Home
Last updated: 2023-08-30
Checks: 1 1
Knit directory: duplex_sequencing_screen/
This reproducible R Markdown analysis was created with workflowr (version 1.6.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of
the R Markdown file created these results, you’ll want to first commit
it to the Git repo. If you’re still working on the analysis, you can
ignore this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version f9eea37. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: data/Consensus_Data/.Rhistory
Ignored: data/Consensus_Data/Novogene_lane11/sample1/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample1/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample1/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample1/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample1/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample1/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample2/archive/sscs_aligned_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample2/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample2/duplex/variant_caller_outputs/archive_ensembl/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample2/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample2/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample2/sscs/variant_caller_outputs/archive_ensembl/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample2/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample3/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample3/duplex/variant_caller_outputs/archive_ensembl/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample3/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample3/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample3/sscs/variant_caller_outputs/archive ensembl/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample3/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample4/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample4/duplex/variant_caller_outputs/archive ensembl/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample4/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample4/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample4/sscs/variant_caller_outputs/archive ensembl/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample4/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample5/variant_caller_outputs/sscs_L858R_aligned_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample5/variant_caller_outputs/sscs_L858R_aligned_filtered_sample5.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample6/archive/sscs_aligned_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample6/sscs_L858R_aligned_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample6/variant_caller_outputs/variants_ann_sample6.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample7/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample7/sscs/variant_caller_outputs/archive ensembl/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane11/sample7/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane12/sample1/low_sscscounts/sscs_aligned_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane12/sample1/low_sscscounts/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane12/sample1/sscs_aligned_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane12/sample1/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane12/sample3/sscs_combined_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane12/sample3/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane12/sample5/sscs_combined_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane12/sample5/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane12/sample7/sscs_combined_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane12/sample7/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane12/sample9/sscs_combined_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane12/sample9/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample1/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample1/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample1/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample1/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample10/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample10/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample10/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample10/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample10/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample10/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample11/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample11/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample11/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample11/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample11/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample11/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample12/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample12/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample12/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample12/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample12/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample12/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample2/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample2/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample3/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample3/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample4/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample4/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample5/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample5/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample6/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample6/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample7/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample7/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample7/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample7/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample7/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample7/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample8/
Ignored: data/Consensus_Data/Novogene_lane13/sample9/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample9/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample9/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample9/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample9/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane13/sample9/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample10_combined/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample10_combined/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample10_combined/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample10_combined/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample10_combined/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample10_combined/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample11/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample11/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample11/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample11/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample11/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample11/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample12/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample12/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample12/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample12/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample12/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample12/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample13/
Ignored: data/Consensus_Data/Novogene_lane14/sample14_combined/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample14_combined/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample14_combined/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample14_combined/sscs.filt_1.fa.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample14_combined/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample14_combined/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample14_combined/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample14b/
Ignored: data/Consensus_Data/Novogene_lane14/sample15/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample15/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample15/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample15/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample15/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample15/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample16/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample16/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample16/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample16/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample16/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample16/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample17/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample17/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample17/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample17/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample17/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample17/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample18/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample18/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample18/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample18/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample18/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample18/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample1_combined/
Ignored: data/Consensus_Data/Novogene_lane14/sample2_combined/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample2_combined/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample3/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample3/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample4/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample4/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample5/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample5/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample6/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample6/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample7/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample7/variant_caller_outputs/duplex/
Ignored: data/Consensus_Data/Novogene_lane14/sample7/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample8/
Ignored: data/Consensus_Data/Novogene_lane14/sample9/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample9/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample9/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample9/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample9/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane14/sample9/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/Novogene_lane2/
Ignored: data/Consensus_Data/Novogene_lane3/
Ignored: data/Consensus_Data/Novogene_lane4/
Ignored: data/Consensus_Data/Novogene_lane5/
Ignored: data/Consensus_Data/Novogene_lane6/
Ignored: data/Consensus_Data/Novogene_lane7/
Ignored: data/Consensus_Data/Ranomics_Pooled/
Ignored: data/Consensus_Data/archive/
Ignored: data/Consensus_Data/novogene_lane15/sample_1/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_1/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_1/firstrun(lowsequencing)/
Ignored: data/Consensus_Data/novogene_lane15/sample_1/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_2/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_2/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_2/firstrun(lowsequencing)/sscs/
