Variable Coding
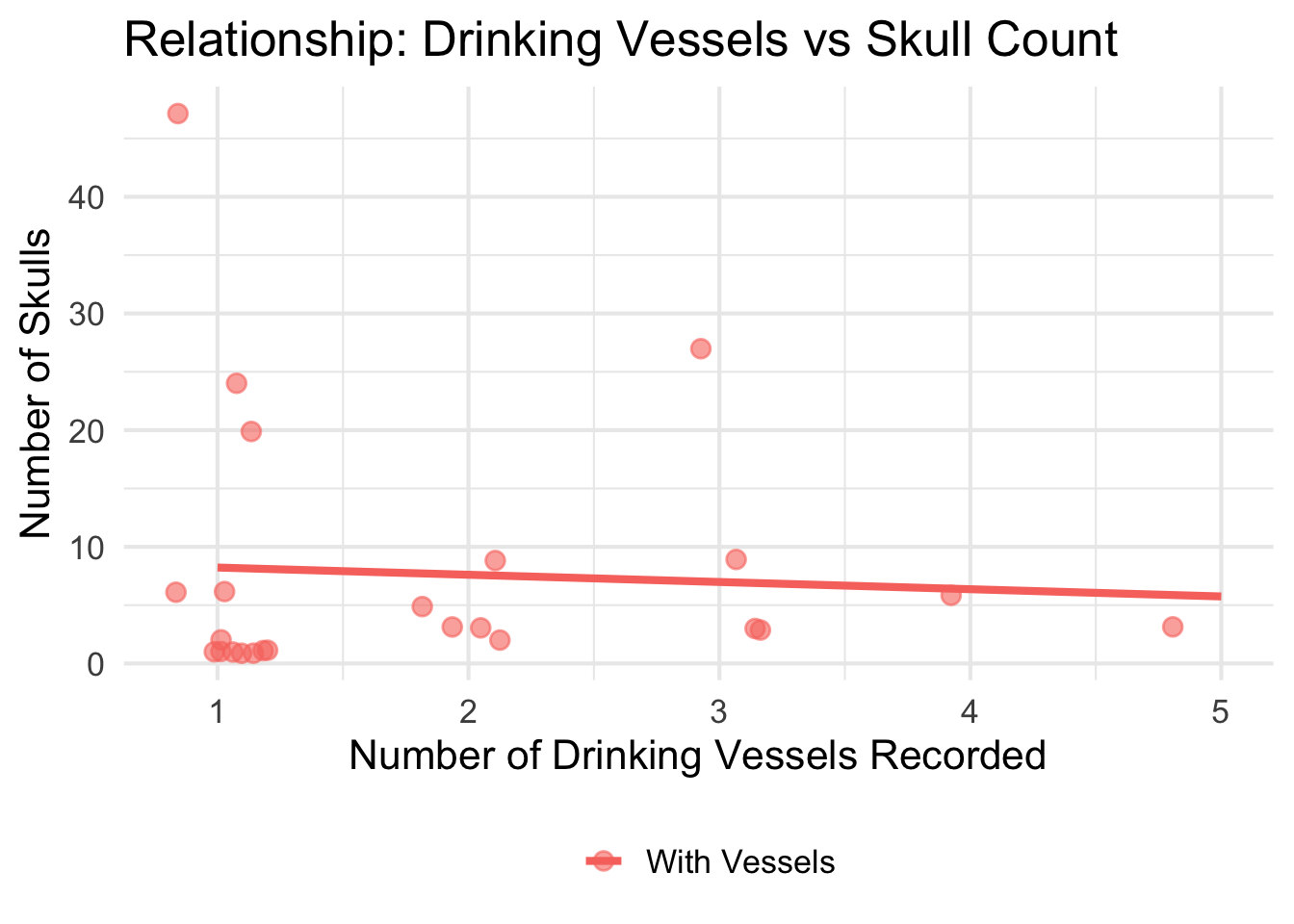
Raw text fields were recoded into analysis-ready variables. Drinking
vessel presence, skeletal remains, and hair pin presence were each
recorded as binary yes/no fields in the spreadsheet and converted to
logical values (TRUE/FALSE); entries that
could not be classified were retained as NA and excluded
from relevant calculations. Primary and secondary burial were coded from
“present”/“absent” fields in the same way.
