS-GBS-main/
Ignored: output/Matrizes/Geno.all.rds
Ignored: output/Matrizes/NIR.all.rds
Ignored: output/Matrizes/figures/
Ignored: output/adjust_models.rar
Ignored: output/climate_results/additional_climate_plots.tiff
Ignored: output/climate_results/combined_additional_climate_plots.tiff
Ignored: output/climate_results/combined_climate_plot.tiff
Ignored: output/cv-schemes.tiff
Untracked files:
Untracked: output/figures/
Untracked: output/results/
Untracked: output/tables/
Untracked: output/variance_components/
Unstaged changes:
Modified: analysis/analysis_prediction.Rmd
Modified: analysis/analysis_prediction_pt.Rmd
Modified: analysis/analysis_prediction_run_outputs.Rmd
Modified: analysis/analysis_prediction_run_outputs_pt.Rmd
Modified: analysis/variance_components.Rmd
Modified: analysis/variance_components_pt.Rmd
Modified: analysis/visualization.Rmd
Modified: analysis/visualization_pt.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (analysis/variance_components.Rmd)
and HTML (docs/variance_components.html) files. If you’ve
configured a remote Git repository (see ?wflow_git_remote),
click on the hyperlinks in the table below to view the files as they
were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 2d2b88e | WevertonGomesCosta | 2026-04-06 | update |
| Rmd | 4fb14e1 | WevertonGomesCosta | 2026-04-06 | update variance components |
| html | 4fb14e1 | WevertonGomesCosta | 2026-04-06 | update variance components |
| Rmd | d1beaba | WevertonGomesCosta | 2026-04-06 | update |
| html | d1beaba | WevertonGomesCosta | 2026-04-06 | update |
| html | a7d077a | WevertonGomesCosta | 2026-04-03 | update |
| Rmd | b3df8b2 | WevertonGomesCosta | 2026-04-03 | update |
| Rmd | 0a6059c | WevertonGomesCosta | 2026-04-03 | update |
| html | f6d5560 | WevertonGomesCosta | 2026-04-03 | update files |
| html | 005c393 | WevertonGomesCosta | 2026-04-02 | update |
| html | d54afd7 | WevertonGomesCosta | 2026-04-01 | update |
| Rmd | 00ef3ef | WevertonGomesCosta | 2026-04-01 | update |
| html | 00ef3ef | WevertonGomesCosta | 2026-04-01 | update |
| Rmd | 1915912 | WevertonGomesCosta | 2026-03-31 | update |
| Rmd | 9827bcc | WevertonGomesCosta | 2026-03-31 | update scripts rmd |
| html | 4e5af9a | WevertonGomesCosta | 2026-03-27 | update |
| Rmd | 640dfbe | WevertonGomesCosta | 2026-03-27 | update |
| Rmd | 0a6715e | WevertonGomesCosta | 2026-03-27 | update |
| html | 3573a52 | WevertonGomesCosta | 2026-03-27 | update |
| Rmd | 717d6b9 | WevertonGomesCosta | 2026-02-24 | update |
| html | 717d6b9 | WevertonGomesCosta | 2026-02-24 | update |
| Rmd | c996e1e | WevertonGomesCosta | 2026-02-24 | update |
| Rmd | 5b10ff7 | WevertonGomesCosta | 2026-02-24 | update |
| Rmd | 1a46586 | WevertonGomesCosta | 2025-11-04 | update variance_components .rmd and .html |
| html | 1a46586 | WevertonGomesCosta | 2025-11-04 | update variance_components .rmd and .html |
| Rmd | f18937a | WevertonGomesCosta | 2025-11-04 | chance components_variance to variance_components |
The document starts by setting knitting options so that tables, code, and figures are displayed consistently throughout the tutorial.
Language / Idioma: English | Português
1. Introduction
This tutorial summarizes variance components for the predictive models used in this study. In the current workflow, this file is used mainly to consolidate and interpret results that were already estimated previously, although it still keeps the code structure needed to fit the models when that step must be rerun.
The goal is to quantify how much of the total phenotypic variance is attributed to the main effects and interaction terms included in each model. In practical terms, this script helps answer questions such as:
- How much variance is associated with the categorical environment effect (E)?
- How much is captured by the genomic kernels?
- How much is captured by the phenomic/NIRS kernels (P)?
- How much is associated with the weather kernel (W), which is used mainly in models Eta10-Eta18?
- How relevant are the interaction terms such as G × E, G × W, P × E, and P × W?
This script is organized into four stages:
- Load kernels and phenotypic data produced in
matrizes.Rmd. - Define the 18 variance-component models.
- Read or rerun the BGLR outputs, depending on whether the saved posterior summaries already exist.
- Process and visualize the variance decomposition for Grain Yield (GY) and Kernel Weight (KW).
A high variance share for a given source should not be interpreted automatically as evidence of biological complementarity or better predictive performance. This point is especially important for P, because NIRS-based similarity may reflect strong physiological response to the environment in addition to any stable biological signal, and for W, because the weather kernel complements but does not replace the categorical environment effect.
Inputs, outputs and next step
Inputs of this step - matrices and kernels generated
by matrizes.Rmd - the reduced catalog of 18 models used in
the project
Outputs of this step - raw variance-component results - processed variance-component summaries - percentage tables and diagnostic figures
Next step - this stage complements the structural
interpretation of the models and should be read together with the
predictive results generated later in
analysis_prediction.Rmd.
2. R Environment and Required Inputs
First, we load the packages used to manipulate the data, fit the Bayesian models, and generate the final article-quality figures.
The next step loads the packages needed for reading the saved artifacts, reshaping the results, plotting summaries, and running the heavy estimation stage when required.
library(dplyr)
library(tidyr)
library(readr)
library(stringr)
library(tibble)
library(ggplot2)
library(BGLR)
library(parallel)
library(doParallel)
library(foreach)
library(ggthemes)
library(scales)
library(fs)
library(grid)
library(knitr)
library(kableExtra)2.1. File paths
All kernels and phenotypic objects are expected to be available in
output/Matrizes/, which is the output directory created in
matrizes.Rmd. The variance-component results generated here
are stored in output/variance_components/.