Ignored: data/Consensus_Data/novogene_lane15/sample_2/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_2/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/firstrun(lowsequencing)/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/firstrun(lowsequencing)/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/firstrun(lowsequencing)/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/firstrun(lowsequencing)/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/ngs/Sample3_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/ngs/sample3a(firsthalf)/Sample3_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/ngs/sample3a(firsthalf)/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/ngs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_3/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_4/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_4/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_4/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_4/firstrun(lowsequencing)/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_4/firstrun(lowsequencing)/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_4/firstrun(lowsequencing)/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_4/firstrun(lowsequencing)/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_4/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_4/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_4/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_5/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_5/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_5/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_5/firstrun(lowsequencing)/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_5/firstrun(lowsequencing)/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_5/firstrun(lowsequencing)/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_5/firstrun(lowsequencing)/sscs/variant_caller_outputs/.empty/
Ignored: data/Consensus_Data/novogene_lane15/sample_5/firstrun(lowsequencing)/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_5/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_5/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_5/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_6/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_6/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_6/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_6/firstrun(lowsequencing)/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_6/firstrun(lowsequencing)/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_6/firstrun(lowsequencing)/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_6/firstrun(lowsequencing)/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_6/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_6/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_6/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_7/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_7/duplex/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_7/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_7/firstrun(lowsequencing)/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_7/firstrun(lowsequencing)/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_7/firstrun(lowsequencing)/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_7/firstrun(lowsequencing)/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_7/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_7/sscs/variant_caller_outputs/archive/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane15/sample_7/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample10/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample10/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample10/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample10/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample11/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample11/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample11/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample11/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample12/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample12/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample12/sscs/
Ignored: data/Consensus_Data/novogene_lane16a/Sample13/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample13/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample13/sscs/
Ignored: data/Consensus_Data/novogene_lane16a/Sample14/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample14/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample14/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample14/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample1_combined/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample1_combined/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample1_combined/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample1_combined/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample2/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample2/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample2/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample3/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample3/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample3/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample3/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample4/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample4/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample4/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample4/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample5/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample5/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample5/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample5/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample6/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample6/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample6/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample6/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample7/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample7/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample7/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample7/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample8/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample8/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample8/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample8/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample9/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample9/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample9/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16a/Sample9/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16a/duplex/
Ignored: data/Consensus_Data/novogene_lane16b/Sample10/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample10/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample10/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample10/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample11/
Ignored: data/Consensus_Data/novogene_lane16b/Sample15/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample15/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample15/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample15/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample1_combined/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample1_combined/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample1_combined/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample1_combined/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample2/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample2/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample2/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample2/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample3/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample3/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample3/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample3/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample4/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample4/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample4/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample4/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample5/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample5/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample5/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample5/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample6/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample6/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample6/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample6/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample7_combined/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample7_combined/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample7_combined/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample7_combined/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample8_combined/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample8_combined/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample8_combined/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample8_combined/sscs/variant_caller_outputs/archive/
Ignored: data/Consensus_Data/novogene_lane16b/Sample8_combined/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample9/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample9/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample9/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane16b/Sample9/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample10/duplex/
Ignored: data/Consensus_Data/novogene_lane17/sample10/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample10/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample11/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample11/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample11/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample11/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample1_combined/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample1_combined/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample1_combined/low_depth/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample1_combined/low_depth/duplex/low_depth/
Ignored: data/Consensus_Data/novogene_lane17/sample1_combined/low_depth/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample1_combined/low_depth/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample1_combined/low_depth/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample1_combined/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample1_combined/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample2/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample2/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample2/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample2/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample3/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample3/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample3/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample3/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample4/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample4/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample4/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample4/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample5/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample5/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample5/low_seq_depth/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample5/low_seq_depth/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample5/low_seq_depth/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample5/low_seq_depth/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample5/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample5/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample6/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample6/low_seq_depths/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample6/low_seq_depths/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample6/low_seq_depths/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample6/low_seq_depths/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample6/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample6/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample7/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample7/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample7/low_seq_depths/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample7/low_seq_depths/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample7/low_seq_depths/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample7/low_seq_depths/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample7/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample7/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample8/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample8/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample8/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample9/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample9/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample9/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17/sample9/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17b/Sample1 copy 2/
Ignored: data/Consensus_Data/novogene_lane17b/Sample1 copy 3/