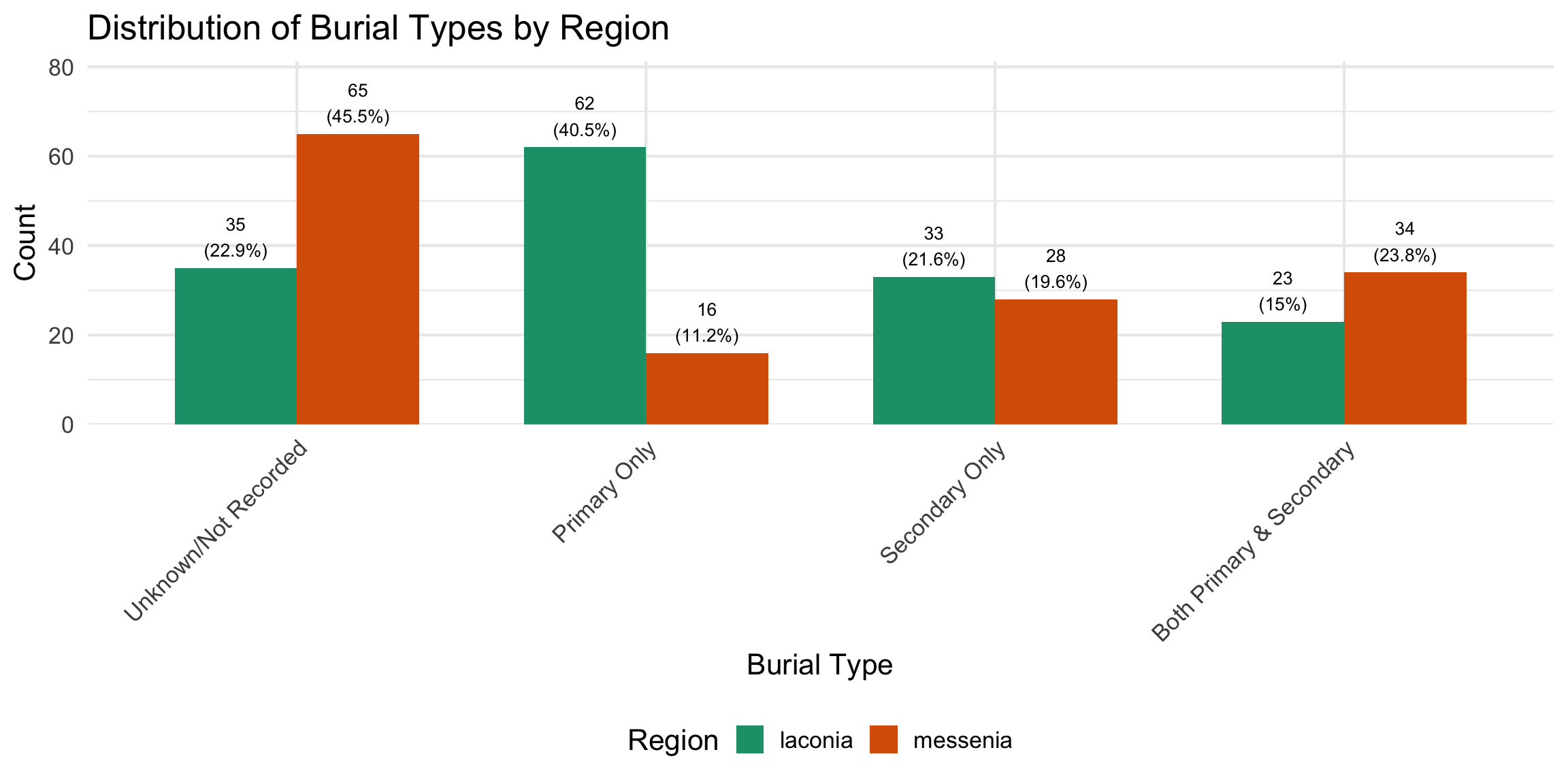
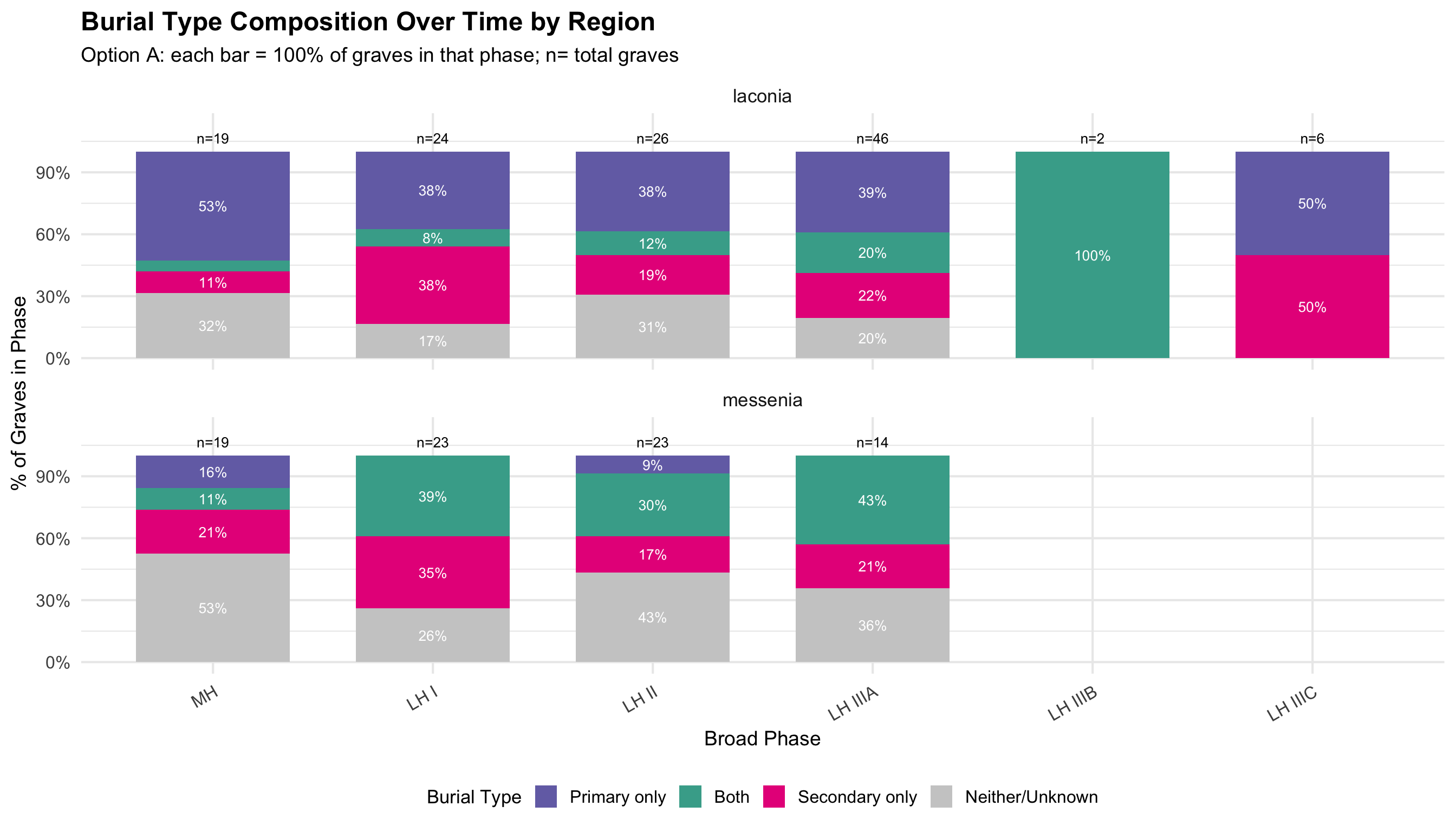
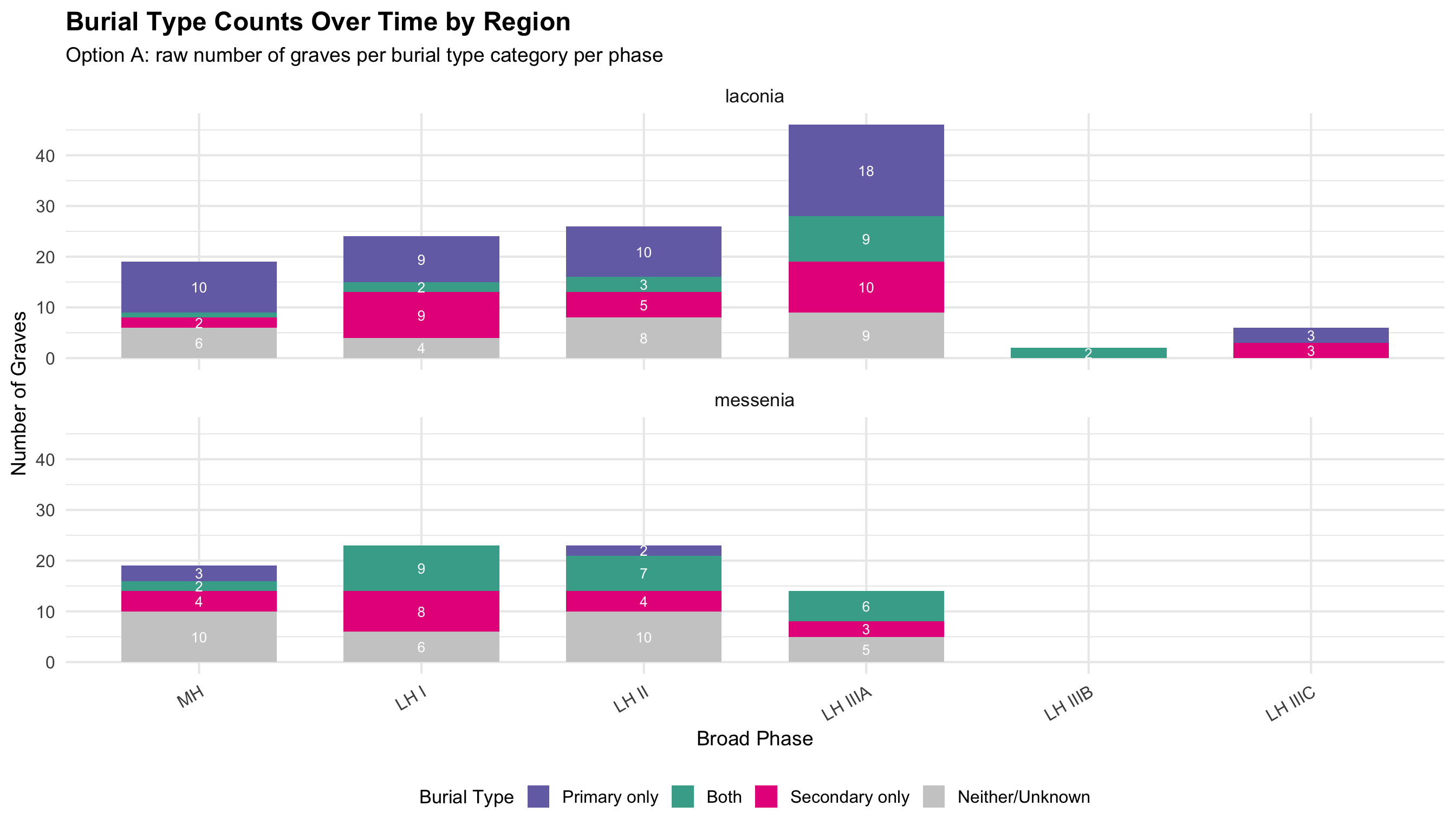
Burial contexts were then classified into four mutually exclusive
categories based on the combination of primary and secondary burial
evidence: Primary Only, Secondary Only, Both
Primary and Secondary, and Unknown/Not Recorded. Tomb
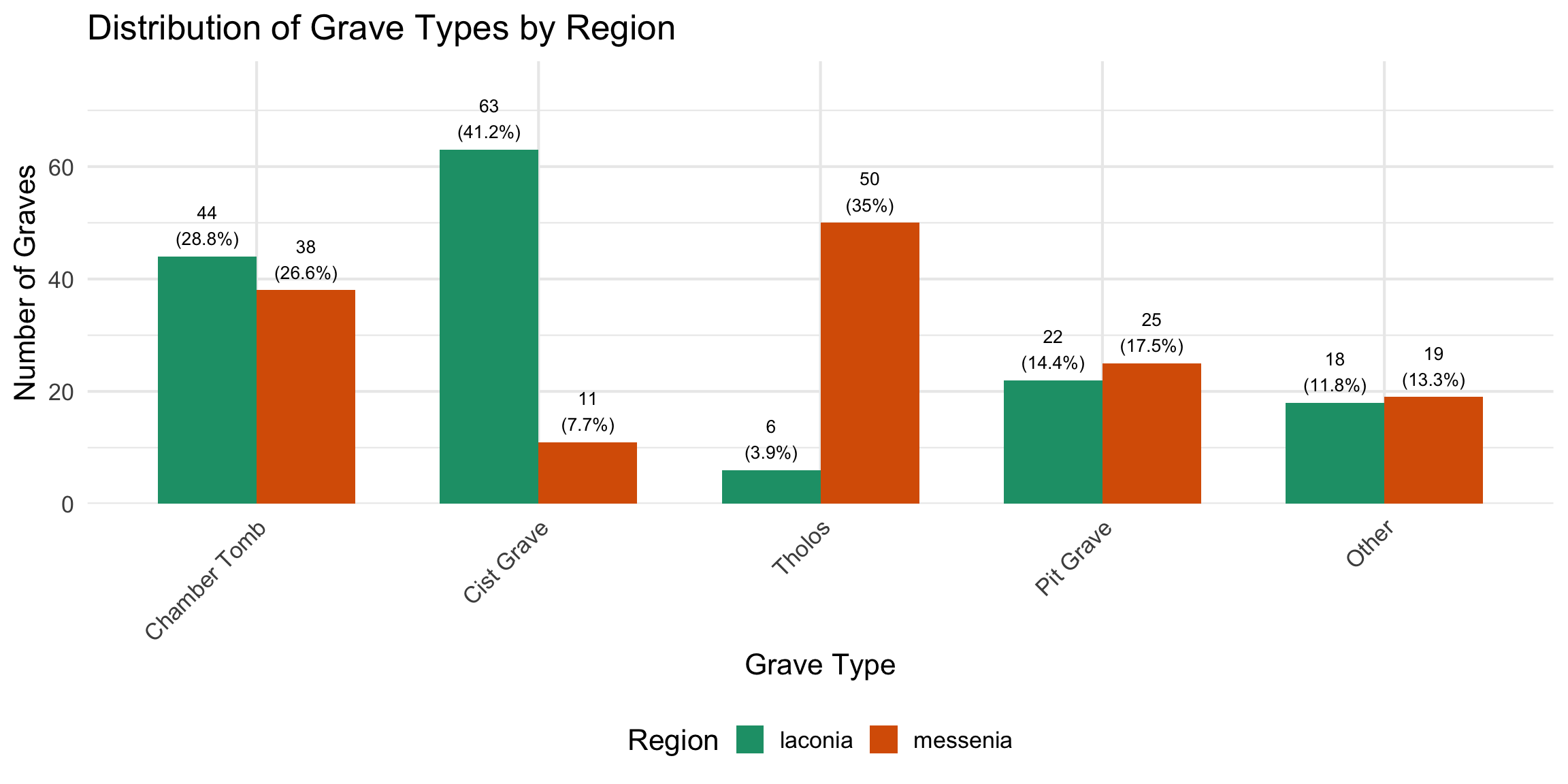
types were simplified from their original descriptive labels into five
structural categories — Tholos, Chamber Tomb, Cist Grave, Pit Grave, and
Dromos Only — to allow comparison across sites where nomenclature
varies.
cups_skulls_clean <- cups_skulls %>%
# Clean column names
janitor::clean_names() %>%
# Trim whitespace from character columns
mutate(across(where(is.character), str_trim)) %>%
# Standardize yes/no values in drinking vessel column
mutate(
has_drinking_vessel = case_when(
tolower(drinking_vessel) == "yes" ~ TRUE,
tolower(drinking_vessel) == "no" ~ FALSE,
TRUE ~ NA
),
has_remains = case_when(
tolower(remains) == "yes" ~ TRUE,
tolower(remains) == "no" ~ FALSE,
TRUE ~ NA
),
primary_burial = case_when(
tolower(primary) == "present" ~ TRUE,
tolower(primary) == "absent" ~ FALSE,
TRUE ~ NA
),
secondary_burial = case_when(
tolower(secondary) == "present" ~ TRUE,
tolower(secondary) == "absent" ~ FALSE,
TRUE ~ NA
),
has_pin = case_when(