The following lines define the project directories used to find matrices, store outputs, and organize the saved variance-component files.
kernel_dir <- "output/Matrizes"
results_dir <- "output/variance_components"
figure_dir <- file.path(results_dir, "figures")
bglr_runs_dir <- file.path(results_dir, "bglr_runs")
dir.create(results_dir, recursive = TRUE, showWarnings = FALSE)
dir.create(bglr_runs_dir, recursive = TRUE, showWarnings = FALSE)
dir.create(figure_dir, recursive = TRUE, showWarnings = FALSE)2.2. Input checks
Before reading the files, we check whether the expected
.rds objects are present in the matrices output folder.
This makes the tutorial easier to follow because the script stops
immediately if a required object is missing.
Before moving into model definition or result processing, the script checks that the required input files and directories are available.
required_files <- c(
"ZG.rds", "ZP.rds", "ZE.rds", "ZEZE.rds", "ZW.rds",
"ZGZE.rds", "ZPZE.rds", "ZGZW.rds", "ZPZW.rds",
"GGK.rds", "PGK.rds", "GAK.rds", "PAK.rds",
"GGKE.rds", "PGKE.rds", "GGKW.rds", "PGKW.rds",
"GAKE.rds", "PAKE.rds", "GAKW.rds", "PAKW.rds",
"Pheno.rds", "Pedigree.rds"
)
missing_files <- required_files[!file.exists(file.path(kernel_dir, required_files))]
if (length(missing_files) > 0) {
stop(
"The following required files were not found in output/Matrizes/: ",
paste(missing_files, collapse = ", "),
". Run matrizes.Rmd first."
)
}2.3. Labels used in the final tables and figures
These vectors define how the variance components will appear in the final outputs. Keeping these labels visible in the main script helps the reader understand how internal model names are translated into article-ready labels.
Here the script defines the labels and naming conventions that will be reused in tables, output files, and figures.
component_display <- c(
E = "Environment (E)",
G = "Genomic linear (G)",
P = "Phenomic linear (P)",
W = "Weather kernel (W)",
GE = "G × E",
PE = "P × E",
GW = "G × W",
PW = "P × W",
GGK = "Genomic Gaussian (GGK)",
PGK = "Phenomic Gaussian (PGK)",
GAK = "Genomic arc-cosine (GAK)",
PAK = "Phenomic arc-cosine (PAK)",
GGKE = "GGK × E",
PGKE = "PGK × E",
GAKE = "GAK × E",
PAKE = "PAK × E",
GGKW = "GGK × W",
PGKW = "PGK × W",
GAKW = "GAK × W",
PAKW = "PAK × W",
Residual = "Residual"
)
component_order <- c(
"Environment (E)",
"Genomic linear (G)",
"Phenomic linear (P)",
"Weather kernel (W)",
"G × E",
"P × E",
"G × W",
"P × W",
"Genomic Gaussian (GGK)",
"Phenomic Gaussian (PGK)",
"Genomic arc-cosine (GAK)",
"Phenomic arc-cosine (PAK)",
"GGK × E",
"PGK × E",
"GAK × E",
"PAK × E",
"GGK × W",
"PGK × W",
"GAK × W",
"PAKW × W",
"Residual"
)
component_order[component_order == "PAKW × W"] <- "PAK × W"
normalize_component_name <- function(component_name, model_value = NA_integer_) {
component_name <- as.character(component_name)
if (length(model_value) == 1L && length(component_name) > 1L) {
model_value <- rep(model_value, length(component_name))
}
if (length(model_value) != length(component_name)) {
stop("normalize_component_name() requires 'component_name' and 'model_value' to have the same length, or a scalar 'model_value'.")
}
vapply(seq_along(component_name), function(i) {
current_component <- component_name[i]
current_model <- model_value[i]
if (is.factor(current_model)) {
current_model <- as.character(current_model)
}
if (is.character(current_model)) {
current_model <- suppressWarnings(as.integer(sub("^Eta", "", current_model)))
}
if (length(current_component) == 0 || is.na(current_component) || !nzchar(current_component)) {
return(current_component)
}
if (length(current_model) == 1L && !is.na(current_model) && grepl("^ETA_[0-9]+$", current_component)) {
effect_index <- suppressWarnings(as.integer(sub("^ETA_", "", current_component)))
if (!is.na(effect_index) && current_model >= 1 && current_model <= length(models)) {
model_names <- names(models[[current_model]])
if (!is.null(model_names) && effect_index >= 1 && effect_index <= length(model_names)) {
current_component <- model_names[effect_index]
}
}
}
current_component <- sub("_(BRR|RKHS)$", "", current_component)
current_component
}, character(1))
}3. Loading the Kernels and Phenotypic Objects
This section loads the matrices generated in
matrizes.Rmd. These objects are the core inputs for the
variance-component analysis.
At this point, the script reads the matrices and kernels generated in the previous module so the variance-component models use the same aligned inputs as the prediction stage.