Ignored: data/Consensus_Data/novogene_lane17b/Sample1/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17b/Sample1/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17b/Sample1/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17b/Sample1/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17b/Sample2/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane17b/Sample2/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane17b/Sample2/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample1/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample1/l298l/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample1/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample1/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample1/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample1/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample1/nol298l/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample1/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample1/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample1/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample1/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample1/sscs/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample10/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample10/l298l/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample10/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample10/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample10/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample10/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample10/ngs/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample10/ngs/Sample_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample10/ngs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample10/nol298l/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample10/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample10/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample10/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample10/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample10/sscs/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample11/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample11/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample11/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample11/l298l/sscs/variant_caller_outputs/slow_variantcaller_outputs_forcomparison/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample11/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample11/ngs/Sample_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample11/ngs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample11/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample11/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample11/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample11/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample12/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample12/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample12/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample12/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample12/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample12/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample12/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample12/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample13/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample13/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample13/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample13/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample13/ngs/Sample_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample13/ngs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample13/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample13/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample13/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample13/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample14/duplex/
Ignored: data/Consensus_Data/novogene_lane18/sample14/l298l/duplex/
Ignored: data/Consensus_Data/novogene_lane18/sample14/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample14/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample14/ngs/Sample_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample14/ngs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample14/nol298l/duplex/
Ignored: data/Consensus_Data/novogene_lane18/sample14/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample14/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample15/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample15/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample15/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample15/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample15/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample15/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample15/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample15/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample16/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample16/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample16/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample16/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample16/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample16/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample16/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample16/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample17/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample17/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample17/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample17/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample17/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample17/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample17/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample17/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample18/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample18/l298l/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample18/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample18/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample18/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample18/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample18/nol298l/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample18/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample18/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample18/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample18/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample2/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample2/l298l/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample2/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample2/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample2/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample2/l298l/sscs/variant_caller_outputs/fastercaller/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample2/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample2/nol298l/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample2/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample2/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample2/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample2/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample2/sscs/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample3/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample3/l298l/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample3/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample3/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample3/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample3/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample3/nol298l/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample3/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample3/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample3/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample3/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample3/sscs/.DS_Store
Ignored: data/Consensus_Data/novogene_lane18/sample4/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample4/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample4/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample4/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample4/ngs/Sample_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample4/ngs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample4/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample4/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample4/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample4/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample5/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample5/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample5/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample5/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample5/ngs/Sample_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample5/ngs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample5/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample5/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample5/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample5/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample6/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample6/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample6/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample6/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample6/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample6/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample6/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample6/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample7/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample7/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample7/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample7/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample7/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample7/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample7/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample7/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample8/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample8/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample8/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample8/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample8/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample8/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample8/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample8/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample9/l298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample9/l298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample9/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample9/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample9/nol298l/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample9/nol298l/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample9/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/sample9/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample3/duplex/
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample3/l298l/duplex/
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample3/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample3/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample3/nol298l/duplex/
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample3/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample3/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample3/sscs/
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample5/duplex/
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample5/l298l/duplex/
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample5/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample5/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample5/nol298l/duplex/
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample5/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample5/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample5/sscs/
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample6/duplex/
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample6/l298l/duplex/
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample6/l298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample6/l298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample6/nol298l/duplex/