tolower(hair_pin) == "yes" ~ TRUE,
tolower(hair_pin) == "no" ~ FALSE,
TRUE ~ NA
)
) %>%
# Create burial category
mutate(
burial_category = case_when(
primary_burial & secondary_burial ~ "Both Primary & Secondary",
primary_burial & !secondary_burial ~ "Primary Only",
!primary_burial & secondary_burial ~ "Secondary Only",
TRUE ~ "Unknown/Not Recorded"
)
) %>%
# Simplify grave type
# ** YOU CAN CUSTOMIZE THIS SECTION **
# If you want different groupings, edit the patterns here
# Current grouping is based on archaeological complexity
mutate(
grave_type_simple = case_when(
str_detect(tolower(type), "tholos") ~ "Tholos",
str_detect(tolower(type), "chamber") ~ "Chamber Tomb",
str_detect(tolower(type), "cist") ~ "Cist Grave",
str_detect(tolower(type), "pit") ~ "Pit Grave",
str_detect(tolower(type), "dromos") ~ "Dromos Only",
TRUE ~ "Other"
)
)
# Show cleaned data structure
glimpse(cups_skulls_clean)
Rows: 296
Columns: 22
$ grave_id <chr> "vapheio_tholos", "arkynes_tholos_A", "arkyn…
$ site <chr> "Vapheio", "Arkynes", "Arkynes", "Arkynes", …
$ type <chr> "tholos", "tholos", "tholos", "tholos", "tho…
$ period <chr> "LHIIA-LHIIIA1", "LHIIIA- LHIIIB", "LHIII", …
$ secondary <chr> "absent", "absent", "absent", "present", "pr…
$ primary <chr> "absent", "absent", "present", "absent", "pr…
$ dimensions <chr> "83.28m^2. dromos: L 29.8m x w3.18-3.45, sto…