ZG <- readRDS(file.path(kernel_dir, "ZG.rds"))
ZP <- readRDS(file.path(kernel_dir, "ZP.rds"))
ZE <- readRDS(file.path(kernel_dir, "ZE.rds"))
ZEZE <- readRDS(file.path(kernel_dir, "ZEZE.rds"))
ZW <- readRDS(file.path(kernel_dir, "ZW.rds"))
ZGZE <- readRDS(file.path(kernel_dir, "ZGZE.rds"))
ZPZE <- readRDS(file.path(kernel_dir, "ZPZE.rds"))
ZGZW <- readRDS(file.path(kernel_dir, "ZGZW.rds"))
ZPZW <- readRDS(file.path(kernel_dir, "ZPZW.rds"))
GGK <- readRDS(file.path(kernel_dir, "GGK.rds"))
PGK <- readRDS(file.path(kernel_dir, "PGK.rds"))
GAK <- readRDS(file.path(kernel_dir, "GAK.rds"))
PAK <- readRDS(file.path(kernel_dir, "PAK.rds"))
GGKE <- readRDS(file.path(kernel_dir, "GGKE.rds"))
PGKE <- readRDS(file.path(kernel_dir, "PGKE.rds"))
GGKW <- readRDS(file.path(kernel_dir, "GGKW.rds"))
PGKW <- readRDS(file.path(kernel_dir, "PGKW.rds"))
GAKE <- readRDS(file.path(kernel_dir, "GAKE.rds"))
PAKE <- readRDS(file.path(kernel_dir, "PAKE.rds"))
GAKW <- readRDS(file.path(kernel_dir, "GAKW.rds"))
PAKW <- readRDS(file.path(kernel_dir, "PAKW.rds"))
Pheno <- readRDS(file.path(kernel_dir, "Pheno.rds"))
Pedigree <- readRDS(file.path(kernel_dir, "Pedigree.rds"))
n_obs <- nrow(Pheno)
cat("Number of phenotypic observations:", n_obs, "\n")Number of phenotypic observations: 982 cat("Number of pedigrees:", length(Pedigree), "\n")Number of pedigrees: 329 3.1. Alignment diagnostics
Before fitting the models, it is important to verify that the kernels
are square and aligned to the same observation order used in
Pheno. In a tutorial context, this is an important check
because many downstream errors are caused by mismatched row and column
orders.
The loaded kernels are then checked for compatibility in dimension and ordering before they are combined in the Bayesian models.
reference_ids <- Pheno$ObsID
ZE <- ZE[match(reference_ids, rownames(ZE)), , drop = FALSE]
ZG <- ZG[reference_ids, reference_ids, drop = FALSE]
ZP <- ZP[reference_ids, reference_ids, drop = FALSE]
ZEZE <- ZEZE[reference_ids, reference_ids, drop = FALSE]
ZW <- ZW[reference_ids, reference_ids, drop = FALSE]
ZGZE <- ZGZE[reference_ids, reference_ids, drop = FALSE]
ZPZE <- ZPZE[reference_ids, reference_ids, drop = FALSE]
ZGZW <- ZGZW[reference_ids, reference_ids, drop = FALSE]
ZPZW <- ZPZW[reference_ids, reference_ids, drop = FALSE]
GGK <- GGK[reference_ids, reference_ids, drop = FALSE]
PGK <- PGK[reference_ids, reference_ids, drop = FALSE]
GAK <- GAK[reference_ids, reference_ids, drop = FALSE]
PAK <- PAK[reference_ids, reference_ids, drop = FALSE]
GGKE <- GGKE[reference_ids, reference_ids, drop = FALSE]
PGKE <- PGKE[reference_ids, reference_ids, drop = FALSE]
GGKW <- GGKW[reference_ids, reference_ids, drop = FALSE]
PGKW <- PGKW[reference_ids, reference_ids, drop = FALSE]
GAKE <- GAKE[reference_ids, reference_ids, drop = FALSE]
PAKE <- PAKE[reference_ids, reference_ids, drop = FALSE]
GAKW <- GAKW[reference_ids, reference_ids, drop = FALSE]
PAKW <- PAKW[reference_ids, reference_ids, drop = FALSE]
kernel_inventory <- tibble(
kernel = c("ZG", "ZP", "ZEZE", "ZW", "GGK", "PGK", "GAK", "PAK"),
n_rows = c(nrow(ZG), nrow(ZP), nrow(ZEZE), nrow(ZW), nrow(GGK), nrow(PGK), nrow(GAK), nrow(PAK)),
n_cols = c(ncol(ZG), ncol(ZP), ncol(ZEZE), ncol(ZW), ncol(GGK), ncol(PGK), ncol(GAK), ncol(PAK)),
is_square = c(
nrow(ZG) == ncol(ZG),
nrow(ZP) == ncol(ZP),
nrow(ZEZE) == ncol(ZEZE),
nrow(ZW) == ncol(ZW),
nrow(GGK) == ncol(GGK),
nrow(PGK) == ncol(PGK),
nrow(GAK) == ncol(GAK),
nrow(PAK) == ncol(PAK)
),
symmetric_names = c(
identical(rownames(ZG), colnames(ZG)),
identical(rownames(ZP), colnames(ZP)),
identical(rownames(ZEZE), colnames(ZEZE)),
identical(rownames(ZW), colnames(ZW)),
identical(rownames(GGK), colnames(GGK)),
identical(rownames(PGK), colnames(PGK)),
identical(rownames(GAK), colnames(GAK)),
identical(rownames(PAK), colnames(PAK))
),
aligned_with_reference = c(
identical(rownames(ZG), reference_ids),
identical(rownames(ZP), reference_ids),
identical(rownames(ZEZE), reference_ids),
identical(rownames(ZW), reference_ids),
identical(rownames(GGK), reference_ids),
identical(rownames(PGK), reference_ids),
identical(rownames(GAK), reference_ids),
identical(rownames(PAK), reference_ids)
)
)
kernel_inventory %>%
dplyr::mutate(
n_rows = format(n_rows, big.mark = ","),
n_cols = format(n_cols, big.mark = ",")
) %>%
knitr::kable(
caption = "Diagnostic summary of the main kernels used in the variance-component analysis after explicit alignment to the phenotypic observation order.",
align = "lcccccc"
) %>%
kableExtra::kable_styling(
bootstrap_options = c("striped", "hover", "condensed", "responsive"),
full_width = FALSE,
position = "left"
) %>%
kableExtra::scroll_box(width = "100%", height = "260px")| kernel | n_rows | n_cols | is_square | symmetric_names | aligned_with_reference |
|---|---|---|---|---|---|
| ZG | 982 | 982 | TRUE | TRUE | TRUE |
| ZP | 982 | 982 | TRUE | TRUE | TRUE |
| ZEZE | 982 | 982 | TRUE | TRUE | TRUE |
| ZW | 982 | 982 | TRUE | TRUE | TRUE |
| GGK | 982 | 982 | TRUE | TRUE | TRUE |
| PGK | 982 | 982 | TRUE | TRUE | TRUE |
| GAK | 982 | 982 | TRUE | TRUE | TRUE |
| PAK | 982 | 982 | TRUE | TRUE | TRUE |
4. Definition of the 18 Predictive Models
Each model is defined as a list in the format required by the
ETA argument of BGLR. The environment term
(E) is included as a fixed-effect design matrix with
model = "BRR", while the kernels are fitted as random
effects with model = "RKHS".