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample6/nol298l/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample6/nol298l/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane18/tlane18a_sample6/sscs/
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample1/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample1/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample1/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample1/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample10/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample10/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample10/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample10/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample11/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample11/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample11/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample11/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample12/duplex/
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample12/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample12/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample2/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample2/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample2/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample2/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample3/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample3/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample3/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample3/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample4/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample4/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample4/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample4/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample5/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample5/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample5/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample5/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample6/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample6/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample6/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample6/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample7/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample7/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample7/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample7/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample8/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample8/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample8/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample8/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample9/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample9/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample9/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19a_Sample9/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample1/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample1/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample1/sscs/
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample10/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample10/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample10/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample10/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample11/duplex/
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample11/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample11/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample12/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample12/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample12/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample12/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample13/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample13/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample13/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample13/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample14/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample14/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample14/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample14/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample15/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample15/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample15/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample15/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample16/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample16/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample16/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample16/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample2/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample2/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample2/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample2/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample3/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample3/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample3/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample3/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample4/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample4/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample4/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample4/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample5/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample5/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample5/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample5/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample6/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample6/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample6/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample6/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample7/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample7/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample7/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample7/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample8/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample8/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample8/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample8/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample9/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample9/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample9/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19/Ln19b_Sample9/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample10/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample10/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample10/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample10/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample11/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample11/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample11/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample11/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample2/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample2/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample2/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample2/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample3/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample3/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample3/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample3/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample4/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample4/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample4/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample4/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample5/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample5/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample5/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample5/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample6/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample6/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample6/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample6/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample7/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample7/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample7/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample7/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample8/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample8/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample8/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample8/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample9/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample9/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample9/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19a_Sample9/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19b_Sample9/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19b_Sample9/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19b_Sample9/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane19_20/Ln19b_Sample9/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane22/.DS_Store
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample1/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample1/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample1/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample1/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample2/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample2/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample2/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample2/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample3/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample3/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample3/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample3/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample4/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample4/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample4/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample4/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample5/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample5/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample5/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample5/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample6/duplex/
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample6/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample6/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample7/
Ignored: data/Consensus_Data/novogene_lane22/Lane22_Sample8/
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample13/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample13/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample13/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample13/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample14/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample14/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample14/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample14/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample15/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample15/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample15/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample15/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample16/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample16/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample16/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample16/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample17/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample17/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample17/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample17/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample18/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample18/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample18/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample18/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample19/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample19/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample19/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample19/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample20/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample20/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample20/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample20/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample21/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample21/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample21/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample21/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample22/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample22/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample22/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample22/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample23/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample23/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample23/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample23/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample24/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample24/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample24/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample24/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample25/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample25/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample25/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample25/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample26/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample26/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample26/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample26/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample27/duplex/duplex_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample27/duplex/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample27/sscs/sscs_sorted_filtered.tsv.gz
