Signatures of selection: Did locusts or specific genes experience positive selection?
Maeva Techer
2025-09-17
Last updated: 2025-09-17
Checks: 4 3
Knit directory:
locust-comparative-genomics/
This reproducible R Markdown analysis was created with workflowr (version 1.7.1). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of
the R Markdown file created these results, you’ll want to first commit
it to the Git repo. If you’re still working on the analysis, you can
ignore this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20221025) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
- absrel parsing
- absrel-plot
- busted unlabel parsing
- busted-enrichment
- busted-label pruned results
- busted-label results
- busted-results2
- busted-results3
- busted-results4
- busted-unlabel pruned results2
- busted-unlabel results
- busted-unlabel results2
- combined full tree plot
- combined plot
- deg and PSGs full
- deg and PSGs pruned
- GO enrich selection
- relax parsing
- relax-busted labelled
- relax-busted
- run pathway selected busted1
- run pathway selected busted2
- unnamed-chunk-21
- venn intensification
- venn relaxation
To ensure reproducibility of the results, delete the cache directory
2_signatures-selection_cache and re-run the analysis. To
have workflowr automatically delete the cache directory prior to
building the file, set delete_cache = TRUE when running
wflow_build() or wflow_publish().
Using absolute paths to the files within your workflowr project makes it difficult for you and others to run your code on a different machine. Change the absolute path(s) below to the suggested relative path(s) to make your code more reproducible.
| absolute | relative |
|---|---|
| /Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data/HYPHY_selection/combined_dnds_table.csv | data/HYPHY_selection/combined_dnds_table.csv |
| /Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data/orthofinder/Polyneoptera/Results_I2_iqtree/Orthogroups/Orthogroups_CladeAssignment_WithCopyStatus_cleaned.csv | data/orthofinder/Polyneoptera/Results_I2_iqtree/Orthogroups/Orthogroups_CladeAssignment_WithCopyStatus_cleaned.csv |
| /Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data/orthofinder/Polyneoptera | data/orthofinder/Polyneoptera |
| /Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data | data |
| /Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data/HYPHY_selection | data/HYPHY_selection |
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version b7729ad. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .Rhistory
Ignored: analysis/.DS_Store
Ignored: analysis/.Rhistory
Ignored: analysis/2_signatures-selection_cache/
Ignored: analysis/figure/
Ignored: code/.DS_Store
Ignored: code/scripts/.DS_Store
Ignored: code/scripts/pal2nal.v14/.DS_Store
Ignored: data/.DS_Store
Ignored: data/DEG_results/.DS_Store
Ignored: data/DEG_results/Bulk_RNAseq/.DS_Store
Ignored: data/DEG_results/Bulk_RNAseq/americana/.DS_Store
Ignored: data/DEG_results/Bulk_RNAseq/cancellata/.DS_Store
Ignored: data/DEG_results/Bulk_RNAseq/cubense/.DS_Store
Ignored: data/DEG_results/Bulk_RNAseq/gregaria/.DS_Store
Ignored: data/DEG_results/Bulk_RNAseq/nitens/.DS_Store
Ignored: data/HYPHY_selection/.DS_Store
Ignored: data/HYPHY_selection/ParsedABSRELResults_unlabeled/.DS_Store
Ignored: data/HYPHY_selection/pathway_enrichment/.DS_Store
Ignored: data/HYPHY_selection/pathway_enrichment/americana/
Ignored: data/HYPHY_selection/pathway_enrichment/cancellata/
Ignored: data/HYPHY_selection/pathway_enrichment/cubense/
Ignored: data/HYPHY_selection/pathway_enrichment/nitens/
Ignored: data/HYPHY_selection/pathway_enrichment/piceifrons/
Ignored: data/WGCNA/.DS_Store
Ignored: data/WGCNA/input/.DS_Store
Ignored: data/WGCNA/input/Bulk_RNAseq/.DS_Store
Ignored: data/WGCNA/output/.DS_Store
Ignored: data/WGCNA/output/Bulk_RNAseq/.DS_Store
Ignored: data/WGCNA/output/Bulk_RNAseq/gregaria/.DS_Store
Ignored: data/behavioral_data/.DS_Store
Ignored: data/behavioral_data/Raw_data/.DS_Store
Ignored: data/cafe5_results/.DS_Store
Ignored: data/list/.DS_Store
Ignored: data/list/Bulk_RNAseq/.DS_Store
Ignored: data/list/GO_Annotations/.DS_Store
Ignored: data/list/GO_Annotations/DesertLocustR/.DS_Store
Ignored: data/list/excluded_loci/.DS_Store
Ignored: data/orthofinder/.DS_Store
Ignored: data/orthofinder/Polyneoptera/.DS_Store
Ignored: data/orthofinder/Polyneoptera/Results_I2_iqtree/.DS_Store
Ignored: data/orthofinder/Polyneoptera/Results_I2_iqtree/Orthogroups/.DS_Store
Ignored: data/orthofinder/Polyneoptera/Results_I2_withDaust/.DS_Store
Ignored: data/orthofinder/Polyneoptera/Results_I2_withDaust/Orthogroups/.DS_Store
Ignored: data/orthofinder/Schistocerca/.DS_Store
Ignored: data/orthofinder/Schistocerca/Results_I2/.DS_Store
Ignored: data/orthofinder/Schistocerca/Results_I2/Orthogroups/.DS_Store
Ignored: data/overlap/.DS_Store
Ignored: data/pathway_enrichment/.DS_Store
Ignored: data/pathway_enrichment/OLD/.DS_Store
Ignored: data/pathway_enrichment/OLD/custom_sgregaria_orgdb/.DS_Store
Ignored: data/pathway_enrichment/REVIGO_results/.DS_Store
Ignored: data/pathway_enrichment/REVIGO_results/BP/.DS_Store
Ignored: data/pathway_enrichment/REVIGO_results/CC/.DS_Store
Ignored: data/pathway_enrichment/REVIGO_results/MF/.DS_Store
Ignored: data/pathway_enrichment/americana/.DS_Store
Ignored: data/pathway_enrichment/cancellata/.DS_Store
Ignored: data/pathway_enrichment/gregaria/.DS_Store
Ignored: data/pathway_enrichment/nitens/Thorax/
Ignored: data/pathway_enrichment/piceifrons/.DS_Store
Ignored: data/readcounts/.DS_Store
Ignored: data/readcounts/Bulk_RNAseq/.DS_Store
Ignored: data/readcounts/RNAi/.DS_Store
Untracked files:
Untracked: analysis/KEGG_expansion_diagnostics.csv
Untracked: data/HYPHY_selection/functional_pathways/
Untracked: data/RefSeq/
Untracked: data/cafe5_results/Base_change_FILE/
Untracked: data/cafe5_results/Gene_count_FILE/
Unstaged changes:
Modified: analysis/2_gene-family.Rmd
Modified: analysis/2_psmc-analysis.Rmd
Modified: analysis/2_signatures-selection.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown
(analysis/2_signatures-selection.Rmd) and HTML
(docs/2_signatures-selection.html) files. If you’ve
configured a remote Git repository (see ?wflow_git_remote),
click on the hyperlinks in the table below to view the files as they
were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | b7729ad | Maeva TECHER | 2025-09-10 | adding selction part |
| html | b7729ad | Maeva TECHER | 2025-09-10 | adding selction part |
| Rmd | 6367e32 | Maeva TECHER | 2025-09-04 | Updates PSMC |
| html | 6367e32 | Maeva TECHER | 2025-09-04 | Updates PSMC |
| html | 05239ca | Maeva TECHER | 2025-07-02 | Build site. |
| Rmd | b6a3e83 | Maeva TECHER | 2025-07-02 | workflowr::wflow_publish("analysis/2_signatures-selection.Rmd") |
| html | 6146883 | Maeva TECHER | 2025-07-01 | Build site. |
| Rmd | a2d2955 | Maeva TECHER | 2025-07-01 | Updated wgcna and compiling |
| html | a2d2955 | Maeva TECHER | 2025-07-01 | Updated wgcna and compiling |
| html | 0168e2b | Maeva TECHER | 2025-06-05 | Build site. |
| html | 9a03ca6 | Maeva TECHER | 2025-06-05 | Update website |
| html | 17484e8 | Maeva TECHER | 2025-06-05 | Build site. |
| html | 3e696d6 | Maeva TECHER | 2025-06-05 | Adding ortho heatmap |
| Rmd | 4e391c3 | Maeva TECHER | 2025-05-30 | add new analysis orthology, synteny |
| html | 4e391c3 | Maeva TECHER | 2025-05-30 | add new analysis orthology, synteny |
| Rmd | cacc1db | Maeva TECHER | 2025-05-02 | updates files |
| html | cacc1db | Maeva TECHER | 2025-05-02 | updates files |
| Rmd | b982319 | Maeva TECHER | 2025-03-03 | update font |
| html | b982319 | Maeva TECHER | 2025-03-03 | update font |
| html | f6a4762 | Maeva TECHER | 2025-02-27 | Build site. |
| Rmd | e55bac6 | Maeva TECHER | 2025-01-26 | Updating the github |
| html | e55bac6 | Maeva TECHER | 2025-01-26 | Updating the github |
| html | faf2db3 | Maeva TECHER | 2025-01-13 | update markdown |
| html | 6954b9b | Maeva TECHER | 2025-01-13 | Build site. |
| Rmd | 8df3d7c | Maeva TECHER | 2025-01-13 | changes |
| Rmd | b80db34 | Maeva TECHER | 2025-01-13 | Adding selection analysis part |
| html | b80db34 | Maeva TECHER | 2025-01-13 | Adding selection analysis part |
| html | 3fa8e62 | Maeva TECHER | 2024-11-09 | updated analysis |
| html | edb70fe | Maeva TECHER | 2024-11-08 | overlap and deg results created |
| html | ba35b82 | Maeva A. TECHER | 2024-06-20 | Build site. |
| html | acfa0db | Maeva A. TECHER | 2024-05-14 | Build site. |
| Rmd | 2c5b31c | Maeva A. TECHER | 2024-05-14 | wflow_publish("analysis/2_signatures-selection.Rmd") |
| html | 0837617 | Maeva A. TECHER | 2024-01-30 | Build site. |
| html | f701a01 | Maeva A. TECHER | 2024-01-30 | reupdate |
| html | 6e878be | Maeva A. TECHER | 2024-01-24 | Build site. |
| html | 1b09cbe | Maeva A. TECHER | 2024-01-24 | remove |
| html | 4ae7db7 | Maeva A. TECHER | 2023-12-18 | Build site. |
| Rmd | 53877fa | Maeva A. TECHER | 2023-12-18 | add pages |
Testing for signatures of selection using HyPhy: aBSREL, BUSTED and RELAX
Note: We used OrthoFinder results, PAL2NAL and HyPhy to identify signatures of selection in orthologous genes. For this part, refers to the well curated pipeline FormicidaeMolecularEvolution by Megan Barkdull (Assistant Curator of Entomology at the Natural History Museum of Los Angeles County). We describe below the modifications made and mostly copied the workflow from her Github.
We will be running three methods on our tree:
Has a gene experienced positive selection at any site in a locust species or group of species? To answer this question, we will apply BUSTED (Branch-Site Unrestricted Statistical Test for Episodic Diversification). This method works well for datasets with fewer than 10 taxa and helps identify positive selection events associated with species or groups.
Are certain species in the Schistocerca phylogeny subject to episodic (at a subset of sites) positive or purifying selection? For this analysis, we will use aBSREL (adaptive Branch-Site Random Effects Likelihood), the preferred method for detecting episodic selection on individual branches within the locust phylogeny.
Have selection pressures on genes been relaxed or intensified in a subset of Schistocerca species? For this, we will use RELAX which is not designed to detect positive selection but rather to determine whether selection pressures have been relaxed or intensified along a specified set of “test” branches.
1. Parsing orthogroups files for PAL2NAL
The script written by M. Barkdull remains unchanged; however, it
requires R with the phylotools package installed. This step
ensures that the OrthoFinder FASTA file is reordered. Instead of having
one file per orthogroup, this process consolidates the data into
species-specific files, with all orthogroups combined and properly
reordered. These files will be input for PAL2NAL, which is
a program that converts a multiple sequence alignement of proteins and
the corresponding DNA sequences (here cds) into a codon alignment.
srun --ntasks 1 --cpus-per-task 8 --mem 50G --time 04:00:00 --pty bash
ml GCC/13.2.0 OpenMPI/4.1.6 R_tamu/4.4.1 MCScanX/2024.19.19
export R_LIBS=$SCRATCH/R_LIBS_USER/
# Example for Schistocerca only
./scripts/DataMSA.R ./scripts/inputurls_Schistocerca_Jan2025.txt /scratch/group/songlab/maeva/LocustsGenomeEvolution/Schistocerca_I2/5_OrthoFinder/fasta/Results_Jan15_I2/MultipleSequenceAlignments/
# Example for Polyneoptera
./scripts/DataMSA.R ./scripts/inputurls_13polyneoptera_May2025.txt /scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/MultipleSequenceAlignments/This is the final messages we get when it is successful.
Once we have obtained all the files, we go to the next step which is
filtering the protein alignment files to contain only the subset of
genes that will be called by PAL2NAL. This is due to the
fact that certain genes were not classified in orthogroups.
ml GCC/13.2.0 OpenMPI/4.1.6 R_tamu/4.4.1
export R_LIBS=$SCRATCH/R_LIBS_USER/
# Example for Schistocerca only
./scripts/FilteringCDSbyMSA.R ./scripts/inputurls_Schistocerca_Jan2025.txt
# Example for Polyneoptera
./scripts/FilteringCDSbyMSA.R ./scripts/inputurls_13polyneoptera_May2025.txt Some of the files seems to have discrepancy of one “>” entry line
between the protein and cds file (due to a concatenation error that I
could not troubleshoot) so we are going to run the script
doublecheckCDSbyMAS which I created to remove extra
line.
# Example for Schistocerca only
./scripts/doublecheckCDSbyMAS ./scripts/inputurls_Schistocerca_Jan2025.txt
# Example for Polyneoptera
./scripts/doublecheckCDSbyMAS ./scripts/inputurls_13polyneoptera_May2025.txt
# You can also check if there is a difference with the following quick steps
grep ">" ./6_1_SpeciesMSA/proteins_Sscub.fasta | sort > proteins_Sscub_names.txt
grep ">" ./6_2_FilteredCDS/filtered_Sscub_cds.fasta | sort > cds_Sscub_names.txt
diff proteins_Sscub_names.txt cds_Sscub_names.txtThe following is the content of doublecheckCDSbyMAS:
#!/bin/bash
# Check if input file is provided
if [ "$#" -lt 1 ]; then
echo "Usage: $0 <input_file>"
exit 1
fi
# Input file containing species information
input_file=$1
# Directories
protein_dir="./6_1_SpeciesMSA"
cds_dir="./6_2_FilteredCDS"
backup_dir="$protein_dir/backup"
log_file="./cleaning_check.log"
# Create necessary directories
mkdir -p "$backup_dir"
rm -f "$log_file" # Clear previous logs
# Extract species abbreviations (no header in the file)
species_list=$(awk -F',' '{print $4}' "$input_file")
# Loop through each species
for species in $species_list; do
protein_file="$protein_dir/proteins_${species}.fasta"
cds_file="$cds_dir/filtered_${species}_cds.fasta"
cleaned_protein_file="$protein_dir/proteins_${species}_cleaned.fasta"
cleaned_cds_file="$cds_dir/filtered_${species}_cds_cleaned.fasta"
echo "Processing species: $species"
# Check if protein and CDS files exist
if [[ -f "$protein_file" && -f "$cds_file" ]]; then
# Backup the original protein file
cp "$protein_file" "$backup_dir/proteins_${species}.fasta.bak"
echo "Backup created for: $protein_file -> $backup_dir/proteins_${species}.fasta.bak"
# Cleaning Step: Align sequence headers between protein and CDS files
grep ">" "$protein_file" | sort > proteins_names.txt
grep ">" "$cds_file" | sort > cds_names.txt
# Identify common sequence headers
comm -12 proteins_names.txt cds_names.txt > common_names.txt
# Check if common_names.txt is empty (indicating no matching headers)
if [[ ! -s common_names.txt ]]; then
echo "ERROR: No common sequence headers found for species: $species" >> "$log_file"
echo "ERROR: Cleaning failed for species: $species due to no matching sequence headers."
continue
fi
# Filter protein file
grep -A 1 -Ff common_names.txt "$protein_file" > "$cleaned_protein_file" || {
echo "ERROR: Failed to clean protein file for species: $species" >> "$log_file"
continue
}
# Filter CDS file
grep -A 1 -Ff common_names.txt "$cds_file" > "$cleaned_cds_file" || {
echo "ERROR: Failed to clean CDS file for species: $species" >> "$log_file"
continue
}
# Replace the original files with cleaned versions
mv "$cleaned_protein_file" "$protein_file"
mv "$cleaned_cds_file" "$cds_file"
# Perform grep check to validate cleaning
grep ">" "$protein_file" | sort > proteins_names_cleaned.txt
grep ">" "$cds_file" | sort > cds_names_cleaned.txt
diff_output=$(diff proteins_names_cleaned.txt cds_names_cleaned.txt)
if [[ -z "$diff_output" ]]; then
echo "Check passed for species: $species" >> "$log_file"
echo "Protein and CDS sequence names match for species: $species."
else
echo "Check failed for species: $species" >> "$log_file"
echo "Protein and CDS sequence names mismatch for species: $species." >> "$log_file"
echo "$diff_output" >> "$log_file"
fi
else
echo "ERROR: Missing files for species: $species" >> "$log_file"
echo "ERROR: Protein or CDS file missing for species: $species. Skipping."
fi
done
# Cleanup temporary files
rm -f proteins_names.txt cds_names.txt common_names.txt proteins_names_cleaned.txt cds_names_cleaned.txt
echo "All species processed. Logs saved to $log_file."2. Generating codon-aware nucleotide alignments
PAL2NAL is installed on Grace as a module but the same
version is available in the script of this repository. We will use the
inputs generated in the previous step to obtain codon-aware
alignments.
# Example for Schistocerca only
./scripts/DataRunPAL2NAL ./scripts/inputurls_Schistocerca_Jan2025.txt
# Example for Polyneoptera
./scripts/DataRunPAL2NAL ./scripts/inputurls_13polyneoptera_May2025.txt 3. Assembling nucleotide sequence orthogroups for input in HyPHY
From M. Bardull: For some models like BUSTED, we need files that
contain orthologous nucleotide sequences from each species. Therefore,
we must recombine our codon-aware alignments in a step that is the
inverse of previous steps. To do this, use the R script
./scripts/DataSubsetCDS.R. Run with the command:
ml GCC/13.2.0 OpenMPI/4.1.6 R_tamu/4.4.1
export R_LIBS=$SCRATCH/R_LIBS_USER/
# Example for Schistocerca only
./scripts/DataSubsetCDS.R ./scripts/inputurls_Schistocerca_Jan2025.txt /scratch/group/songlab/maeva/LocustsGenomeEvolution/Schistocerca_I2/5_OrthoFinder/fasta/Results_Jan15_I2/MultipleSequenceAlignments/
# Example for Polyneoptera
./scripts/DataSubsetCDS.R ./scripts/inputurls_13polyneoptera_May2025.txt /scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/MultipleSequenceAlignments/From M. Bardull: BUSTED will not run on sequences which contain stop
codons, even if these are reasonable, terminal stop codons.
HypPhy includes a utility which will mask these these
terminal stop codons in the orthogroups (there should be few-to-no other
stop codons, because our alignments are codon-aware). To execute this
step, use the following:
module purge
ml GCC/13.3.0 OpenMPI/5.0.3 HyPhy/2.5.71
./scripts/DataRemoveStopCodons
# for large groups launch it with sbatch
sbatch ./scripts/DataRemoveStopCodons4. Preparing labeled phylogenies
Before performing a signature of selection analysis using HyPhy, it is important to note that some methods such as RELAX, require the phylogeny to have labeled branches to define branches. These labels define branch sets for selection testing and allow to compare selection pressures.
So we modify the script LabellingPhylogeniesHYPHY.R
ml GCC/13.2.0 OpenMPI/4.1.6 R_tamu/4.4.1
export R_LIBS=$SCRATCH/R_LIBS_USER/
# Example for Schistocerca only
./scripts/LabellingPhylogeniesHYPHY.R /scratch/group/songlab/maeva/LocustsGenomeEvolution/Schistocerca_I2/5_OrthoFinder/fasta/Results_Jan15_I2/Resolved_Gene_Trees/ Locusts.txt Locusts
# Example for Polyneoptera
./scripts/LabellingPhylogeniesHYPHY.R /scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Resolved_Gene_Trees/ Locusts.txt LocustsHow it appears when it is successful. You can see that the locust species are labelled with {Foreground}.
After running BUSTED one time, I realized that I want to check the
signal of selection only on Schistocerca and Locusta.
For that, I am pruning the trees and the fasta files of Polyneoptera if
I want to keep it with the following script
PruningLabellingPhylogeniesHYPHY.R:
ml GCC/13.2.0 OpenMPI/4.1.6 R_tamu/4.4.1
export R_LIBS=$SCRATCH/R_LIBS_USER/
# Example for Polyneoptera
./scripts/PruningLabellingPhylogeniesHYPHY.R /scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Resolved_Gene_Trees/ Locusts.txt Species2keep.txt LocustsHow the input files should look:
[maeva-techer@grace1 Polyneoptera_FINAL]$ cat Species2keep.txt
Samer
Sscub
Snite
Sgreg
Scanc
Spice
LmigrDetails of the PruningLabellingPhylogeniesHYPHY.R are
pasted below. If we used these folders, we will have to modify our
Run{HYPHYMETHOD} to reflect the source of the new fasta sequences in
8_2_RemovedStops_Pruned:
[maeva-techer@grace1 Polyneoptera_FINAL]$ cat scripts/PruningLabellingPhylogeniesHYPHY.R
#!/usr/bin/env Rscript
# ============================= #
# LOAD LIBRARIES #
# ============================= #
suppressPackageStartupMessages({
library(fs)
library(Biostrings)
library(ape)
library(tidyverse)
library(purrr)
})
# ============================= #
# READ ARGUMENTS #
# ============================= #
args <- commandArgs(trailingOnly = TRUE)
if (length(args) < 3) {
stop("Usage: Rscript LabelAndPruneTreesHYPHY.R <tree_dir> <foreground_species.txt> <retained_species.txt>", call. = FALSE)
}
tree_dir <- args[1]
fg_species_file <- args[2]
retained_species_file <- args[3]
output_prefix <- ifelse(length(args) >= 4, args[4], "labelled")
# ============================= #
# READ SPECIES FILES #
# ============================= #
foreground_species <- read_lines(fg_species_file) %>% str_trim()
retained_species <- read_lines(retained_species_file) %>% str_trim()
# ============================= #
# OUTPUT SETUP #
# ============================= #
tree_output_dir <- file.path("9_1_LabelledPhylogenies_Pruned", output_prefix)
fasta_input_dir <- "8_2_RemovedStops"
fasta_output_dir <- "8_2_RemovedStops_Pruned"
dir_create(tree_output_dir)
dir_create(fasta_output_dir)
# ============================= #
# LABEL + PRUNE FUNC #
# ============================= #
multiTreeLabelAndPrune <- function(tree_path, retained_sp, foreground_sp, export_path) {
tree <- read.tree(tree_path)
message("🌳 Processing tree: ", basename(tree_path))
# Get tip abbreviations
tip_species <- sapply(strsplit(tree$tip.label, "_"), `[`, 1)
keep_tips <- tree$tip.label[tip_species %in% retained_sp]
if (length(keep_tips) < 4) {
message("⚠️ Skipping ", basename(tree_path), " — fewer than 4 retained tips.")
return(NULL)
}
pruned_tree <- drop.tip(tree, setdiff(tree$tip.label, keep_tips))
# Label foreground tips
pruned_tree$tip.label <- map_chr(pruned_tree$tip.label, function(label) {
sp_abbr <- strsplit(label, "_")[[1]][1]
if (sp_abbr %in% foreground_sp) paste0(label, "{Foreground}") else label
})
# Label nodes leading to foreground
fg_tips <- grep("\\{Foreground\\}", pruned_tree$tip.label)
if (length(fg_tips) > 0) {
pruned_tree$node.label <- rep("", pruned_tree$Nnode)
ancestor_nodes <- pruned_tree$edge[pruned_tree$edge[, 2] %in% fg_tips, 1]
pruned_tree$node.label[ancestor_nodes - length(pruned_tree$tip.label)] <- "{Foreground}"
}
write.tree(pruned_tree, file = export_path)
message("✅ Tree saved to: ", export_path)
}
# ============================= #
# FASTA PRUNE FUNC #
# ============================= #
pruneFastaBySpecies <- function(fasta_path, retained_sp, export_path) {
message("🧬 Processing FASTA: ", basename(fasta_path))
fasta <- readDNAStringSet(fasta_path)
keep_idx <- vapply(names(fasta), function(x) {
sp_abbr <- strsplit(x, "_")[[1]][1]
sp_abbr %in% retained_sp
}, logical(1))
pruned_fasta <- fasta[keep_idx]
if (length(pruned_fasta) == 0) {
message("⚠️ No retained sequences in: ", basename(fasta_path))
return(NULL)
}
writeXStringSet(pruned_fasta, filepath = export_path)
message("✅ FASTA saved to: ", export_path)
}
# ============================= #
# MAIN LOOP #
# ============================= #
# Prune + label trees
tree_files <- dir_ls(tree_dir, regexp = "\\.txt$|\\.treefile$|\\.nwk$")
walk(tree_files, function(tree_file) {
og_name <- path_file(tree_file)
export_name <- file.path(tree_output_dir, paste0(output_prefix, "Labelled_", og_name))
multiTreeLabelAndPrune(tree_file, retained_species, foreground_species, export_name)
})
# Prune FASTA files
fasta_files <- dir_ls(fasta_input_dir, glob = "*.fasta")
walk(fasta_files, function(fa_file) {
out_fa <- file.path(fasta_output_dir, path_file(fa_file))
pruneFastaBySpecies(fa_file, retained_species, out_fa)
})
message("🎉 All trees and FASTA files processed and saved.")