The reduced set of 18 Bayesian multi-kernel models is defined below in the form required for the variance-component workflow.
models <- list()
models[[1]] <- list(
E = list(X = ZE, model = "BRR"),
G = list(K = ZG, model = "RKHS"),
GE = list(K = ZGZE, model = "RKHS")
)
models[[2]] <- list(
E = list(X = ZE, model = "BRR"),
P = list(K = ZP, model = "RKHS"),
PE = list(K = ZPZE, model = "RKHS")
)
models[[3]] <- list(
E = list(X = ZE, model = "BRR"),
GGK = list(K = GGK, model = "RKHS"),
GGKE = list(K = GGKE, model = "RKHS")
)
models[[4]] <- list(
E = list(X = ZE, model = "BRR"),
PGK = list(K = PGK, model = "RKHS"),
PGKE = list(K = PGKE, model = "RKHS")
)
models[[5]] <- list(
E = list(X = ZE, model = "BRR"),
GAK = list(K = GAK, model = "RKHS"),
GAKE = list(K = GAKE, model = "RKHS")
)
models[[6]] <- list(
E = list(X = ZE, model = "BRR"),
PAK = list(K = PAK, model = "RKHS"),
PAKE = list(K = PAKE, model = "RKHS")
)
models[[7]] <- list(
E = list(X = ZE, model = "BRR"),
G = list(K = ZG, model = "RKHS"),
P = list(K = ZP, model = "RKHS"),
GE = list(K = ZGZE, model = "RKHS"),
PE = list(K = ZPZE, model = "RKHS")
)
models[[8]] <- list(
E = list(X = ZE, model = "BRR"),
GGK = list(K = GGK, model = "RKHS"),
PGK = list(K = PGK, model = "RKHS"),
GGKE = list(K = GGKE, model = "RKHS"),
PGKE = list(K = PGKE, model = "RKHS")
)
models[[9]] <- list(
E = list(X = ZE, model = "BRR"),
GAK = list(K = GAK, model = "RKHS"),
PAK = list(K = PAK, model = "RKHS"),
GAKE = list(K = GAKE, model = "RKHS"),
PAKE = list(K = PAKE, model = "RKHS")
)
models[[10]] <- list(
E = list(X = ZE, model = "BRR"),
W = list(K = ZW, model = "RKHS"),
G = list(K = ZG, model = "RKHS"),
GE = list(K = ZGZE, model = "RKHS"),
GW = list(K = ZGZW, model = "RKHS")
)
models[[11]] <- list(
E = list(X = ZE, model = "BRR"),
W = list(K = ZW, model = "RKHS"),
P = list(K = ZP, model = "RKHS"),
PE = list(K = ZPZE, model = "RKHS"),
PW = list(K = ZPZW, model = "RKHS")
)
models[[12]] <- list(
E = list(X = ZE, model = "BRR"),
W = list(K = ZW, model = "RKHS"),
G = list(K = ZG, model = "RKHS"),
P = list(K = ZP, model = "RKHS"),
GE = list(K = ZGZE, model = "RKHS"),
PE = list(K = ZPZE, model = "RKHS"),
GW = list(K = ZGZW, model = "RKHS"),
PW = list(K = ZPZW, model = "RKHS")
)
models[[13]] <- list(
E = list(X = ZE, model = "BRR"),
W = list(K = ZW, model = "RKHS"),
GGK = list(K = GGK, model = "RKHS"),
GGKE = list(K = GGKE, model = "RKHS"),
GGKW = list(K = GGKW, model = "RKHS")
)
models[[14]] <- list(
E = list(X = ZE, model = "BRR"),
W = list(K = ZW, model = "RKHS"),
PGK = list(K = PGK, model = "RKHS"),
PGKE = list(K = PGKE, model = "RKHS"),
PGKW = list(K = PGKW, model = "RKHS")
)
models[[15]] <- list(
E = list(X = ZE, model = "BRR"),
W = list(K = ZW, model = "RKHS"),
GGK = list(K = GGK, model = "RKHS"),
PGK = list(K = PGK, model = "RKHS"),
GGKE = list(K = GGKE, model = "RKHS"),
PGKE = list(K = PGKE, model = "RKHS"),
GGKW = list(K = GGKW, model = "RKHS"),
PGKW = list(K = PGKW, model = "RKHS")
)
models[[16]] <- list(
E = list(X = ZE, model = "BRR"),
W = list(K = ZW, model = "RKHS"),
GAK = list(K = GAK, model = "RKHS"),
GAKE = list(K = GAKE, model = "RKHS"),
GAKW = list(K = GAKW, model = "RKHS")
)
models[[17]] <- list(
E = list(X = ZE, model = "BRR"),
W = list(K = ZW, model = "RKHS"),
PAK = list(K = PAK, model = "RKHS"),
PAKE = list(K = PAKE, model = "RKHS"),
PAKW = list(K = PAKW, model = "RKHS")
)
models[[18]] <- list(
E = list(X = ZE, model = "BRR"),
W = list(K = ZW, model = "RKHS"),
GAK = list(K = GAK, model = "RKHS"),
PAK = list(K = PAK, model = "RKHS"),
GAKE = list(K = GAKE, model = "RKHS"),
PAKE = list(K = PAKE, model = "RKHS"),
GAKW = list(K = GAKW, model = "RKHS"),
PAKW = list(K = PAKW, model = "RKHS")
)
names(models) <- sprintf("Eta%02d", seq_along(models))4.1. Model catalog
This table is useful both for internal checks and for linking figure labels to the formal description of each model.