Ignored: data/Consensus_Data/novogene_lane23/Lane23_Sample27/sscs/variant_caller_outputs/variants_ann.csv.gz
Ignored: data/Consensus_Data/sscs_dcs_comparisons/
Ignored: output/ABLEnrichmentScreens/ABL_Region1_Lane18_Comparisons/NA/
Ignored: output/ABLEnrichmentScreens/ABL_Region1_Lane18_Comparisons/baf3_IL3_low_rep1vsrep2/
Ignored: output/ABLEnrichmentScreens/ABL_Region1_Lane18_Comparisons/baf3_Imat_Lowvsk562_Imat_Medium/
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane3/il3_indep_1.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane3/il3_indep_1.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane3/il3_indep_2.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane3/il3_indep_2.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane3/il3_indep_3.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane3/il3_indep_3.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane3/sorted_1.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane3/sorted_1.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane3/sorted_2.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane3/sorted_2.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane3/sorted_3.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane3/sorted_3.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/il3_indep_1.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/il3_indep_1.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/il3_indep_2.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/il3_indep_2.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/il3_indep_3.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/il3_indep_3.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/il3_indep_4.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/il3_indep_4.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/il3_indep_5.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/il3_indep_5.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/sorted_1.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/sorted_1.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/sorted_2.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/sorted_2.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/sorted_3.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/sorted_3.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/sorted_4.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/sorted_4.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/sorted_5.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/sorted_5.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/sorted_6.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane4/sorted_6.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP4_Im_High_D2.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP4_Im_High_D2.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP4_Im_High_D4.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP4_Im_High_D4.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP4_Im_Low_D2.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP4_Im_Low_D2.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP4_Im_Low_D4.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP4_Im_Low_D4.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP4_Im_Medium_D2.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP4_Im_Medium_D2.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP4_Im_Medium_D4.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP4_Im_Medium_D4.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP5_Im_High_D2.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP5_Im_High_D2.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP5_Im_High_D4.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP5_Im_High_D4.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP5_Im_Low_D2.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP5_Im_Low_D2.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP5_Im_Low_D4.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP5_Im_Low_D4.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP5_Im_Medium_D2.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP5_Im_Medium_D2.1.consensus.variant-calls.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP5_Im_Medium_D4.1.consensus.variant-calls.genome.vcf.gz
Ignored: output/Twinstrand/ABL1AppOutput/Novogene_Lane5/RP5_Im_Medium_D4.1.consensus.variant-calls.vcf.gz
Untracked files:
Untracked: analysis/ABL_sheared_libraries.Rmd
Untracked: output/ABLEnrichmentScreens/ABL_Region1_Lane18_Comparisons/cross-replicate/baf3_IL3_rep1vsrep2_ft/screen_comparison_baf3_IL3_low_rep1vsrep2ft.csv
Unstaged changes:
Modified: .DS_Store
Modified: analysis/.DS_Store
Modified: analysis/ABL_SM_CRISPR_Cut_Analyses.Rmd
Modified: analysis/ABL_cosmic_analysis.Rmd
Modified: analysis/ABL_kinasefunction_data_parser.Rmd
Deleted: analysis/ABL_unevenness_analysis.Rmd
Modified: analysis/BCRABL_FunctionalKinaseAnalysis.Rmd
Modified: analysis/index.Rmd
Modified: code/compare_screens.R
Modified: data/.DS_Store
Modified: data/Consensus_Data/.DS_Store
Modified: data/Consensus_Data/novogene_lane18/tlane18a_sample6/.DS_Store
Modified: output/.DS_Store
Modified: output/ABLEnrichmentScreens/.DS_Store
Modified: output/ABLEnrichmentScreens/ABL_Region1_Lane18_Comparisons/.DS_Store
Modified: output/ABLEnrichmentScreens/ABL_Region1_Lane18_Comparisons/cross-replicate/.DS_Store
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (analysis/index.Rmd) and HTML
(docs/index.html) files. If you’ve configured a remote Git
repository (see ?wflow_git_remote), click on the hyperlinks
in the table below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| html | f9eea37 | haiderinam | 2023-08-30 | 2023 Updates |
| Rmd | 8202ef7 | haiderinam | 2023-04-10 | Added BE vs SM analyses on putative mutant |
| html | 8202ef7 | haiderinam | 2023-04-10 | Added BE vs SM analyses on putative mutant |
| Rmd | 60b906b | haiderinam | 2023-04-10 | Added ROC analyses on BE-SM data |
| html | 60b906b | haiderinam | 2023-04-10 | Added ROC analyses on BE-SM data |
| Rmd | a33eea5 | haiderinam | 2023-03-25 | Added ABL CDF Analyses of evenness |
| html | a33eea5 | haiderinam | 2023-03-25 | Added ABL CDF Analyses of evenness |
| Rmd | 81a5af3 | haiderinam | 2023-02-25 | Added shortest codon finder function |
| Rmd | fa7baf1 | haiderinam | 2022-12-15 | Updates to the variant caller |
| html | fabfb30 | haiderinam | 2022-12-09 | Build site. |
| Rmd | 077edd0 | haiderinam | 2022-12-09 | Added K562 Webpage |
| html | 077edd0 | haiderinam | 2022-12-09 | Added K562 Webpage |
| Rmd | cec2e31 | haiderinam | 2022-12-08 | Added error rate analyses |
| html | cec2e31 | haiderinam | 2022-12-08 | Added error rate analyses |
| html | 404fc72 | haiderinam | 2022-11-04 | Build site. |
| Rmd | ff4b4e8 | haiderinam | 2022-11-04 | wflow_publish("analysis/index.Rmd") |
| Rmd | 3a2f887 | haiderinam | 2022-11-04 | Added analyses of IL3 independence |
| Rmd | 6f54acf | haiderinam | 2022-11-04 | Added analyses of SSCS vs DCS error rates |
| html | 28519e7 | haiderinam | 2022-10-29 | Build site. |
| Rmd | ba25cb7 | haiderinam | 2022-10-29 | wflow_publish("analysis/index.Rmd") |
| Rmd | 7ed0af8 | haiderinam | 2022-08-22 | August 2022 Updates |
| html | 86ce8ca | haiderinam | 2022-08-22 | August 2022 Update |
| html | f368371 | haiderinam | 2020-08-13 | Build site. |
| html | 7fe0f9b | haiderinam | 2020-07-06 | Build site. |
| Rmd | e4a901d | haiderinam | 2020-07-06 | wflow_publish("analysis/index.Rmd") |
| html | 231d21d | haiderinam | 2020-06-07 | Build site. |
| html | 8f50f99 | haiderinam | 2020-06-07 | Build site. |
| Rmd | 5da3af0 | haiderinam | 2020-06-07 | wflow_publish("analysis/index.Rmd") |
| html | 75ddcb1 | haiderinam | 2020-06-02 | Build site. |
| Rmd | a173683 | haiderinam | 2020-06-02 | wflow_publish("analysis/index.Rmd") |
| html | bd2dec5 | haiderinam | 2020-06-02 | Build site. |
| Rmd | 1bb5940 | haiderinam | 2020-06-02 | wflow_publish("analysis/index.Rmd") |
| html | c43acef | haiderinam | 2020-06-02 | Build site. |
| Rmd | d5c9f64 | haiderinam | 2020-06-02 | wflow_publish("analysis/index.Rmd") |
| html | eaca616 | haiderinam | 2020-04-20 | Build site. |
| Rmd | 2bba93e | haiderinam | 2020-04-20 | wflow_publish("analysis/*.Rmd") |
| html | 05e3df6 | haiderinam | 2020-04-07 | Build site. |
| html | 1c86f69 | haiderinam | 2020-04-07 | Build site. |
| Rmd | e731909 | haiderinam | 2020-04-07 | wflow_publish("analysis/index.Rmd") |
| html | 4516a7f | haiderinam | 2020-04-06 | Build site. |
| Rmd | 3a57291 | haiderinam | 2020-04-06 | wflow_publish(files = "analysis/index.Rmd") |
| html | c2930d5 | haiderinam | 2020-04-03 | Build site. |
| Rmd | 52f6884 | haiderinam | 2020-04-03 | wflow_publish("analysis/*.Rmd") |
| html | 6af2cdc | haiderinam | 2020-04-03 | Build site. |
| Rmd | 5cda9d3 | haiderinam | 2020-04-03 | wflow_publish("analysis/*.Rmd") |
| html | 0b9b87b | haiderinam | 2020-04-02 | Build site. |
| Rmd | fc5b9c0 | haiderinam | 2020-04-02 | wflow_publish("analysis/*.Rmd") |
| html | 2bce927 | haiderinam | 2020-04-02 | Build site. |
| html | 4ed9b35 | haiderinam | 2020-04-02 | Build site. |
| Rmd | e31836a | haiderinam | 2020-04-02 | wflow_publish("analysis/*.Rmd") |
| html | 99181d5 | haiderinam | 2020-04-02 | Build site. |
| Rmd | 17c01d4 | haiderinam | 2020-04-02 | wflow_publish("analysis/index.Rmd") |
| html | 1e2f469 | haiderinam | 2020-04-02 | Build site. |
| html | e11eec5 | haiderinam | 2020-04-02 | Build site. |
| Rmd | 0053f96 | haiderinam | 2020-04-02 | wflow_publish("analysis/*.Rmd") |
| html | b1cbbfa | haiderinam | 2020-04-02 | Build site. |
| Rmd | 703346a | haiderinam | 2020-04-02 | Start workflowr project. |
Duplex Sequencing Spike-Ins
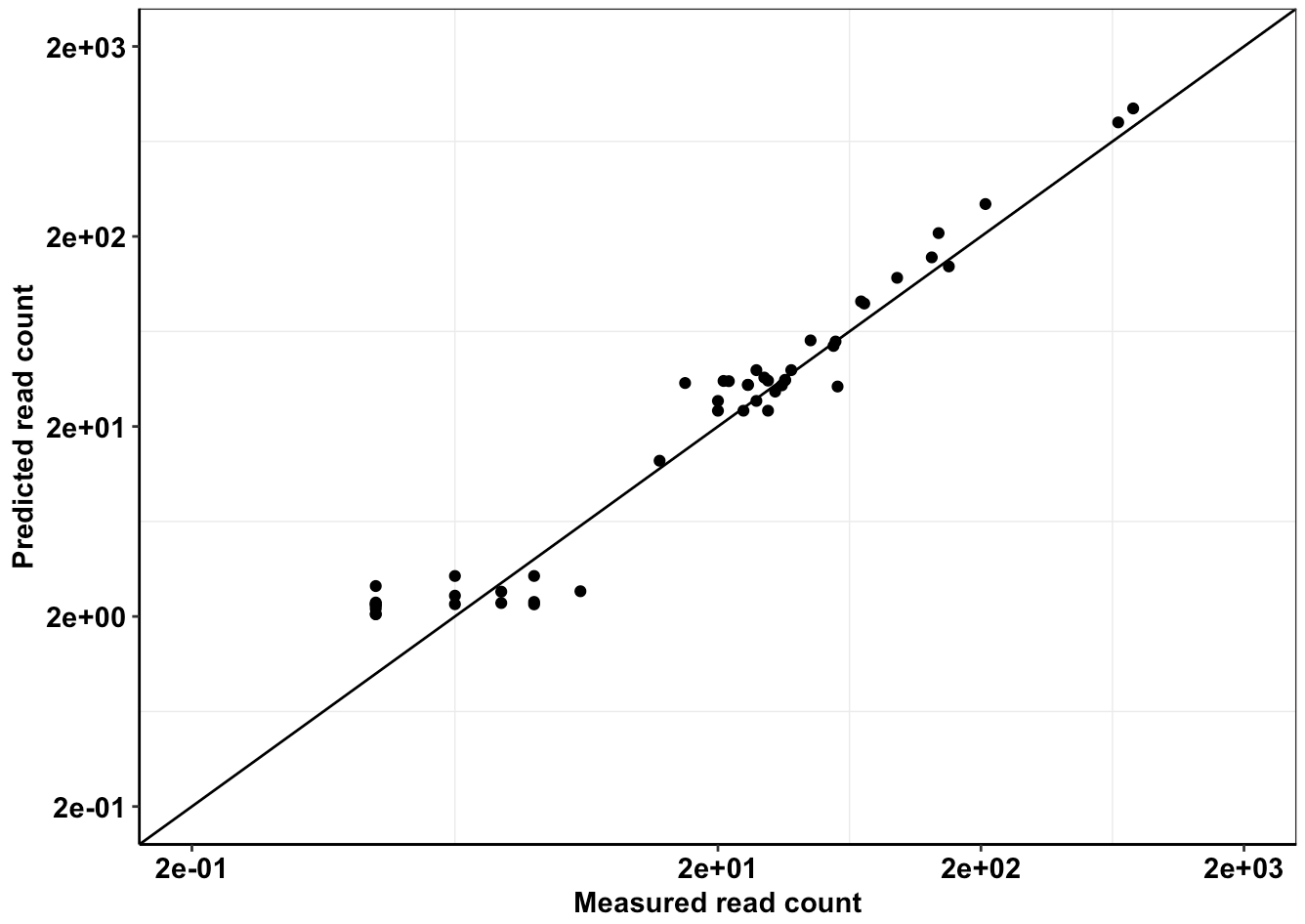
Click here for the script that reads BCRABL dose response data, and fits it to a 4-parameter dose response curve.
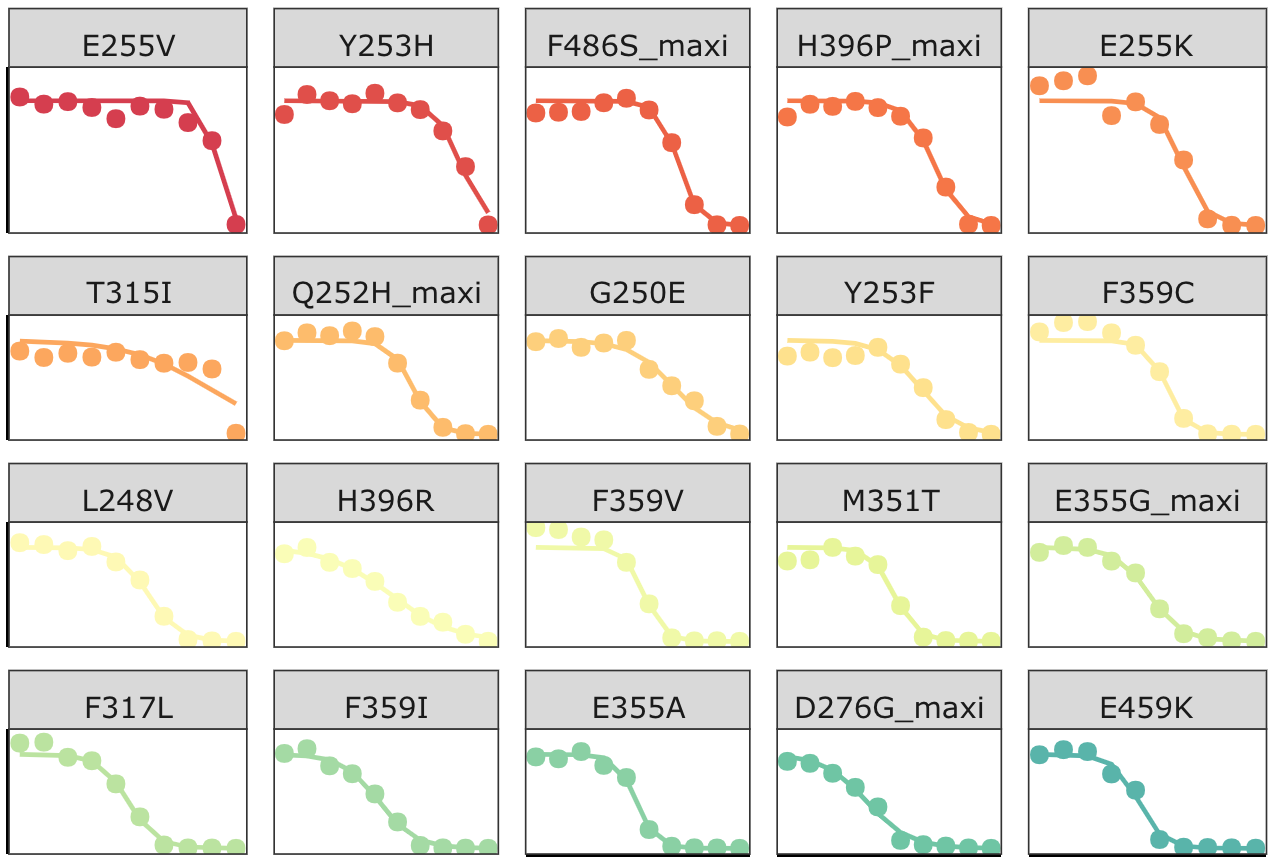
Click here for the script that reads mutation annotation data and generates dataframes with annotated variants and the correct cell counts
Click here for initial growth rate plots
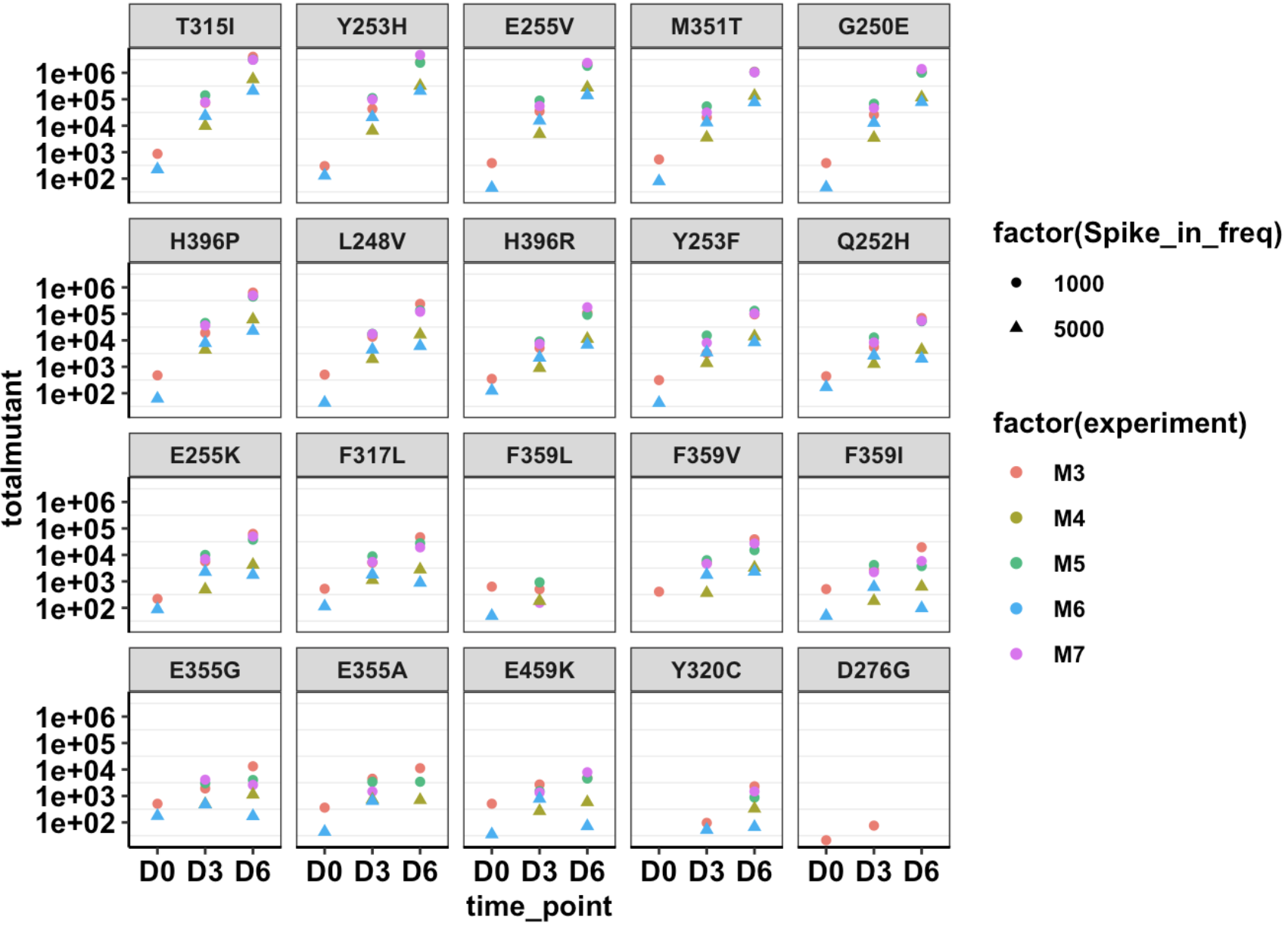
Click here for growth rate plots for ENU mutants
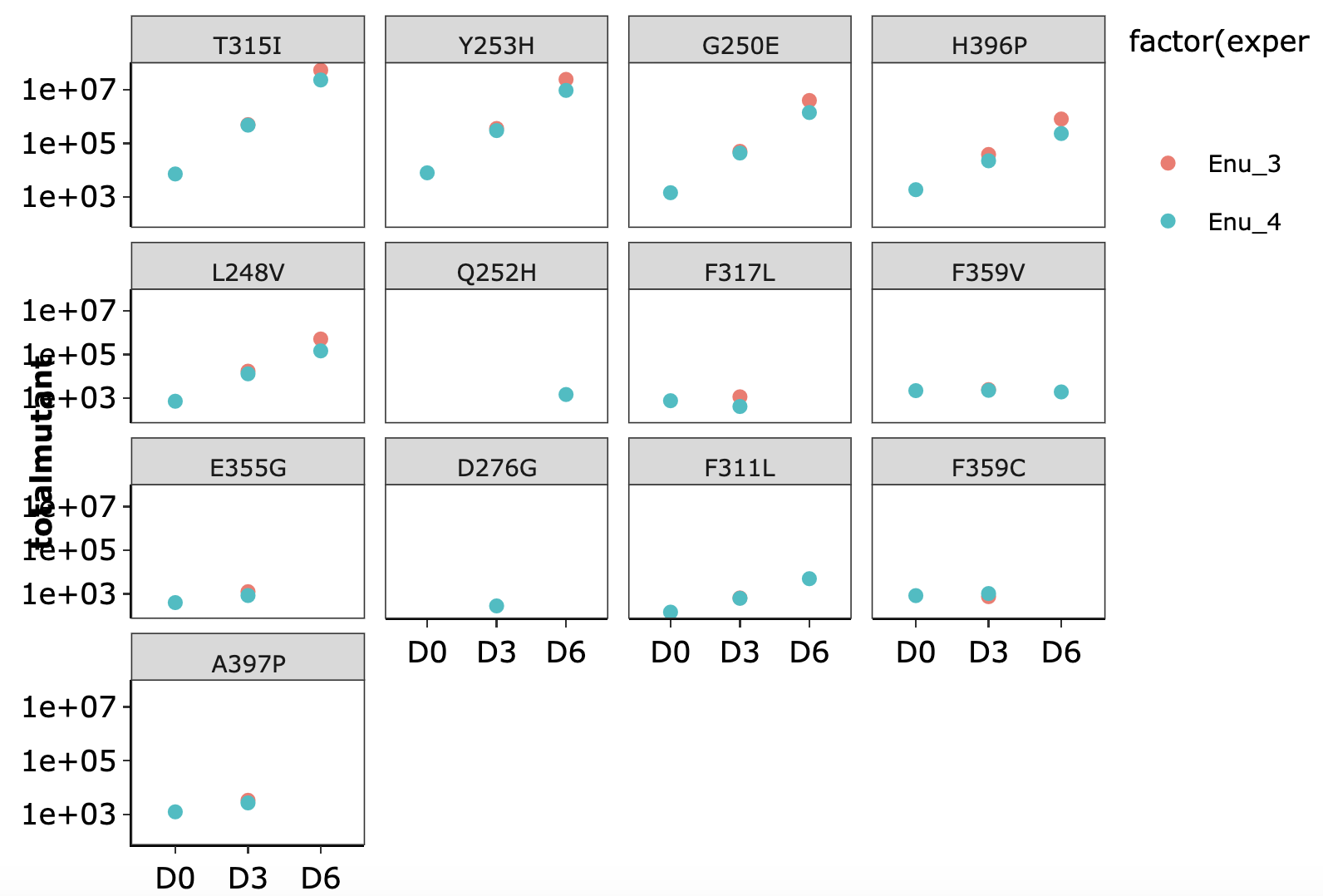
Click here for analysis looking at whether we achieved our desired depth of coverages
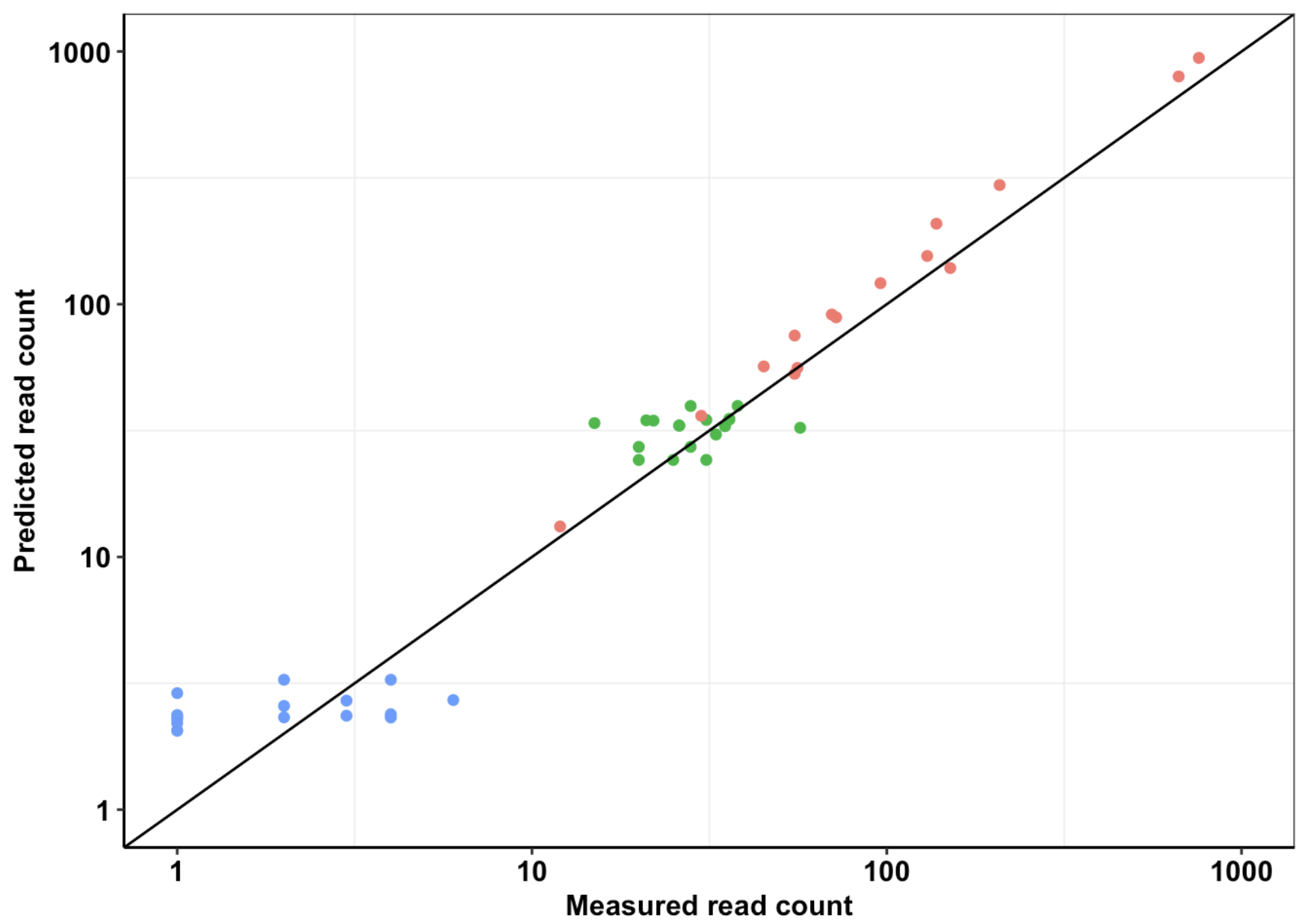
Click here for analysis looking at how well our data predicts BCRABL clinical abundance compared to conventional IC50s
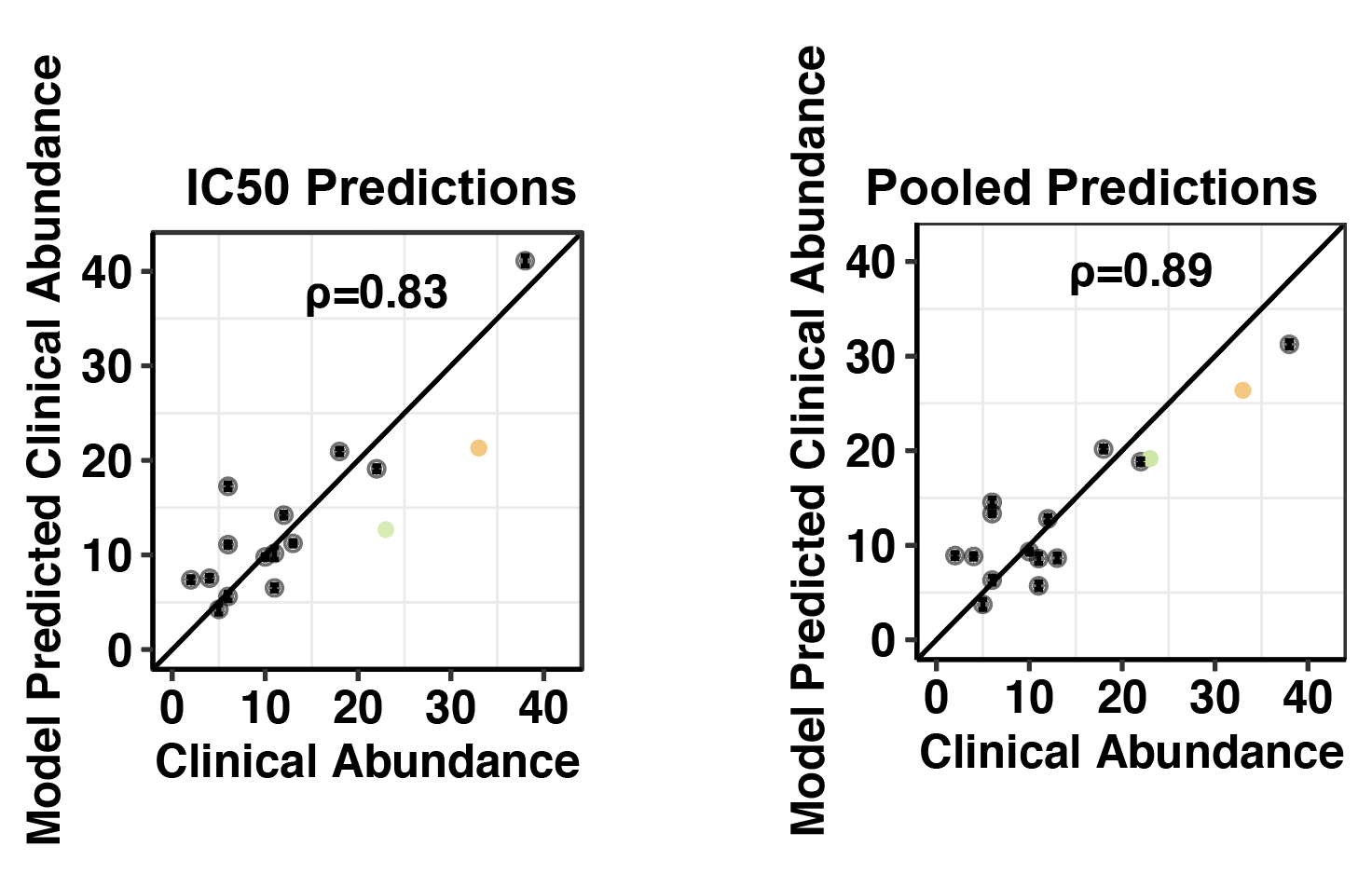
Click here for method to obtain confidence intervals on growth rates from IC50 measurements. Plotted alongside observed growth rate data, these confidence intervals show how well our pooled data matches predictions based off of IC50s. (Needs updates in methodology)
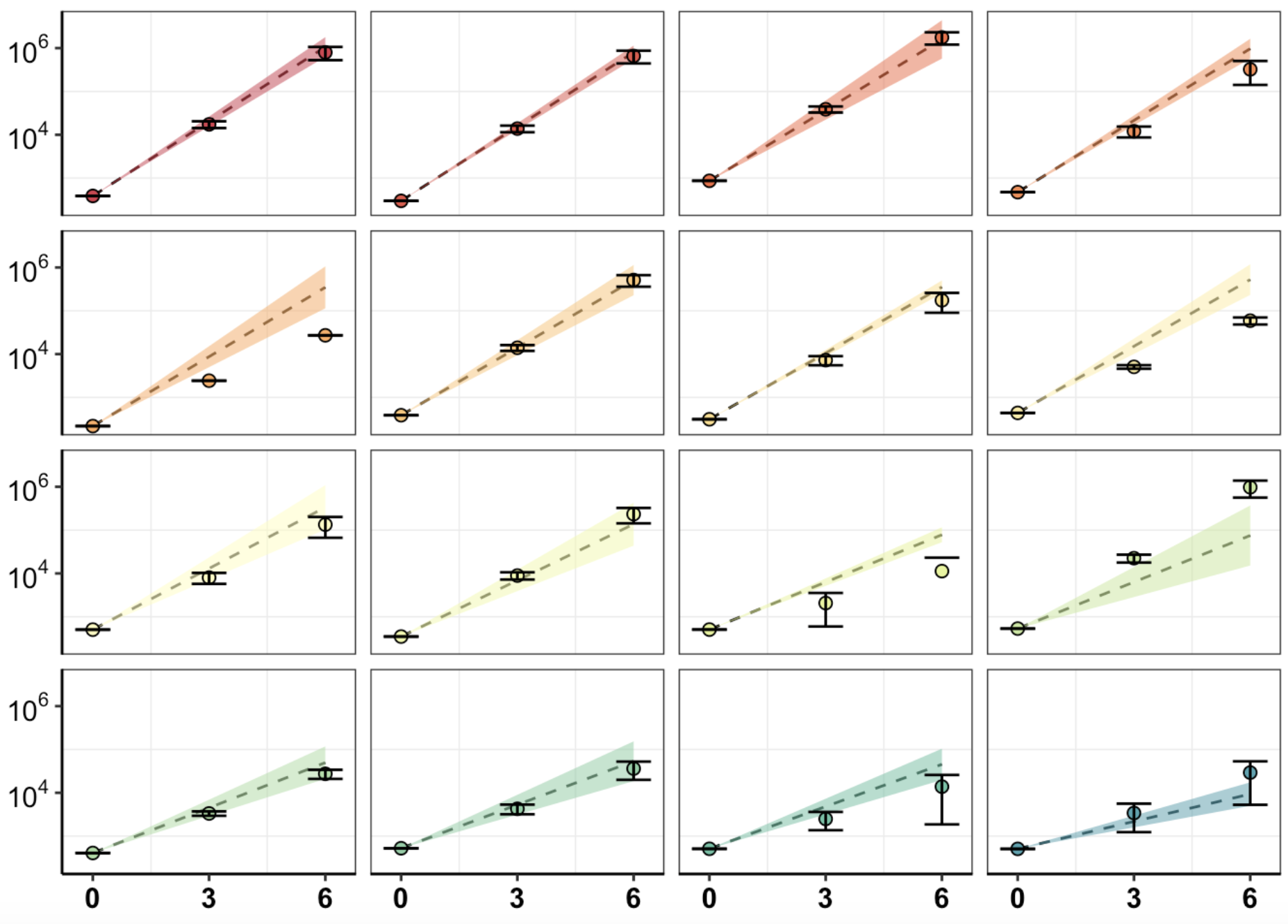
Click here for analyses of replicate to replicate agreement between the spike in replicates. Also includes this analysis for the mutagenesis replicates. Also includes the agreement in observed vs predicted depth of coverages for all sequencing pools. Note: this analysis does not include dosage corrections that improves replicate to replicate heterogeneity.