$ orientation <chr> "NE-SW", "SE-NW", NA, NA, "E-W", "N-S", "W-E…
$ drinking_vessel <chr> "yes", "no", "no", "no", "no", "no", "no", "…
$ remains <chr> "yes", "yes", "yes", "yes", "yes", "no", "no…
$ drinking_vessel_number <dbl> 6, NA, NA, NA, NA, NA, NA, 4, NA, NA, 5, NA,…
$ skulls_number <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, …
$ area_of_chamber_estimated <dbl> 84.130, 17.350, NA, 28.274, 7.793, NA, 18.32…
$ region <chr> "laconia", "laconia", "laconia", "laconia", …
$ hair_pin <chr> "yes", "no", "no", "yes", "no", "no", "no", …
$ has_drinking_vessel <lgl> TRUE, FALSE, FALSE, FALSE, FALSE, FALSE, FAL…
$ has_remains <lgl> TRUE, TRUE, TRUE, TRUE, TRUE, FALSE, FALSE, …
$ primary_burial <lgl> FALSE, FALSE, TRUE, FALSE, TRUE, FALSE, FALS…
$ secondary_burial <lgl> FALSE, FALSE, FALSE, TRUE, TRUE, FALSE, FALS…
$ has_pin <lgl> TRUE, FALSE, FALSE, TRUE, FALSE, FALSE, FALS…
$ burial_category <chr> "Unknown/Not Recorded", "Unknown/Not Recorde…
$ grave_type_simple <chr> "Tholos", "Tholos", "Tholos", "Tholos", "Tho…
Chronological Classification
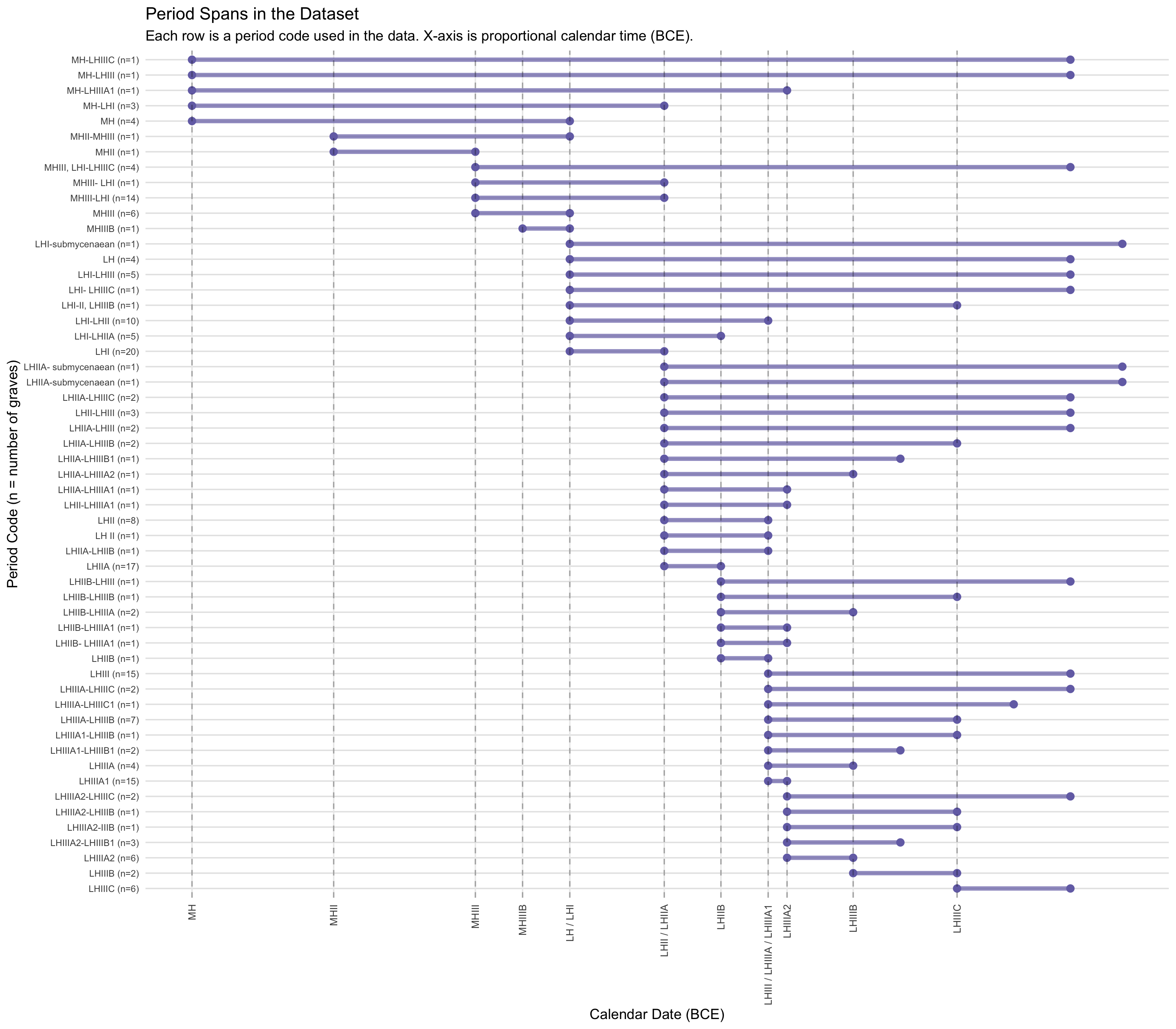
Period strings as recorded in the original dataset (e.g., “LHIIA,”
“LHI–LHIII”) were matched against a period dictionary that assigns each
code an earliest and latest component phase. Calendar date ranges were
then attached by joining against a lookup table of BCE start and end