The code below builds a compact model catalog linking each model identifier to its category and descriptive label for later summaries.
model_info <- tibble(
ModelID = 1:18,
Model = paste0("Eta", 1:18),
ETA = names(models),
Category = c(
rep("Single-source + E", 6),
rep("Integrated G/P + E", 3),
rep("Weather-augmented", 9)
),
Description = c(
"E + G + GE",
"E + P + PE",
"E + GGK + GGKE",
"E + PGK + PGKE",
"E + GAK + GAKE",
"E + PAK + PAKE",
"E + G + P + GE + PE",
"E + GGK + PGK + GGKE + PGKE",
"E + GAK + PAK + GAKE + PAKE",
"E + W + G + GE + GW",
"E + W + P + PE + PW",
"E + W + G + P + GE + PE + GW + PW",
"E + W + GGK + GGKE + GGKW",
"E + W + PGK + PGKE + PGKW",
"E + W + GGK + PGK + GGKE + PGKE + GGKW + PGKW",
"E + W + GAK + GAKE + GAKW",
"E + W + PAK + PAKE + PAKW",
"E + W + GAK + PAK + GAKE + PAKE + GAKW + PAKW"
)
)
readr::write_csv(model_info, file.path(results_dir, "model_catalog.csv"))
model_info %>%
knitr::kable(
caption = "Catalog of the 18 models evaluated in the variance-component analysis.",
align = "cll"
) %>%
kableExtra::kable_styling(
bootstrap_options = c("striped", "hover", "condensed", "responsive"),
full_width = FALSE,
position = "left"
) %>%
kableExtra::scroll_box(width = "100%", height = "340px")| ModelID | Model | ETA | Category | Description |
|---|---|---|---|---|
| 1 | Eta1 | Eta01 | Single-source + E | E + G + GE |
| 2 | Eta2 | Eta02 | Single-source + E | E + P + PE |
| 3 | Eta3 | Eta03 | Single-source + E | E + GGK + GGKE |
| 4 | Eta4 | Eta04 | Single-source + E | E + PGK + PGKE |
| 5 | Eta5 | Eta05 | Single-source + E | E + GAK + GAKE |
| 6 | Eta6 | Eta06 | Single-source + E | E + PAK + PAKE |
| 7 | Eta7 | Eta07 | Integrated G/P + E | E + G + P + GE + PE |
| 8 | Eta8 | Eta08 | Integrated G/P + E | E + GGK + PGK + GGKE + PGKE |
| 9 | Eta9 | Eta09 | Integrated G/P + E | E + GAK + PAK + GAKE + PAKE |
| 10 | Eta10 | Eta10 | Weather-augmented | E + W + G + GE + GW |
| 11 | Eta11 | Eta11 | Weather-augmented | E + W + P + PE + PW |
| 12 | Eta12 | Eta12 | Weather-augmented | E + W + G + P + GE + PE + GW + PW |
| 13 | Eta13 | Eta13 | Weather-augmented | E + W + GGK + GGKE + GGKW |
| 14 | Eta14 | Eta14 | Weather-augmented | E + W + PGK + PGKE + PGKW |
| 15 | Eta15 | Eta15 | Weather-augmented | E + W + GGK + PGK + GGKE + PGKE + GGKW + PGKW |
| 16 | Eta16 | Eta16 | Weather-augmented | E + W + GAK + GAKE + GAKW |
| 17 | Eta17 | Eta17 | Weather-augmented | E + W + PAK + PAKE + PAKW |
| 18 | Eta18 | Eta18 | Weather-augmented | E + W + GAK + PAK + GAKE + PAKE + GAKW + PAKW |
5. MCMC Settings
These values define the Gibbs sampler used by BGLR.
Adjust them here if a longer chain is required for convergence
diagnostics or sensitivity analyses.
This section defines the MCMC settings and execution controls that regulate the estimation stage when the heavy computation is turned on.
nIter <- 5000
burnIn <- 1000
thin <- 10
num_rep <- 10
traits <- c("GY", "KW")6. Running BGLR for Each Model
This is the computationally expensive step. The chunk is kept with
eval=FALSE so that the tutorial can be rendered without
rerunning all Bayesian models every time. When you need updated variance
components, change eval to TRUE and run the
chunk.
The next block contains the computationally expensive parallel estimation stage that fits the variance-component models, preserves the iterative BGLR artifacts in persistent folders, and saves the raw posterior summaries.