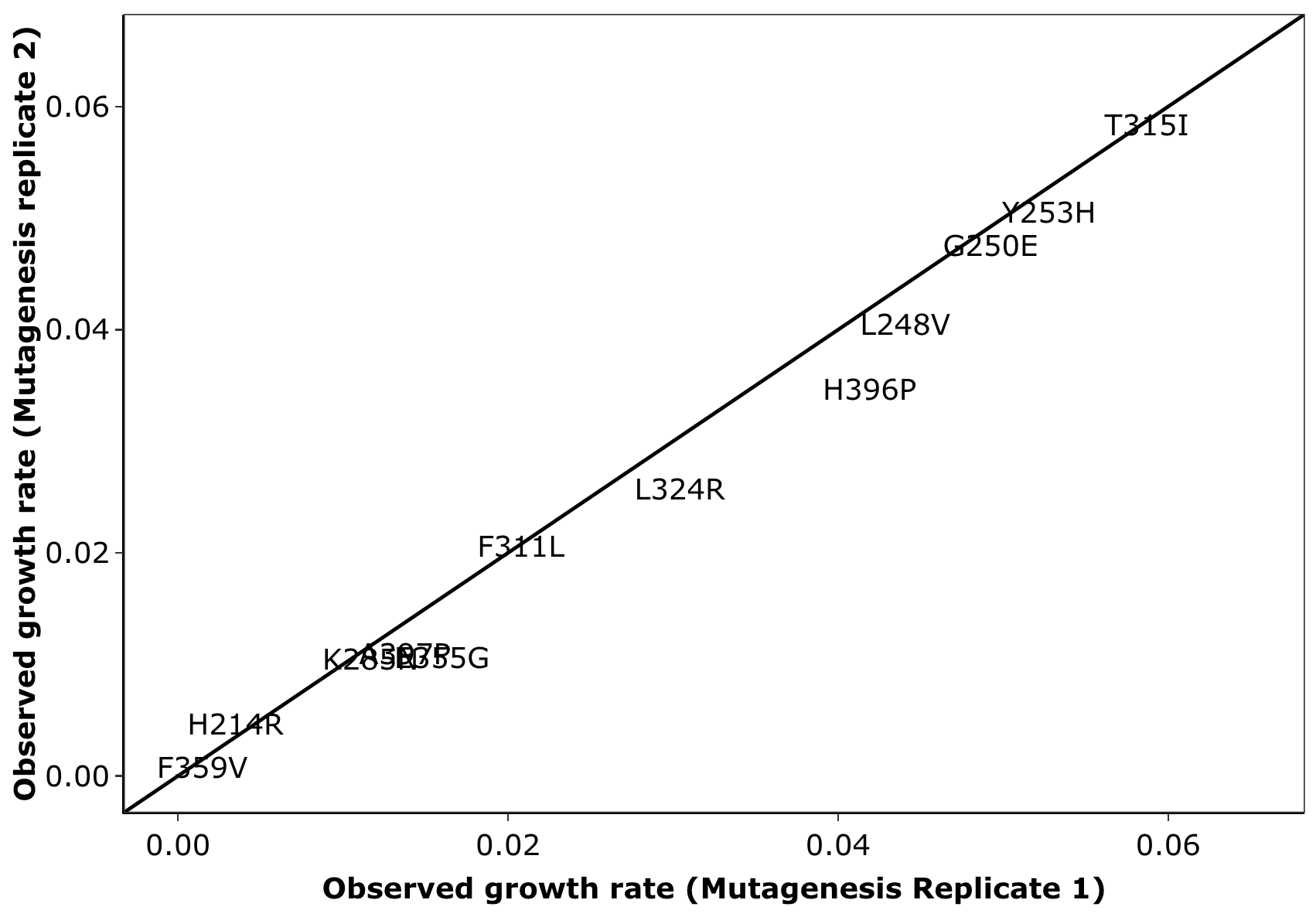
Click here to see analysis of predictions based on Enrich2 vs Shendure vs our method
Click here for our dosing normalization strategy
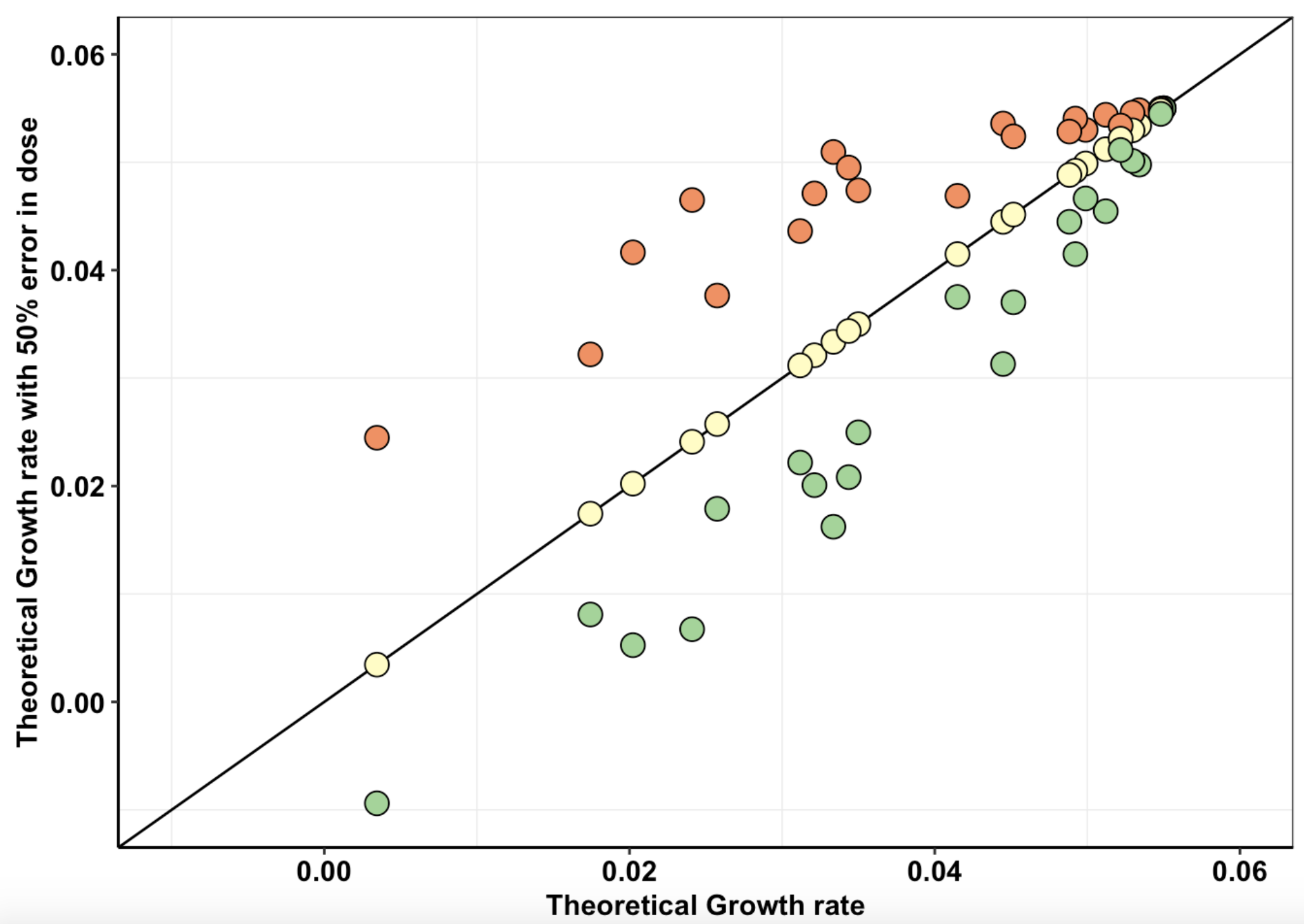
Click here for simulations showing enrichment and depletion events in a 3 mutant pool.
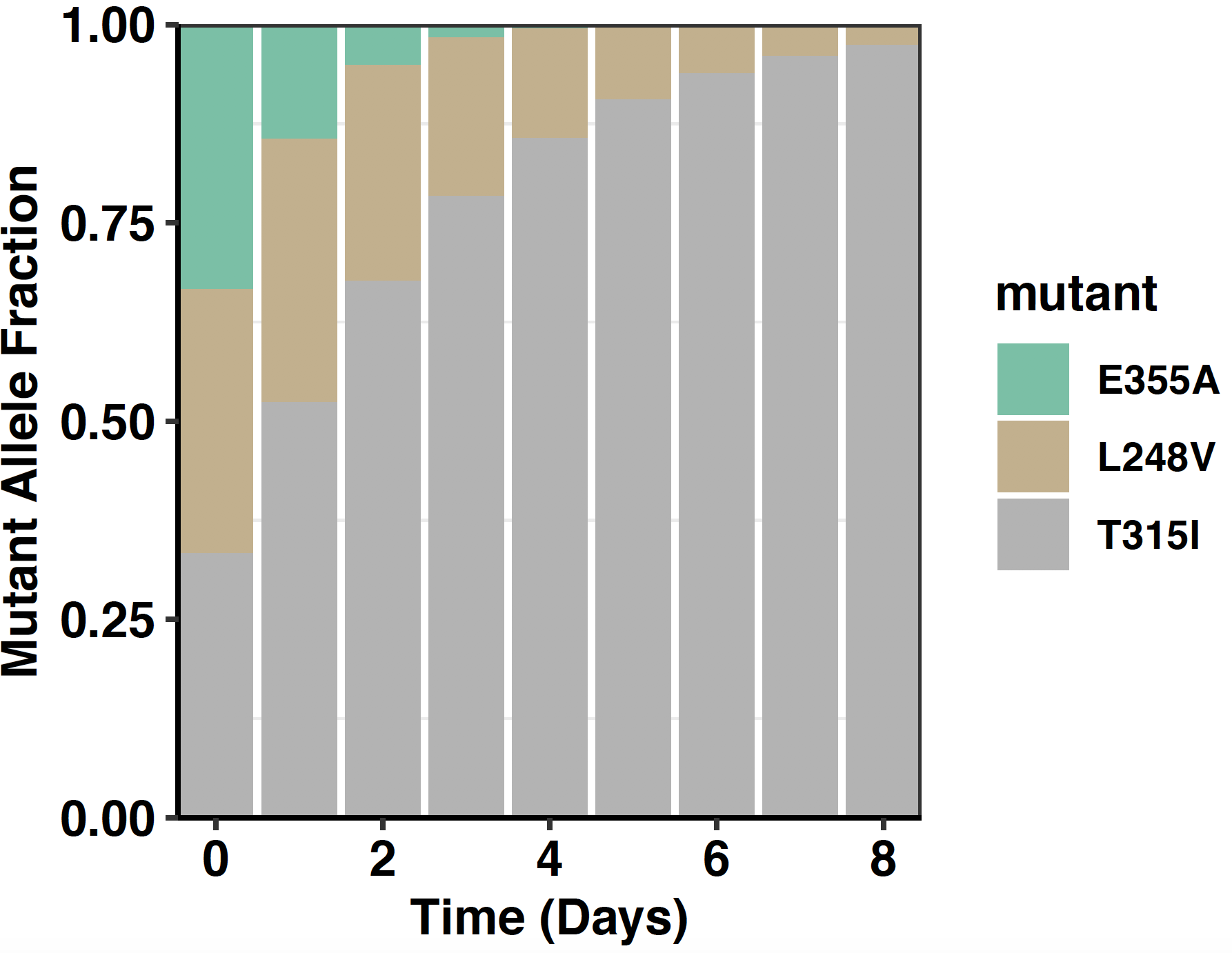
Click here showing the conformity between the growth rates in the presence of drug measured at a low MAF via FACs, sequencing, and via IC50 studies for a single mutant.