5. Annotating proteins with InterProScan and Orthogroups with KinFin
As part of the process, we want to make sure that the genes under selection have meaningful biological interpretations through functional annotation and GO enrichment analysis. To achieve this, we will use InterProScan to annotate individual genes and KinFin to generate gene-level annotations, assigning functional categories to entire orthogroups. This approach aligns with the orthogroup-level focus of our analyses in aBSREL, BUSTED, and RELAX, providing insights into the functional relevance of selective pressures.
For that we run the following command:
# Example for Schistocerca only
./scripts/RunningInterProScan_modif ./scripts/inputurls_Schistocerca_Jan2025.txt /scratch/group/songlab/maeva/LocustsGenomeEvolution/Schistocerca_I2/5_OrthoFinder/fasta/
# Example for Polyneoptera
./scripts/RunningInterProScan_modif ./scripts/inputurls_13polyneoptera_Jan2025.txt /scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_I2/5_OrthoFinder/fasta/
# we replace the version of interproscan to the most recent: interproscan-5.72-103.0
# we also comment out ax.set_facecolor('white')' on lines 681 and 1754 of ./kinfin/src/kinfin.pyHere is the details of
./scripts/RunningInterProScan_modif
#!/bin/bash
## SLURM Job Specifications
#SBATCH --job-name=interproscan # Set the job name
#SBATCH --time=4-00:00:00 # Set the wall clock limit to 4 days
#SBATCH --ntasks=1 # Request 1 task
#SBATCH --cpus-per-task=12 # Request 12 CPUs for the task
#SBATCH --mem=100G # Request 100GB memory
#SBATCH --output=interproscan_%j.out # Standard output log
#SBATCH --error=interproscan_%j.err # Standard error log
# Ensure the script receives correct arguments
if [ "$#" -ne 2 ]; then
echo "Usage: $0 <input_file> <path_to_proteins_directory>"
exit 1
fi
input_file=$1
proteins_dir=$2
# Load necessary modules
ml Java/11.0.2
ml WebProxy
export http_proxy=http://10.73.132.63:8080
export https_proxy=http://10.73.132.63:8080
# Main working directories
interpro_dir="./11_InterProScan/interproscan-5.72-103.0"
output_dir="$interpro_dir/out"
backup_dir="./11_InterProScan/backup"
# Create necessary directories
mkdir -p "$output_dir"
mkdir -p "$backup_dir"
# Iterate through the input file to process each species
while read -r line; do
# Extract the species abbreviation
name=$(echo "$line" | awk -F',' '{print $4}')
protein_name="${name}_filteredTranscripts.fasta"
echo "Processing species: $name"
# Check if the protein file exists
protein_path="$proteins_dir/$protein_name"
if [ ! -f "$protein_path" ]; then
echo "Protein file $protein_name not found in $proteins_dir. Skipping."
continue
fi
# Check if the species has already been annotated
annotated_file="$output_dir/${protein_name}.tsv"
if [ -f "$annotated_file" ]; then
echo "$annotated_file exists; skipping $name."
continue
fi
# Backup original protein file and clean it
cp "$protein_path" "$backup_dir/${protein_name}.bak"
cp "$protein_path" "$interpro_dir/$protein_name"
sed -i'.original' -e "s|\*||g" "$interpro_dir/$protein_name"
rm "$interpro_dir/${protein_name}.original"
# Run InterProScan
echo "Running InterProScan for $protein_name..."
cd "$interpro_dir"
./interproscan.sh -i "$protein_name" -d out/ -t p --goterms -appl Pfam -f TSV
cd - > /dev/null
done < "$input_file"
# Combine all annotated results into a single file
cat "$output_dir"/*.tsv > "$interpro_dir/all_proteins.tsv"
echo "Annotation completed. Combined results stored in $interpro_dir/all_proteins.tsv."
# KinFin Preparation
kinfin_dir="./11_InterProScan/kinfin"
if [ ! -d "$kinfin_dir" ]; then
echo "KinFin not installed. Please install KinFin and rerun this step."
exit 1
fi
# Convert InterProScan results to KinFin-compatible format
echo "Preparing InterProScan results for KinFin..."
"$kinfin_dir/scripts/iprs2table.py" -i "$interpro_dir/all_proteins.tsv" --domain_sources Pfam
# Copy Orthofinder files to KinFin directory
cp 5_OrthoFinder/fasta/OrthoFinder/Results*/Orthogroups/Orthogroups.txt "$kinfin_dir/"
cp 5_OrthoFinder/fasta/OrthoFinder/Results*/WorkingDirectory/SequenceIDs.txt "$kinfin_dir/"
cp 5_OrthoFinder/fasta/OrthoFinder/Results*/WorkingDirectory/SpeciesIDs.txt "$kinfin_dir/"
# Create KinFin configuration file
echo '#IDX,TAXON' > "$kinfin_dir/config.txt"
sed 's/: /,/g' "$kinfin_dir/SpeciesIDs.txt" | cut -f 1 -d"." >> "$kinfin_dir/config.txt"
# Run KinFin functional annotation
echo "Running KinFin functional annotation..."
"$kinfin_dir/kinfin" --cluster_file "$kinfin_dir/Orthogroups.txt" \
--config_file "$kinfin_dir/config.txt" \
--sequence_ids_file "$kinfin_dir/SequenceIDs.txt" \
--functional_annotation functional_annotation.txt
echo "Functional annotation completed."6. dN/dS FitMG94
We will compute per gene and per branch the dN/dS score using HYPHY and the model FitMG94 as this will help us compute a mean dN/dS per branch. This analysis fits the Muse Gaut (+ GTR) model of codon substitution to an alignment and a tree and reports parameter estimates and trees scaled on expected number of synonymous and non synonymous substitutions per nucleotide site.
NB: Make sure to download the FitMG94.bf and place it in
the TemplateBatchFiles folder for HypHy otherwise it will
not work.
STEP 1. To run our analysis, we will use a loop for all orthogroups alignment and trees as follow:
#!/bin/bash
# Define paths
ALIGN_DIR="/Users/maevatecher/Desktop/Polyneoptera_FINAL/8_2_RemovedStops"
TREE_DIR="/Users/maevatecher/Desktop/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Resolved_Gene_Trees"
HYPHY="/Users/maevatecher/miniconda3/envs/hyphy_env/bin/hyphy"
TEMPLATE="/Users/maevatecher/miniconda3/envs/hyphy_env/share/hyphy/TemplateBatchFiles/FitMG94.bf"
# Loop through all cleaned CDS alignments
for aln in ${ALIGN_DIR}/cleaned_OG*_cds.fasta; do
# Get OG ID from filename
og=$(basename "$aln" _cds.fasta | sed 's/^cleaned_//')
# Define tree path and output file
tree="${TREE_DIR}/${og}_tree.txt"
out_json="${aln}.FITTER.json"
# Check if tree exists
if [[ -f "$tree" ]]; then
echo "Running FitMG94 for $og..."
$HYPHY "$TEMPLATE" \
--alignment "$aln" \
--tree "$tree" \
--type local \
--output "$out_json"
else
echo "Warning: Tree file not found for $og, skipping."
fi
done
STEP 2. We parse the dN/dS scores computed with HyPhy in the *json file as follow:
#!/bin/bash
# Output file
output_file="combined_dnds_table.csv"
# Write header
echo "Orthogroup,Branch,Species,dN/dS" > "$output_file"
# Loop through each JSON file
for json in 8_2_RemovedStops/*.FITTER.json; do
if [[ -f "$json" ]]; then
# Get orthogroup name
og=$(basename "$json" | sed 's/cleaned_//' | cut -d_ -f1)
# Use jq to extract Branch, Species, and dN/dS
jq -r --arg og "$og" '
.["branch attributes"]["0"]
| to_entries
| map([
$og,
.key,
(.key | capture("^(?<sp>[A-Z][a-z]{4})").sp // "internal"),
.value["Confidence Intervals"].MLE
])
| .[]
| @csv
' "$json" >> "$output_file"
fi
done
STEP 3. Plotting the dN/dS per Species
library(ggplot2)
library(dplyr)
Attaching package: 'dplyr'The following objects are masked from 'package:stats':
filter, lagThe following objects are masked from 'package:base':
intersect, setdiff, setequal, unionlibrary(readr)
# Load data
dnds_data <- read_csv("/Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data/HYPHY_selection/combined_dnds_table.csv") %>% rename(orthogroup = Orthogroup)Rows: 309886 Columns: 4── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
chr (3): Orthogroup, Branch, Species
dbl (1): dN/dS
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.relax_data <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults/joint_busted_relax_annotated.csv")Rows: 5339 Columns: 53
── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
chr (11): orthogroup, selection_status, file, input_file, selection_type, As...
dbl (39): pval_foreground, padj_foreground, pval_background, padj_background...
lgl (3): suspect_result.x, suspect_result.y, significant
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.# Load the list of orthogroups (one per line)
single_copy_OGs <- read_lines("/Users/maevatecher/Desktop/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Orthogroups/Orthogroups_SingleCopyOrthologues_selanalysiswide.txt")
# Extract PSG
psg_df <- relax_data %>%
select(orthogroup, selection_status, suspect_result.x) %>%
filter(suspect_result.x == "FALSE") %>%
mutate(PSG = selection_status %in% c("Foreground only", "Both foreground and background")) %>%
distinct()
# Merge and clean
dnds_data <- dnds_data %>%
left_join(psg_df, by = "orthogroup") %>%
mutate(
PSG = ifelse(is.na(PSG), FALSE, PSG),
Species = factor(Species, levels = c("Lmigr", "Sgreg", "Scanc", "Spice", "Samer", "Sscub", "Snite")),
`dN/dS` = as.numeric(`dN/dS`),
locust_group = ifelse(Species %in% c("Lmigr", "Sgreg", "Scanc", "Spice"), "Locust", "Other")
) %>%
filter(!is.na(`dN/dS`), !is.na(Species), `dN/dS` <= 1) %>%
filter(orthogroup %in% single_copy_OGs)
# PLOT
ggplot(dnds_data, aes(x = Species, y = `dN/dS`, fill = locust_group)) +
geom_violin(
aes(group = Species),
scale = "area",
trim = FALSE,
adjust = 1.2,
alpha = 0.7,
color = "black"
) +
# geom_jitter(
# aes(color = PSG),
# width = 0.15,
# size = 0.8,
# alpha = 0.5
# ) +
# Add red mean lines
stat_summary(
fun = mean,
geom = "crossbar",
width = 0.3,
color = "red",
fatten = 3
) +
facet_wrap(
~ PSG,
labeller = labeller(PSG = c(`TRUE` = "PSG Orthogroups", `FALSE` = "Non-PSG Orthogroups"))
) +
scale_fill_manual(values = c("Locust" = "gray60", "Other" = "white")) +
scale_color_manual(values = c("TRUE" = "#D95F02", "FALSE" = "gray80")) +
coord_cartesian(ylim = c(0, 1)) +
labs(
title = "Raincloud Violin Plot of dN/dS per Species",
subtitle = "Faceted by PSG Status; Locusts shaded in gray; Red line = mean",
x = "Species",
y = "dN/dS"
) +
theme_minimal(base_size = 14) +
theme(
strip.text = element_text(face = "bold"),
axis.text.x = element_text(angle = 45, hjust = 1)
)Warning: No shared levels found between `names(values)` of the manual scale and the
data's colour values.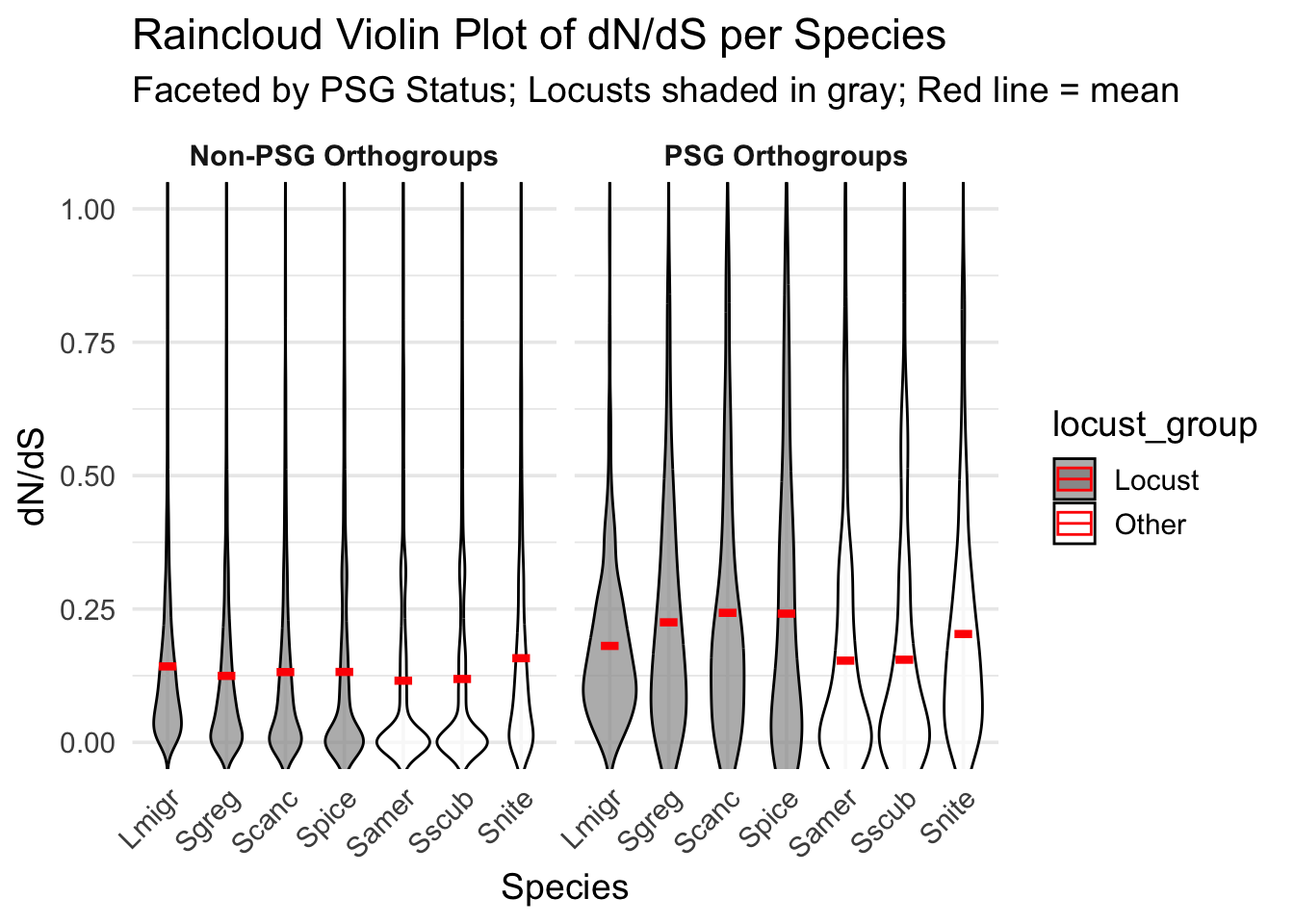
ggplot(dnds_data, aes(x = Species, y = `dN/dS`, fill = PSG)) +
geom_violin(
aes(group = interaction(Species, PSG)),
position = position_dodge(width = 0.6), # smaller = less gap
scale = "width",
width = 0.7, # narrower violins
trim = FALSE,
adjust = 1.2,
alpha = 0.7,
color = "black"
) +
stat_summary(
aes(group = PSG),
fun = mean,
geom = "crossbar",
width = 0.2,
color = "red",
fatten = 3,
position = position_dodge(width = 0.6)
) +
scale_fill_manual(values = c(`TRUE` = "#D95F02", `FALSE` = "gray70"),
labels = c(`TRUE` = "PSG", `FALSE` = "Non-PSG")) +
coord_cartesian(ylim = c(0, 1)) +
labs(
title = "dN/dS per Species",
subtitle = "PSGs vs Non-PSGs side by side",
x = "Species",
y = "dN/dS",
fill = "Category"
) +
theme_minimal(base_size = 14) +
theme(
axis.text.x = element_text(angle = 45, hjust = 1)
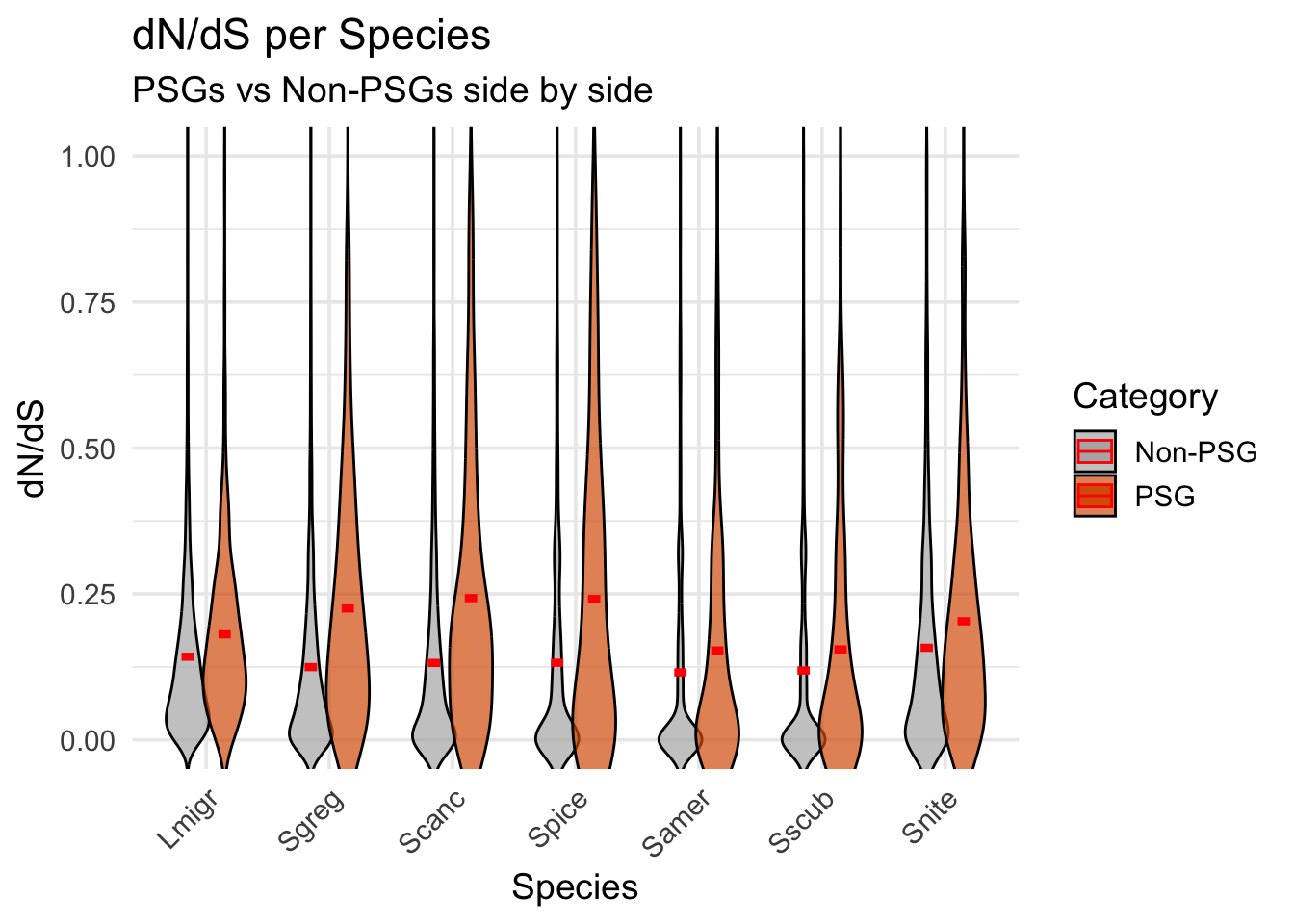
)
7. aBSREL
Running the analysis
We will perform aBSREL analysis using both unlabeled and labelled phylogeny.
# For Polyneoptera
# For unlabelled phylogeny
sbatch ./scripts/RunaBSREL_May2025.sh \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Resolved_Gene_Trees/ \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Orthogroups/Orthogroups_SingleCopyOrthologues_selanalysiswide.txt
# For labelled phylogeny
sbatch ./scripts/RunaBSREL_labeled_May2025.sh \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/9_1_LabelledPhylogenies/Locusts \
Locusts \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Orthogroups/Orthogroups_SingleCopyOrthologues_selanalysiswide.txt
# For labelled phylogeny PRUNED
sbatch ./scripts/RunaBSREL_labeled_June2025.sh \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/9_1_LabelledPhylogenies_Pruned/Locusts \
Locusts \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Orthogroups/Orthogroups_SingleCopyOrthologues_selanalysiswide.txt
Unlabeled
Parsing the json files
For parsing the results, you just do:
srun --ntasks 1 --cpus-per-task 16 --mem 50G --time 05:00:00 --pty bash
ml GCC/13.2.0 OpenMPI/4.1.6 R_tamu/4.4.1
export R_LIBS=$SCRATCH/R_LIBS_USER/
Rscript ./scripts/Parsing_aBSRELresulsr_unlabel_June2025.RBelow is the detail of the parsing aBSREL script
./scripts/Parsing_aBSRELresulsr_unlabel_June2025.R:
# ===========================
# A. LOAD LIBRARIES
# ===========================
library(jsonlite)
library(dplyr)
Attaching package: 'dplyr'The following objects are masked from 'package:stats':
filter, lagThe following objects are masked from 'package:base':
intersect, setdiff, setequal, unionlibrary(stringr)
library(readr)
library(tibble)
# ===========================
# B. DEFINE PATHS
# ===========================
input_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/9_2_ABSRELResults_unlabeled/"
output_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedABSRELResults_unlabeled/"
orthotable_path <- "/Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data/orthofinder/Polyneoptera/Results_I2_iqtree/Orthogroups/Orthogroups_CladeAssignment_WithCopyStatus_cleaned.csv"
# ===========================
# C. INITIALIZE
# ===========================
files <- list.files(path = input_dir, pattern = "\\.json$", full.names = TRUE)
file_all <- file.path(output_dir, "parsed_absrel_full.csv")
file_sig <- file.path(output_dir, "parsed_absrel_significant_full.csv")
all_results <- data.frame()
sig_results <- data.frame()
# ===========================
# D. PARSE JSON FILES
# ===========================
for (file in files) {
try({
json <- fromJSON(file)
orthogroup <- str_extract(basename(file), "OG[0-9]+")
# Check required fields exist
if (!"branch attributes" %in% names(json)) next
if (!"0" %in% names(json$`branch attributes`)) next
branches <- json$`branch attributes`$`0`
# Pull alignment stats once per file
aln_length <- json$input$`number of sites`
seq_count <- json$input$`number of sequences`
for (branch_name in names(branches)) {
entry <- branches[[branch_name]]
rates <- entry$`Rate Distributions`
n <- if (!is.null(rates)) nrow(as.data.frame(rates)) else 0
row <- data.frame(
Orthogroup = orthogroup,
Branch = branch_name,
Species = substr(branch_name, 1, 5),
Baseline_MG94xREV = entry$`Baseline MG94xREV`,
Baseline_omega = entry$`Baseline MG94xREV omega ratio`,
Full_adaptive_model = entry$`Full adaptive model`,
Full_adaptive_model_nonsyn = entry$`Full adaptive model (non-synonymous subs/site)`,
Full_adaptive_model_syn = entry$`Full adaptive model (synonymous subs/site)`,
LRT = entry$`LRT`,
Nucleotide_GTR = entry$`Nucleotide GTR`,
Rate_classes = entry$`Rate classes`,
Uncorrected_P = entry$`Uncorrected P-value`,
Corrected_P = entry$`Corrected P-value`,
aln_length = aln_length,
seq_count = seq_count,
Omega1 = NA, Percent1 = NA,
Omega2 = NA, Percent2 = NA,
Omega3 = NA, Percent3 = NA,
stringsAsFactors = FALSE
)
# Add omega/proportion values
if (!is.null(rates)) {
df <- as.data.frame(rates)
for (i in 1:min(n, 3)) {
row[[paste0("Omega", i)]] <- df[i, 1]
row[[paste0("Percent", i)]] <- df[i, 2]
}
}
all_results <- bind_rows(all_results, row)
if (!is.null(row$Corrected_P) && !is.na(row$Corrected_P) && row$Corrected_P <= 0.05) {
sig_results <- bind_rows(sig_results, row)
}
}
}, silent = TRUE)
}
# ===========================
# E. FLAG SUSPECT BRANCHES
# ===========================
all_results <- all_results %>%
mutate(
Baseline_omega = as.numeric(Baseline_omega),
aln_length = as.numeric(aln_length),
seq_count = as.numeric(seq_count),
Suspect_Result = is.na(Baseline_omega) | Baseline_omega > 100 | aln_length < 100 | seq_count < 4
)
sig_results <- sig_results %>%
mutate(
Baseline_omega = as.numeric(Baseline_omega),
aln_length = as.numeric(aln_length),
seq_count = as.numeric(seq_count),
Suspect_Result = is.na(Baseline_omega) | Baseline_omega > 100 | aln_length < 100 | seq_count < 4
)
# ===========================
# F. WRITE OUTPUT
# ===========================
if (!dir.exists(output_dir)) dir.create(output_dir, recursive = TRUE)
write_csv(all_results, file_all)
write_csv(sig_results, file_sig)
cat("✅ Full parsing complete.\n")✅ Full parsing complete.# ===========================
# G. CREATE SUMMARY TABLE
# ===========================
createSummaryTable <- function(results_df) {
results_df %>%
mutate(across(starts_with("Omega"), as.numeric),
Corrected_P = as.numeric(Corrected_P),
Significant = Corrected_P <= 0.05) %>%
rowwise() %>%
mutate(
Mean_omega = mean(c_across(starts_with("Omega")), na.rm = TRUE),
Max_omega = max(c_across(starts_with("Omega")), na.rm = TRUE)
) %>%
ungroup() %>%
group_by(Orthogroup) %>%
summarise(
Total_Branches = n(),
Significant_Branches = sum(Significant),
Proportion_Significant = Significant_Branches / Total_Branches,
Suspect_Branches = sum(Suspect_Result),
Positive_Species = paste0(Species[Significant], collapse = ";"),
Mean_omega = mean(Mean_omega, na.rm = TRUE),
Max_omega = max(Max_omega, na.rm = TRUE),
.groups = "drop"
)
}
summary_table <- createSummaryTable(all_results)
write_csv(summary_table, file.path(output_dir, "absrel_summary_table.csv"))
# ===========================
# H. ANNOTATE WITH ORTHO INFO
# ===========================
orthologtable <- read_csv(orthotable_path)Rows: 20027 Columns: 21── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
chr (7): Orthogroup, Assigned_Clade, Orthogroup_Type, SelAnalysis, SelAnaly...
dbl (14): Brsri, Csecu, Pamer, Asimp, Gbima, Glong, Lmigr, Samer, Scanc, Sgr...