dates for each component phase, based on conventional Aegean Bronze Age
chronology. The period dictionary and calendar date table are loaded and
applied below.
# NOTE: Set eval=TRUE once the student has completed the dictionaries
# Load period dictionary
period_dict <- read_csv(here::here("data", "period_dictionary.csv")) %>%
mutate(across(where(is.character), str_trim))
# Load tomb type dictionary
tomb_dict <- read_csv(here::here("data", "tomb_type_dictionary.csv")) %>%
mutate(tomb_type = str_trim(tomb_type))
# Join period dictionary and extract period components
cups_skulls_clean <- cups_skulls_clean %>%
left_join(
period_dict %>% select(period_code, chronological_order, earliest_component, latest_component),
by = c("period" = "period_code")
) %>%
# Get ordinal values for earliest component
left_join(
period_dict %>% select(period_code, chronological_order) %>% rename(start_order = chronological_order),
by = c("earliest_component" = "period_code")
) %>%
# Get ordinal values for latest component
left_join(
period_dict %>% select(period_code, chronological_order) %>% rename(end_order = chronological_order),
by = c("latest_component" = "period_code")
) %>%
# For single periods (where start = end), use that value
# For ranges, create midpoint
mutate(
period_start_order = start_order,
period_end_order = end_order,
period_midpoint_order = (start_order + end_order) / 2