numCores <- parallel::detectCores()
useCores <- max(1, numCores - 1)
cl <- parallel::makeCluster(useCores)
doParallel::registerDoParallel(cl)
combos <- expand.grid(
Model = 1:18,
REP = 1:num_rep,
Trait = traits,
stringsAsFactors = FALSE
)
results_list <- foreach(
i = 1:nrow(combos),
.packages = c("BGLR", "dplyr", "tibble")
) %dopar% {
Model <- combos$Model[i]
rep_num <- combos$REP[i]
Trait <- combos$Trait[i]
rep_label <- sprintf("rep%02d", rep_num)
eta_label <- paste0("Eta", Model)
pheno <- data.frame(Pheno)
y_data <- as.numeric(pheno[[Trait]])
out_dir <- file.path(bglr_runs_dir, Trait, rep_label, eta_label)
dir.create(out_dir, recursive = TRUE, showWarnings = FALSE)
save_prefix <- file.path(
out_dir,
paste0("VC_", Trait, "_", eta_label, "_", rep_label, "_")
)
start_time <- Sys.time()
fit <- BGLR(
y = y_data,
ETA = models[[Model]],
nIter = nIter,
burnIn = burnIn,
thin = thin,
saveAt = save_prefix
)
end_time <- Sys.time()
effect_names <- names(models[[Model]])
effect_models <- vapply(models[[Model]], function(x) x$model, character(1))
effect_labels <- paste0(effect_names, "_", effect_models)
generated_files <- list.files(
out_dir,
pattern = paste0("^", basename(save_prefix)),
full.names = TRUE
)
if (length(generated_files) > 0) {
for (current_file in generated_files) {
current_name <- basename(current_file)
renamed_name <- current_name
for (j in seq_along(effect_labels)) {
renamed_name <- sub(
paste0("ETA_", j),
effect_labels[j],
renamed_name,
fixed = TRUE
)
}
if (!identical(current_name, renamed_name)) {
file.rename(current_file, file.path(out_dir, renamed_name))
}
}
}
variance_samples <- list()
if (!is.null(fit$ETA)) {
if (is.null(effect_names)) {
effect_names <- paste0("Effect_", seq_along(fit$ETA))
}
for (j in seq_along(fit$ETA)) {
parameter_names <- names(fit$ETA[[j]])
var_parameter <- parameter_names[grepl("^var", parameter_names)]
if (length(var_parameter) > 0) {
variance_samples[[effect_names[j]]] <- fit$ETA[[j]][[var_parameter[1]]]
}
}
}
metrics_object <- list(
varE_samples = fit$varE,
variances_samples = variance_samples,
DIC = fit$fit$DIC,
runtime_sec = as.numeric(difftime(end_time, start_time, units = "secs"))
)
saveRDS(metrics_object, file.path(out_dir, "variance_metrics.rds"))
result_table <- tibble(
Component = "Residual",
MeanVar = mean(metrics_object$varE_samples, na.rm = TRUE)
)
if (length(metrics_object$variances_samples) > 0) {
for (nm in names(metrics_object$variances_samples)) {
result_table <- bind_rows(
result_table,
tibble(
Component = nm,
MeanVar = mean(metrics_object$variances_samples[[nm]], na.rm = TRUE)
)
)
}
}
rm(fit)
gc()
result_table %>%
mutate(Model = Model, Rep = rep_num, Trait = Trait)
}
stopCluster(cl)
all_variance <- bind_rows(results_list)
write_csv(all_variance, file.path(results_dir, "variance_components_raw.csv"))
saveRDS(all_variance, file.path(results_dir, "variance_components_raw.rds"))7. Consolidating Saved Posterior Summaries
In the current version of the workflow, this is the section used most
often. Instead of refitting all models every time, the script reads the
previously saved variance_metrics.rds files from
output/variance_components/bglr_runs/ and consolidates them
into one object. This keeps the tutorial practical and reproducible,
while making it clear that this page is mainly a
result-consolidation step unless the user intentionally
reruns the full BGLR estimation stage.
The saved outputs produced by the estimation stage are consolidated below into a single structure that can be processed more easily later.
rds_files <- list.files(
path = bglr_runs_dir,
pattern = "(variance_metrics|_(metrics|metricas))\\.rds$",
recursive = TRUE,
full.names = TRUE
)
if (length(rds_files) > 0) {
all_tables <- vector("list", length(rds_files))
for (i in seq_along(rds_files)) {
current_file <- normalizePath(rds_files[i], winslash = "/", mustWork = FALSE)
path_parts <- str_split(current_file, "/")[[1]]
rep_part <- path_parts[str_detect(path_parts, "^rep[0-9]+$|^rep_[0-9]+$")][1]
eta_part <- path_parts[str_detect(path_parts, "^Eta")][1]
trait_part <- path_parts[path_parts %in% traits][1]
rep_value <- as.integer(str_remove(rep_part, "^rep_?0*"))
model_value <- as.integer(str_remove(eta_part, "^Eta0*"))
trait_value <- trait_part
current_object <- readRDS(current_file)
current_table <- tibble(
Rep = rep_value,
Model = model_value,
Trait = trait_value,
Component = "Residual",
MeanVar = mean(current_object$varE_samples, na.rm = TRUE),
DIC = current_object$DIC,
runtime_sec = current_object$runtime_sec
)
if (length(current_object$variances_samples) > 0) {
for (nm in names(current_object$variances_samples)) {
current_table <- bind_rows(
current_table,
tibble(
Rep = rep_value,
Model = model_value,
Trait = trait_value,
Component = normalize_component_name(nm, model_value = model_value),
MeanVar = mean(current_object$variances_samples[[nm]], na.rm = TRUE),
DIC = current_object$DIC,
runtime_sec = current_object$runtime_sec
)
)
}
}
all_tables[[i]] <- current_table
}
all_variance <- bind_rows(all_tables)
saveRDS(all_variance, file.path(results_dir, "variance_components_all.rds"))
write_csv(all_variance, file.path(results_dir, "variance_components_raw.csv"))
}8. Loading the Consolidated Variance-Component Object
This section loads the consolidated results and prepares them for summarization. If the file does not exist, the script stops and instructs the user to execute the model-fitting stage first.
The document now loads the already consolidated variance-component results so that it can summarize and visualize them without repeating the expensive estimation.
raw_file <- file.path(results_dir, "variance_components_all.rds")
if (!file.exists(raw_file)) {
stop(
"The file '", raw_file, "' was not found. ",
"Run the Bayesian fitting stage first or keep the previously saved _metrics.rds files in output/variance_components/."
)
}
all_variance <- readRDS(raw_file)
all_variance <- all_variance %>%
mutate(
Trait = str_extract(Trait, "GY|KW"),
Trait = ifelse(is.na(Trait), Trait, Trait),
Model = as.integer(Model)
)
cat("Number of rows in consolidated variance object:", nrow(all_variance), "\n")Number of rows in consolidated variance object: 2100 9. Summarizing the Variance Components
Now we average the posterior means across repetitions for each model, trait, and component. Then we convert the results to percentages of total variance, which makes the models directly comparable.