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.absrel_df <- left_join(all_results, orthologtable, by = "Orthogroup") %>%
mutate(
Selection = as.numeric(Corrected_P) <= 0.05,
Category = case_when(
SelAnalysisStrict == "Included" ~ "1:1 Polyneoptera",
SelAnalysisLocusts == "Included" ~ "1:1 Caelifera only",
SelAnalysisMixed == "Included" ~ "1:1 Mixed",
TRUE ~ "Other"
)
) %>%
relocate(Orthogroup, Branch, Species, Category, Selection, Corrected_P, starts_with("Omega"))
write_csv(absrel_df, file.path(output_dir, "parsed_absrel_annotated.csv"))
# ===========================
# I. SUMMARY PER SPECIES × CATEGORY
# ===========================
absrel_species_summary <- absrel_df %>%
group_by(Species, Category) %>%
summarise(
Significant_Branches = sum(Selection),
NonSignificant_Branches = sum(!Selection),
Suspect_Branches = sum(Suspect_Result),
Total_Branches = n(),
.groups = "drop"
) %>%
arrange(Species, Category)
knitr::kable(absrel_species_summary, caption = "Summary of aBSREL results: per species and category")| Species | Category | Significant_Branches | NonSignificant_Branches | Suspect_Branches | Total_Branches |
|---|---|---|---|---|---|
| Asimp | 1:1 Mixed | 31 | 1255 | 45 | 1286 |
| Asimp | 1:1 Polyneoptera | 54 | 3038 | 63 | 3092 |
| Brsri | 1:1 Mixed | 108 | 956 | 13 | 1064 |
| Brsri | 1:1 Polyneoptera | 147 | 2945 | 13 | 3092 |
| Csecu | 1:1 Mixed | 25 | 1177 | 67 | 1202 |
| Csecu | 1:1 Polyneoptera | 67 | 3025 | 134 | 3092 |
| Gbima | 1:1 Mixed | 131 | 584 | 66 | 715 |
| Gbima | 1:1 Polyneoptera | 724 | 2368 | 434 | 3092 |
| Glong | 1:1 Mixed | 62 | 670 | 71 | 732 |
| Glong | 1:1 Polyneoptera | 167 | 2925 | 526 | 3092 |
| Lmigr | 1:1 Caelifera only | 94 | 667 | 0 | 761 |
| Lmigr | 1:1 Mixed | 120 | 1370 | 173 | 1490 |
| Lmigr | 1:1 Polyneoptera | 125 | 2967 | 414 | 3092 |
| Pamer | 1:1 Mixed | 33 | 1192 | 65 | 1225 |
| Pamer | 1:1 Polyneoptera | 82 | 3010 | 114 | 3092 |
| Samer | 1:1 Caelifera only | 51 | 710 | 78 | 761 |
| Samer | 1:1 Mixed | 100 | 1390 | 115 | 1490 |
| Samer | 1:1 Polyneoptera | 86 | 3006 | 191 | 3092 |
| Scanc | 1:1 Caelifera only | 63 | 698 | 19 | 761 |
| Scanc | 1:1 Mixed | 111 | 1379 | 29 | 1490 |
| Scanc | 1:1 Polyneoptera | 107 | 2985 | 40 | 3092 |
| Sgreg | 1:1 Caelifera only | 51 | 710 | 5 | 761 |
| Sgreg | 1:1 Mixed | 70 | 1420 | 17 | 1490 |
| Sgreg | 1:1 Polyneoptera | 108 | 2984 | 26 | 3092 |
| Snite | 1:1 Caelifera only | 44 | 717 | 15 | 761 |
| Snite | 1:1 Mixed | 130 | 1360 | 23 | 1490 |
| Snite | 1:1 Polyneoptera | 114 | 2978 | 69 | 3092 |
| Spice | 1:1 Caelifera only | 73 | 688 | 30 | 761 |
| Spice | 1:1 Mixed | 122 | 1368 | 73 | 1490 |
| Spice | 1:1 Polyneoptera | 156 | 2936 | 122 | 3092 |
| Sscub | 1:1 Caelifera only | 41 | 720 | 52 | 761 |
| Sscub | 1:1 Mixed | 93 | 1397 | 105 | 1490 |
| Sscub | 1:1 Polyneoptera | 80 | 3012 | 181 | 3092 |
| n10 | 1:1 Mixed | 46 | 652 | 201 | 698 |
| n10 | 1:1 Polyneoptera | 145 | 2511 | 968 | 2656 |
| n11 | 1:1 Polyneoptera | 115 | 2603 | 176 | 2718 |
| n2 | 1:1 Caelifera only | 35 | 669 | 200 | 704 |
| n2 | 1:1 Mixed | 71 | 1401 | 216 | 1472 |
| n2 | 1:1 Polyneoptera | 171 | 2910 | 268 | 3081 |
| n3 | 1:1 Caelifera only | 23 | 645 | 136 | 668 |
| n3 | 1:1 Mixed | 72 | 1383 | 377 | 1455 |
| n3 | 1:1 Polyneoptera | 114 | 2942 | 1126 | 3056 |
| n4 | 1:1 Caelifera only | 31 | 610 | 125 | 641 |
| n4 | 1:1 Mixed | 114 | 1310 | 242 | 1424 |
| n4 | 1:1 Polyneoptera | 167 | 2890 | 395 | 3057 |
| n5 | 1:1 Caelifera only | 43 | 578 | 138 | 621 |
| n5 | 1:1 Mixed | 126 | 1207 | 203 | 1333 |
| n5 | 1:1 Polyneoptera | 111 | 2831 | 547 | 2942 |
| n6 | 1:1 Mixed | 79 | 1132 | 233 | 1211 |
| n6 | 1:1 Polyneoptera | 113 | 2492 | 322 | 2605 |
| n7 | 1:1 Mixed | 68 | 979 | 157 | 1047 |
| n7 | 1:1 Polyneoptera | 79 | 2204 | 382 | 2283 |
| n8 | 1:1 Mixed | 54 | 851 | 174 | 905 |
| n8 | 1:1 Polyneoptera | 74 | 2059 | 364 | 2133 |
| n9 | 1:1 Mixed | 77 | 777 | 235 | 854 |
| n9 | 1:1 Polyneoptera | 82 | 1893 | 446 | 1975 |
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Plotting the results
library("ggplot2")
# Load annotated aBSREL results
absrel_annotated_df <- read_csv(
"/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedABSRELResults_unlabeled/parsed_absrel_annotated.csv"
)
# Ensure Selection is logical and Category is not NA
absrel_clean <- absrel_annotated_df %>%
filter(!is.na(Category)) %>%
mutate(
Selection = as.logical(Selection),
Category = factor(Category, levels = c("1:1 Polyneoptera", "1:1 Caelifera only", "1:1 Mixed")),
Species = factor(Species)
)
# Create summary count table per species × category × selection status
absrel_summary_counts <- absrel_clean %>%
group_by(Species, Category, Selection) %>%
summarise(N = n(), .groups = "drop")
# Set factor levels for consistent fill order
absrel_summary_counts$Selection <- factor(
absrel_summary_counts$Selection,
levels = c(FALSE, TRUE),
labels = c("Not Selected", "Selected")
)
# Reorder Species factor
absrel_summary_counts <- absrel_summary_counts %>%
filter(!Species %in% paste0("n", 1:11)) %>% # Clean internal nodes in one line
mutate(Species = factor(Species, levels = c("Pamer", "Csecu", "Brsri",
"Glong","Gbima", "Asimp", "Sscub","Samer", "Spice", "Scanc", "Snite", "Sgreg",
"Lmigr"
)))
# Plot
ggplot(absrel_summary_counts, aes(x = Species, y = N, fill = Selection)) +
geom_bar(stat = "identity", position = "stack") +
coord_flip() + # Flip to horizontal
facet_wrap(~ Category, ncol = 1, scales = "free_y") +
scale_fill_manual(values = c("Not Selected" = "#d9d9d9", "Selected" = "#1b9e77")) +
labs(
title = "aBSREL Selection Summary per Species and Orthogroup Category",
x = "Species",
y = "Number of Branches",
fill = "Selection Status"
) +
theme_minimal(base_size = 14) +
theme(
axis.text.y = element_text(face = "italic"),
strip.text = element_text(face = "bold")
)
category_colors <- c(
"1:1 Polyneoptera" = "gold",
"1:1 Mixed" = "black",
"1:1 Caelifera only" = "green3",
"Not Selected" = "gray90"
)
# Get Not Selected totals
not_selected <- absrel_summary_counts %>%
filter(Selection == "Not Selected") %>%
group_by(Species) %>%
summarise(N = sum(N), Category = "Not Selected", .groups = "drop")
# Bind with selected data
combined_bar_data <- bind_rows(
absrel_summary_counts %>% filter(Selection == "Selected"),
not_selected
)
ggplot(combined_bar_data,
aes(x = Species, y = N, fill = Category)) +
geom_bar(stat = "identity", position = "stack") +
coord_flip() +
scale_fill_manual(values = category_colors) +
labs(
title = "Selected Branches per Species",
subtitle = "Colored by Orthogroup Category",
x = "Species",
y = "Number of Selected Branches",
fill = "Orthogroup Category"
) +
theme_minimal(base_size = 14) +
theme(
axis.text.y = element_text(face = "italic"),
strip.text = element_text(face = "bold")
)
ggplot(absrel_summary_counts %>% filter(Selection == "Selected"),
aes(x = Species, y = N, fill = Category)) +
geom_bar(stat = "identity", position = "stack") +
coord_flip() +
scale_fill_manual(values = category_colors) +
labs(
title = "Selected Branches per Species",
subtitle = "Colored by Orthogroup Category",
x = "Species",
y = "Number of Selected Branches",
fill = "Orthogroup Category"
) +
theme_minimal(base_size = 14) +
theme(
axis.text.y = element_text(face = "italic"),
strip.text = element_text(face = "bold")
)
### tree
library(pheatmap)
library(tidyverse)
# Filter significant only
significant_mat <- absrel_annotated_df %>%
filter(`Corrected_P` <= 0.05) %>%
mutate(Significant = 1) %>%
distinct(Orthogroup, Species, Significant) %>%
pivot_wider(names_from = Orthogroup, values_from = Significant, values_fill = 0) %>%
column_to_rownames("Species") %>%
as.matrix()
# Save heatmap
pdf(file.path(output_dir, "heatmap_significant_orthogroups.pdf"), width = 9, height = 6)
pheatmap(significant_mat,
cluster_rows = TRUE,
cluster_cols = TRUE,
color = c("white", "darkred"),
main = "aBSREL: Positive Selection Heatmap")
dev.off()
library(tidyverse)
library(igraph)
# Define helper function for pairwise combinations
pairwise_combinations <- function(df, col) {
col <- rlang::ensym(col)
df %>%
group_by(!!col) %>%
filter(n() > 1) %>%
summarise(pairs = list(t(combn(unique(.[[deparse(col)]]), 2))), .groups = "drop") %>%
unnest_wider(pairs, names_sep = "_") %>%
rename(from = pairs_1, to = pairs_2)
}
# Build edge list from significant orthogroups with more than 1 species
#edges <- all_results %>%
# filter(Corrected_P <= 0.05) %>%
# distinct(Species, Orthogroup) %>%
# pairwise_combinations(Orthogroup)
# Create graph object
#g <- graph_from_data_frame(edges, directed = FALSE)
# Optional: plot it
#pdf(file.path(output_dir, "network_positive_selection_species.pdf"), width = 8, height = 8)
#plot(
# g,
# vertex.size = 30,
# vertex.label.cex = 0.9,
# vertex.label.color = "black",
# vertex.color = "skyblue",
# edge.width = ,
# main = "Network of Species Co-selected in aBSREL Orthogroups"
#)
#dev.off()
# ===========================
# Load Libraries
# ===========================
library(ape)
library(viridis)
library(tidyverse)
# ===========================
# File Paths
# ===========================
input_results <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedABSRELResults_unlabeled/parsed_absrel_full.csv"
tree_file <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Species_Tree/SpeciesTree_rooted_node_labels.txt"
output_file <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedABSRELResults_unlabeled/tree_colored_by_omega3_allbranches_FINAL.pdf"
trusted_orthogroups <- readLines("/Users/maevatecher/Desktop/Polyneoptera_FINAL/trusted_ogs_v2.txt")
# ===========================
# Load Data
# ===========================
all_results <- read_csv(input_results, show_col_types = FALSE)
filtered_results <- all_results %>%
filter(Suspect_Result == "FALSE") %>%
filter(Orthogroup %in% trusted_orthogroups)
# filter(Orthogroup %in% trusted_orthogroups & !is.na(Baseline_omega))
tree <- read.tree(tree_file)
desired_order <- c(
"Pamer_filteredTranscripts", "Csecu_filteredTranscripts",
"Sgreg_filteredTranscripts", "Snite_filteredTranscripts", "Scanc_filteredTranscripts",
"Spice_filteredTranscripts", "Sscub_filteredTranscripts", "Samer_filteredTranscripts",
"Lmigr_filteredTranscripts", "Asimp_filteredTranscripts", "Glong_filteredTranscripts",
"Gbima_filteredTranscripts", "Brsri_filteredTranscripts"
)
# Ensure all desired tips are in the tree
stopifnot(all(desired_order %in% tree$tip.label))
# Ladderize and reorder the tip labels by desired order
tree <- ladderize(tree, right = FALSE)
# Reorder tree$tip.label visually via `plot.phylo()` call
tip_order <- match(tree$tip.label, desired_order)
# ===========================
# Harmonize Labels
# ===========================
# Create node label lookup: node index → cleaned label
node_labels <- c(tree$tip.label, tree$node.label)
names(node_labels) <- 1:(length(tree$tip.label) + tree$Nnode)
# Remove "_filteredTranscripts..." and lowercase
node_to_label <- tolower(gsub("_filteredTranscripts.*", "", node_labels))
names(node_to_label) <- names(node_labels)
# Clean Branch names from all_results
omega_df <- filtered_results %>%
filter(!is.na(Omega1)) %>%
mutate(label = tolower(gsub("_filteredTranscripts.*", "", Branch))) %>%
group_by(label) %>%
summarize(mean_omega = mean(as.numeric(Omega3), na.rm = TRUE)) %>%
ungroup()
omega_df <- omega_df %>% filter(label != "n2") %>%
filter(label != "n3") %>%
filter(label != "n4") %>%
filter(label != "n5") %>%
filter(label != "n6") %>%
filter(label != "n7") %>%
filter(label != "n8") %>%
filter(label != "n9") %>%
filter(label != "n10") %>%
filter(label != "n11") %>%
filter(label != "asimp") %>%
filter(label != "brsri") %>%
filter(label != "csecu") %>%
filter(label != "gbima") %>%
filter(label != "glong") %>%
filter(label != "pamer")
# ===========================
# Map omega3 to Tree Branches
# ===========================
omega_vals <- rep(NA, nrow(tree$edge))
for (i in seq_len(nrow(tree$edge))) {
child_node <- tree$edge[i, 2]
label <- node_to_label[as.character(child_node)]
if (!is.na(label) && label %in% omega_df$label) {
omega_vals[i] <- omega_df$mean_omega[omega_df$label == label]
}
}
# ===========================
# Generate Colors
# ===========================
color_scale <- viridis(100)
if (all(is.na(omega_vals))) {
warning("No omega values matched any tree node labels.")
edge_colors <- rep("grey", length(omega_vals))
} else {
omega_vals <- as.numeric(omega_vals)
cut_omega <- cut(omega_vals, breaks = 100)
edge_colors <- color_scale[as.numeric(cut_omega)]
edge_colors[is.na(edge_colors)] <- "grey"
}
# ===========================
# Plot Tree with Edge Colors
# ===========================
pdf(output_file, width = 9, height = 7)
par(mar = c(5, 4, 4, 6)) # leave space for legend
plot(tree,
edge.color = edge_colors,
edge.width = 4,
cex = 1,
main = "Mean Omega3 per Branch (Tips + Internal)",
show.tip.label = TRUE,
use.edge.length = FALSE,
tip.order = tip_order)
# Continuous legend (manual)
zlim_vals <- range(omega_vals, na.rm = TRUE)
legend_vals <- pretty(zlim_vals, n = 5)
legend_colors <- color_scale[as.numeric(cut(legend_vals, breaks = 100))]
legend("topright",
legend = round(legend_vals, 2),
fill = legend_colors,
border = NA,
title = "omega")
dev.off()
message("✅ Tree plot saved to: ", output_file)
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Annotating OGs under selection
Here we are working with Orthogroups number under selection per branch, and we want to have a functional enrichment of the genes associated per species to allow comparison. First we need to extract the positively selected genes/orthogroups for each species:
library(dplyr)
library(readr)
# Load aBSREL results
absrel_df <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedABSRELResults_unlabeled/parsed_absrel_annotated.csv")
# Filter PSGs: significant, non-suspect
psg_ogs <- absrel_df %>%
filter(Selection == TRUE, Suspect_Result == FALSE) %>%
dplyr::select(Species, Orthogroup)
ortho_dir <- "/Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data/orthofinder/Polyneoptera"
input_file <- file.path(ortho_dir, "Results_I2_iqtree/Orthogroups_genesproteinbiotype_13species_annotated_May2025.csv")
ortho_map <- read.csv(input_file, header = TRUE, stringsAsFactors = FALSE)
head(ortho_map) Orthogroup SpeciesID protein_id GeneID
1 OG0000000 Asimp_filteredTranscripts XP_067003642.2 LOC136874043
2 OG0000000 Asimp_filteredTranscripts XP_067004661.1 LOC136874869
3 OG0000000 Asimp_filteredTranscripts XP_067015293.1 LOC136886419
4 OG0000000 Asimp_filteredTranscripts XP_067015651.2 LOC136886746
5 OG0000000 Asimp_filteredTranscripts XP_068085770.1 LOC137496902
6 OG0000000 Asimp_filteredTranscripts XP_068087037.1 LOC137503369
Description Species GeneType
1 farnesol dehydrogenase isoform X1 Asimp protein-coding
2 dehydrogenase/reductase SDR family member 11 Asimp protein-coding
3 farnesol dehydrogenase Asimp protein-coding
4 farnesol dehydrogenase Asimp protein-coding
5 farnesol dehydrogenase-like Asimp protein-coding
6 farnesol dehydrogenase-like Asimp protein-coding
Accession Begin End Orthogroup_Type
1 NC_090269.1 316521124 316596183 MultiCopy
2 NC_090269.1 316408775 316497937 MultiCopy
3 NC_090279.1 165240845 165292312 MultiCopy
4 NC_090279.1 164532314 164617967 MultiCopy
5 NC_090275.1 60074551 60101839 MultiCopy
6 NC_090279.1 164618824 164719067 MultiCopy# Join PSG orthogroups with gene IDs, species-aware
psg_genes <- psg_ogs %>%
inner_join(ortho_map, by = c("Orthogroup","Species")) %>%
dplyr::select(Species, Orthogroup, GeneID)Then we use the same function we used for DEGs GO terms and KEGG enrichment:
# === Paths and Constants ===
workDir <- "/Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data"
GODir <- file.path(workDir, "list", "GO_Annotations")
RefDir <- file.path(workDir, "RefSeq")
enrichDir <- file.path(workDir, "HYPHY_selection/functional_pathways/aBSREL")
selListDir <- file.path(workDir, "HYPHY_selection/ParsedABSRELResults_unlabeled")
species_list <- c("gregaria", "cancellata", "piceifrons", "americana", "cubense", "nitens")
# === Load Required Libraries ===
library(data.table)
library(dplyr)
library(readr)
library(clusterProfiler)
library(GO.db)
library(rtracklayer)
library(DesertLocustR) # Local installation
gff_map <- c(
gregaria = "GCF_023897955.1_iqSchGreg1.2_genomic.gff",
cancellata = "GCF_023864275.1_iqSchCanc2.1_genomic.gff",
piceifrons = "GCF_021461385.2_iqSchPice1.1_genomic.gff",
americana = "GCF_021461395.2_iqSchAmer2.1_genomic.gff",
cubense = "GCF_023864345.2_iqSchSeri2.2_genomic.gff",
nitens = "GCF_023898315.1_iqSchNite1.1_genomic.gff"
)
annot_map <- c(
gregaria = "EggNog_Arthropoda_one2one.emapper.annotations",
cancellata = "GCF_023864275.1_iqSchCanc2.1_Arthopoda_one2one.emapper.annotations",
piceifrons = "GCF_021461385.2_iqSchPice1.1_Arthopoda_one2one.emapper.annotations",
americana = "GCF_021461395.2_iqSchAmer2.1_Arthopoda_one2one.emapper.annotations",
cubense = "GCF_023864345.2_iqSchSeri2.2_Arthopoda_one2one.emapper.annotations",
nitens = "GCF_023898315.1_iqSchNite1.1_Arthopoda_one2one.emapper.annotations"
)
# GO enrichment
enrich_GO <- function(dge_genes.df, term2gene, term2name, pval, qval){
genes <- rownames(dge_genes.df)
enricher(genes, TERM2GENE = term2gene, TERM2NAME = term2name, pvalueCutoff = pval,
pAdjustMethod = "BH", qvalueCutoff = qval)
}
# KEGG preparation
assign_kegg_ids <- function(sig_genes.df){
if (is.vector(sig_genes.df)) {
sig_genes.df <- data.frame(X.query = sig_genes.df, stringsAsFactors = FALSE)
} else {
sig_genes.df$X.query <- rownames(sig_genes.df)
}
dge_with_kegg_ids <- left_join(sig_genes.df, kegg_final, by = "X.query")
dge_with_kegg_ids$KEGG_ko[grepl("^K", dge_with_kegg_ids$KEGG_ko)]
}
# KEGG enrichment
enrich_KEGG <- function(dge_genes.df, pval_cutoff = 0.05, qval_cutoff = 0.05) {
gene_with_kegg_ids <- assign_kegg_ids(dge_genes.df)
enrichKEGG(
gene = gene_with_kegg_ids,
organism = "ko",
pvalueCutoff = pval_cutoff,
qvalueCutoff = qval_cutoff,
pAdjustMethod = "BH"
)
}
run_GO_enrichment_selected <- function(
gene_list,
go_table,
term2name,
species,
suffix,
ontology,
output_dir,
show_n = 30,
top_n = 30
) {
if (length(gene_list) == 0) return(NULL)
if (!dir.exists(output_dir)) {
dir.create(output_dir, recursive = TRUE)
}
# Make sure column names are correct for clusterProfiler::enricher()
go_table_fixed <- go_table[, 1:2]
colnames(go_table_fixed) <- c("go_id", "gene_id")
term2name_fixed <- term2name[, 1:2]
colnames(term2name_fixed) <- c("go_id", "name")
# Run enrichment
go_result <- enricher(
gene = gene_list,
TERM2GENE = go_table_fixed,
TERM2NAME = term2name_fixed,
pvalueCutoff = 0.05,
qvalueCutoff = 0.05
)
if (!is.null(go_result) &&
inherits(go_result, "enrichResult") &&
nrow(go_result@result) > 0 &&
sum(!is.na(go_result@result$Description)) > 0) {
# Save dotplot
try({
pdf(file = file.path(output_dir, paste0("GO_", ontology, "_dotplot_", species, "_", suffix, ".pdf")),
width = 8, height = 6)
print(dotplot(go_result, showCategory = min(show_n, nrow(go_result@result))) +
ggtitle(paste(ontology, suffix)))
dev.off()
}, silent = TRUE)
# Export top terms with log10(p)
species_enrich_ready <- go_result@result[, c("ID", "p.adjust")]
species_enrich_ready$logp <- -log10(species_enrich_ready$p.adjust)
species_enrich_ready <- species_enrich_ready[order(-species_enrich_ready$logp), ]
species_enrich_ready <- head(species_enrich_ready, n = top_n)[, c("ID", "logp")]
write.table(species_enrich_ready,
file = file.path(output_dir, paste0("enrich_", ontology, "_GOs_", species, "_", suffix, ".txt")),
sep = "\t", quote = FALSE, row.names = FALSE, col.names = FALSE)
# Also export the full table if needed
write.csv(go_result@result,
file = file.path(output_dir, paste0("GO_enrichment_full_", ontology, "_", species, "_", suffix, ".csv")),
row.names = FALSE)
} else {
message(paste0("⚠️ No GO enrichment result to plot/export for ", species, " - ", suffix))
}
}
run_KEGG_enrichment_selected <- function(gene_list, species, suffix, output_dir,
show_n = 40, top_n = 40) {
if (length(gene_list) == 0) return(NULL)
kegg_result <- enrich_KEGG(gene_list, pval_cutoff = 0.05, qval_cutoff = 0.05)
if (!is.null(kegg_result) && inherits(kegg_result, "enrichResult") &&
nrow(kegg_result@result) > 0) {
try({
pdf(file = file.path(output_dir, paste0("KEGG_dotplot_", species, "_", suffix, ".pdf")),
width = 8, height = 6)
print(dotplot(kegg_result, showCategory = min(show_n, nrow(kegg_result@result))) +
ggtitle(paste("KEGG", suffix)))
dev.off()
}, silent = TRUE)
# Full result
write.csv(kegg_result@result,
file = file.path(output_dir, paste0("KEGG_enrichment_", species, "_", suffix, ".csv")),
row.names = FALSE)
# Top KEGG terms
species_enrich_kegg <- kegg_result@result[, c("ID", "p.adjust")]
species_enrich_kegg$logp <- -log10(species_enrich_kegg$p.adjust)
species_enrich_kegg <- species_enrich_kegg[order(-species_enrich_kegg$logp), ][1:min(nrow(species_enrich_kegg), top_n), ]
species_enrich_kegg <- species_enrich_kegg[, c("ID", "logp")]
write.table(species_enrich_kegg,
file = file.path(output_dir, paste0("enrich_KEGG_", species, "_", suffix, ".txt")),
sep = "\t", quote = FALSE, row.names = FALSE, col.names = FALSE)
} else {
message(paste("⚠️ No KEGG enrichment result to plot/export for", species, "-", suffix))
}
}
GO_terms_list <- list()
ontologies_list <- list()
term2name_list <- list()
kegg_final_list <- list()
# Mapping external species names to internal codes in ortho_map
species_translate <- c(
gregaria = "Sgreg",
cancellata = "Scanc",
piceifrons = "Spice",
americana = "Samer", # double-check this is correct
cubense = "Sscub",
nitens = "Snite"
)
for (sp in species_list) {
message("Preparing annotations for ", sp)
sp_code <- species_translate[sp]
eggnog_path <- file.path(GODir, annot_map[[sp]])
gff_path <- file.path(RefDir, gff_map[[sp]])
output_dir <- file.path(enrichDir, sp)
dir.create(output_dir, recursive = TRUE, showWarnings = FALSE)
eggnog_annots <- read.delim(eggnog_path, sep = "\t", skip = 4)
eggnog_annots <- eggnog_annots[1:(nrow(eggnog_annots) - 3), ]
gff.df <- as.data.frame(import(gff_path))
protein_2_gene <- unique(gff.df[c("Name", "gene")])
protein_2_gene_df <- subset(protein_2_gene, grepl("^XP", protein_2_gene$Name))
eggnog_annots$Name <- eggnog_annots$X.query
eggnog_annots <- left_join(eggnog_annots, protein_2_gene_df, by = "Name")
eggnog_annots$X.query <- eggnog_annots$gene
# GO
GO_terms <- data.table(eggnog_annots[, c("X.query", "GOs")])
GO_terms <- GO_terms[, .(GOs = unlist(strsplit(GOs, ","))), by = X.query]
term2name <- GO_terms[, .(GOs, X.query)]
term2name$Names <- mapIds(GO.db, keys = term2name$GOs, column = "TERM", keytype = "GOID")
term2name$Ontology <- mapIds(GO.db, keys = term2name$GOs, column = "ONTOLOGY", keytype = "GOID")
term2name <- as.data.frame(term2name)
go_bp <- term2name[term2name$Ontology == "BP", c("GOs", "X.query")]
go_mf <- term2name[term2name$Ontology == "MF", c("GOs", "X.query")]
go_cc <- term2name[term2name$Ontology == "CC", c("GOs", "X.query")]
term2name_filtered <- term2name[!is.na(term2name$Names), c("GOs", "Names")]
ontologies <- list(BP = go_bp, MF = go_mf, CC = go_cc)
# KEGG
KO_terms <- data.table(eggnog_annots[, c("X.query", "KEGG_ko")])
KO_terms$KEGG_ko <- gsub("ko:", "", as.character(KO_terms$KEGG_ko))
KO_terms <- KO_terms[, .(KEGG_ko = unlist(strsplit(KEGG_ko, ","))), by = X.query]
kegg_final <- KO_terms[, .(KEGG_ko, X.query)]
# Store per species
GO_terms_list[[sp]] <- GO_terms
ontologies_list[[sp]] <- ontologies
term2name_list[[sp]] <- term2name_filtered
kegg_final_list[[sp]] <- kegg_final
}We run the loop using the the list of selected genes extracted in step 1:
# ===== Set up parameters =====
species_list <- c("gregaria", "cancellata", "piceifrons", "americana", "cubense", "nitens")
suffix <- "aBSREL"
# Mapping external species names to internal codes in ortho_map
species_translate <- c(
gregaria = "Sgreg",
cancellata = "Scanc",
piceifrons = "Spice",
americana = "Samer", # double-check this is correct
cubense = "Sscub",
nitens = "Snite"
)
# ===== Loop through each species =====
go_results_all <- list()
kegg_results_all <- list()
# initialize diagnostic log
diagnostic_log <- data.frame(
Species = character(),
PSG_genes = integer(),
GO_overlap = integer(),
KEGG_overlap = integer(),
stringsAsFactors = FALSE
)
for (sp in species_list) {
message("Processing ", sp)
sp_code <- species_translate[sp]
output_dir <- file.path(enrichDir, sp)
dir.create(output_dir, recursive = TRUE, showWarnings = FALSE)
# 1. Extract PSG GeneIDs for this species
species_genes <- psg_genes %>%
filter(Species == sp_code) %>%
pull(GeneID) %>%
unique()
n_psg <- length(species_genes)
message("→ ", n_psg, " PSG genes in ", sp)
# 2. Check overlaps with GO and KEGG annotations
go_annot <- GO_terms_list[[sp]]$X.query
kegg_annot <- kegg_final_list[[sp]]$X.query # ✅ use the correct list
go_overlap <- sum(species_genes %in% go_annot)
kegg_overlap <- sum(species_genes %in% kegg_annot)
message("→ Overlap counts: GO = ", go_overlap, ", KEGG = ", kegg_overlap)
if (kegg_overlap == 0) {
message(" ⚠️ No KEGG matches for ", sp,
". Example missing IDs: ",
paste(head(setdiff(species_genes, kegg_annot), 5), collapse = ", "))
}
# Add to diagnostic log
diagnostic_log <- rbind(diagnostic_log, data.frame(
Species = sp,
PSG_genes = n_psg,
GO_overlap = go_overlap,
KEGG_overlap = kegg_overlap
))
# 3. Run GO enrichment (by ontology)
selected_genes_go <- species_genes[species_genes %in% go_annot]
go_by_onto <- list()
for (onto in names(ontologies_list[[sp]])) {
go_by_onto[[onto]] <- run_GO_enrichment_selected(
gene_list = selected_genes_go,
go_table = ontologies_list[[sp]][[onto]],
term2name = term2name_list[[sp]],
species = sp,
suffix = suffix,
ontology = onto,
output_dir = output_dir
)
}
go_results_all[[sp]] <- go_by_onto
# 4. Run KEGG enrichment
selected_genes_kegg <- species_genes[species_genes %in% kegg_annot]
kegg_final <<- kegg_final_list[[sp]] # required by assign_kegg_ids()
kegg_results_all[[sp]] <- run_KEGG_enrichment_selected(
gene_list = selected_genes_kegg,
species = sp,
suffix = suffix,
output_dir = output_dir
)
}
# save diagnostic log
write.csv(diagnostic_log, file.path(enrichDir, "diagnostic_overlap_log.csv"), row.names = FALSE)
print(diagnostic_log) Species PSG_genes GO_overlap KEGG_overlap
1 gregaria 229 197 197
2 cancellata 280 252 252
3 piceifrons 347 321 321
4 americana 222 197 197
5 cubense 206 188 188
6 nitens 285 256 2568. BUSTED
We will perform BUSTED analysis using both unlabeled and labelled phylogeny.