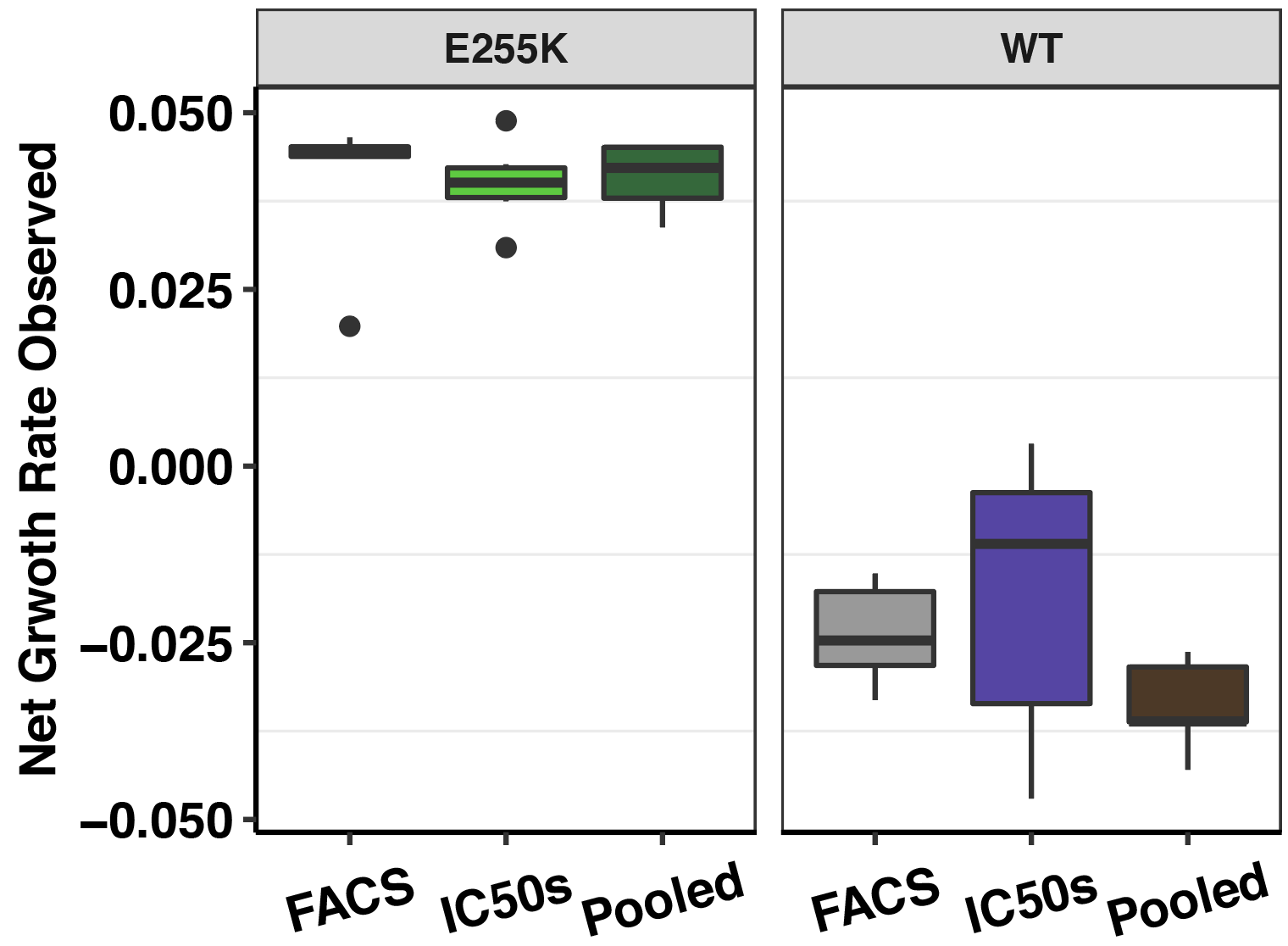
Click here to view BCRABL Imatinib heatmaps generated using CRISPR-cut libraries
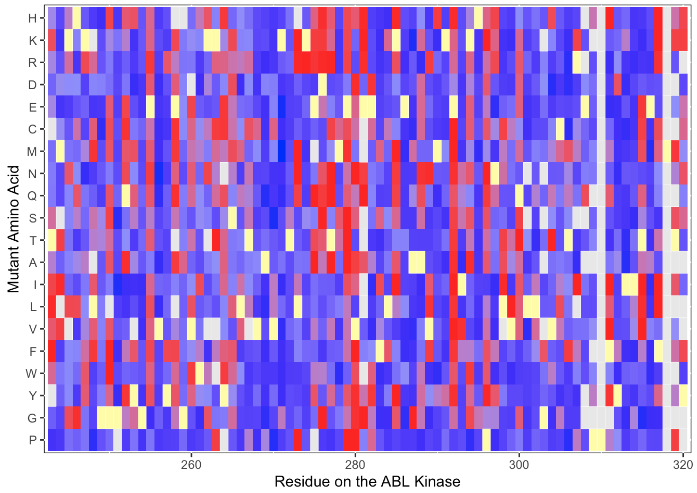
Click here to view preliminary BCRABL Imatinib and Asciminib heatmaps made with enzymatically sheared libraries
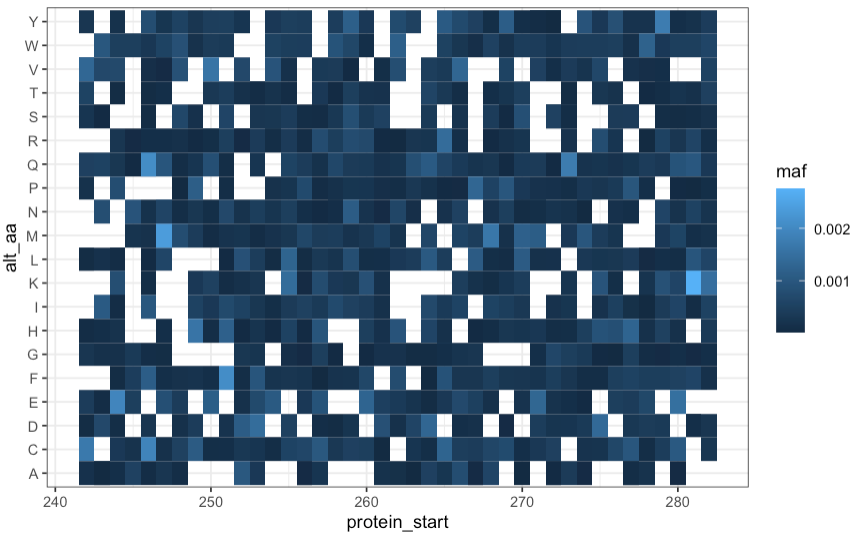
Click here and to view iL3 independence data and here to view iL3 independence heatmap and here to view imatinib heatmaps on enzymatically sheared data
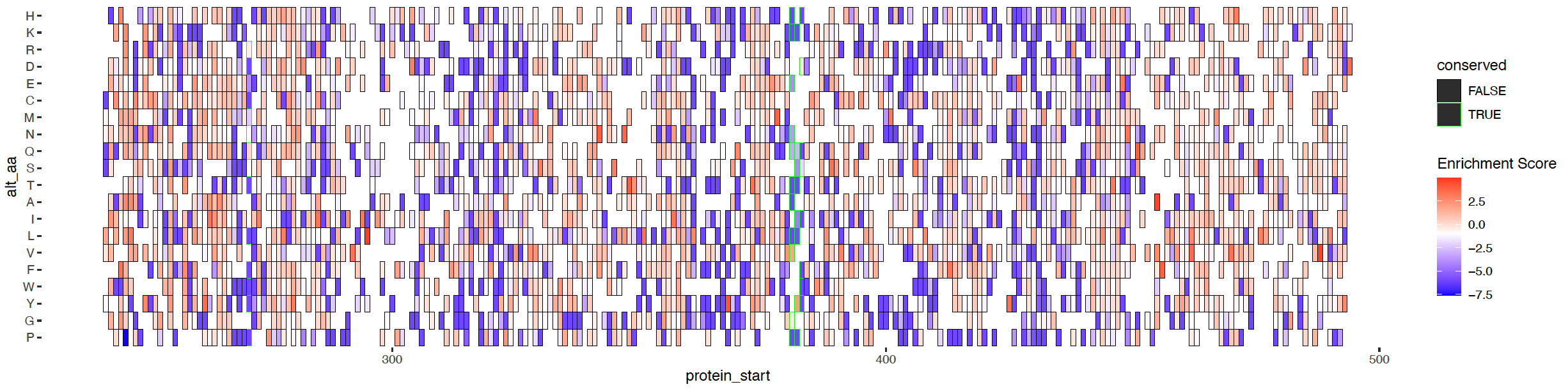
Click here to view error rates without barcodes, with single strand consensi, and with duplex consensus sequencing.
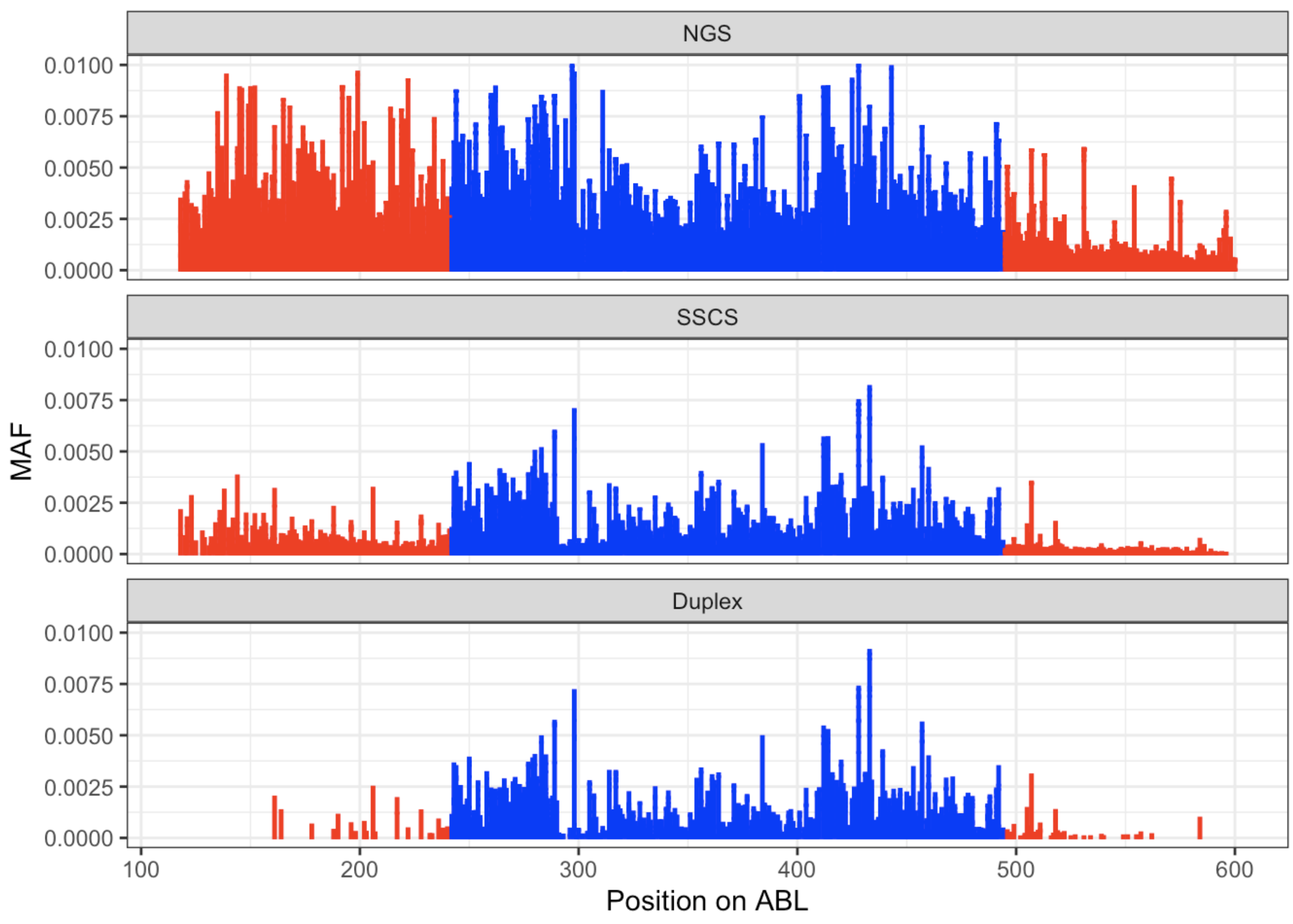
Click here to view the results of the BCRABL Library transduced in K562 Cells
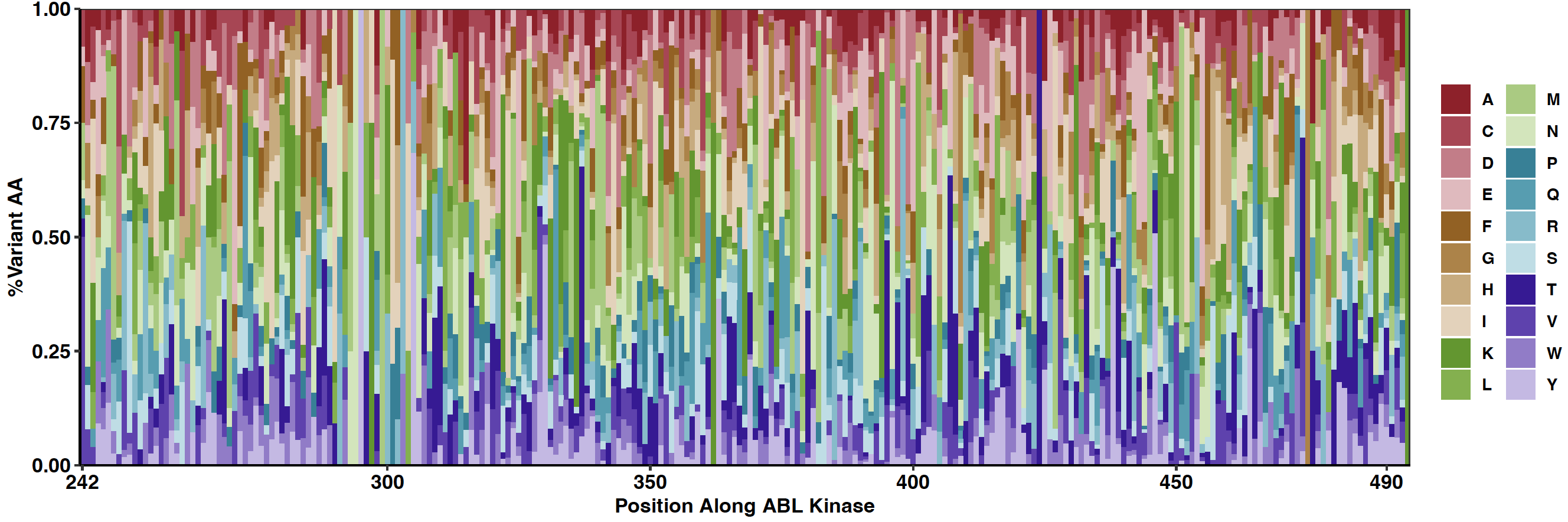
Click here to view analyses of the CDFs of our saturating mutagenesis libraries. These show how even or uneven the libraries are at different stages of the screening process.