The loaded results are then reshaped and annotated so that posterior means can be compared across traits, models, and component categories.
variance_summary <- all_variance %>%
group_by(Model, Trait, Component) %>%
summarise(
AvgVar = mean(MeanVar, na.rm = TRUE),
MeanDIC = mean(DIC, na.rm = TRUE),
MeanRuntimeSec = mean(runtime_sec, na.rm = TRUE),
.groups = "drop"
) %>%
filter(!is.na(AvgVar), AvgVar > 0) %>%
group_by(Model, Trait) %>%
mutate(
TotalVar = sum(AvgVar),
Percentage = AvgVar / TotalVar
) %>%
ungroup() %>%
left_join(model_info, by = c("Model" = "ModelID")) %>%
mutate(
ModelNumber = Model,
Model = factor(paste0("Eta", ModelNumber), levels = paste0("Eta", 1:18)),
Component = normalize_component_name(Component, model_value = ModelNumber),
ComponentLabel = dplyr::recode(Component, !!!component_display),
ComponentLabel = factor(ComponentLabel, levels = component_order),
Trait = factor(Trait, levels = c("GY", "KW"))
) %>%
dplyr::select(-ModelNumber)
readr::write_csv(variance_summary, file.path(results_dir, "variance_components_processed.csv"))9.1. Wide-format summary table
This table is helpful when you need a compact numerical summary for the manuscript or supplementary material.
The next table summarizes the relative contribution of each variance component across models.
variance_table <- variance_summary %>%
dplyr::select(Model, Trait, ComponentLabel, Percentage) %>%
dplyr::mutate(Percentage = round(100 * Percentage, 2)) %>%
tidyr::pivot_wider(names_from = ComponentLabel, values_from = Percentage)
readr::write_csv(variance_table, file.path(results_dir, "variance_components_table_percent.csv"))
head(variance_table, 12) %>%
knitr::kable(
caption = "Preview of the processed variance-component table (percent of total variance).",
align = "lcccccccccccc"
) %>%
kableExtra::kable_styling(
bootstrap_options = c("striped", "hover", "condensed", "responsive"),
full_width = FALSE,
position = "left"
) %>%
kableExtra::scroll_box(width = "100%", height = "320px")| Model | Trait | Environment (E) | Genomic linear (G) | G × E | Residual | Phenomic linear (P) | P × E | Genomic Gaussian (GGK) | GGK × E | Phenomic Gaussian (PGK) | PGK × E | Genomic arc-cosine (GAK) | GAK × E | Phenomic arc-cosine (PAK) | PAK × E | G × W | Weather kernel (W) | P × W | GGK × W | PGK × W | GAK × W | PAK × W |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Eta1 | GY | 83.59 | 11.96 | 1.70 | 2.75 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA |
| Eta1 | KW | 75.42 | 20.80 | 1.46 | 2.33 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA |
| Eta2 | GY | 0.00 | NA | NA | 0.00 | 100 | 0 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA |
| Eta2 | KW | 0.00 | NA | NA | 0.00 | 100 | 0 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA |
| Eta3 | GY | 49.52 | NA | NA | 1.92 | NA | NA | 45.13 | 3.43 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA |
| Eta3 | KW | 35.42 | NA | NA | 1.26 | NA | NA | 61.28 | 2.04 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA |
| Eta4 | GY | 78.05 | NA | NA | 10.87 | NA | NA | NA | NA | 7.54 | 3.53 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA |
| Eta4 | KW | 73.38 | NA | NA | 11.51 | NA | NA | NA | NA | 11.07 | 4.04 | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA | NA |
| Eta5 | GY | 5.49 | NA | NA | 0.26 | NA | NA | NA | NA | NA | NA | 92.52 | 1.72 | NA | NA | NA | NA | NA | NA | NA | NA | NA |
| Eta5 | KW | 3.38 | NA | NA | 0.16 | NA | NA | NA | NA | NA | NA | 95.71 | 0.75 | NA | NA | NA | NA | NA | NA | NA | NA | NA |
| Eta6 | GY | 18.38 | NA | NA | 3.26 | NA | NA | NA | NA | NA | NA | NA | NA | 77.06 | 1.30 | NA | NA | NA | NA | NA | NA | NA |
| Eta6 | KW | 10.60 | NA | NA | 2.14 | NA | NA | NA | NA | NA | NA | NA | NA | 86.37 | 0.89 | NA | NA | NA | NA | NA | NA | NA |
10. Figure for the Article
The next figure summarizes the percentage contribution of each
variance component across the 18 models for both traits. To keep the
x-axis readable, the models are displayed as Eta1 to
Eta18, and the detailed descriptions remain available in
the model catalog saved above.
This figure should be read as a structural summary of the fitted models, not as direct proof that combining more data sources automatically leads to stronger biological complementarity or better predictive performance.
The final figure turns the processed table into a compact visual summary for interpretation.