The unlabeled gene tree phylogeny will allow for an exploratory analysis, testing all Polyneoptera for positive selection. While this approach provides a broad overview, it comes at the cost of reduced statistical power due to multiple testing.
In contrast, the labelled gene tree phylogeny will focus specifically on migratory locust species compared to all other species or to non-migratory grasshoppers, enabling us to associate traits with selective pressures.
Note: The new version of OrthoFinder
makes a list of SingleCopy Orthologues by adding a N0:H
before the orthogroup name. N0.HOG0000086 N0.HOG0000090 N0.HOG0000212
N0.HOG0000220 N0.HOG0000478 N0.HOG0000479 N0.HOG0000503
N0.HOG0000505
So we need to clean that up before running our files using the command
sed 's/^N0\.HOG/OG/' Orthogroups_SingleCopyOrthologues.txt > Orthogroups_SingleCopyOrthologues_renamed.txtRunning the analysis
We will perform BUSTED analysis using both unlabeled and labelled gene tree phylogeny. The unlabeled phylogeny will allow for a gene-wide exploratory analysis treating the entire tree of Polyneoptera as foreground.
# For Polyneoptera
# For unlabelled phylogeny
sbatch scripts/RunBUSTED_May2025.sh \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Resolved_Gene_Trees/ \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Orthogroups/Orthogroups_SingleCopyOrthologues_selanalysiswide.txt
# For labelled phylogeny
sbatch ./scripts/RunBUSTED_labeled_May2025.sh \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/9_1_LabelledPhylogenies/Locusts \
Locusts \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Orthogroups/Orthogroups_SingleCopyOrthologues_selanalysiswide.txt
# For labelled phylogeny PRUNED
sbatch ./scripts/RunBUSTED_labeled_June2025.sh \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/9_1_LabelledPhylogenies_Pruned/Locusts \
Locusts \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Orthogroups/Orthogroups_SingleCopyOrthologues_selanalysiswide.txt If we want to run R interactively on the cluster:
srun --ntasks 1 --cpus-per-task 16 --mem 50G --time 05:00:00 --pty bash
ml GCC/13.2.0 OpenMPI/4.1.6 R_tamu/4.4.1
ml WebProxy
export R_LIBS=$SCRATCH/R_LIBS_USER/
Rscript ./scripts/Parsing_BUSTEDresulsr_unlabel_June2025.RUnlabelled
Parsing the json files
To parse all the details from the BUSTED by testing all branches to
see if we have selection pressures
./scripts/Parsing_BUSTEDresulsr_unlabel_June2025.R:
library(jsonlite)
library(tidyverse)── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
✔ forcats 1.0.0 ✔ purrr 1.0.4
✔ ggplot2 3.5.2 ✔ tidyr 1.3.1
✔ lubridate 1.9.4
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ purrr::flatten() masks jsonlite::flatten()
✖ dplyr::lag() masks stats::lag()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorslibrary(hexbin)
# ============ SETTINGS ============
input_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/8_3_BustedResults"
single_copy_file <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Orthogroups/Orthogroups_SingleCopyOrthologues_selanalysiswide.txt"
output_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedBUSTEDResults_unlabeled"
dir.create(output_dir, showWarnings = FALSE, recursive = TRUE)
# ============ JSON PARSER ============
parse_busted <- function(file) {
tryCatch({
busted <- jsonlite::fromJSON(file)
og <- stringr::str_extract(basename(file), "^OG[0-9]+")
busted_input_file <- busted[["input"]][["file name"]]
busted_pval <- as.numeric(busted[["test results"]][["p-value"]])
# Corrected metadata fields
aln_length <- busted[["input"]][["number of sites"]]
seq_count <- busted[["input"]][["number of sequences"]]
# Rate model info: safely extract from 'Test'
rate_info <- busted[["fits"]][["Unconstrained model"]][["Rate Distributions"]][["Test"]]
omega_vals <- sapply(rate_info, function(x) as.numeric(x[["omega"]]))
prop_vals <- sapply(rate_info, function(x) as.numeric(x[["proportion"]]))
# Optional: if missing, use NA
omega3 <- ifelse(length(omega_vals) >= 3, omega_vals[3], NA)
prop3 <- ifelse(length(prop_vals) >= 3, prop_vals[3], NA)
# Flag potential overfitting
suspect <- is.na(omega3) || omega3 > 1000 || prop3 < 0.001 || aln_length < 100 || seq_count < 4
tibble::tibble(
file = file,
input_file = busted_input_file,
orthogroup = og,
seq_count = seq_count,
aln_length = aln_length,
omega1 = omega_vals[1],
prop1 = prop_vals[1],
omega2 = omega_vals[2],
prop2 = prop_vals[2],
omega3 = omega3,
prop_sites = prop3,
pval = busted_pval,
padj = p.adjust(busted_pval, method = "BH"),
significant = p.adjust(busted_pval, method = "BH") < 0.05,
suspect_result = suspect
)
}, error = function(e) {
message("⚠️ Error parsing: ", file, " -> ", e$message)
return(NULL)
})
}
# ============ Parse All JSON Files ============
json_files <- list.files(input_dir, pattern = "\\.json$", full.names = TRUE)
parsed <- map_dfr(json_files, parse_busted)
# ============ Save All Results ============
write_csv(parsed, file.path(output_dir, "BUSTED_results_all.csv"))
write_csv(filter(parsed, significant), file.path(output_dir, "BUSTED_results_significant.csv"))
# ============ Optional: Filter for Single-Copy Orthogroups ============
if (file.exists(single_copy_file)) {
sc_ogs <- read_lines(single_copy_file) %>% str_trim()
parsed_sc <- parsed %>% filter(orthogroup %in% sc_ogs)
write_csv(parsed_sc, file.path(output_dir, "BUSTED_results_singlecopy.csv"))
write_csv(filter(parsed_sc, significant), file.path(output_dir, "BUSTED_results_singlecopy_significant.csv"))
}
# ============ Quick Summary ============
message("✅ Parsed: ", nrow(parsed), " orthogroups")✅ Parsed: 5343 orthogroupsmessage("🧬 Positive selection (FDR < 0.05): ", sum(parsed$significant))🧬 Positive selection (FDR < 0.05): 2183library(ggplot2)
# Create hexbin plot for significant results
p <- ggplot(filter(parsed, significant), aes(x = prop_sites, y = omega3)) +
geom_hex(bins = 40) +
scale_fill_gradient(trans = "log10", low = "#ccf0ed", high = "#014a44") +
scale_x_log10() +
scale_y_log10() +
labs(
title = "Selection Landscape for Positively Selected Genes",
x = "Proportion of Sites Under Selection (log10)",
y = "Strength of Selection (omega, log10)",
fill = "Number of Genes"
) +
theme_bw()
ggsave(filename = file.path(output_dir, "busted_hexbin_plot.pdf"), plot = p, width = 7, height = 6)
p2 <- ggplot(parsed, aes(x = omega3, y = -log10(padj), color = significant)) +
geom_point(alpha = 0.8) +
scale_color_manual(values = c("FALSE" = "grey60", "TRUE" = "red")) +
labs(
x = "Strength of Selection (omega3)",
y = expression(-log[10]~"(FDR-adjusted p-value)"),
color = "Significant"
) +
theme_minimal()
ggsave(file.path(output_dir, "busted_volcano_plot.pdf"), p2, width = 7, height = 6)
p3 <- parsed %>%
filter(significant) %>%
arrange(padj) %>%
slice_head(n = 20) %>%
ggplot(aes(x = reorder(orthogroup, -padj), y = -log10(padj))) +
geom_col(fill = "steelblue") +
coord_flip() +
labs(
x = "Orthogroup",
y = expression(-log[10]~"(FDR-adjusted p-value)"),
title = "Top 20 Positively Selected Orthogroups"
) +
theme_classic()
ggsave(file.path(output_dir, "busted_top20_barplot.pdf"), p3, width = 8, height = 6)
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Extracting the selected genes
We run BUSTED analysis on a total of 5,347 single copy
orthogroups for which:
3,094 orthogroups with 1:1 orthologs for all species (SelecAnalysisStrict = Included).
763 orthogroups with 1:1 orthologs for Caelifera species (SelAnalysisLocusts = Included).
1,490 orthogroups with mixed 1:1 orthologs with all Caelifera species (SelAnalysisMixed = Included).
We will show the results by category.
library(ggplot2)
library(dplyr)
library(readr)
library(ggnewscale)
ortho_dir <- "/Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data/orthofinder/Polyneoptera"
input_file <- file.path(ortho_dir, "Results_I2_iqtree/Orthogroups/Orthogroups_CladeAssignment_WithCopyStatus_cleaned.csv")
orthologtable <- read.csv(input_file, header = TRUE, stringsAsFactors = FALSE)
hyphy_dir <- "/Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data/HYPHY_selection"
input_file2 <- file.path(hyphy_dir, "ParsedBUSTEDResults_unlabeled/BUSTED_results_all.csv")
bustedtable <- read.csv(input_file2, header = TRUE, stringsAsFactors = FALSE) %>%
select(-input_file, -file)
busted_df <- left_join(orthologtable, bustedtable, by = c("Orthogroup" = "orthogroup"))
# Save as CSV
workDir <- "/Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data"
output_file <- file.path(workDir, "HYPHY_selection/ParsedBUSTEDResults_unlabeled/busted_compiled.csv")
write.csv(busted_df, output_file, row.names = FALSE)
busted_SingleStrict <- busted_df %>%
filter(SelAnalysisStrict == "Included")
busted_locust <- busted_df %>%
filter(SelAnalysisLocusts == "Included")
busted_mixed <- busted_df %>%
filter(SelAnalysisMixed == "Included")
library(dplyr)
library(tibble)
summary_table <- tibble(
Category = c("1:1 Polyneoptera", "1:1 Caelifera only", "1:1 Mixed"),
Total = c(
sum(busted_df$SelAnalysisStrict == "Included", na.rm = TRUE),
sum(busted_df$SelAnalysisLocusts == "Included", na.rm = TRUE),
sum(busted_df$SelAnalysisMixed == "Included", na.rm = TRUE)
),
Significant = c(
sum(busted_df$SelAnalysisStrict == "Included" & busted_df$padj < 0.05, na.rm = TRUE),
sum(busted_df$SelAnalysisLocusts == "Included" & busted_df$padj < 0.05, na.rm = TRUE),
sum(busted_df$SelAnalysisMixed == "Included" & busted_df$padj < 0.05, na.rm = TRUE)
),
Suspect = c(
sum(busted_df$SelAnalysisStrict == "Included" & busted_df$padj < 0.05 & busted_df$suspect_result == TRUE, na.rm = TRUE),
sum(busted_df$SelAnalysisLocusts == "Included" & busted_df$padj < 0.05 & busted_df$suspect_result == TRUE, na.rm = TRUE),
sum(busted_df$SelAnalysisMixed == "Included" & busted_df$padj < 0.05 & busted_df$suspect_result == TRUE, na.rm = TRUE)
)
)%>%
mutate(True_Selected = Significant - Suspect)
knitr::kable(summary_table, caption = "Summary of BUSTED results per orthogroup category")| Category | Total | Significant | Suspect | True_Selected |
|---|---|---|---|---|
| 1:1 Polyneoptera | 3094 | 1414 | 280 | 1134 |
| 1:1 Caelifera only | 763 | 210 | 59 | 151 |
| 1:1 Mixed | 1490 | 559 | 110 | 449 |
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
A total of 2,183 orthogroups showed signature of selection but only 1,734 showed omega3 values that were not suspect (due to model over fitting, short alignment, low divergence).
Below is the selection landscape for all orthogroups with corrected (BH) p-value < 0.05.
# Filter for significant genes
parsed <- busted_df %>%
filter(!is.na(padj), padj < 0.05) %>%
filter(!is.na(prop_sites), !is.na(omega3)) # <- ensure both axes are numeric
# Separate the suspect results
suspect_points <- parsed %>%
filter(suspect_result == TRUE)
# Base: All significant data
base_data <- parsed %>% filter(suspect_result == FALSE)
suspect_data <- parsed %>% filter(suspect_result == TRUE)
p <- ggplot() +
geom_hex(data = base_data, aes(x = prop_sites, y = omega3), bins = 40) +
scale_fill_gradient(trans = "log10", low = "#ccf0ed", high = "#014a44") +
new_scale_fill() + # Needed from ggh4x or ggnewscale to add a second fill scale
geom_hex(data = suspect_data, aes(x = prop_sites, y = omega3), bins = 40, inherit.aes = FALSE) +
scale_fill_gradient(trans = "log10", low = "mistyrose", high = "red", name = "Suspect Count") +
scale_x_log10() +
scale_y_log10() +
labs(
title = "Selection Landscape: Suspect Results in Red Hexes",
x = "Proportion of Sites Under Selection (log10)",
y = "Strength of Selection (omega, log10)"
) +
theme_bw()
p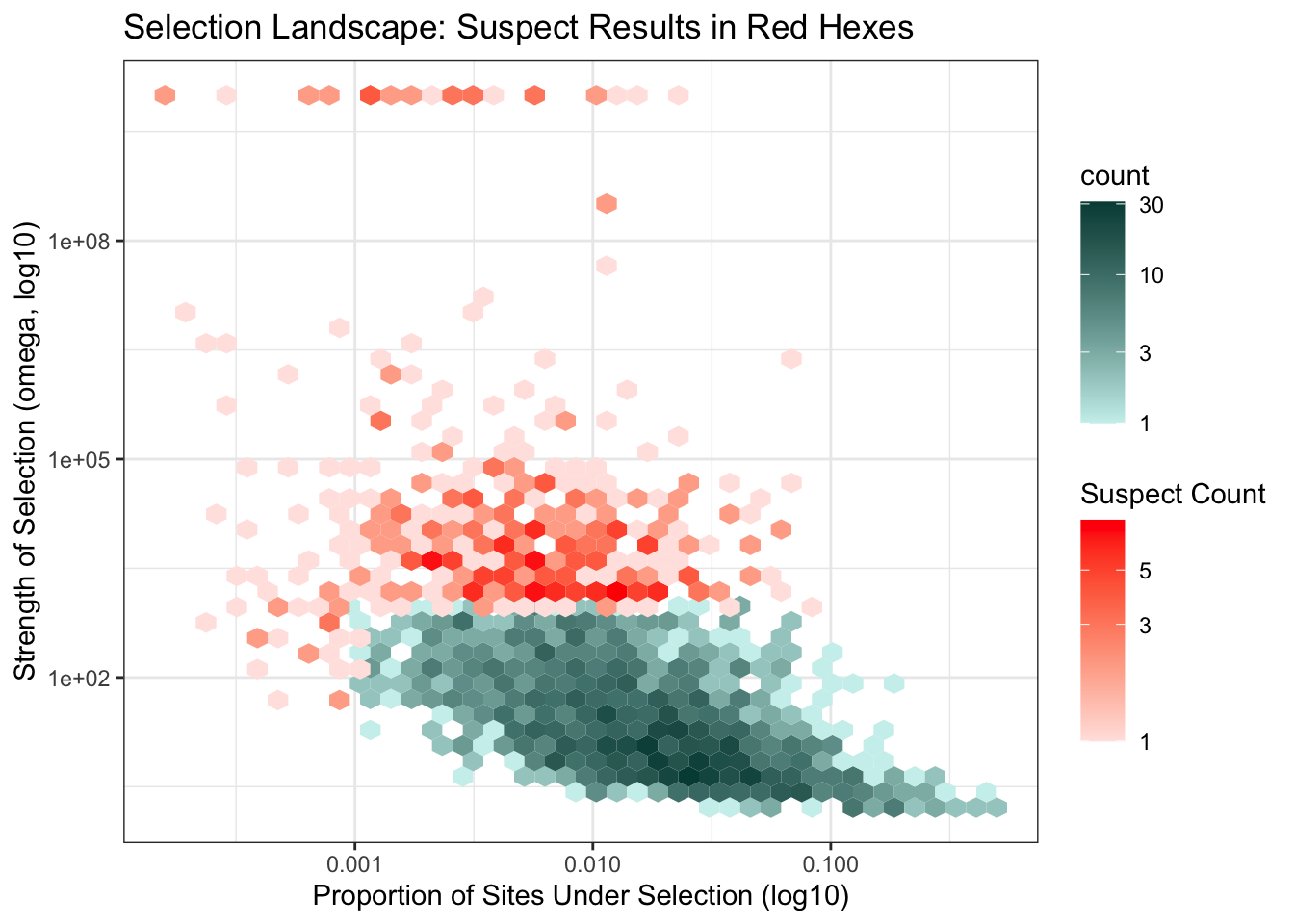
| Version | Author | Date |
|---|---|---|
| a2d2955 | Maeva TECHER | 2025-07-01 |
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Below we show the hex bin graphs for only the 1:1 Polyneoptera and 1:1 Caelifera selection landscapes.