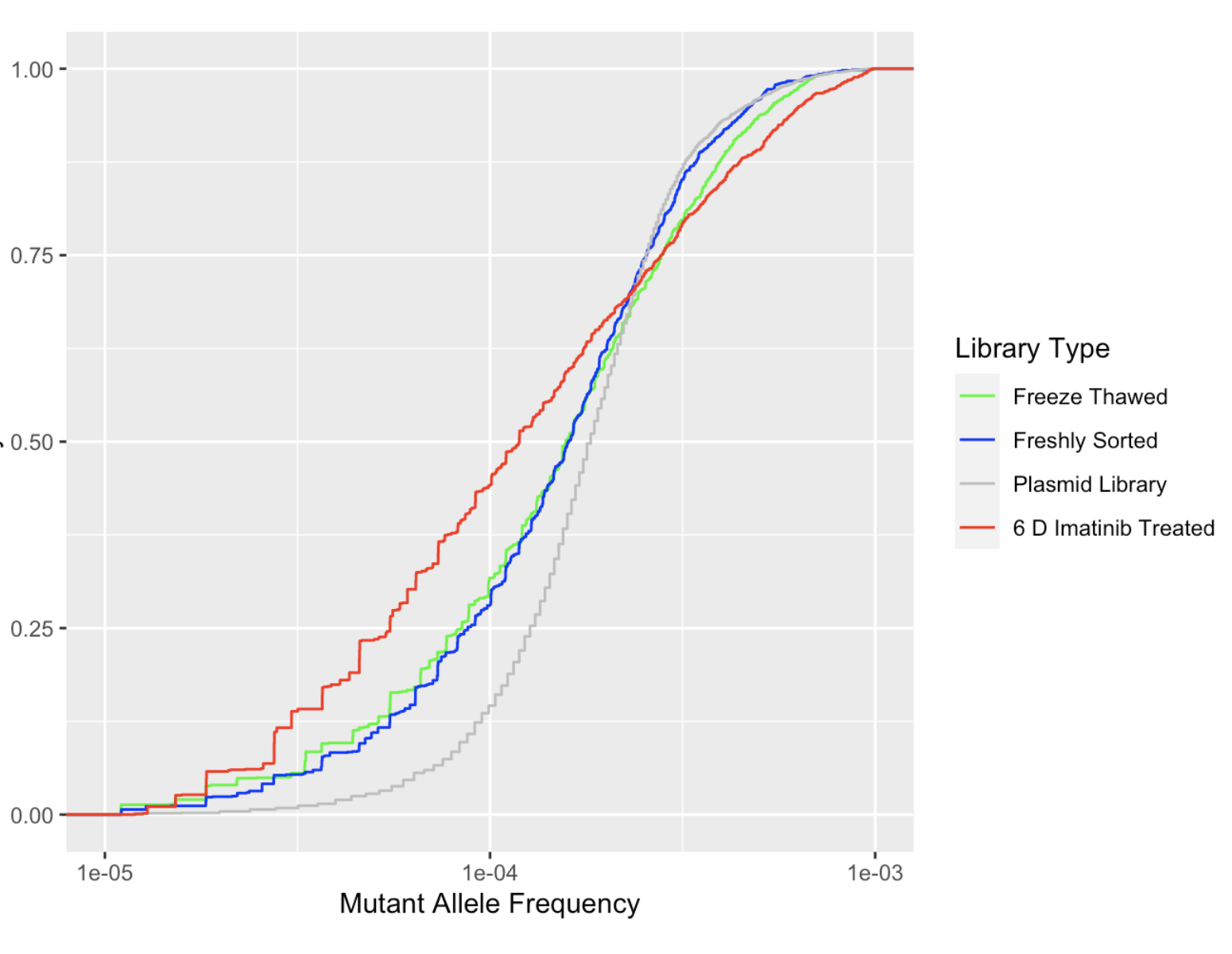
Click here to view analyses in which we compared base editing and saturating mutagenesis data on ABL. This is followed by analysis in which we decide on filter cutoffs for enrichment scores to detect signficantly depleting mutants that drop out for IL3 independence in BaF3s.
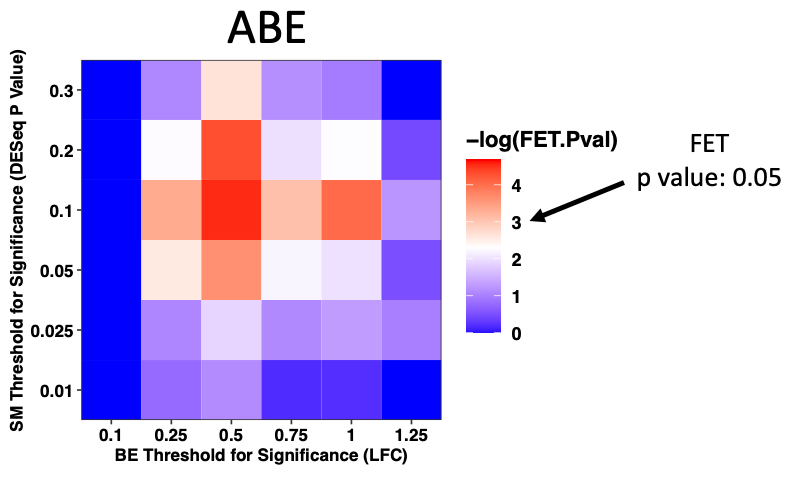
Click here to view analyses in which we predicted admixtures of mutants made by sgRNAs in Base Editing Screens. We then used the saturating mutagenesis data to predict the fitness of these sgRNAs.
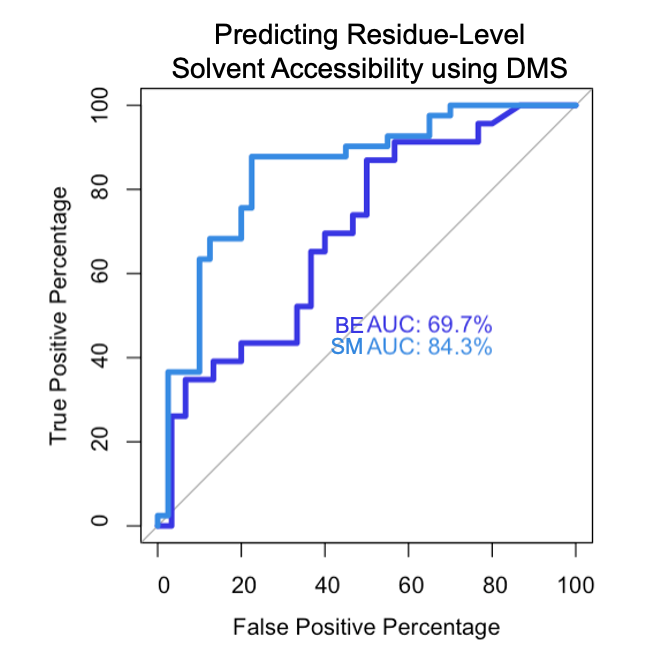
Click here to view initial kinase function analyses.
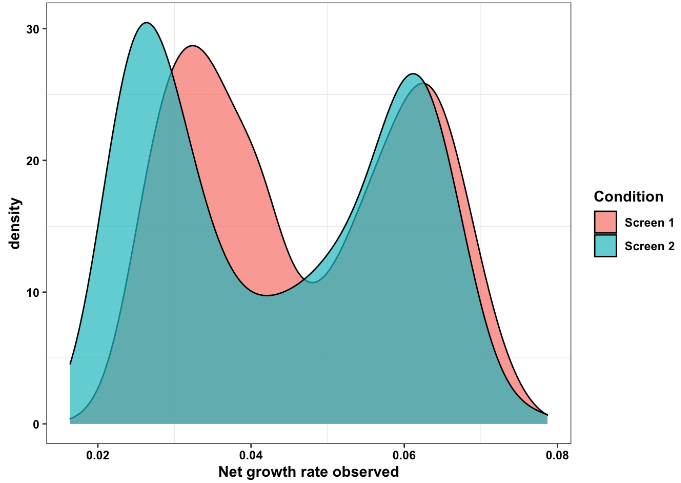
Click here to view our analyses of whether mutants in the cosmic/sanger database are resistant according to our screens.
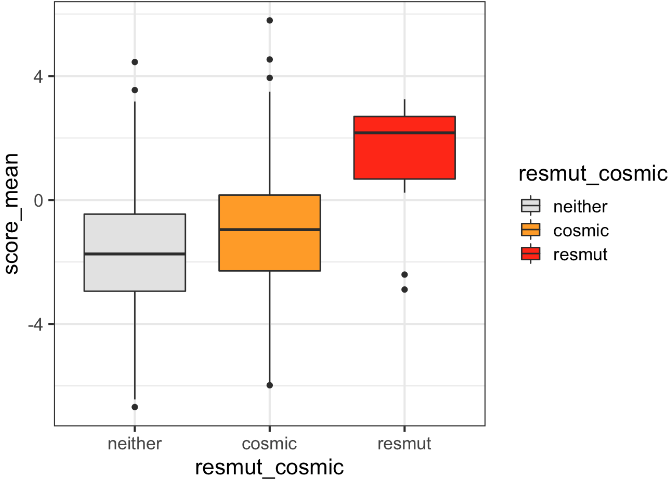 .
.
Click here to be redirected to the github page that contains all the data and analysis rmd files.
What is in each directory:
- Data: contains data downloaded prior to
analyses.
- Output: contains data that the code makes
- Analysis: contains your Rmarkdown files with the
code.
- Docs: contain the html output from the Rmd files in
the analysis directory.
- Code: contains .R files that are functions that the Rmd files in the analysis folder use.