)
# Calendar year lookup for each single-component period code
# Sources: user-supplied dates; sub-phases (MHIIIB, LHIIIB1, LHIIIC1) estimated
component_cal <- tribble(
~component, ~cal_start_bce, ~cal_end_bce,
"MH", 2000, 1600,
"MHII", 1850, 1700,
"MHIII", 1700, 1600,
"MHIIIB", 1650, 1600,
"LH", 1600, 1070,
"LHI", 1600, 1500,
"LHII", 1500, 1390,
"LHIIA", 1500, 1440,
"LHIIB", 1440, 1390,
"LHIII", 1390, 1070,
"LHIIIA", 1390, 1300,
"LHIIIA1", 1390, 1370,
"LHIIIA2", 1370, 1300,
"LHIIIB", 1300, 1190,
"LHIIIB1", 1300, 1250,
"LHIIIC", 1190, 1070,
"LHIIIC1", 1190, 1130,
"submycenaean", 1070, 1015
)
# Add calendar year columns: start = start of earliest component,
# end = end of latest component
cups_skulls_clean <- cups_skulls_clean %>%
left_join(
component_cal %>% select(component, cal_start_bce),
by = c("earliest_component" = "component")
) %>%
rename(period_start_cal = cal_start_bce) %>%
left_join(
component_cal %>% select(component, cal_end_bce),
by = c("latest_component" = "component")
) %>%
rename(period_end_cal = cal_end_bce) %>%
mutate(period_midpoint_cal = (period_start_cal + period_end_cal) / 2)
# Broad phase groupings with calendar spans for temporal plots
phase_cal <- tribble(
~broad_phase, ~phase_start_bce, ~phase_end_bce,
"MH", 2000, 1600,
"LH I", 1600, 1500,
"LH II", 1500, 1390,
"LH IIIA", 1390, 1300,