# Filter for significant genes
parsed_polyneoptera <- busted_SingleStrict %>%
filter(!is.na(padj), padj < 0.05) %>%
filter(!is.na(prop_sites), !is.na(omega3)) %>% # <- ensure both axes are numeric
filter(suspect_result == FALSE)
# Plot
p <- ggplot(parsed_polyneoptera, aes(x = prop_sites, y = omega3)) +
geom_hex(bins = 40) +
scale_fill_gradient(trans = "log10", low = "#ccf0ed", high = "#014a44") +
scale_x_log10() +
scale_y_log10() +
labs(
title = "Selection Landscape for Positively Selected Genes (1:1 Polyneoptera)",
x = "Proportion of Sites Under Selection (log10)",
y = "Strength of Selection (omega, log10)",
fill = "Number of Orthogroups"
) +
theme_bw()
p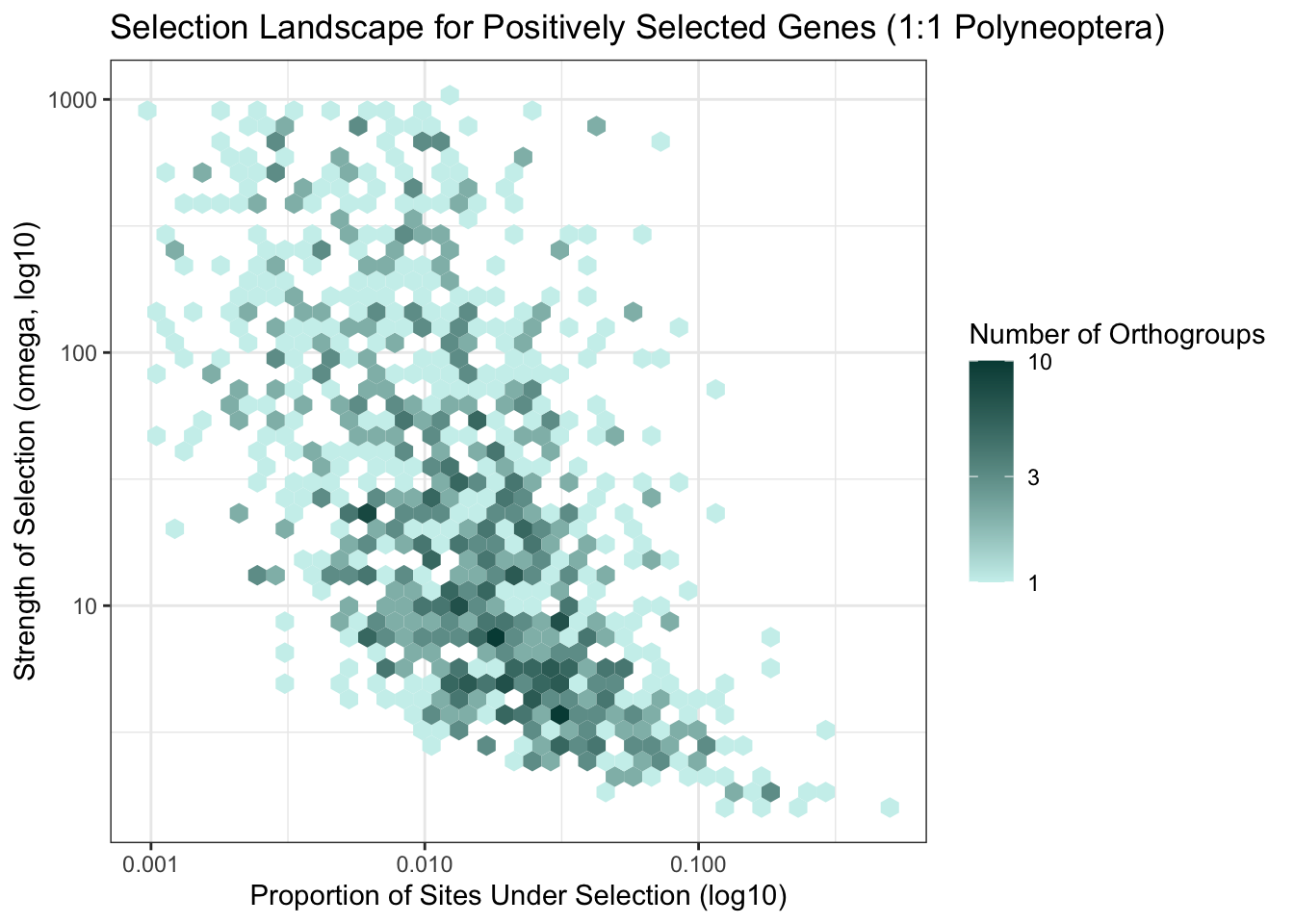
| Version | Author | Date |
|---|---|---|
| a2d2955 | Maeva TECHER | 2025-07-01 |
# Filter for significant genes
parsed_locust <- busted_locust %>%
filter(!is.na(padj), padj < 0.05) %>%
filter(!is.na(prop_sites), !is.na(omega3)) %>% # <- ensure both axes are numeric
filter(suspect_result == FALSE)
# We make a hexbin graph only for genes that are locust only
ggplot(parsed_locust, aes(x = prop_sites, y = omega3)) +
geom_hex(bins = 40) +
scale_fill_gradient(trans = "log10", low = "#e6f2ff", high = "#084594") +
scale_x_log10() +
scale_y_log10() +
labs(
title = "Selection on Strict Single-Copy Genes (1:1 Caelifera only)",
x = "Proportion of Sites Under Selection (log10)",
y = "Strength of Selection (omega, log10)",
fill = "Number of Orthogroups"
) +
theme_bw()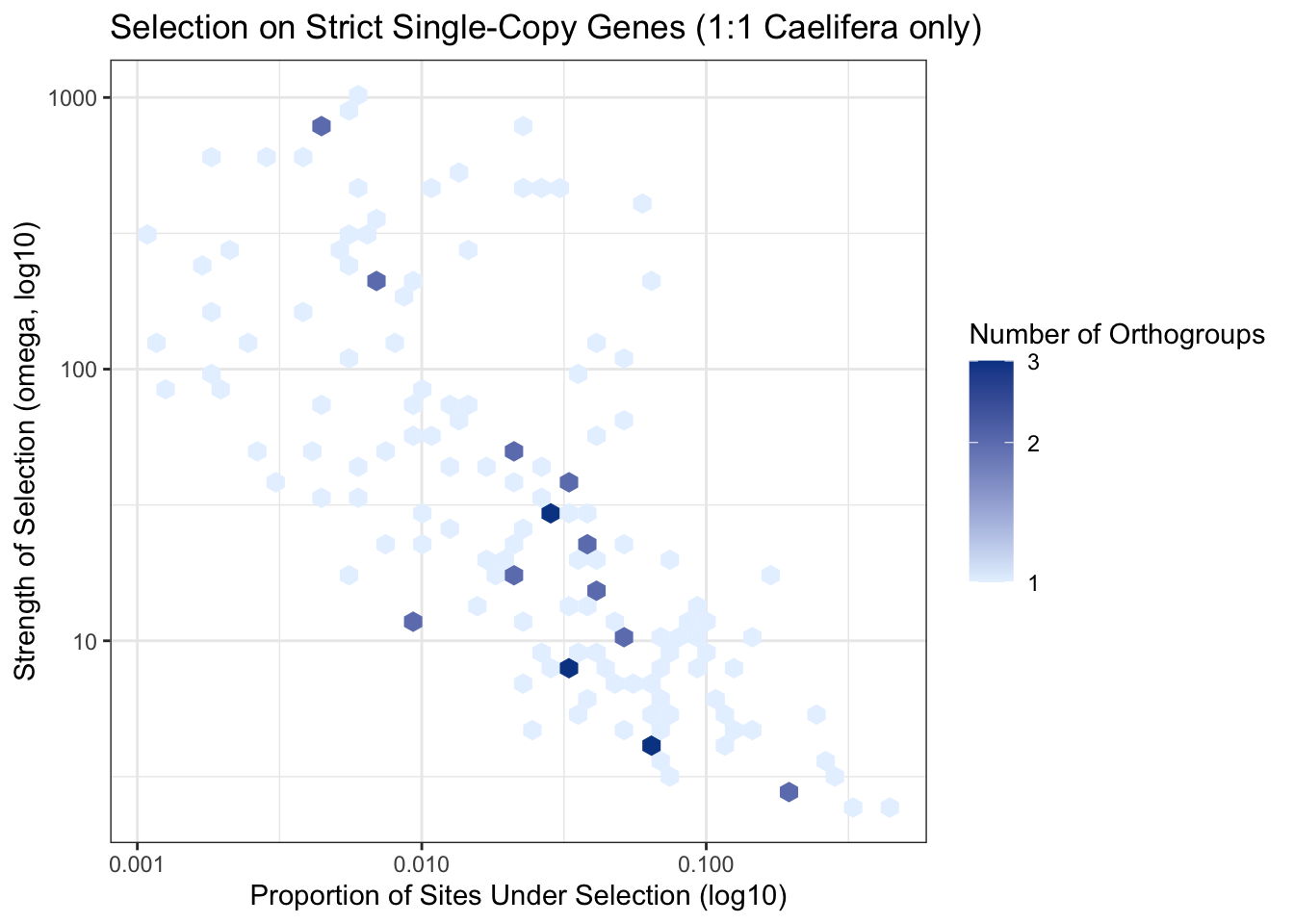
| Version | Author | Date |
|---|---|---|
| a2d2955 | Maeva TECHER | 2025-07-01 |
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
To explore more clearly which orthogroups are showing high selective pressure, we made an interactive version of the hex bin plot below:
library(plotly)
# Prepare your data (already filtered for single-copy, etc.)
plot_data <- parsed_locust %>%
mutate(
log_omega3 = log10(omega3),
log_prop_sites = log10(prop_sites)
)
p <- ggplot(parsed_locust, aes(x = log10(prop_sites), y = log10(omega3))) +
geom_hex(bins = 40, aes(fill = ..count..)) +
geom_point(aes(text = Orthogroup), alpha = 0.1, color = "black") + # invisible overlay for tooltip
scale_fill_viridis_c(trans = "log10") +
labs(
x = "log10(Proportion of Sites Under Selection)",
y = "log10(Omega3)",
fill = "Orthogroup Count",
title = "Interactive Hexbin with Orthogroup Hover (1:1 genes Caelifera only)"
) +
theme_minimal()
ggplotly(p, tooltip = "text")
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Annotating OGs under selection
Now we will enrich the pathways for which the genes under selection were found:
ortho_dir <- "/Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data/orthofinder/Polyneoptera"
input_file <- file.path(ortho_dir, "Results_I2_iqtree/Orthogroups_genesproteinbiotype_13species_annotated_May2025.csv")
ortho_map <- read.csv(input_file, header = TRUE, stringsAsFactors = FALSE)
head(ortho_map) Orthogroup SpeciesID protein_id GeneID
1 OG0000000 Asimp_filteredTranscripts XP_067003642.2 LOC136874043
2 OG0000000 Asimp_filteredTranscripts XP_067004661.1 LOC136874869
3 OG0000000 Asimp_filteredTranscripts XP_067015293.1 LOC136886419
4 OG0000000 Asimp_filteredTranscripts XP_067015651.2 LOC136886746
5 OG0000000 Asimp_filteredTranscripts XP_068085770.1 LOC137496902
6 OG0000000 Asimp_filteredTranscripts XP_068087037.1 LOC137503369
Description Species GeneType
1 farnesol dehydrogenase isoform X1 Asimp protein-coding
2 dehydrogenase/reductase SDR family member 11 Asimp protein-coding
3 farnesol dehydrogenase Asimp protein-coding
4 farnesol dehydrogenase Asimp protein-coding
5 farnesol dehydrogenase-like Asimp protein-coding
6 farnesol dehydrogenase-like Asimp protein-coding
Accession Begin End Orthogroup_Type
1 NC_090269.1 316521124 316596183 MultiCopy
2 NC_090269.1 316408775 316497937 MultiCopy
3 NC_090279.1 165240845 165292312 MultiCopy
4 NC_090279.1 164532314 164617967 MultiCopy
5 NC_090275.1 60074551 60101839 MultiCopy
6 NC_090279.1 164618824 164719067 MultiCopy# Extract the orthogroup names
selected_orthogroups <- parsed_locust %>%
filter(!is.na(padj), padj < 0.05) %>%
filter(!is.na(prop_sites), !is.na(omega3)) %>% # <- ensure both axes are numeric
filter(suspect_result == FALSE) %>%
pull(Orthogroup) %>% unique()
# Get corresponding GeneID
selected_genes <- ortho_map %>%
filter(Orthogroup %in% selected_orthogroups) %>%
pull(GeneID) %>% unique()
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
We can use the same pipeline and functions as in our section 3: GO enrichment for DEGs.
# === Paths and Constants ===
workDir <- "/Users/maevatecher/Documents/GitHub/locust-comparative-genomics/data"
GODir <- file.path(workDir, "list", "GO_Annotations")
RefDir <- file.path(workDir, "RefSeq")
enrichDir <- file.path(workDir, "HYPHY_selection/functional_pathways/BUSTED_unlabeled")
selListDir <- file.path(workDir, "HYPHY_selection/ParsedBUSTEDResults_unlabeled")
species_list <- c("gregaria", "cancellata", "piceifrons", "americana", "cubense", "nitens")
# === Load Required Libraries ===
library(data.table)
library(dplyr)
library(readr)
library(clusterProfiler)
library(GO.db)
library(rtracklayer)
library(DesertLocustR) # Local installation
gff_map <- c(
gregaria = "GCF_023897955.1_iqSchGreg1.2_genomic.gff",
cancellata = "GCF_023864275.1_iqSchCanc2.1_genomic.gff",
piceifrons = "GCF_021461385.2_iqSchPice1.1_genomic.gff",
americana = "GCF_021461395.2_iqSchAmer2.1_genomic.gff",
cubense = "GCF_023864345.2_iqSchSeri2.2_genomic.gff",
nitens = "GCF_023898315.1_iqSchNite1.1_genomic.gff"
)
annot_map <- c(
gregaria = "EggNog_Arthropoda_one2one.emapper.annotations",
cancellata = "GCF_023864275.1_iqSchCanc2.1_Arthopoda_one2one.emapper.annotations",
piceifrons = "GCF_021461385.2_iqSchPice1.1_Arthopoda_one2one.emapper.annotations",
americana = "GCF_021461395.2_iqSchAmer2.1_Arthopoda_one2one.emapper.annotations",
cubense = "GCF_023864345.2_iqSchSeri2.2_Arthopoda_one2one.emapper.annotations",
nitens = "GCF_023898315.1_iqSchNite1.1_Arthopoda_one2one.emapper.annotations"
)
# GO enrichment
enrich_GO <- function(dge_genes.df, term2gene, term2name, pval, qval){
genes <- rownames(dge_genes.df)
enricher(genes, TERM2GENE = term2gene, TERM2NAME = term2name, pvalueCutoff = pval,
pAdjustMethod = "BH", qvalueCutoff = qval)
}
# KEGG preparation
assign_kegg_ids <- function(sig_genes.df){
if (is.vector(sig_genes.df)) {
sig_genes.df <- data.frame(X.query = sig_genes.df, stringsAsFactors = FALSE)
} else {
sig_genes.df$X.query <- rownames(sig_genes.df)
}
dge_with_kegg_ids <- left_join(sig_genes.df, kegg_final, by = "X.query")
dge_with_kegg_ids$KEGG_ko[grepl("^K", dge_with_kegg_ids$KEGG_ko)]
}
# KEGG enrichment
enrich_KEGG <- function(dge_genes.df, pval_cutoff = 0.05, qval_cutoff = 0.05) {
gene_with_kegg_ids <- assign_kegg_ids(dge_genes.df)
enrichKEGG(
gene = gene_with_kegg_ids,
organism = "ko",
pvalueCutoff = pval_cutoff,
qvalueCutoff = qval_cutoff,
pAdjustMethod = "BH"
)
}
run_GO_enrichment_selected <- function(
gene_list,
go_table,
term2name,
species,
suffix,
ontology,
output_dir,
show_n = 30,
top_n = 30
) {
if (length(gene_list) == 0) return(NULL)
if (!dir.exists(output_dir)) {
dir.create(output_dir, recursive = TRUE)
}
# Make sure column names are correct for clusterProfiler::enricher()
go_table_fixed <- go_table[, 1:2]
colnames(go_table_fixed) <- c("go_id", "gene_id")
term2name_fixed <- term2name[, 1:2]
colnames(term2name_fixed) <- c("go_id", "name")
# Run enrichment
go_result <- enricher(
gene = gene_list,
TERM2GENE = go_table_fixed,
TERM2NAME = term2name_fixed,
pvalueCutoff = 0.05,
qvalueCutoff = 0.05
)
if (!is.null(go_result) &&
inherits(go_result, "enrichResult") &&
nrow(go_result@result) > 0 &&
sum(!is.na(go_result@result$Description)) > 0) {
# Save dotplot
try({
pdf(file = file.path(output_dir, paste0("GO_", ontology, "_dotplot_", species, "_", suffix, ".pdf")),
width = 8, height = 6)
print(dotplot(go_result, showCategory = min(show_n, nrow(go_result@result))) +
ggtitle(paste(ontology, suffix)))
dev.off()
}, silent = TRUE)
# Export top terms with log10(p)
species_enrich_ready <- go_result@result[, c("ID", "p.adjust")]
species_enrich_ready$logp <- -log10(species_enrich_ready$p.adjust)
species_enrich_ready <- species_enrich_ready[order(-species_enrich_ready$logp), ]
species_enrich_ready <- head(species_enrich_ready, n = top_n)[, c("ID", "logp")]
write.table(species_enrich_ready,
file = file.path(output_dir, paste0("enrich_", ontology, "_GOs_", species, "_", suffix, ".txt")),
sep = "\t", quote = FALSE, row.names = FALSE, col.names = FALSE)
# Also export the full table if needed
write.csv(go_result@result,
file = file.path(output_dir, paste0("GO_enrichment_full_", ontology, "_", species, "_", suffix, ".csv")),
row.names = FALSE)
} else {
message(paste0("⚠️ No GO enrichment result to plot/export for ", species, " - ", suffix))
}
}
run_KEGG_enrichment_selected <- function(gene_list, species, suffix, output_dir,
show_n = 40, top_n = 40) {
if (length(gene_list) == 0) return(NULL)
kegg_result <- enrich_KEGG(gene_list, pval_cutoff = 0.05, qval_cutoff = 0.05)
if (!is.null(kegg_result) && inherits(kegg_result, "enrichResult") &&
nrow(kegg_result@result) > 0) {
try({
pdf(file = file.path(output_dir, paste0("KEGG_dotplot_", species, "_", suffix, ".pdf")),
width = 8, height = 6)
print(dotplot(kegg_result, showCategory = min(show_n, nrow(kegg_result@result))) +
ggtitle(paste("KEGG", suffix)))
dev.off()
}, silent = TRUE)
# Full result
write.csv(kegg_result@result,
file = file.path(output_dir, paste0("KEGG_enrichment_", species, "_", suffix, ".csv")),
row.names = FALSE)
# Top KEGG terms
species_enrich_kegg <- kegg_result@result[, c("ID", "p.adjust")]
species_enrich_kegg$logp <- -log10(species_enrich_kegg$p.adjust)
species_enrich_kegg <- species_enrich_kegg[order(-species_enrich_kegg$logp), ][1:min(nrow(species_enrich_kegg), top_n), ]
species_enrich_kegg <- species_enrich_kegg[, c("ID", "logp")]
write.table(species_enrich_kegg,
file = file.path(output_dir, paste0("enrich_KEGG_", species, "_", suffix, ".txt")),
sep = "\t", quote = FALSE, row.names = FALSE, col.names = FALSE)
} else {
message(paste("⚠️ No KEGG enrichment result to plot/export for", species, "-", suffix))
}
}
GO_terms_list <- list()
ontologies_list <- list()
term2name_list <- list()
kegg_final_list <- list()
# Mapping external species names to internal codes in ortho_map
species_translate <- c(
gregaria = "Sgreg",
cancellata = "Scanc",
piceifrons = "Spice",
americana = "Samer", # double-check this is correct
cubense = "Sscub",
nitens = "Snite"
)
for (sp in species_list) {
message("Preparing annotations for ", sp)
sp_code <- species_translate[sp]
eggnog_path <- file.path(GODir, annot_map[[sp]])
gff_path <- file.path(RefDir, gff_map[[sp]])
output_dir <- file.path(enrichDir, sp)
dir.create(output_dir, recursive = TRUE, showWarnings = FALSE)
eggnog_annots <- read.delim(eggnog_path, sep = "\t", skip = 4)
eggnog_annots <- eggnog_annots[1:(nrow(eggnog_annots) - 3), ]
gff.df <- as.data.frame(import(gff_path))
protein_2_gene <- unique(gff.df[c("Name", "gene")])
protein_2_gene_df <- subset(protein_2_gene, grepl("^XP", protein_2_gene$Name))
eggnog_annots$Name <- eggnog_annots$X.query
eggnog_annots <- left_join(eggnog_annots, protein_2_gene_df, by = "Name")
eggnog_annots$X.query <- eggnog_annots$gene
# GO
GO_terms <- data.table(eggnog_annots[, c("X.query", "GOs")])
GO_terms <- GO_terms[, .(GOs = unlist(strsplit(GOs, ","))), by = X.query]
term2name <- GO_terms[, .(GOs, X.query)]
term2name$Names <- mapIds(GO.db, keys = term2name$GOs, column = "TERM", keytype = "GOID")
term2name$Ontology <- mapIds(GO.db, keys = term2name$GOs, column = "ONTOLOGY", keytype = "GOID")
term2name <- as.data.frame(term2name)
go_bp <- term2name[term2name$Ontology == "BP", c("GOs", "X.query")]
go_mf <- term2name[term2name$Ontology == "MF", c("GOs", "X.query")]
go_cc <- term2name[term2name$Ontology == "CC", c("GOs", "X.query")]
term2name_filtered <- term2name[!is.na(term2name$Names), c("GOs", "Names")]
ontologies <- list(BP = go_bp, MF = go_mf, CC = go_cc)
# KEGG
KO_terms <- data.table(eggnog_annots[, c("X.query", "KEGG_ko")])
KO_terms$KEGG_ko <- gsub("ko:", "", KO_terms$KEGG_ko)
KO_terms <- KO_terms[, .(KEGG_ko = unlist(strsplit(KEGG_ko, ","))), by = X.query]
kegg_final <- KO_terms[, .(KEGG_ko, X.query)]
# Store per species
GO_terms_list[[sp]] <- GO_terms
ontologies_list[[sp]] <- ontologies
term2name_list[[sp]] <- term2name_filtered
kegg_final_list[[sp]] <- kegg_final
}
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
For the Polyneoptera genes: we enrich only for S. gregaria as own model since the genes are orthologs.
# ===== Prepare list of selected genes from orthogroups ====
selected_orthogroups <- parsed_polyneoptera %>%
filter(!is.na(padj), padj < 0.05) %>%
filter(!is.na(prop_sites), !is.na(omega3)) %>% # <- ensure both axes are numeric
filter(suspect_result == FALSE) %>%
pull(Orthogroup) %>% unique()
selected_genes <- ortho_map %>%
filter(Orthogroup %in% selected_orthogroups) %>%
pull(GeneID) %>%
unique()
# ===== Set up parameters =====
#species_list <- c("gregaria", "cancellata", "piceifrons", "americana", "cubense", "nitens")
species_list <- c("gregaria")
suffix <- "BUSTED_POLYNEOPTERA"
# Mapping external species names to internal codes in ortho_map
species_translate <- c(
gregaria = "Sgreg",
cancellata = "Scanc",
piceifrons = "Spice",
americana = "Samer", # double-check this is correct
cubense = "Sscub",
nitens = "Snite"
)
go_results_all <- list()
kegg_results_all <- list()
# ===== Loop through each species =====
for (sp in species_list) {
message("Processing ", sp)
sp_code <- species_translate[sp]
output_dir <- file.path(enrichDir, sp)
species_genes <- ortho_map %>%
filter(Orthogroup %in% selected_orthogroups, Species == sp_code) %>%
pull(GeneID) %>%
unique()
# Get species-specific GO terms
selected_genes_annot <- species_genes[species_genes %in% GO_terms_list[[sp]]$X.query]
message("→ ", length(selected_genes_annot), " genes for GO enrichment in ", sp)
# GO enrichment
go_by_onto <- list()
for (onto in names(ontologies_list[[sp]])) {
go_by_onto[[onto]] <- run_GO_enrichment_selected(
gene_list = selected_genes_annot,
go_table = ontologies_list[[sp]][[onto]],
term2name = term2name_list[[sp]],
species = sp,
suffix = suffix,
ontology = onto,
output_dir = output_dir
)
}
go_results_all[[sp]] <- go_by_onto
# KEGG enrichment
kegg_final <<- kegg_final_list[[sp]] # used inside assign_kegg_ids
kegg_results_all[[sp]] <- run_KEGG_enrichment_selected(
gene_list = selected_genes_annot,
species = sp,
suffix = suffix,
output_dir = output_dir
)
}
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Now we check the genes under selection only in Schistocerca clade, only for S. gregaria as own model since the genes are orthologs.
# ===== Prepare list of selected genes from orthogroups =====
selected_orthogroups <- parsed_locust %>%
filter(!is.na(padj), padj < 0.05) %>%
filter(!is.na(prop_sites), !is.na(omega3)) %>% # <- ensure both axes are numeric
filter(suspect_result == FALSE) %>%
pull(Orthogroup) %>% unique()
selected_genes <- ortho_map %>%
filter(Orthogroup %in% selected_orthogroups) %>%
pull(GeneID) %>%
unique()
# ===== Set up parameters =====
#species_list <- c("gregaria", "cancellata", "piceifrons", "americana", "cubense", "nitens")
species_list <- c("gregaria")
suffix <- "BUSTED_CAELIFERA"
# Mapping external species names to internal codes in ortho_map
species_translate <- c(
gregaria = "Sgreg",
cancellata = "Scanc",
piceifrons = "Spice",
americana = "Samer", # double-check this is correct
cubense = "Sscub",
nitens = "Snite"
)
go_results_all <- list()
kegg_results_all <- list()
# ===== Loop through each species =====
for (sp in species_list) {
message("Processing ", sp)
sp_code <- species_translate[sp]
output_dir <- file.path(enrichDir, sp)
species_genes <- ortho_map %>%
filter(Orthogroup %in% selected_orthogroups, Species == sp_code) %>%
pull(GeneID) %>%
unique()
# Get species-specific GO terms
selected_genes_annot <- species_genes[species_genes %in% GO_terms_list[[sp]]$X.query]
message("→ ", length(selected_genes_annot), " genes for GO enrichment in ", sp)
# GO enrichment
go_by_onto <- list()
for (onto in names(ontologies_list[[sp]])) {
go_by_onto[[onto]] <- run_GO_enrichment_selected(
gene_list = selected_genes_annot,
go_table = ontologies_list[[sp]][[onto]],
term2name = term2name_list[[sp]],
species = sp,
suffix = suffix,
ontology = onto,
output_dir = output_dir
)
}
go_results_all[[sp]] <- go_by_onto
# KEGG enrichment
kegg_final <<- kegg_final_list[[sp]] # used inside assign_kegg_ids
kegg_results_all[[sp]] <- run_KEGG_enrichment_selected(
gene_list = selected_genes_annot,
species = sp,
suffix = suffix,
output_dir = output_dir
)
}
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Labelled
Parsing the json files
To parse the results from the *json files from the BUSTED-PH with
migratory locusts as foreground branches we run this
./scripts/Parsing_BUSTEDresulsr_labelled_June2025.R:
#!/usr/bin/env Rscript
# ============================= #
# LOAD LIBRARIES #
# ============================= #
suppressPackageStartupMessages({
library(tidyverse)
library(jsonlite)
library(fs)
})
# ============================= #
# PARSE ONE FILE #
# ============================= #
parse_busted <- function(file) {
tryCatch({
busted <- jsonlite::fromJSON(file)
og <- stringr::str_extract(basename(file), "^OG[0-9]+")
busted_input_file <- busted[["input"]][["file name"]]
busted_pval <- as.numeric(busted[["test results"]][["p-value"]])
# Metadata
aln_length <- busted[["input"]][["number of sites"]]
seq_count <- busted[["input"]][["number of sequences"]]
# Rate model info (Unconstrained model → Test)
rate_info <- busted[["fits"]][["Unconstrained model"]][["Rate Distributions"]][["Test"]]
omega_vals <- sapply(rate_info, function(x) as.numeric(x[["omega"]]))
prop_vals <- sapply(rate_info, function(x) as.numeric(x[["proportion"]]))
omega3 <- ifelse(length(omega_vals) >= 3, omega_vals[3], NA)
prop3 <- ifelse(length(prop_vals) >= 3, prop_vals[3], NA)
suspect <- is.na(omega3) || omega3 > 1000 || prop3 < 0.001 || aln_length < 100 || seq_count < 4
tibble(
file = basename(file),
input_file = busted_input_file,
orthogroup = og,
seq_count = seq_count,
aln_length = aln_length,
omega1 = omega_vals[1],
prop1 = prop_vals[1],
omega2 = omega_vals[2],
prop2 = prop_vals[2],
omega3 = omega3,
prop_sites = prop3,
pval = busted_pval,
padj = p.adjust(busted_pval, method = "BH"),
significant = p.adjust(busted_pval, method = "BH") < 0.05,
suspect_result = suspect
)
}, error = function(e) {
message("⚠️ Error parsing: ", file, " → ", e$message)
return(NULL)
})
}
# ============================= #
# RUN SCRIPT #
# ============================= #
# Define your directories
foreground_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/8_4_BustedResults_labeled/Locusts/foreground/"
background_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/8_4_BustedResults_labeled/Locusts/background/"
output_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedBUSTEDResults_labeled"
# Create output folder
dir_create(output_dir)
# Find all files
fg_files <- dir_ls(foreground_dir, glob = "*.json")
bg_files <- dir_ls(background_dir, glob = "*.json")
# Parse all
parsed_fg <- purrr::map_dfr(fg_files, parse_busted) %>% mutate(dataset = "foreground")
parsed_bg <- purrr::map_dfr(bg_files, parse_busted) %>% mutate(dataset = "background")
# Merge
results_all <- bind_rows(parsed_fg, parsed_bg)
# Save
write_csv(results_all, file.path(output_dir, "busted_results_labeled_full.csv"))
message("✅ Parsed labeled BUSTED results for ", nrow(results_all), " genes.")