trait_labels <- c(
GY = "A. Grain yield (GY)",
KW = "B. Kernel weight (KW)"
)
variance_plot <- variance_summary %>%
ggplot(aes(x = Model, y = Percentage, fill = ComponentLabel)) +
geom_col(color = "white", linewidth = 0.15, width = 0.82) +
facet_wrap(~ Trait, ncol = 1, labeller = as_labeller(trait_labels)) +
ggthemes::scale_fill_gdocs(name = "Variance component", drop = FALSE) +
scale_y_continuous(
labels = scales::percent_format(accuracy = 1),
expand = expansion(mult = c(0, 0.02))
) +
labs(
x = "Model ID",
y = "Proportion of total variance"
) +
theme_minimal(base_size = 11) +
theme(
plot.title = element_text(face = "bold", size = 13, hjust = 0.5),
strip.text = element_text(face = "bold", size = 11),
axis.title = element_text(face = "bold"),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1, size = 8),
axis.text.y = element_text(size = 9),
legend.position = "right",
legend.box = "vertical",
legend.title = element_text(face = "bold", size = 10),
legend.text = element_text(size = 8),
legend.key.height = unit(0.45, "cm"),
legend.key.width = unit(0.55, "cm"),
panel.grid.major.x = element_blank(),
panel.grid.minor = element_blank()
) +
guides(fill = guide_legend(ncol = 2, byrow = TRUE))
print(variance_plot)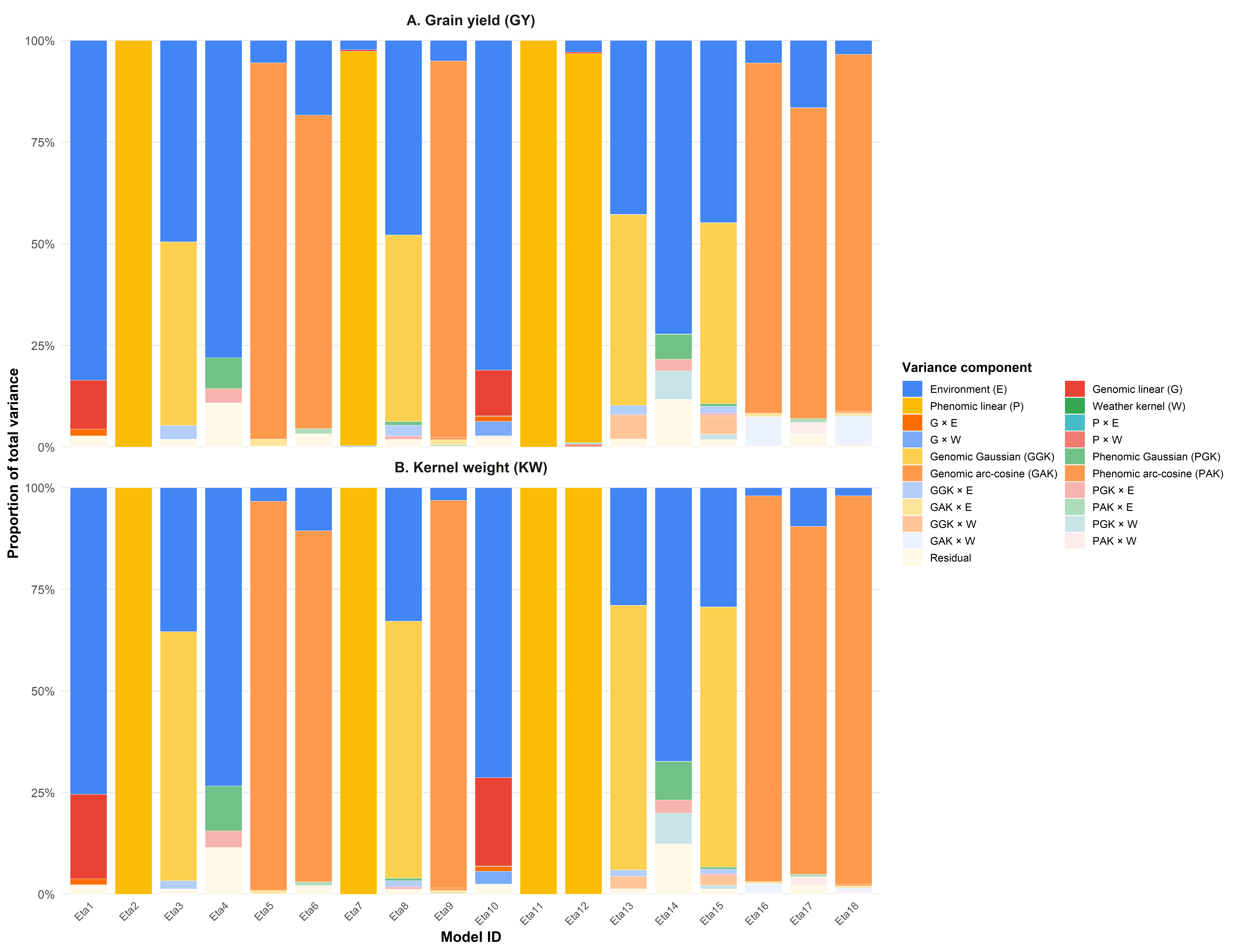
Figure 3. Posterior mean percentage of total variance attributed to each model component across the 18 predictive models for grain yield (GY) and kernel weight (KW). Each stacked bar represents one model (Eta1-Eta18), averaged across repetitions, and colors identify the variance components included in the model structure. The weather kernel (W) and its associated interaction terms appear mainly in models Eta10-Eta18. Variance decomposition should be interpreted as a description of model structure and not automatically as evidence of stronger biological complementarity among information sources.
ggsave(
filename = file.path(figure_dir, "variance_components_article.tiff"),
plot = variance_plot,
width = 14,
height = 10,
dpi = 300,
compression = "lzw",
bg = "white"
)11. Interpretation Guide
This final section clarifies how the figure should be interpreted.
- A large Environment (E) segment indicates that the categorical macro-environment effect dominates the variance partition.
- Large genomic terms such as G, GGK, or GAK indicate that genomic similarity captures an important share of the trait variability.
- Large phenomic terms such as P, PGK, or PAK indicate stronger contribution of the NIRS-derived similarity, but these terms should be interpreted with caution because NIRS may also reflect strong physiological response to the environment.
- Large weather-related terms such as W, G × W, or P × W indicate that weather-derived covariates explain additional structure. These terms appear mainly in models Eta10-Eta18 and should be interpreted as complementary to the categorical environment effect, not as a replacement for E.
- A large Residual segment indicates that important sources of variability remain unexplained by the fitted model.
In the context of this article, these variance decompositions complement the predictive analyses by showing which information sources contribute to model structure. However, a large variance share for a component should not be read automatically as evidence of stronger biological complementarity or of better predictive ability.