"LH IIIB", 1300, 1190,
"LH IIIC", 1190, 1070,
"SM", 1070, 1015
) %>% mutate(
phase_mid_bce = (phase_start_bce + phase_end_bce) / 2,
phase_duration = phase_start_bce - phase_end_bce,
broad_phase = factor(broad_phase,
levels = c("MH", "LH I", "LH II", "LH IIIA", "LH IIIB", "LH IIIC", "SM"),
ordered = TRUE)
)
# Assign each grave to broadest phase based on its earliest component
cups_skulls_clean <- cups_skulls_clean %>%
mutate(
broad_phase = case_when(
earliest_component %in% c("MH", "MHII", "MHIII", "MHIIIB") ~ "MH",
earliest_component %in% c("LH", "LHI") ~ "LH I",
earliest_component %in% c("LHII", "LHIIA", "LHIIB") ~ "LH II",
earliest_component %in% c("LHIII", "LHIIIA", "LHIIIA1", "LHIIIA2") ~ "LH IIIA",
earliest_component %in% c("LHIIIB", "LHIIIB1") ~ "LH IIIB",
earliest_component %in% c("LHIIIC", "LHIIIC1") ~ "LH IIIC",
earliest_component == "submycenaean" ~ "SM",
TRUE ~ NA_character_
),
broad_phase = factor(broad_phase,
levels = c("MH", "LH I", "LH II", "LH IIIA", "LH IIIB", "LH IIIC", "SM"),
ordered = TRUE)
)
# Map grave_type_simple names to tomb_type dictionary names
grave_to_tomb_map <- tibble(
grave_type_simple = c("Tholos", "Chamber Tomb", "Cist Grave", "Pit Grave", "Dromos Only", "Other"),
tomb_type_dict = c("tholos", "chamber tomb", "cist", "pit", "dromos of tholos", NA)
)
# Join tomb type dictionary via mapping
cups_skulls_clean <- cups_skulls_clean %>%