library(tidyverse)
# Load the parsed results
busted_df <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedBUSTEDResults_labeled/busted_results_labeled_full.csv")
# Pivot all needed columns
busted_summary <- busted_df %>%
dplyr::select(
orthogroup, dataset,
significant, suspect_result,
seq_count, aln_length,
omega1, omega2, omega3,
prop1, prop2, prop_sites,
pval, padj
) %>%
distinct() %>%
pivot_wider(
names_from = dataset,
values_from = c(
significant, suspect_result,
seq_count, aln_length,
omega1, omega2, omega3,
prop1, prop2, prop_sites,
pval, padj
),
names_glue = "{.value}_{dataset}",
values_fill = NA
) %>%
mutate(
selection_status = case_when(
significant_foreground & !significant_background ~ "Foreground only",
!significant_foreground & significant_background ~ "Background only",
significant_foreground & significant_background ~ "Both foreground and background",
TRUE ~ "No selection"
),
suspect_result = suspect_result_foreground | suspect_result_background
) %>%
dplyr::select(
orthogroup, selection_status, suspect_result,
pval_foreground, padj_foreground,
pval_background, padj_background,
starts_with("seq_count"), starts_with("aln_length"),
starts_with("omega"), starts_with("prop")
)
# View or write to file
print(busted_summary)
write_csv(
busted_summary,
"/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedBUSTEDResults_labeled/busted_orthogroup_summary.csv"
)
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Plotting fg vs bg
# ========== Load Libraries ==========
library(tidyverse)
# ========== Load Data ==========
busted_df <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedBUSTEDResults_labeled/busted_results_labeled_full.csv")
# ========== Data Preparation ==========
# Pivot to wide format
plot_df <- busted_df %>%
dplyr::select(orthogroup, dataset, omega3, significant, suspect_result) %>%
pivot_wider(
names_from = dataset,
values_from = c(omega3, significant, suspect_result),
names_sep = "_"
) %>%
# Filter out rows with missing omega3 or suspect results
filter(
!is.na(omega3_foreground),
!is.na(omega3_background),
suspect_result_foreground == FALSE,
suspect_result_background == FALSE
) %>%
# Log-transform omega3
mutate(
log_omega3_fore = log10(omega3_foreground + 1e-3),
log_omega3_back = log10(omega3_background + 1e-3),
color_status = case_when(
significant_foreground ~ "Positive Selection",
TRUE ~ "Not Significant"
)
)
# ========== Plot ==========
plot <- ggplot(plot_df, aes(x = log_omega3_fore, y = log_omega3_back)) +
geom_point(aes(color = color_status), size = 3, alpha = 0.8) +
scale_color_manual(values = c("Positive Selection" = "#9B59B6", "Not Significant" = "gray80")) +
labs(
x = expression(log[10]~omega[3]~"(foreground branches)"),
y = expression(log[10]~omega[3]~"(background branches)"),
color = "Selection"
) +
coord_cartesian(xlim = c(-2, 4), ylim = c(-2, 4)) +
theme_minimal(base_size = 14) +
theme(legend.position = "top")
# ========== Export ==========
ggsave("bustedPH_logomega3_scatter_nosuspect.pdf", plot, width = 8, height = 6)
# Preview
print(plot)
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Labelled Pruned
Parsing the json files
#!/usr/bin/env Rscript
# ============================= #
# LOAD LIBRARIES #
# ============================= #
suppressPackageStartupMessages({
library(tidyverse)
library(jsonlite)
library(fs)
})
# ============================= #
# PARSE ONE FILE #
# ============================= #
parse_busted <- function(file) {
tryCatch({
busted <- jsonlite::fromJSON(file)
og <- stringr::str_extract(basename(file), "^OG[0-9]+")
busted_input_file <- busted[["input"]][["file name"]]
busted_pval <- as.numeric(busted[["test results"]][["p-value"]])
# Metadata
aln_length <- busted[["input"]][["number of sites"]]
seq_count <- busted[["input"]][["number of sequences"]]
# Rate model info (Unconstrained model → Test)
rate_info <- busted[["fits"]][["Unconstrained model"]][["Rate Distributions"]][["Test"]]
omega_vals <- sapply(rate_info, function(x) as.numeric(x[["omega"]]))
prop_vals <- sapply(rate_info, function(x) as.numeric(x[["proportion"]]))
omega3 <- ifelse(length(omega_vals) >= 3, omega_vals[3], NA)
prop3 <- ifelse(length(prop_vals) >= 3, prop_vals[3], NA)
suspect <- is.na(omega3) || omega3 > 1000 || prop3 < 0.001 || aln_length < 100 || seq_count < 4
tibble(
file = basename(file),
input_file = busted_input_file,
orthogroup = og,
seq_count = seq_count,
aln_length = aln_length,
omega1 = omega_vals[1],
prop1 = prop_vals[1],
omega2 = omega_vals[2],
prop2 = prop_vals[2],
omega3 = omega3,
prop_sites = prop3,
pval = busted_pval,
padj = p.adjust(busted_pval, method = "BH"),
significant = p.adjust(busted_pval, method = "BH") < 0.05,
suspect_result = suspect
)
}, error = function(e) {
message("⚠️ Error parsing: ", file, " → ", e$message)
return(NULL)
})
}
# ============================= #
# RUN SCRIPT #
# ============================= #
# Define your directories
foreground_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/8_4_BustedResults_labeled_Pruned/Locusts/foreground/"
background_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/8_4_BustedResults_labeled_Pruned/Locusts/background/"
output_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedBUSTEDResults_labeled_Pruned"
# Create output folder
dir_create(output_dir)
# Find all files
fg_files <- dir_ls(foreground_dir, glob = "*.json")
bg_files <- dir_ls(background_dir, glob = "*.json")
# Parse all
parsed_fg <- purrr::map_dfr(fg_files, parse_busted) %>% mutate(dataset = "foreground")
parsed_bg <- purrr::map_dfr(bg_files, parse_busted) %>% mutate(dataset = "background")
# Merge
results_all <- bind_rows(parsed_fg, parsed_bg)
# Save
write_csv(results_all, file.path(output_dir, "busted_results_labeled_full.csv"))
message("✅ Parsed labeled BUSTED results for ", nrow(results_all), " genes.")
library(tidyverse)
# Load the parsed results
busted_df <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedBUSTEDResults_labeled_Pruned/busted_results_labeled_full.csv")
# Pivot all needed columns
busted_summary <- busted_df %>%
dplyr::select(
orthogroup, dataset,
significant, suspect_result,
seq_count, aln_length,
omega1, omega2, omega3,
prop1, prop2, prop_sites,
pval, padj
) %>%
distinct() %>%
pivot_wider(
names_from = dataset,
values_from = c(
significant, suspect_result,
seq_count, aln_length,
omega1, omega2, omega3,
prop1, prop2, prop_sites,
pval, padj
),
names_glue = "{.value}_{dataset}",
values_fill = NA
) %>%
mutate(
selection_status = case_when(
significant_foreground & !significant_background ~ "Foreground only",
!significant_foreground & significant_background ~ "Background only",
significant_foreground & significant_background ~ "Both foreground and background",
TRUE ~ "No selection"
),
suspect_result = suspect_result_foreground | suspect_result_background
) %>%
dplyr::select(
orthogroup, selection_status, suspect_result,
pval_foreground, padj_foreground,
pval_background, padj_background,
starts_with("seq_count"), starts_with("aln_length"),
starts_with("omega"), starts_with("prop")
)
# View or write to file
print(busted_summary)
write_csv(
busted_summary,
"/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedBUSTEDResults_labeled_Pruned/busted_orthogroup_summary.csv"
)
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Plotting fg vs bg
# ========== Load Libraries ==========
library(tidyverse)
# ========== Load Data ==========
busted_df <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedBUSTEDResults_labeled_Pruned/busted_results_labeled_full.csv")
# ========== Data Preparation ==========
# Pivot to wide format
plot_df <- busted_df %>%
dplyr::select(orthogroup, dataset, omega3, significant, suspect_result) %>%
pivot_wider(
names_from = dataset,
values_from = c(omega3, significant, suspect_result),
names_sep = "_"
) %>%
# Filter out rows with missing omega3 or suspect results
filter(
!is.na(omega3_foreground),
!is.na(omega3_background),
suspect_result_foreground == FALSE,
suspect_result_background == FALSE
) %>%
# Log-transform omega3
mutate(
log_omega3_fore = log10(omega3_foreground + 1e-3),
log_omega3_back = log10(omega3_background + 1e-3),
color_status = case_when(
significant_foreground ~ "Positive Selection",
TRUE ~ "Not Significant"
)
)
# ========== Plot ==========
plot <- ggplot(plot_df, aes(x = log_omega3_fore, y = log_omega3_back)) +
geom_point(aes(color = color_status), size = 3, alpha = 0.8) +
scale_color_manual(values = c("Positive Selection" = "#9B59B6", "Not Significant" = "gray80")) +
labs(
x = expression(log[10]~omega[3]~"(foreground branches)"),
y = expression(log[10]~omega[3]~"(background branches)"),
color = "Selection"
) +
coord_cartesian(xlim = c(-2, 4), ylim = c(-2, 4)) +
theme_minimal(base_size = 14) +
theme(legend.position = "top")
# ========== Export ==========
ggsave("bustedPH_logomega3_scatter_nosuspect.pdf", plot, width = 8, height = 6)
# Preview
print(plot)
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
9. RELAX
Running the analysis
We will perform RELAX analysis to check whether the foreground branches that have experienced selection were intensifying or relaxing:
# For Polyneoptera
# For labelled phylogeny
sbatch ./scripts/RunRELAX_labeled_May2025.sh \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/9_1_LabelledPhylogenies/Locusts \
Locusts \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Orthogroups/Orthogroups_SingleCopyOrthologues_selanalysiswide.txt
# For labelled phylogeny PRUNED
sbatch ./scripts/RunRELAX_labeled_June2025.sh \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/9_1_LabelledPhylogenies_Pruned/Locusts \
Locusts \
/scratch/group/songlab/maeva/LocustsGenomeEvolution/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Orthogroups/Orthogroups_SingleCopyOrthologues_selanalysiswide.txt Labelled
Parsing the json files
For parsing the results, you just do:
# ============ SETTINGS ============
input_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/10_1_RelaxResults"
single_copy_file <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Orthogroups/Orthogroups_SingleCopyOrthologues_selanalysiswide.txt"
output_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults"
dir.create(output_dir, showWarnings = FALSE, recursive = TRUE)
# ============ LIBRARIES ============
library(jsonlite)
library(tidyverse)
# ============ FUNCTIONS ============
parse_relax_json <- function(file) {
tryCatch({
result <- fromJSON(file)
og <- stringr::str_extract(basename(file), "^OG[0-9]+")
input_file <- result[["input"]][["file name"]]
seq_count <- result[["input"]][["number of sequences"]]
aln_length <- result[["input"]][["number of sites"]]
test_results <- result[["test results"]]
pval <- as.numeric(test_results[["p-value"]])
k_val <- as.numeric(test_results[["relaxation or intensification parameter"]])
selection_type <- case_when(
is.na(pval) ~ "Undetermined",
pval < 0.05 & k_val > 1 ~ "Intensification",
pval < 0.05 & k_val < 1 ~ "Relaxation",
TRUE ~ "No significant difference"
)
suspect <- is.na(k_val) || k_val > 10 || k_val < 0.1 || is.na(pval) || aln_length < 100 || seq_count < 4
tibble(
file = file,
input_file = input_file,
orthogroup = og,
seq_count = seq_count,
aln_length = aln_length,
k = k_val,
pval = pval,
padj = NA, # placeholder, will be set after all parsing
selection_type = selection_type,
suspect_result = suspect
)
}, error = function(e) {
message("⚠️ Error parsing ", file, ": ", e$message)
return(NULL)
})
}
relax_results <- parse_all_relax_jsons(input_dir)
if (nrow(relax_results) > 0) {
relax_results <- relax_results %>%
mutate(
padj = p.adjust(pval, method = "BH"),
significant = padj < 0.05
)
}
parse_all_relax_jsons <- function(folder) {
files <- list.files(folder, pattern = "\\.json$", full.names = TRUE)
files <- files[file.info(files)$size > 0]
results <- purrr::map_dfr(files, parse_relax_json)
results <- results %>%
mutate(
padj = p.adjust(pval, method = "BH"),
significant = padj < 0.05
)
return(results)
}
# ============ EXECUTE PARSING ============
relax_results <- parse_all_relax_jsons(input_dir)
# ============ WRITE OUTPUT ============
write_csv(relax_results, file.path(output_dir, "relax_results.csv"))
message("✅ Parsed RELAX results saved to: ", file.path(output_dir, "relax_results_csv"))
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Combined results
# Load required libraries
library(readr)
library(dplyr)
# === Define file paths ===
busted_file <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedBUSTEDResults_labeled/busted_orthogroup_summary.csv"
relax_file <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults/relax_results.csv"
# === Read in both tables ===
busted_df <- read_csv(busted_file, show_col_types = FALSE)
relax_df <- read_csv(relax_file, show_col_types = FALSE)
# === Left join BUSTED + RELAX results ===
combined_df <- left_join(busted_df, relax_df, by = "orthogroup")
# === Preview and save ===
head(combined_df)# A tibble: 6 × 33
orthogroup selection_status suspect_result.x pval_foreground padj_foreground
<chr> <chr> <lgl> <dbl> <dbl>
1 OG0004339 Background only FALSE 0.5 0.5
2 OG0004340 Background only FALSE 0.0744 0.0744
3 OG0004341 Foreground only TRUE 0.0000166 0.0000166
4 OG0004343 No selection FALSE 0.487 0.487
5 OG0004344 No selection TRUE 0.5 0.5
6 OG0004345 No selection FALSE 0.161 0.161
# ℹ 28 more variables: pval_background <dbl>, padj_background <dbl>,
# seq_count_foreground <dbl>, seq_count_background <dbl>,
# aln_length_foreground <dbl>, aln_length_background <dbl>,
# omega1_foreground <dbl>, omega1_background <dbl>, omega2_foreground <dbl>,
# omega2_background <dbl>, omega3_foreground <dbl>, omega3_background <dbl>,
# prop1_foreground <dbl>, prop1_background <dbl>, prop2_foreground <dbl>,
# prop2_background <dbl>, prop_sites_foreground <dbl>, …# Save combined table
write_csv(combined_df, "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults/joint_busted_relax_results.csv")
message("✅ Combined BUSTED + RELAX table saved.")✅ Combined BUSTED + RELAX table saved.
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Labelled pruned
Parsing the json files
For parsing the results, you just do:
# ============ SETTINGS ============
input_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/10_1_RelaxResults_Pruned"
single_copy_file <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/5_OrthoFinder/fasta/Results_May26_iqtree/Orthogroups/Orthogroups_SingleCopyOrthologues_selanalysiswide.txt"
output_dir <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults_Pruned"
dir.create(output_dir, showWarnings = FALSE, recursive = TRUE)
# ============ LIBRARIES ============
library(jsonlite)
library(tidyverse)
# ============ FUNCTIONS ============
parse_relax_json <- function(file) {
tryCatch({
result <- fromJSON(file)
og <- stringr::str_extract(basename(file), "^OG[0-9]+")
input_file <- result[["input"]][["file name"]]
seq_count <- result[["input"]][["number of sequences"]]
aln_length <- result[["input"]][["number of sites"]]
test_results <- result[["test results"]]
pval <- as.numeric(test_results[["p-value"]])
k_val <- as.numeric(test_results[["relaxation or intensification parameter"]])
selection_type <- case_when(
is.na(pval) ~ "Undetermined",
pval < 0.05 & k_val > 1 ~ "Intensification",
pval < 0.05 & k_val < 1 ~ "Relaxation",
TRUE ~ "No significant difference"
)
suspect <- is.na(k_val) || k_val > 10 || k_val < 0.1 || is.na(pval) || aln_length < 100 || seq_count < 4
tibble(
file = file,
input_file = input_file,
orthogroup = og,
seq_count = seq_count,
aln_length = aln_length,
k = k_val,
pval = pval,
padj = NA, # placeholder, will be set after all parsing
selection_type = selection_type,
suspect_result = suspect
)
}, error = function(e) {
message("⚠️ Error parsing ", file, ": ", e$message)
return(NULL)
})
}
relax_results <- parse_all_relax_jsons(input_dir)
if (nrow(relax_results) > 0) {
relax_results <- relax_results %>%
mutate(
padj = p.adjust(pval, method = "BH"),
significant = padj < 0.05
)
}
parse_all_relax_jsons <- function(folder) {
files <- list.files(folder, pattern = "\\.json$", full.names = TRUE)
files <- files[file.info(files)$size > 0]
results <- purrr::map_dfr(files, parse_relax_json)
results <- results %>%
mutate(
padj = p.adjust(pval, method = "BH"),
significant = padj < 0.05
)
return(results)
}
# ============ EXECUTE PARSING ============
relax_results <- parse_all_relax_jsons(input_dir)
# ============ WRITE OUTPUT ============
write_csv(relax_results, file.path(output_dir, "relax_results.csv"))
message("✅ Parsed RELAX results saved to: ", file.path(output_dir, "relax_results_csv"))
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Combined results
# Load required libraries
library(readr)
library(dplyr)
# === Define file paths ===
busted_file <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedBUSTEDResults_labeled_Pruned/busted_orthogroup_summary.csv"
relax_file <- "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults_Pruned/relax_results.csv"
# === Read in both tables ===
busted_df <- read_csv(busted_file, show_col_types = FALSE)
relax_df <- read_csv(relax_file, show_col_types = FALSE)
# === Left join BUSTED + RELAX results ===
combined_df <- left_join(busted_df, relax_df, by = "orthogroup")
# === Preview and save ===
head(combined_df)# A tibble: 6 × 33
orthogroup selection_status suspect_result.x pval_foreground padj_foreground
<chr> <chr> <lgl> <dbl> <dbl>
1 OG0004339 No selection TRUE 0.5 0.5
2 OG0004340 Background only FALSE 0.188 0.188
3 OG0004341 Foreground only TRUE 0.0000218 0.0000218
4 OG0004343 No selection FALSE 0.0943 0.0943
5 OG0004344 No selection TRUE 0.114 0.114
6 OG0004345 Foreground only FALSE 0.0378 0.0378
# ℹ 28 more variables: pval_background <dbl>, padj_background <dbl>,
# seq_count_foreground <dbl>, seq_count_background <dbl>,
# aln_length_foreground <dbl>, aln_length_background <dbl>,
# omega1_foreground <dbl>, omega1_background <dbl>, omega2_foreground <dbl>,
# omega2_background <dbl>, omega3_foreground <dbl>, omega3_background <dbl>,
# prop1_foreground <dbl>, prop1_background <dbl>, prop2_foreground <dbl>,
# prop2_background <dbl>, prop_sites_foreground <dbl>, …# Save combined table
write_csv(combined_df, "/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults_Pruned/joint_busted_relax_results.csv")
message("✅ Combined BUSTED + RELAX table saved.")✅ Combined BUSTED + RELAX table saved.
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
10. Combined BUSTED and RELAX
Full tree
# Load libraries
library(readr)
library(dplyr)
library(ComplexUpset)
library(ggplot2)
# Load your dataset
df <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults/joint_busted_relax_annotated.csv", show_col_types = FALSE)
# Recalculate selection_type from k and pval
df <- df %>%
mutate(
k_val = as.numeric(k),
pval = as.numeric(pval),
padj = as.numeric(padj),
selection_type = case_when(
is.na(pval) ~ "Undetermined",
pval < 0.05 & k_val > 1 ~ "Intensification",
pval < 0.05 & k_val < 1 ~ "Relaxation",
TRUE ~ "No significant difference"
)
)
# Filter non-suspect orthogroups
df_suspect <- df %>%
filter(selection_status != "No selection") %>%
filter(suspect_result.x == FALSE) %>%
mutate(
PSG = selection_status %in% c("Foreground only", "Both foreground and background"),
Relaxation = selection_type == "Relaxation",
Intensification = selection_type == "Intensification",
Insignificant = selection_type == "No significant difference"
) %>%
dplyr::select(orthogroup, PSG, Relaxation, Intensification, Insignificant) %>%
distinct()
# Add color groups: PSG > Relaxation > Other
df_suspect <- df_suspect %>%
mutate(
PSG_color = case_when(
PSG ~ "PSG",
Relaxation ~ "Relaxation",
Intensification ~ "Intensification",
TRUE ~ "Other"
)
)
# Define color palette
custom_colors <- c(
"PSG" = "orchid", # Green
"Relaxation" = "gold", # Orange
"Intensification" = "tomato", # Orange
"Other" = "black" # Default gray
)
# Reorder factor levels BEFORE calling upset()
df_suspect_long <- df_suspect %>%
pivot_longer(cols = c("PSG", "Relaxation", "Intensification", "Insignificant"),
names_to = "set", values_to = "value") %>%
filter(value) %>%
mutate(set = factor(set, levels = c("Insignificant", "Intensification", "Relaxation", "PSG"))) %>%
pivot_wider(names_from = set, values_from = value, values_fill = FALSE)
# Add color groups again
df_suspect_long <- df_suspect_long %>%
mutate(
PSG_color = case_when(
PSG ~ "PSG",
Relaxation ~ "Relaxation",
Intensification ~ "Intensification",
TRUE ~ "Other"
)
)
ComplexUpset::upset(
df_suspect_long,
intersect = c("Insignificant", "Intensification", "Relaxation", "PSG"),
name = "Selection Category",
sort_sets = FALSE,
sort_intersections = FALSE,
intersections=list(
c('PSG', 'Relaxation'),
c('PSG', 'Intensification'),
c('PSG', 'Insignificant'),
'Relaxation',
'Intensification',
'Insignificant'
),
# sort_intersections_by = "degree",
set_sizes = upset_set_size() +
ggtitle("Set Size") +
theme_minimal(),
base_annotations = list(
'Intersection size' = intersection_size(
mapping = aes(fill = PSG_color),
text = list(size = 3),
bar_number_threshold = 10
) +
scale_fill_manual(values = custom_colors, guide = "none")
),
queries=list(
upset_query(
intersect=c('PSG', 'Relaxation'),
color='orchid',
fill='orchid',
only_components=c('intersections_matrix', 'Intersection size')
),
upset_query(
intersect=c('PSG', 'Intensification'),
color='orchid',
fill='orchid',
only_components=c('intersections_matrix', 'Intersection size')
),
upset_query(
intersect=c('PSG', 'Insignificant'),
color='orchid',
fill='orchid',
only_components=c('intersections_matrix', 'Intersection size')
),
upset_query(
intersect=c('Relaxation'),
color='gold',
fill='gold',
only_components=c('intersections_matrix', 'Intersection size')
),
upset_query(
intersect=c('Intensification'),
color='tomato',
fill='tomato',
only_components=c('intersections_matrix', 'Intersection size')
),
upset_query(set='PSG', fill='orchid'),
upset_query(set='Relaxation', fill='gold'),
upset_query(set='Intensification', fill='tomato'),
upset_query(set='Insignificant', fill='black')
)) +
ggtitle("Genes under positive selection in swarming species (Full tree)") +
theme_minimal(base_size = 12)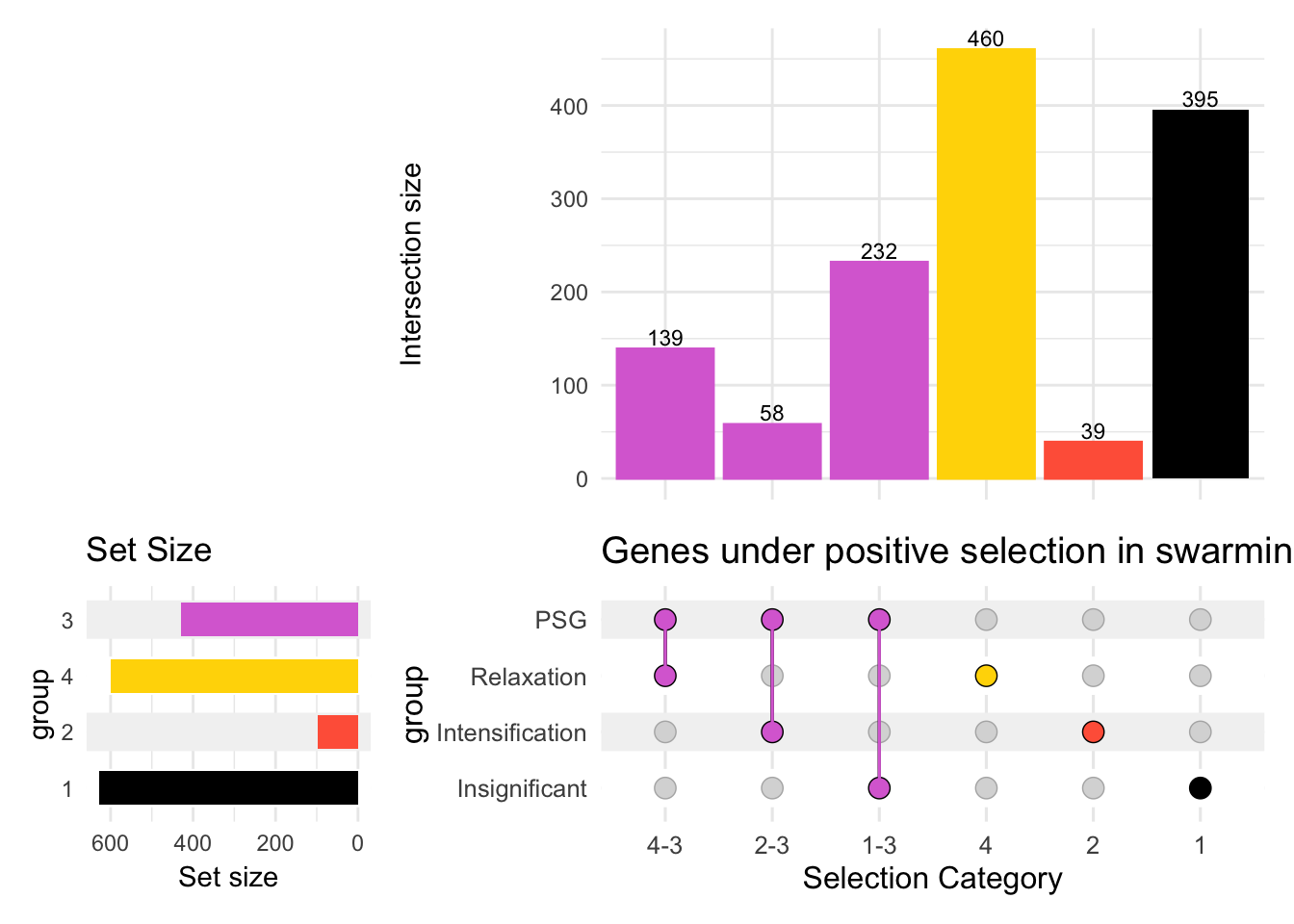
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Pruned tree
# Load libraries
library(readr)
library(dplyr)
library(ComplexUpset)
library(ggplot2)
# Load your dataset
df <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults_Pruned/joint_busted_relax_annotated.csv", show_col_types = FALSE)
# Recalculate selection_type from k and pval
df <- df %>%
mutate(
k_val = as.numeric(k),
pval = as.numeric(pval),
padj = as.numeric(padj),
selection_type = case_when(
is.na(pval) ~ "Undetermined",
pval < 0.05 & k_val > 1 ~ "Intensification",
pval < 0.05 & k_val < 1 ~ "Relaxation",
TRUE ~ "No significant difference"
)
)
# Filter non-suspect orthogroups
df_suspect <- df %>%
filter(selection_status != "No selection") %>%
filter(suspect_result.x == FALSE) %>%
mutate(
PSG = selection_status %in% c("Foreground only", "Both foreground and background"),
Relaxation = selection_type == "Relaxation",
Intensification = selection_type == "Intensification",
Insignificant = selection_type == "No significant difference"
) %>%
dplyr::select(orthogroup, PSG, Relaxation, Intensification, Insignificant) %>%
distinct()
# Add color groups: PSG > Relaxation > Other
df_suspect <- df_suspect %>%
mutate(
PSG_color = case_when(
PSG ~ "PSG",
Relaxation ~ "Relaxation",
Intensification ~ "Intensification",
TRUE ~ "Other"
)
)
# Define color palette
custom_colors <- c(
"PSG" = "orchid", # Green
"Relaxation" = "gold", # Orange
"Intensification" = "tomato", # Orange
"Other" = "black" # Default gray
)
# Reorder factor levels BEFORE calling upset()
df_suspect_long <- df_suspect %>%
pivot_longer(cols = c("PSG", "Relaxation", "Intensification", "Insignificant"),
names_to = "set", values_to = "value") %>%
filter(value) %>%
mutate(set = factor(set, levels = c("Insignificant", "Intensification", "Relaxation", "PSG"))) %>%
pivot_wider(names_from = set, values_from = value, values_fill = FALSE)
# Add color groups again
df_suspect_long <- df_suspect_long %>%
mutate(
PSG_color = case_when(
PSG ~ "PSG",
Relaxation ~ "Relaxation",
Intensification ~ "Intensification",
TRUE ~ "Other"
)
)
ComplexUpset::upset(
df_suspect_long,
intersect = c("Insignificant", "Intensification", "Relaxation", "PSG"),
name = "Selection Category",
sort_sets = FALSE,
sort_intersections = FALSE,
intersections=list(
c('PSG', 'Relaxation'),
c('PSG', 'Intensification'),
c('PSG', 'Insignificant'),
'Relaxation',
'Intensification',
'Insignificant'
),
# sort_intersections_by = "degree",
set_sizes = upset_set_size() +
ggtitle("Set Size") +
theme_minimal(),
base_annotations = list(
'Intersection size' = intersection_size(
mapping = aes(fill = PSG_color),
text = list(size = 3),
bar_number_threshold = 10
) +
scale_fill_manual(values = custom_colors, guide = "none")
),
queries=list(
upset_query(
intersect=c('PSG', 'Relaxation'),
color='orchid',
fill='orchid',
only_components=c('intersections_matrix', 'Intersection size')
),
upset_query(
intersect=c('PSG', 'Intensification'),
color='orchid',
fill='orchid',
only_components=c('intersections_matrix', 'Intersection size')
),
upset_query(
intersect=c('PSG', 'Insignificant'),
color='orchid',
fill='orchid',
only_components=c('intersections_matrix', 'Intersection size')
),
upset_query(
intersect=c('Relaxation'),
color='gold',
fill='gold',
only_components=c('intersections_matrix', 'Intersection size')
),
upset_query(
intersect=c('Intensification'),
color='tomato',
fill='tomato',
only_components=c('intersections_matrix', 'Intersection size')
),
upset_query(set='PSG', fill='orchid'),
upset_query(set='Relaxation', fill='gold'),
upset_query(set='Intensification', fill='tomato'),
upset_query(set='Insignificant', fill='black')
)) +
ggtitle("Genes under positive selection in swarming species (Pruned tree)") +
theme_minimal(base_size = 12)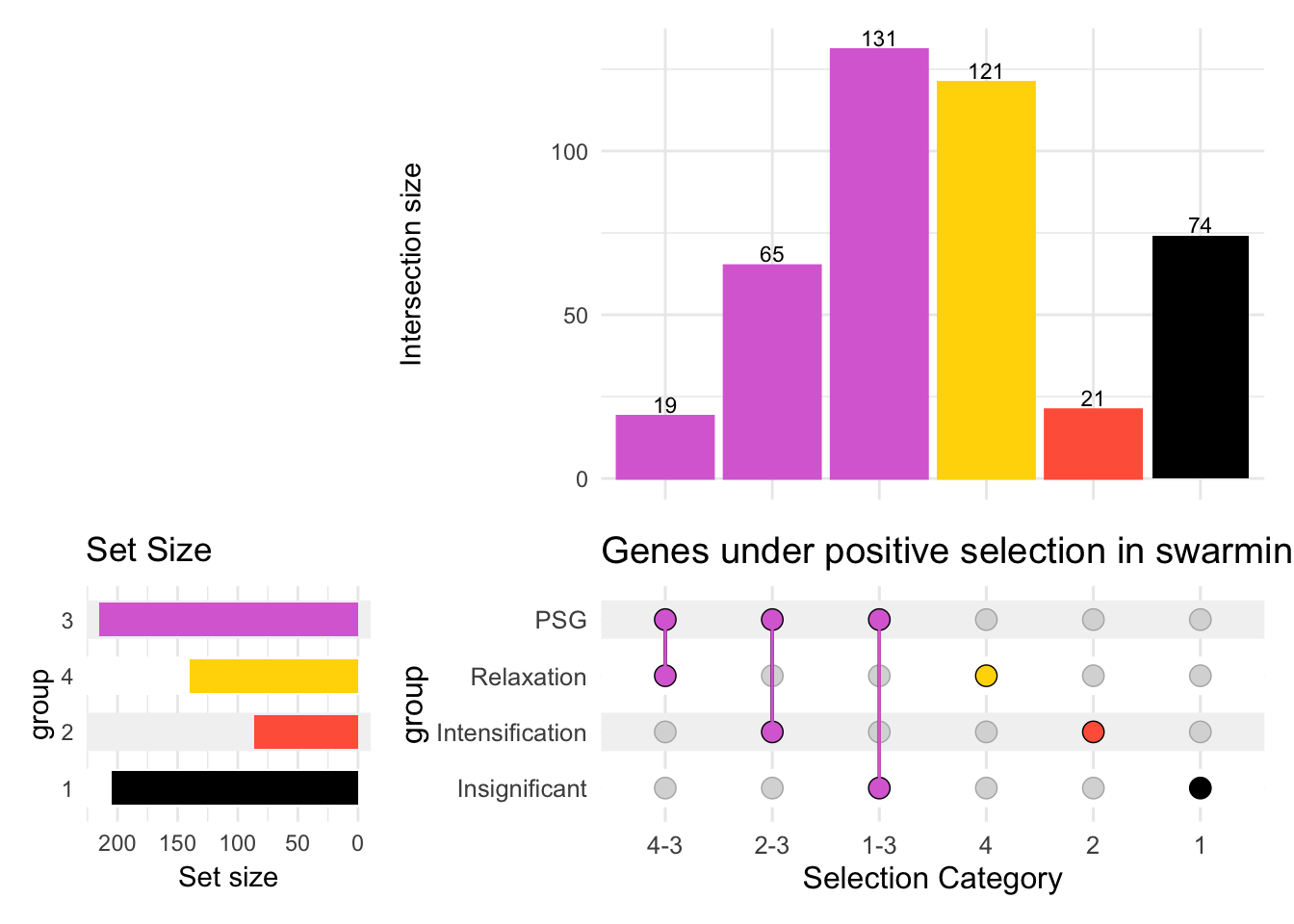
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Venn diagram
# Load required packages
library(readr)
library(dplyr)
library(ggvenn)Loading required package: grid
Attaching package: 'grid'The following object is masked from 'package:Biostrings':
pattern# Load Full tree dataset
df_full <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults/joint_busted_relax_annotated.csv", show_col_types = FALSE)
# Load Pruned tree dataset
df_pruned <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults_Pruned/joint_busted_relax_annotated.csv", show_col_types = FALSE)
# Define helper function to get PSG + Intensification orthogroups
get_psg_intensified <- function(df) {
df %>%
mutate(
k_val = as.numeric(k),
pval = as.numeric(pval),
selection_type = case_when(
is.na(pval) ~ "Undetermined",
pval < 0.05 & k_val > 1 ~ "Intensification",
pval < 0.05 & k_val < 1 ~ "Relaxation",
TRUE ~ "No significant difference"
)
) %>%
filter(suspect_result.x == FALSE) %>%
filter(selection_status %in% c("Foreground only", "Both foreground and background")) %>%
filter(selection_type == "Intensification") %>%
pull(orthogroup) %>%
unique()
}
# Get orthogroup sets
psg_intens_full <- get_psg_intensified(df_full)
psg_intens_pruned <- get_psg_intensified(df_pruned)
# Create named list for Venn input
venn_input <- list(
FullTree = psg_intens_full,
PrunedTree = psg_intens_pruned
)
# Plot Venn diagram
ggvenn(venn_input,
fill_color = c("#66c2a5", "#fc8d62"),
stroke_size = 0.7,
set_name_size = 5,
text_size = 4,
show_percentage = FALSE) +
ggtitle("PSGs under Intensification in swarming locusts (Full vs Pruned Tree)")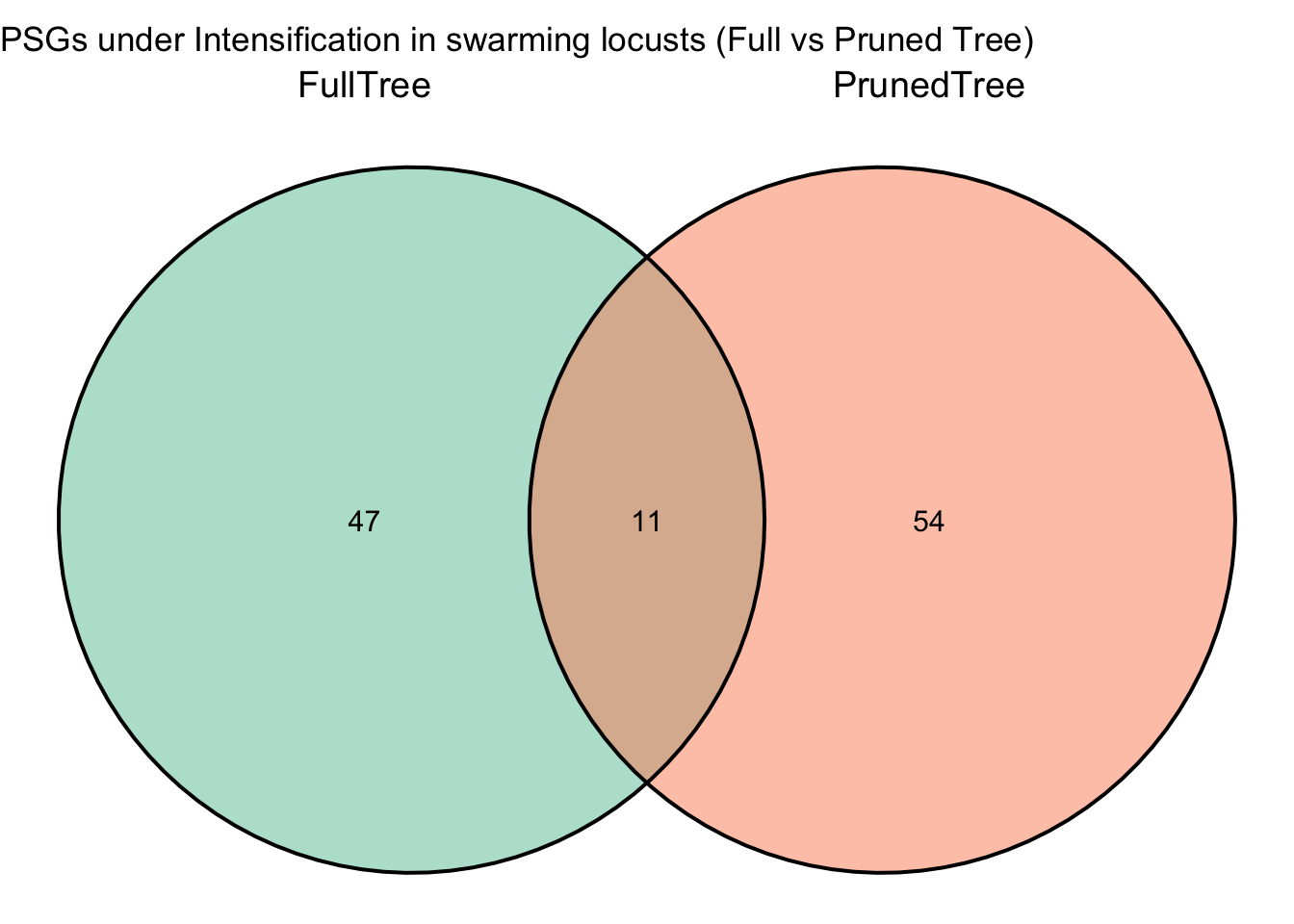
# Get the intersection of PSGs under intensification
intersect_psg_intens <- intersect(psg_intens_full, psg_intens_pruned)
# Show the result
length(intersect_psg_intens) # Number of shared orthogroups[1] 11intersect_psg_intens # List of orthogroup IDs [1] "OG0007006" "OG0007326" "OG0011236" "OG0011677" "OG0011869" "OG0011931"
[7] "OG0012017" "OG0012034" "OG0012129" "OG0012163" "OG0012186"
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
For relaxation PSGs:
# Load required packages
library(readr)
library(dplyr)
library(ggvenn)
# Load Full tree dataset
df_full <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults/joint_busted_relax_annotated.csv", show_col_types = FALSE)
# Load Pruned tree dataset
df_pruned <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults_Pruned/joint_busted_relax_annotated.csv", show_col_types = FALSE)
# Define helper function to get PSG + Intensification orthogroups
get_psg_relaxed <- function(df) {
df %>%
mutate(
k_val = as.numeric(k),
pval = as.numeric(pval),
selection_type = case_when(
is.na(pval) ~ "Undetermined",
pval < 0.05 & k_val > 1 ~ "Intensification",
pval < 0.05 & k_val < 1 ~ "Relaxation",
TRUE ~ "No significant difference"
)
) %>%
filter(suspect_result.x == FALSE) %>%
filter(selection_status %in% c("Foreground only", "Both foreground and background")) %>%
filter(selection_type == "Relaxation") %>%
pull(orthogroup) %>%
unique()
}
# Get orthogroup sets
psg_relax_full <- get_psg_relaxed(df_full)
psg_relax_pruned <- get_psg_relaxed(df_pruned)
# Create named list for Venn input
venn_input <- list(
FullTree = psg_relax_full,
PrunedTree = psg_relax_pruned
)
# Plot Venn diagram
ggvenn(venn_input,
fill_color = c("#66c2a5", "#fc8d62"),
stroke_size = 0.7,
set_name_size = 5,
text_size = 4,
show_percentage = FALSE) +
ggtitle("PSGs under Relaxation in swarming locusts (Full vs Pruned Tree)")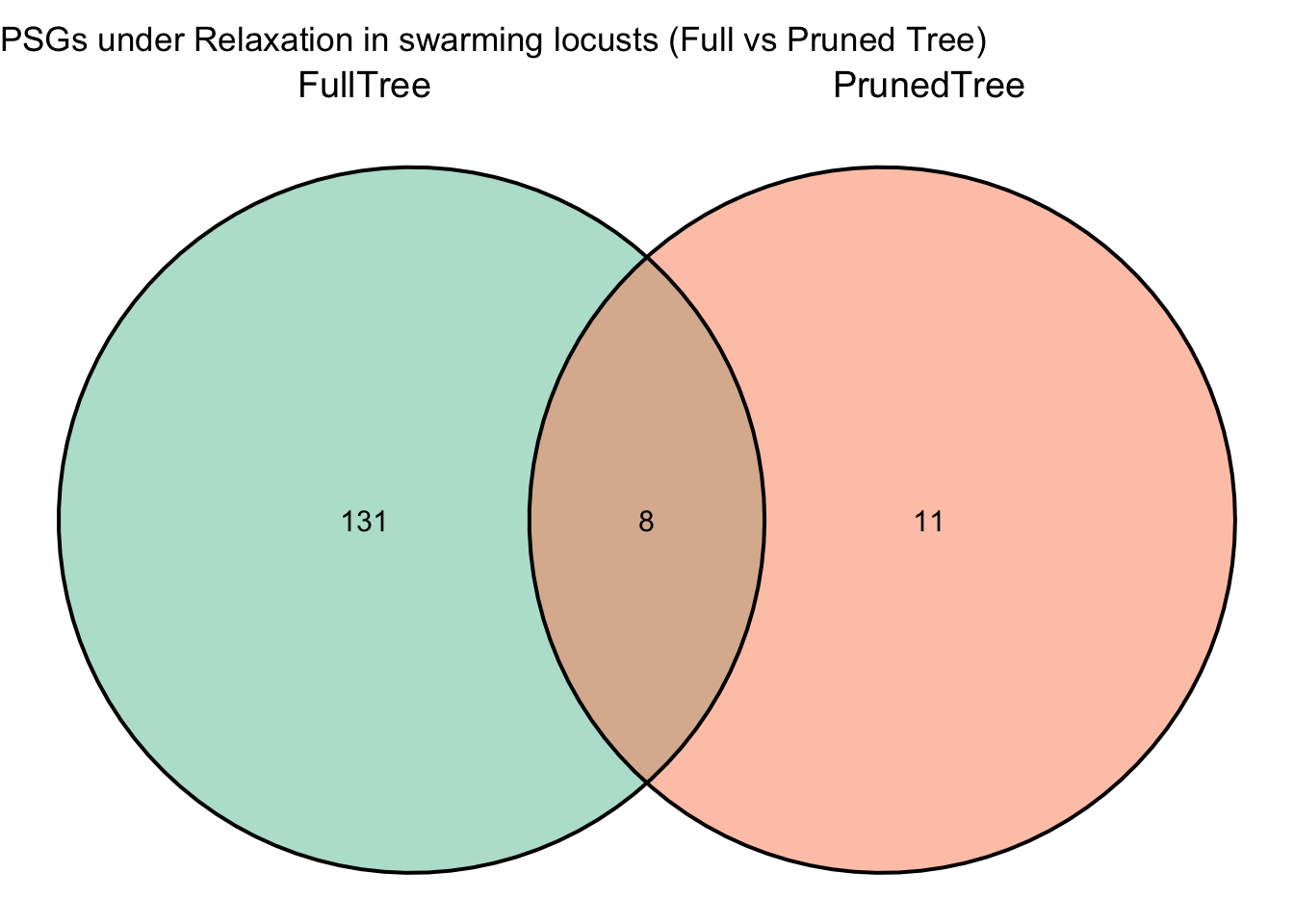
# Get the intersection of PSGs under intensification
intersect_psg_relax <- intersect(psg_relax_full, psg_relax_pruned)
# Show the result
length(intersect_psg_relax) # Number of shared orthogroups[1] 8intersect_psg_relax # List of orthogroup IDs[1] "OG0008483" "OG0008811" "OG0009185" "OG0011592" "OG0011838" "OG0011966"
[7] "OG0012059" "OG0012354"
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
11. DEG and PSGs
Full tree
Here we want to know if the PSG from the BUSTED+RELAX detection share an overlap with DEGs. We extracted the list of Orthogroup DEGs (shared by the three locusts) for this step:
# Install if needed
# install.packages("VennDiagram")
library(VennDiagram)Loading required package: futile.loggerlibrary(dplyr)
library(grid)
thorax <- c(
"OG0000679", "OG0004144", "OG0000104", "OG0000015", "OG0014033", "OG0012058",