left_join(grave_to_tomb_map, by = "grave_type_simple") %>%
left_join(
tomb_dict %>% select(tomb_type, tomb_type_order),
by = c("tomb_type_dict" = "tomb_type")
) %>%
select(-tomb_type_dict) %>%
# Convert to ordered factors
mutate(
tomb_type_ordered = factor(
grave_type_simple,
levels = grave_to_tomb_map$grave_type_simple[
order(tomb_dict$tomb_type_order[match(grave_to_tomb_map$tomb_type_dict, tomb_dict$tomb_type)])
],
ordered = TRUE
),
period_ordered = factor(
period,
levels = arrange(period_dict, chronological_order) %>% pull(period_code),
ordered = TRUE
)
)
cat("✓ Dictionaries successfully loaded and applied!\n")
✓ Dictionaries successfully loaded and applied!
cat("New columns created:\n")
New columns created:
cat(" - period_start_order: chronological order of earliest period component\n")
- period_start_order: chronological order of earliest period component
cat(" - period_end_order: chronological order of latest period component\n")
- period_end_order: chronological order of latest period component
cat(" - period_midpoint_order: midpoint between start and end\n")
- period_midpoint_order: midpoint between start and end
cat(" - tomb_type_ordered: ordered factor for tomb types\n")
- tomb_type_ordered: ordered factor for tomb types
cat(" - period_ordered: ordered factor for periods\n")
- period_ordered: ordered factor for periods