"OG0000357", "OG0000090", "OG0003126", "OG0004306", "OG0000001", "OG0009321",
"OG0015157", "OG0012103", "OG0004309", "OG0000346", "OG0000014", "OG0000796",
"OG0000151", "OG0011174", "OG0013138", "OG0000003", "OG0000391", "OG0007893",
"OG0001029", "OG0010128", "OG0001327", "OG0011184", "OG0000615", "OG0000922",
"OG0000095", "OG0012329", "OG0010429", "OG0010105", "OG0012400", "OG0012312",
"OG0000012", "OG0005229", "OG0000532", "OG0000027", "OG0000017", "OG0009757",
"OG0002737", "OG0002903", "OG0011877", "OG0003752", "OG0007957", "OG0000000",
"OG0009700", "OG0011824", "OG0000043", "OG0011853", "OG0000347", "OG0005741",
"OG0003890", "OG0000334", "OG0000419", "OG0002738", "OG0000065", "OG0000130",
"OG0000327", "OG0000093"
)
head <- c(
"OG0000104", "OG0000015", "OG0000105", "OG0004296", "OG0000371", "OG0003126",
"OG0004306", "OG0012979", "OG0009321", "OG0000073", "OG0002734", "OG0015157",
"OG0000796", "OG0000151", "OG0000222", "OG0000296", "OG0000391", "OG0000196",
"OG0010128", "OG0009808", "OG0000788", "OG0000922", "OG0000120", "OG0000212",
"OG0002726", "OG0007990", "OG0003478", "OG0000027", "OG0002737", "OG0011877",
"OG0007957", "OG0009113", "OG0000334", "OG0000823", "OG0002738", "OG0000093",
"OG0000552", "OG0000553"
)
all_degs <- unique(c(thorax, head))
# Load Pruned tree dataset
df_pruned <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults/joint_busted_relax_annotated.csv", show_col_types = FALSE)
# Recalculate selection_type from k and pval
df <- df_pruned %>%
mutate(
k_val = as.numeric(k),
pval = as.numeric(pval),
padj = as.numeric(padj),
selection_type = case_when(
is.na(pval) ~ "Undetermined",
pval < 0.05 & k_val > 1 ~ "Intensification",
pval < 0.05 & k_val < 1 ~ "Relaxation",
TRUE ~ "No significant difference"
)
)
# Filter non-suspect orthogroups
df_suspect <- df %>%
filter(selection_status != "No selection") %>%
filter(suspect_result.x == FALSE) %>%
mutate(
PSG = selection_status %in% c("Foreground only", "Both foreground and background"),
Relaxation = selection_type == "Relaxation",
Intensification = selection_type == "Intensification",
Insignificant = selection_type == "No significant difference"
) %>%
dplyr::select(orthogroup, PSG, Relaxation, Intensification, Insignificant) %>%
distinct()
# Add color groups: PSG > Relaxation > Other
df_suspect <- df_suspect %>%
mutate(
PSG_color = case_when(
PSG ~ "PSG",
Relaxation ~ "Relaxation",
Intensification ~ "Intensification",
TRUE ~ "Other"
)
)
# Define color palette
custom_colors <- c(
"PSG" = "orchid", # Green
"Relaxation" = "gold", # Orange
"Intensification" = "tomato", # Orange
"Other" = "black" # Default gray
)
# Reorder factor levels BEFORE calling upset()
df_suspect_long <- df_suspect %>%
pivot_longer(cols = c("PSG", "Relaxation", "Intensification", "Insignificant"),
names_to = "set", values_to = "value") %>%
filter(value) %>%
mutate(set = factor(set, levels = c("Insignificant", "Intensification", "Relaxation", "PSG"))) %>%
pivot_wider(names_from = set, values_from = value, values_fill = FALSE)
# Add color groups again
df_suspect_long <- df_suspect_long %>%
mutate(
PSG_color = case_when(
PSG ~ "PSG",
Relaxation ~ "Relaxation",
Intensification ~ "Intensification",
TRUE ~ "Other"
)
)
# Extract PSGs from your annotated dataframe
psgs <- df_suspect_long %>%
filter(PSG) %>%
pull(orthogroup) %>%
unique()
# Draw the Venn diagram
venn.plot <- draw.pairwise.venn(
area1 = length(psgs),
area2 = length(all_degs),
cross.area = length(intersect(psgs, all_degs)),
category = c("PSGs", "DEGs"),
fill = c("orchid", "deepskyblue"),
lty = "blank",
cex = 2,
cat.cex = 2,
cat.pos = c(-20, 20),
cat.dist = c(0.05, 0.05)
)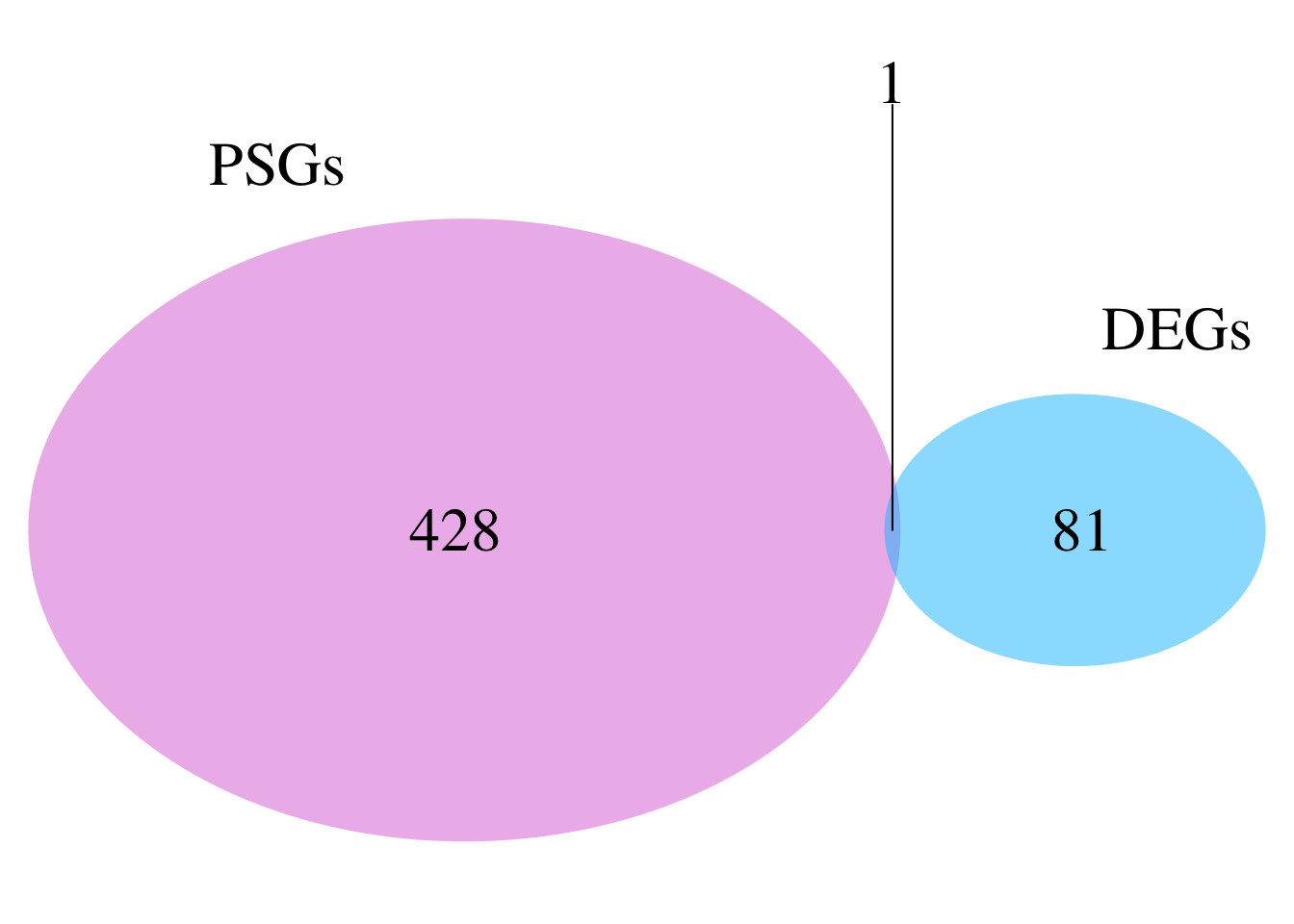
# Display the plot
grid.newpage()
grid.draw(venn.plot)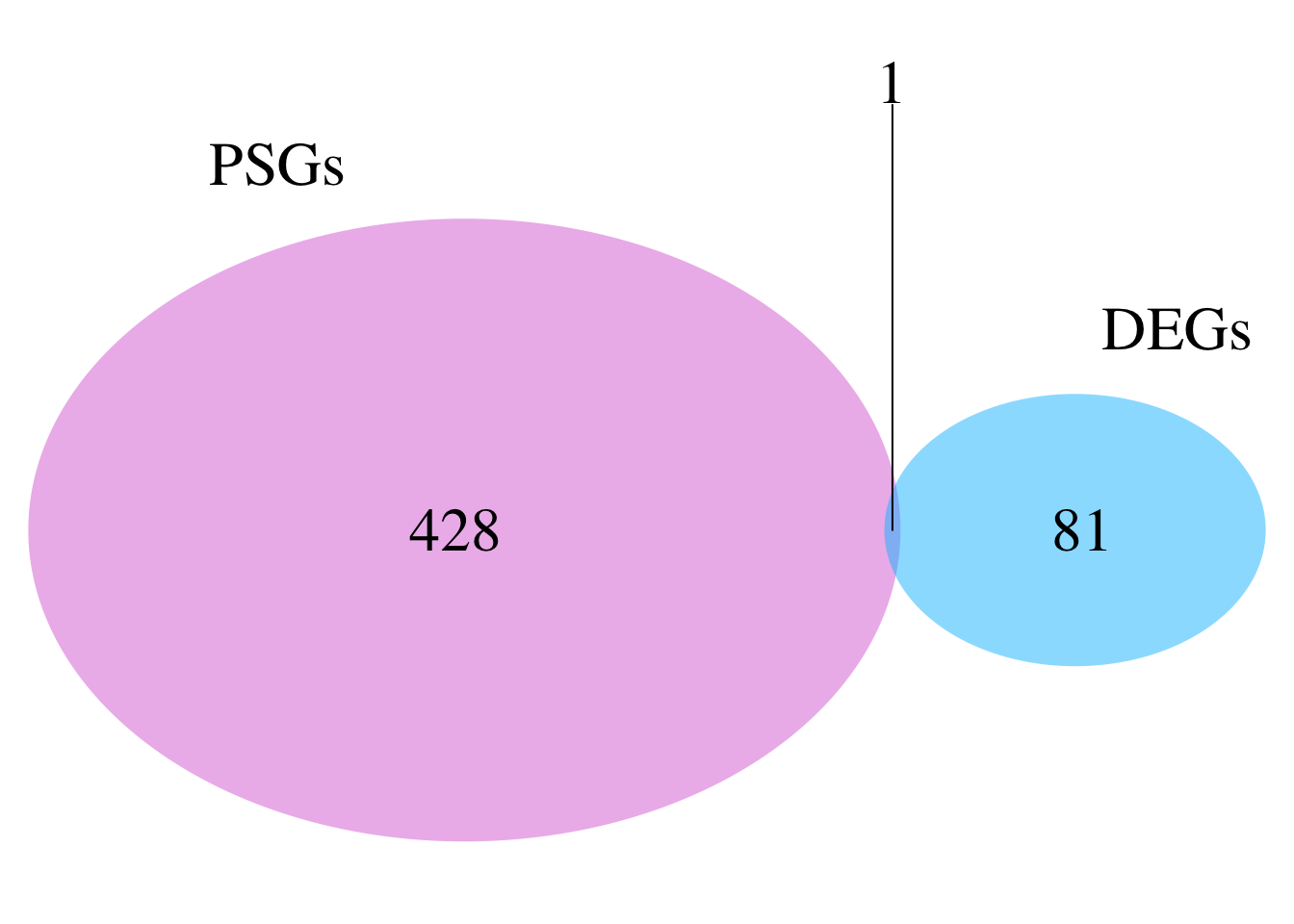
# Get the intersection of PSGs under intensification
intersect_psg_deg <- intersect(psgs, all_degs)
# Show the result
length(intersect_psg_deg) # Number of shared orthogroups[1] 1intersect_psg_deg # List of orthogroup IDs[1] "OG0012312"
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
Pruned tree
Here we want to know if the PSG from the BUSTED+RELAX detection share an overlap with DEGs. We extracted the list of Orthogroup DEGs (shared by the three locusts) for this step:
# Install if needed
# install.packages("VennDiagram")
library(VennDiagram)
library(dplyr)
library(grid)
thorax <- c(
"OG0000679", "OG0004144", "OG0000104", "OG0000015", "OG0014033", "OG0012058",
"OG0000357", "OG0000090", "OG0003126", "OG0004306", "OG0000001", "OG0009321",
"OG0015157", "OG0012103", "OG0004309", "OG0000346", "OG0000014", "OG0000796",
"OG0000151", "OG0011174", "OG0013138", "OG0000003", "OG0000391", "OG0007893",
"OG0001029", "OG0010128", "OG0001327", "OG0011184", "OG0000615", "OG0000922",
"OG0000095", "OG0012329", "OG0010429", "OG0010105", "OG0012400", "OG0012312",
"OG0000012", "OG0005229", "OG0000532", "OG0000027", "OG0000017", "OG0009757",
"OG0002737", "OG0002903", "OG0011877", "OG0003752", "OG0007957", "OG0000000",
"OG0009700", "OG0011824", "OG0000043", "OG0011853", "OG0000347", "OG0005741",
"OG0003890", "OG0000334", "OG0000419", "OG0002738", "OG0000065", "OG0000130",
"OG0000327", "OG0000093"
)
head <- c(
"OG0000104", "OG0000015", "OG0000105", "OG0004296", "OG0000371", "OG0003126",
"OG0004306", "OG0012979", "OG0009321", "OG0000073", "OG0002734", "OG0015157",
"OG0000796", "OG0000151", "OG0000222", "OG0000296", "OG0000391", "OG0000196",
"OG0010128", "OG0009808", "OG0000788", "OG0000922", "OG0000120", "OG0000212",
"OG0002726", "OG0007990", "OG0003478", "OG0000027", "OG0002737", "OG0011877",
"OG0007957", "OG0009113", "OG0000334", "OG0000823", "OG0002738", "OG0000093",
"OG0000552", "OG0000553"
)
all_degs <- unique(c(thorax, head))
# Load Pruned tree dataset
df_pruned <- read_csv("/Users/maevatecher/Desktop/Polyneoptera_FINAL/ParsedRELAXResults_Pruned/joint_busted_relax_annotated.csv", show_col_types = FALSE)
# Recalculate selection_type from k and pval
df <- df_pruned %>%
mutate(
k_val = as.numeric(k),
pval = as.numeric(pval),
padj = as.numeric(padj),
selection_type = case_when(
is.na(pval) ~ "Undetermined",
pval < 0.05 & k_val > 1 ~ "Intensification",
pval < 0.05 & k_val < 1 ~ "Relaxation",
TRUE ~ "No significant difference"
)
)
# Filter non-suspect orthogroups
df_suspect <- df %>%
filter(selection_status != "No selection") %>%
filter(suspect_result.x == FALSE) %>%
mutate(
PSG = selection_status %in% c("Foreground only", "Both foreground and background"),
Relaxation = selection_type == "Relaxation",
Intensification = selection_type == "Intensification",
Insignificant = selection_type == "No significant difference"
) %>%
dplyr::select(orthogroup, PSG, Relaxation, Intensification, Insignificant) %>%
distinct()
# Add color groups: PSG > Relaxation > Other
df_suspect <- df_suspect %>%
mutate(
PSG_color = case_when(
PSG ~ "PSG",
Relaxation ~ "Relaxation",
Intensification ~ "Intensification",
TRUE ~ "Other"
)
)
# Define color palette
custom_colors <- c(
"PSG" = "orchid", # Green
"Relaxation" = "gold", # Orange
"Intensification" = "tomato", # Orange
"Other" = "black" # Default gray
)
# Reorder factor levels BEFORE calling upset()
df_suspect_long <- df_suspect %>%
pivot_longer(cols = c("PSG", "Relaxation", "Intensification", "Insignificant"),
names_to = "set", values_to = "value") %>%
filter(value) %>%
mutate(set = factor(set, levels = c("Insignificant", "Intensification", "Relaxation", "PSG"))) %>%
pivot_wider(names_from = set, values_from = value, values_fill = FALSE)
# Add color groups again
df_suspect_long <- df_suspect_long %>%
mutate(
PSG_color = case_when(
PSG ~ "PSG",
Relaxation ~ "Relaxation",
Intensification ~ "Intensification",
TRUE ~ "Other"
)
)
# Extract PSGs from your annotated dataframe
psgs <- df_suspect_long %>%
filter(PSG) %>%
pull(orthogroup) %>%
unique()
# Draw the Venn diagram
venn.plot <- draw.pairwise.venn(
area1 = length(psgs),
area2 = length(all_degs),
cross.area = length(intersect(psgs, all_degs)),
category = c("PSGs", "DEGs"),
fill = c("orchid", "deepskyblue"),
lty = "blank",
cex = 4,
cat.cex = 2,
cat.pos = c(-20, 20),
cat.dist = c(0.05, 0.05)
)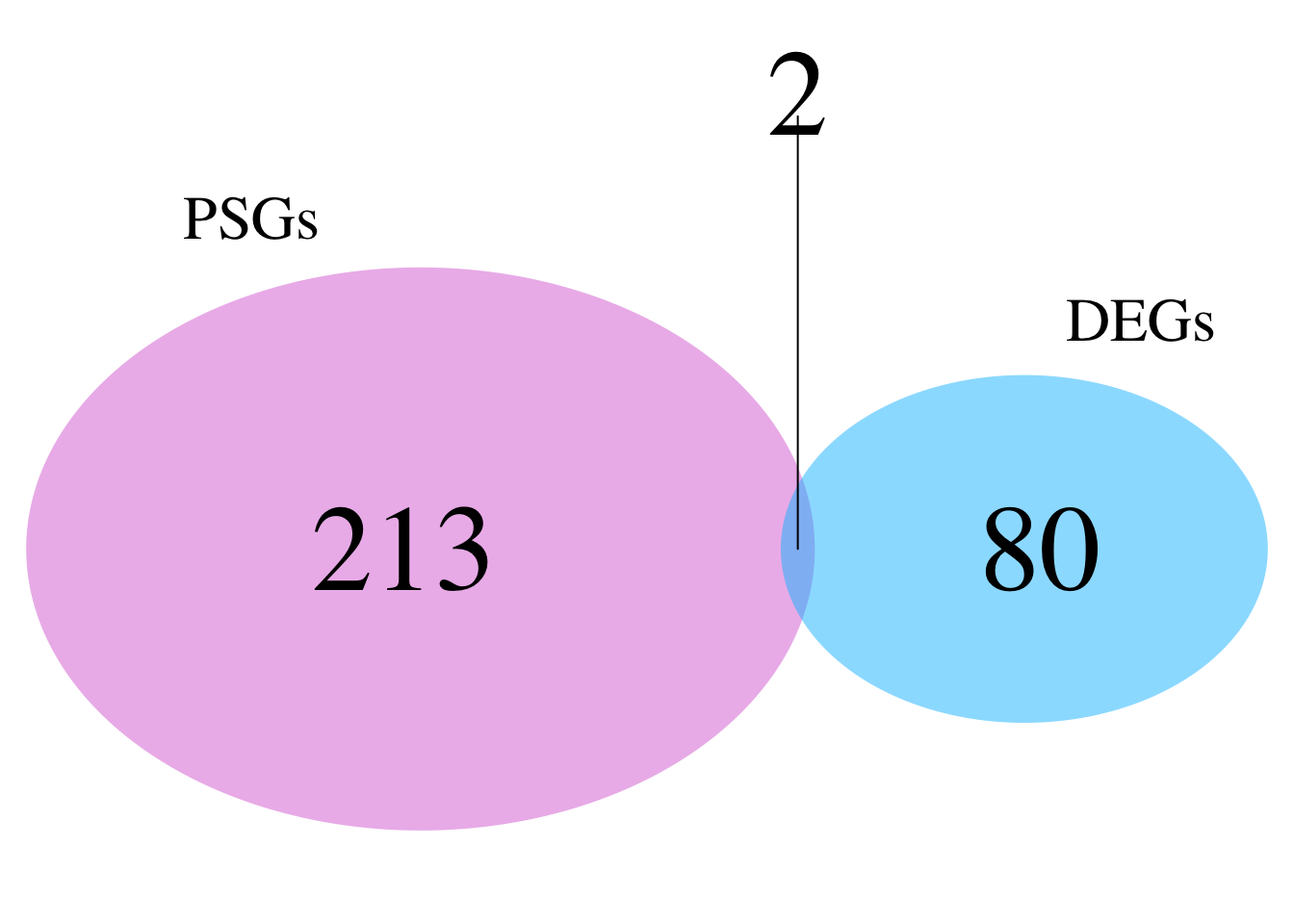
# Display the plot
grid.newpage()
grid.draw(venn.plot)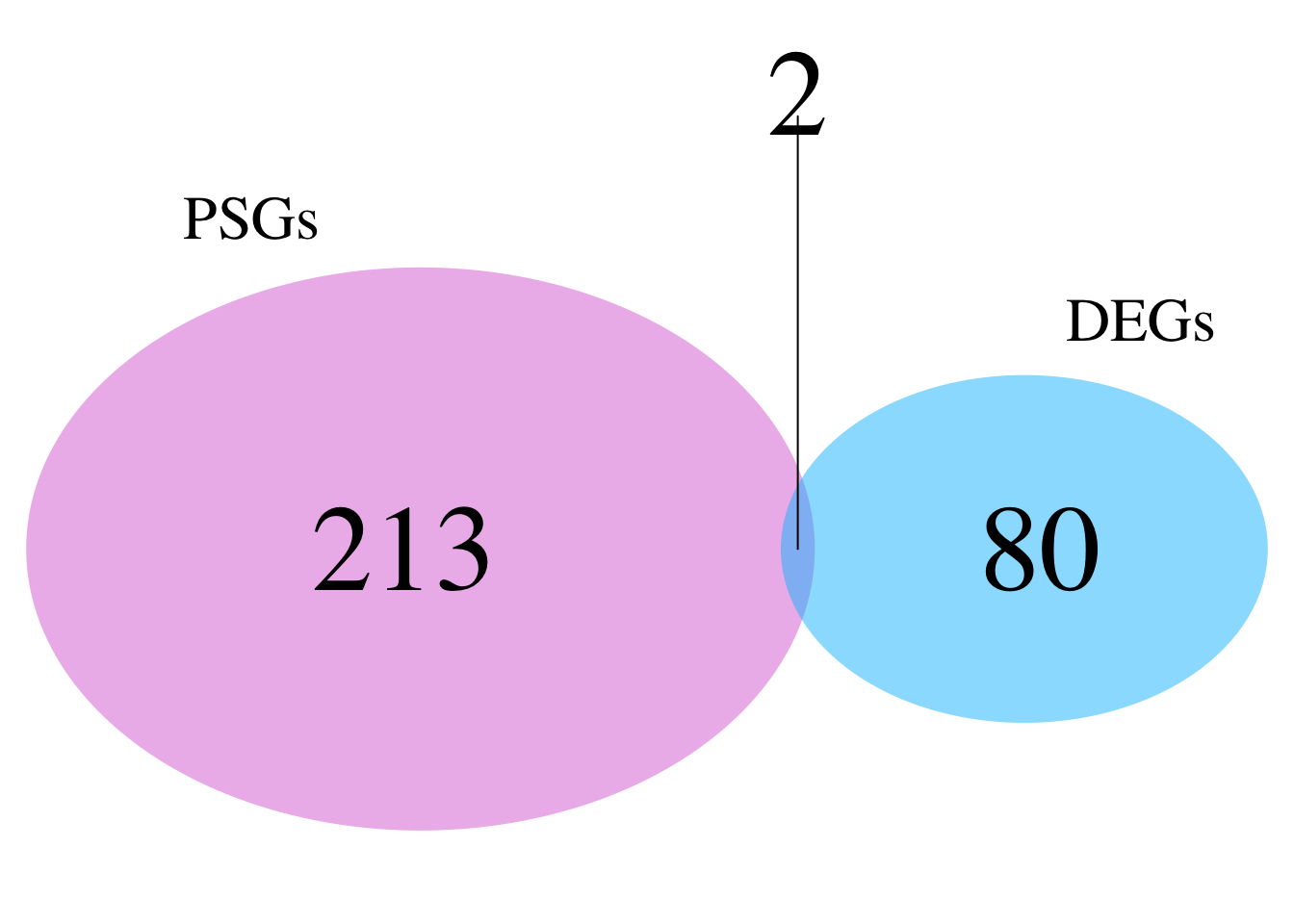
# Get the intersection of PSGs under intensification
intersect_psg_deg <- intersect(psgs, all_degs)
# Show the result
length(intersect_psg_deg) # Number of shared orthogroups[1] 2intersect_psg_deg # List of orthogroup IDs[1] "OG0010128" "OG0012312"
Warning: The above code chunk cached its results, but
it won’t be re-run if previous chunks it depends on are updated. If you
need to use caching, it is highly recommended to also set
knitr::opts_chunk$set(autodep = TRUE) at the top of the
file (in a chunk that is not cached). Alternatively, you can customize
the option dependson for each individual chunk that is
cached. Using either autodep or dependson will
remove this warning. See the
knitr cache options for more details.
sessionInfo()R version 4.4.2 (2024-10-31)
Platform: aarch64-apple-darwin20
Running under: macOS Sequoia 15.6.1
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
time zone: Asia/Tokyo
tzcode source: internal
attached base packages:
[1] grid stats4 stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] VennDiagram_1.7.3 futile.logger_1.4.3 ggvenn_0.1.10
[4] ComplexUpset_1.3.6 plotly_4.11.0 ggnewscale_0.5.2
[7] hexbin_1.28.5 lubridate_1.9.4 forcats_1.0.0
[10] purrr_1.0.4 tidyr_1.3.1 tidyverse_2.0.0
[13] tibble_3.3.0 stringr_1.5.1 jsonlite_2.0.0
[16] DesertLocustR_0.1.0 remotes_2.5.0 Biostrings_2.74.1
[19] XVector_0.46.0 AnnotationHub_3.14.0 BiocFileCache_2.14.0
[22] dbplyr_2.5.0 rtracklayer_1.66.0 GenomicRanges_1.58.0
[25] GenomeInfoDb_1.42.3 GO.db_3.20.0 AnnotationDbi_1.68.0
[28] IRanges_2.40.1 S4Vectors_0.44.0 Biobase_2.66.0
[31] BiocGenerics_0.52.0 clusterProfiler_4.14.6 data.table_1.17.6
[34] readr_2.1.5 dplyr_1.1.4 ggplot2_3.5.2
loaded via a namespace (and not attached):
[1] splines_4.4.2 later_1.4.2
[3] BiocIO_1.16.0 bitops_1.0-9
[5] ggplotify_0.1.2 filelock_1.0.3
[7] R.oo_1.27.1 XML_3.99-0.18
[9] lifecycle_1.0.4 rprojroot_2.0.4
[11] lattice_0.22-7 vroom_1.6.5
[13] magrittr_2.0.3 sass_0.4.10
[15] rmarkdown_2.29 jquerylib_0.1.4
[17] yaml_2.3.10 httpuv_1.6.16
[19] ggtangle_0.0.6 cowplot_1.1.3
[21] DBI_1.2.3 RColorBrewer_1.1-3
[23] abind_1.4-8 zlibbioc_1.52.0
[25] R.utils_2.13.0 RCurl_1.98-1.17
[27] yulab.utils_0.2.0 rappdirs_0.3.3
[29] git2r_0.36.2 GenomeInfoDbData_1.2.13
[31] enrichplot_1.26.6 ggrepel_0.9.6
[33] tidytree_0.4.6 codetools_0.2-20
[35] DelayedArray_0.32.0 DOSE_4.0.1
[37] tidyselect_1.2.1 aplot_0.2.7
[39] UCSC.utils_1.2.0 farver_2.1.2
[41] matrixStats_1.5.0 GenomicAlignments_1.42.0
[43] tools_4.4.2 treeio_1.30.0
[45] Rcpp_1.0.14 glue_1.8.0
[47] SparseArray_1.6.2 xfun_0.52
[49] qvalue_2.38.0 MatrixGenerics_1.18.1
[51] withr_3.0.2 formatR_1.14
[53] BiocManager_1.30.26 fastmap_1.2.0
[55] digest_0.6.37 timechange_0.3.0
[57] R6_2.6.1 gridGraphics_0.5-1
[59] colorspace_2.1-1 dichromat_2.0-0.1
[61] RSQLite_2.4.1 R.methodsS3_1.8.2
[63] generics_0.1.4 httr_1.4.7
[65] htmlwidgets_1.6.4 S4Arrays_1.6.0
[67] whisker_0.4.1 pkgconfig_2.0.3
[69] gtable_0.3.6 blob_1.2.4
[71] workflowr_1.7.1 htmltools_0.5.8.1
[73] fgsea_1.32.4 scales_1.4.0
[75] png_0.1-8 ggfun_0.1.9
[77] knitr_1.50 lambda.r_1.2.4
[79] rstudioapi_0.17.1 tzdb_0.5.0
[81] reshape2_1.4.4 rjson_0.2.23
[83] nlme_3.1-168 curl_6.4.0
[85] cachem_1.1.0 BiocVersion_3.20.0
[87] parallel_4.4.2 restfulr_0.0.15
[89] pillar_1.10.2 vctrs_0.6.5
[91] promises_1.3.3 evaluate_1.0.4
[93] cli_3.6.5 compiler_4.4.2
[95] futile.options_1.0.1 Rsamtools_2.22.0
[97] rlang_1.1.6 crayon_1.5.3
[99] labeling_0.4.3 plyr_1.8.9
[101] fs_1.6.6 stringi_1.8.7
[103] viridisLite_0.4.2 BiocParallel_1.40.2
[105] lazyeval_0.2.2 GOSemSim_2.32.0
[107] Matrix_1.7-3 hms_1.1.3
[109] patchwork_1.3.1 bit64_4.6.0-1
[111] KEGGREST_1.46.0 SummarizedExperiment_1.36.0
[113] igraph_2.1.4 memoise_2.0.1
[115] bslib_0.9.0 ggtree_3.14.0
[117] fastmatch_1.1-6 bit_4.6.0
[119] ape_5.8-1 gson_0.1.0